Abstract
Background
The brown planthopper (BPH), an insect species that feeds on rice plants (Oryza sativa L.), is one of the most destructive insect pests of rice. BPH can be controlled with chemical pesticides, but they are expensive in terms of the cost and environmental hazards. Hence, utilization of resistance genes in resistant varieties is considered as a more economical and eco-friendly effective method for control.
Results
In this study, six dominant BPH-resistance genes (Bph3, Bph14, Bph15, Bph18, Bph20 and Bph21) were introduced into an elite indica rice cultivar Jin 23B using the marker-assisted selection (MAS) method of breeding. One line combining Bph14 and Bph15 and six single gene introgression lines were used to evaluate the gene effects based on three parameters: seedling and tillering resistance of the rice genotypes, honeydew weight, and BPH survival rate. Among all improved lines, combination of Bph14 and Bph15 had the largest effect in conferring resistance to BPH. Bioassays showed that the order of the gene effects was Bph14/Bph15 > Bph15 ≥ Bph14 ≥ Bph20 ≥ Bph21 ≥ Bph3 > Bph18 > none at the rice seedling stage. The pyramided or single-gene introgression lines showed enhanced resistance relative to the control. Furthermore, field trial data demonstrated that yields of improved Jin 23B lines were similar to the control under BPH-free field conditions.
Conclusions
Each of the BPH resistance genes reduced BPH growth and development, and was effective at both the seedling and tillering growth stages. These lines can be used in rice hybrid and production in China.
Similar content being viewed by others
Background
Brown planthopper (Nilaparvata lugens Stål, BPH), is one of the most destructive insect pests of tropical and temperate rice (O. sativa L.) in Asia. BPH extracts phloem saps from host plants using its stylet-type mouthparts, resulting in severe and damaging symptom known as ‘hopper-burn’ (Watanabe and Kitagawa 2000). BPH also transmits viruses (Cha et al. 2008), such as grassy stunt virus and rugged stunt virus to rice plants, causing further yield losses in Asian countries, including China, Japan, Korea and Vietnam (Brar et al. 2010). In rice production, BPH is managed mainly by using insecticides that are expensive in terms of cost and environmental hazards. Therefore, utilization of BPH resistance genes in developing resistant varieties is considered as the most economical and eco-friendly approach to control this pest (Tanaka et al. 2000).
More than 32 BPH resistance loci have been identified in indica rice cultivars and wild rice species (Ali and Chowdhury 2014; Wang et al. 2015). Several of these resistance genes have been assigned to rice chromosomes 2, 3, 4, 6, and 12 (Jena and Kim 2010) and at least ten genes have been fine-mapped to genomic region of less than 200 kb (Hu et al. 2016). Most of the resistance alleles are dominant, but a few are documented as recessive (bph4, bph5, bph7, bph8, bph19,bph25 and bph29). Seven BPH-resistance genes Bph14, Bph26, Bph17, bph29, Bph9, Bph18 and Bph32 have been cloned by map-based cloning (Du et al. 2009; Tamura et al. 2014; Liu et al. 2015; Wang et al. 2015; Zhao et al. 2016; Ji et al. 2016; Ren et al. 2016). Among the resistance loci, the broad-spectrum resistance gene Bph3, identified in cultivars Rathu Heenati and PTB33, was mapped on the short arm of chromosome 6 where it was flanked by SSR markers RM589 and RM588 (Jairin et al. 2007a). Bph14 and Bph15 were previously mapped in introgression line B5 (derived from O. officinalis), with QBph1 (renamed as Bph14) on the long arm of chromosome 3 explaining 26.4% of the variation in BPH response and Qbph2 (renamed as Bph15) on the short arm of chromosome 4 explaining 14.3% of the variation (Huang et al. 2001). Bph14, encoding a coiled-coil, nucleotide-binding, and leucine-rich repeat protein was cloned using a map-based cloning approach (Du et al. 2009). Bph15 was fine mapped to an interval of approximately 50 kb between map positions 6,883,264 and 6,934,420 (Yang et al. 2004). The Bph18 gene, originally derived from O. australiensis, was mapped in indica introgression line IR65482–7–216-1-2, shown to be on the long arm of chromosome 12, and explained 47% of the variation in BPH response (Jena et al. 2006). Map-based cloning and complementation test revealed that Bph18 encodes a CC-NBS-NBS-LRR protein and is a functionally different allele of Bph26 (Ji et al. 2016). Bph20 and Bph21 in introgression line IR71033–121-15 were derived from O. minuta (Accession number 101141), with Bph20 located to a 193.4 kb region on the short arm of chromosome 4, and with Bph21 located to a 194 kb region on the long arm of chromosome 12 (Rahman et al. 2009).
Near-isogenic lines (NILs) and gene combination lines carrying one or several resistance genes have been developed (Sharma et al. 2004; Hu et al. 2012, 2013) and in an effective manner to facilitate the use of the resistance genes or of quantitative trait loci (QTLs) for future rice improvement program. Bph1, Bph3 and Bph4 have already been successfully introduced into some elite cultivars using MAS and backcross breeding programs at the International Rice Research Institute (IRRI) in the Philippines (Khush and Brar 1991; Jena and Kim 2010). Qiu et al. (2011) used MAS to pyramid Bph6 and Bph12 into a japonica line and an indica line. Results of BPH bioassays indicated that the pyramided line showed higher resistance than the Bph6- or Bph12-single introgression line. Hu et al. (2012) incorporated Bph14 and Bph15 genes into Minghui 63 and its hybrid Shanyou 63, and showed that hybrids containing a single BPH resistance gene had enhanced resistance compared to conventional hybrids that lack of resistance gene. Lines combining two genes were more resistant than the respective single gene lines. Hu et al. (2013) incorporated Bph14, Bph15 and Bph18 into cv. 9311 and its hybrids GZ63-4S/9311 and Y58S/9311. Test of BPH response showed that lines with three gene combined were more resistant than lines with two genes or lines with single genes.
The three-line hybrid breeding system is an important method for producing commercial rice hybrids in China and elsewhere. The system involves a cytoplasmic male sterile (CMS) line (A), a fertile maintainer line (B), and a restorer line. The A line is used as a female parent and the restorer line is used as male parent for hybrid seed production. The B line is near-isogenic to the A-line except for its pollen fertility. The B line has viable pollen grains and normal seed setting, so it can pollinate the A line, while the F1 plants from this pollination are male sterile, again. In this way, the male sterility of the A line is maintained, and the A line is reproduced for further use or commercial use in a large scale. Jin 23B is a maintainer line of male-sterile Jin 23A, whereas Jin 23A is an elite indica cytoplasmic male sterile (CMS) line widely used to produce three-line hybrids over the last 20 years in China. More than 100 different hybrids are derived from Jin 23A (http://www.ricedata.cn/), and in 2008 and 2009, Jin 23A-derived hybrid varieties are grown on about 1.7 million ha of rice in China with the highest ranking (Shi et al. 2012). However, these hybrids and their parents have become increasingly susceptible to blast and BPH due to absence of resistance genes and narrow genetic diversity. Blast resistance was recently improved by the introduction of genes Pi1, Pi2 and D12, into Jin 23B by marker assisted backcross breeding (Jiang et al. 2012). To similarly improve BPH resistance, we introduced BPH resistance genes Bph3, Bph14, Bph15, Bph18, Bph20 and Bph21 into Jin 23B with comprehensive evaluation. These lines will contribute to ensuring the security of rice production in China.
Methods
Plant materials and insects
PTB33 (carrying Bph3, abbreviated as 3), served as the donor of Bph3. B5, an introgression line derived from O. officinalis (Huang et al. 2001), was the donor of resistance genes Bph14 and Bph15 (abbreviated as 14 and 15). IR65482–7–216-1-2 (carrying Bph18, abbreviated as 18), an introgression line from O. australiensis (Acc. No. 10082), served as the donor of Bph18. IR71033–121-15 (carrying Bph20 and Bph21, abbreviated as 20 and 21), served as the donor of Bph20 and Bph21. B5 was obtained from College of Life Sciences, Wuhan University, China. IR65482–7–216-1-2, IR71033–121-15 and PTB33 were obtained from the Plant Breeding, Genetics, and Biotechnology Division of the International Rice Research Institute (IRRI), Los Baños, Philippines. Jin 23B, an elite maintainer indica rice obtained from Hunan Hybrid Rice Research Center, Changsha, China, was used as the recurrent parent. PTB33 and Taichung Native 1 (TN1; no resistance) were used as checks in evaluation of BPH resistance.
The BPH used for infestation were collected from rice fields at Wuhan, in 2012, and were maintained in a cage by continuous rearing on the susceptible variety TN1, at the glasshouse facility at Huazhong Agricultural University. Before use in experiments, one- to five-instar BPH nymphs were prepared and maintained on TN1.
Molecular markers used for MAS
SSR markers RM589 and RM19324 (F: GAGGTTGTTTGGATGGATAGATGG and R: AATCCCGTCCTAGAGTTCTTCTACC) were used to detect the presence of Bph3 (Jairin et al. 2007a). The co-segregating insertion/deletion (InDel) marker 76–2 was used for Bph14 identification (Du et al. 2009). The Bph15 gene was identified using simple sequence repeat (SSR) marker RM261 and InDel marker 15–6 (Huang et al. 2001; Yang et al. 2004; Hu et al. 2012). SSR markers RM3331 (F: CCTCCTCCATGAGCTAATGC and R: AGGAGGAGCGGATTTCTCTC), RM28427 (F: CTGTGAGAAGGTTGAGACTTGAAAGG and R: GCAAATGCTCAAGTGAAGTTGG) and RM28561 (F: CTTCAAGACTGGCCCAATATTACTGC and R: TGACTGAAGCCTTCTTCACTTGC), were used to detect the presence of Bph18. SSR marker RM16553 (F: CATAGCCACTTATCGTTGTTACGC and R: TGTCCATCTATGACTGTCCACTACG) and Indel marker HJ34 (F: GCCGAATGGTAAGAAGAG and R: GCGAGTTAACCAATGCTTGG) were used to detected the presence of the Bph20 gene. SSR marker RM28561 (F: CTTCAAGACTGGCCCAATATTACTGC and R: TGACTGAAGCCTTCTTCACTTGC) and PCR-based marker B121 were used to detected the presence of the Bph21 gene (Rahman et al. 2009). SSR analysis was carried out according to the procedures described by McCouch et al. (2002). The Bph3, Bph14, Bph15, Bph18, Bph20, and Bph21 genes were introduced into Jin 23B following a MAS-based backcross breeding strategy. Finally, six single gene introgression lines containing Bph3, Bph14, Bph15, Bph18, Bph20 and Bph21 and a pyramided line containing Bph14 and Bph15 were obtained. The selected lines maintained the excellent quality and yield potential of Jin 23B. A total of seven homozygous gene combinations (designated as 14/15, 3, 14, 15, 18, 20 and 21) were represented.
The whole-genome single nucleotide polymorphism (SNP) array RICE6K was used for genetic background profiling of the introgression lines (Yu et al. 2014). Genomic DNA from each line was extracted from 10 dry seeds. DNA amplification, fragmentation, chip hybridization, single base extension, staining and scanning were conducted by the Life Science and Technology Center, China National Seed Group Co., LTD (Wuhan, China).
Evaluation of BPH response in seedlings and tillering stage of plants
An Experiment was conducted by using a modified bulk method under sophisticated greenhouse condition followed Pathak et al. (1969). PTB33 and TN1 were included as controls in all experiments. Lines were sown in plastic trays (60 cm length × 40 cm width × 10 cm height) with 12 plants per row, 11 rows per tray. When seedlings at 10 days post sowing (three-leaf stage) were infested with second and third instar nymphs (twelve nymphs per seedling). Each experiment was conducted in a randomized complete block design with six replications. Plant responses were recorded seven days after infestation (DAI) when all seedlings of TN1 were almost dead. BPH resistance was evaluated with the six-scale standard of scoring system, described by Huang et al. (2001): 0 = no damage; 1 = very slight damage; 3 = first and second leaves partially yellowing; 5 = pronounced yellowing and stunting; 7 = mostly wilting, the plant still alive; 9 = the plant completely wilted or died, in which higher scores indicate greater susceptibility to BPH.
The recurrent parent, backcross lines, resistant check and susceptible check (TN1) seeds were sown in the field at Wuhan for plant BPH response assessments. Fifteen days later two plants of each line and checks were transplanted into a plastic pot (16.5 cm of bottom diameter × 22.5 cm of top diameter × 19 cm of height) containing pulverized soil in ten replications. All the plants were grown in a cage at the glasshouse facility. At the tillering stage (30 days post transplanting) the plants were infested with second and third instar nymph at a density of 300 nymphs per plant. When the susceptible check was almost dead (approximately 17 days after infestation) all plants were assessed and scored as resistant (R), moderately resistant (MR), moderately susceptible (MS) and highly susceptible (S) (Suh et al. 2011). Scoring occurred a second time about 25 days post infestation.
Quantification of honeydew excretion from adult BPH females
Honeydew collection was accomplished under aseptic condition under greenhouse followed by Pathak et al. (1982) with minor modifications. Seven improved rice families with BPH resistance gene Bph3, Bph14, Bph15, Bph18, Bph20, Bph21 and Bph14/Bph15 and the control Jin 23B and PTB33 were planted in plastic pots. 50 days after transplanting, five-instar BPH female nymph (previously starved for 2 h) were inoculated with a suitable parafilm sheet (3.5 × 4.5 cm) (Incubation rate = two females per sheet) and subjected to the improved lines with suitable check. After 24 h of feeding, insects were removed from the sheet, and honeydew in each sheet were weighted using a 0.1 mg sensitivity balance. An experiment was repeated over four replications.
BPH survival on rice plants
BPH survival measurements were made on the backcross lines and suitable checks following the method of Sebastian et al. (1996). Individual seedlings were grown on a rice liquid culture medium in 500 ml volumetric flasks. Ten insects with second-instar nymph were subsequently put into each volumetric flask, and survivability of BPHs in each flask were recorded in every 2 days of interval for 10 days. The experiment was carried out in greenhouse, and the temperature was maintained at 25–28 °C. BPH survival percentage was calculated as follows. BPH survival percentage = Number of surviving nymph divided by the total number of released nymph. This measurement was repeated for six times.
Pollen fertility of improved lines
Fertility assessments of the backcross lines were made using six to ten selected lines for each genotype. Crosses were made to the male sterile Jin 23A, and the F1 plants were used as female parents in crosses with the corresponding improved lines. Six spikelets per line of each F1 and BC1F1 individual were sampled for assessment of pollen fertility using 1% I2-KI staining (Virmani et al. 1997). The fertility rate of each line was scored as completely sterile (0%), partially sterile (1–30%), partially fertile (> 30–60%) and fertile (> 60%) (Govinda Raj and Virmani 1988).
Evaluation of agronomic traits in the field
Agronomic assessments of the seven backcross lines were carried under natural field conditions, seven homozygous rice families and the control (Jin 23B) were planted in randomized complete block design in Wuhan, China in the autumn of 2013. A trial was performed with six replications, each plot consisted of three rows and each row consisted of 12 plants at a planting density of 17 × 27 cm (17 cm plant to plant and 27 cm row to row distance). Seven plants from central row of each plot were subjected to agronomic trait measurements, including days to heading (DTH) and plant height (PH). After the maturity of rice, the seven plants from central row were harvested individually for panicle number (PN), number of grains per panicle (NGP), spikelet fertility (SF), weight of 1000-grains (GW) and yield per plant (YD) measurement. Filed management followed normal agricultural practices with the nitrogen, phosphorus, potassium fertilizer of 180, 65, and 65 kg/hm2 respectively.
Data analysis
The available RM marker series were searched in the rice genomic database (www.gramene.org). InDel markers were designed based on the references genome of Nipponbare and 9311 reference sequences. Date analysis was performed using one-way ANOVA and Tukey’s tests in Microsoft Excel 2003 or SPSS 17.0.
Results
Development of introgressed and pyramided lines of Jin 23B using marker assisted selection
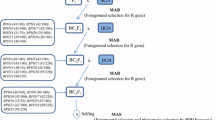
BPH resistance genes were introgressed from donor parents into Jin 23B following a recurrent backcrossing procedure combined with MAS (Fig. 1). F1 progenies obtained from crosses between Jin 23B and each donor, namely PTB33 (carrying Bph3), B5 (carrying Bph14/Bph15), IR65482–7–216-1-2 (carrying Bph18), IR71033–121-15 (carrying Bph20/Bph21). In each BC generation, positive individuals were identified using corresponding markers, and then used for further backcrossing with the recurrent parent Jin 23B. In the BC1F1 population, 19, 27, 21, 22, 24, 23 and 19 individuals were identified for genes Bph3, Bph14, Bph15, Bph18, Bph20, Bph21 and Bph14/Bph15 respectively. In the BC2F1 population, 18, 25, 29, 24, 22, 25 and 18 individuals were identified for gene Bph3, Bph14, Bph15, Bph18, Bph20, Bph21 and Bph14/Bph15. These plants were subsequently backcrossed to Jin 23B by mixing pollen grains. After three generation of backcrossing, homozygous BC3F2 plants were selected (Fig. 1). With dual selection (MAS + BPH bioassay) and further field selection we obtained sets of homozygous lines for Bph3, Bph14, Bph15, Bph18, Bph20, Bph21 and Bph14/15, which were renamed as Jin 23B (3) (16 lines), Jin 23B (14) (28 lines), Jin 23B (15) (26 lines), Jin 23B (18) (23 lines), Jin 23B (20) (16 lines), Jin 23B (21) (25 lines) and Jin 23B (14/15) (15 lines). Following self-pollinating and dual selection, Jin 23B improved lines with enhanced BPH resistance and high yield were developed.
Genetic background examination of the introgressed lines and pyramided line
In this study, RICE6K, a whole-genome single nucleotide polymorphism (SNP) array was used to analyze the genetic background of the introgressed line and pyramided line. This array contained 1111 polymorphic SNP markers between Jin 23B and PTB33, 1040 polymorphic SNP markers between Jin 23B and B5, 682 polymorphic SNP markers between Jin 23B and IR65482–7–216-1-2, and 1050 polymorphic SNP markers between Jin 23B and IR71033–121-15. Populations developed after three backcrosses are expected to have about 87.5% recovery of the recurrent parent genotype. As shown in Fig. 2 the genetic background recoveries of the recurrent parent in Jin 23B (3), Jin 23B (14), Jin 23B (15), Jin 23B (18), Jin 23B (20), Jin 23B (21) and Jin 23B (14/15) were 91.72%, 94.13%, 92.02%, 86.95%, 90.67%, 83.43% and 77.11%, respectively, measured by percentage of the polymorphic marker ratios. Large fragments of donor chromatin were present in the positions of Bph3 in chromosome 6, Bph14 in chromosome 3, Bph15 and Bph20 in chromosome 4, and Bph18 and Bph21 in chromosome 12, because of foreground selection of the target gene in each generation by MAS.
Haplotype maps of genetic background profiling of improved Jin 23B using the RICE6K array. (a) Jin 23B (Bph3), (b) Jin 23B (Bph14), (c) Jin 23B (Bph15), (d) Jin 23B (Bph18), (e) Jin 23B (Bph20), (f) Jin 23B (Bph21), (g) Jin 23B (Bph14/ Bph15). Donor parents included 4 cultivars: PTB33 (Bph3), B5 (Bph14/Bph15), IR65482–7–216-1-2 (Bph18) and IR71033–121-15 (Bph20/Bph21). Arrowheads with red color indicate the positions of introduced gene. Arrowheads with black color indicate the positions of molecular marker used in marker-assisted selection backcross breeding program
BPH responses of the introgression lines at seedling and tillering stages
Introgression lines with homozygous Bph3, Bph14, Bph15, Bph18, Bph20 and Bph21 were used for evaluation of BPH-resistance at both seedling and tillering stage. At the seedling stage (Figs. 3a, 4), the resistance score for Jin 23B harboring Bph14 and Bph15, designated as Jin 23B (14/15), account 2.28, which conferred highest resistance rating among all the selected lines. Introgression lines containing single Bph genes, Jin 23B (15), Jin 23B (14), Jin 23B (20), Jin 23B (21), Jin 23B (3), and Jin 23B (18), with desirable resistance scores were found 3.36, 3.69, 3.75, 3.83, 4.3 and 5.21, respectively, and that for the donor parents PTB33 (carrying Bph3), B5 (carrying Bph14/Bph15), R65482–7–216-1-2 (carrying Bph18) and IR71033–121-15 (carrying Bph20/Bph21) the resistance scores were 3.21, 2.39, 3.86 and 2.67, respectively. In contrast Jin 23B (none) control scored 8.08 (Figs. 3a, 4). For single Bph gene lines, Jin 23B (15) conferred the highest seedling resistance level whereas Jin 23B (18) conferred the lowest seedling resistance level. Furthermore, the pyramided line with two resistance genes (Bph14/Bph15) showed significantly higher resistance compared to the ones with single resistance gene.
a The resistance scores of pyramided line and introgression lines of Jin 23B conferring to BPH at seedling stage. The resistance score was determined by the mean of score from at least 30 lines each homogenous allele combination. b The honeydew weight per BPH females on improved lines of Jin 23B. Number 14/15 represent Jin 23B (Bph14/Bph15), 15 represent Jin 23B (Bph15), 14 represent Jin 23B (Bph14), 20 represent Jin 23B (Bph20), 21 represent Jin 23B (Bph21), 3 represent Jin 23B (Bph3) and 18 represent Jin 23B (Bph18). none (CK): Jin 23B. The different letters above the bars are significantly different at P < 0.05 level
At the tillering stage, all the plants grown in a cage infested with BPH were scored twice to evaluate the resistance degree. Lines Jin 23B (14/15), Jin 23B (15), Jin 23B (14) were classified R to BPH, Jin 23B (20), Jin 23B (21), Jin 23B (3), Jin 23B (18) were MR to BPH, whereas Jin 23B (none) was MS to BPH at 17 days post infestation (Table 1, Fig. 5). Lines Jin 23B (14/15), Jin 23B (15), Jin 23B (14) were classified MR to BPH and Jin 23B (20), Jin 23B (21), Jin 23B (3), Jin 23B (18) were MS to BPH, whereas Jin 23B (none) was S to BPH at the 25 days after infestation (Table 1, Fig. 5). In brief, at tillering stage, Jin 23B (14/15), Jin 23B (15) and Jin 23B (14), showed R to MR to BPH from 17 DAI to 25 DAI, and Jin 23B (20), Jin 23B (21), Jin 23B (3) and Jin 23B (18) showed MR to MS to BPH from 17 DAI to 25 DAI. Whereas Jin 23B (none) showed MS to S to BPH from 17 DAI to 25 DAI.
In general, for pyramided and introgression lines at seedling stage, the order of the gene effect was: Bph14/Bph15 > Bph15 ≥ Bph14 ≥ Bph20 ≥ Bph21 ≥ Bph3 > Bph18 > none (Fig. 3a). At tillering stage, the order of the gene effect was: Bph14/Bph15 = Bph15 = Bph14 > Bph20 = Bph21 = Bph3 = Bph18 > none (Table 1, Fig. 5).
Honeydew weight excreted from BPH
To determine the effects of resistance genes in BPH growth and development, we compared the honeydew weight excreted from BPH after feeding with improved lines (Fig. 3b). For the pyramided or introgression lines, the BPH honeydew weights were gradually increased from 2.11 mg/female on Jin 23B (14/15) to 8.82 mg/female on Jin 23B (18) (Fig. 3b). The honeydew weight for Jin 23B (15), Jin 23B (14), Jin 23B (21), Jin 23B (20), Jin 23B (3) were 3.39, 3.43, 5.19, 6.43 and 7.51 mg/female, respectively (Fig. 3b). In comparison, the negative control, Jin 23B (none), produced the highest honeydew weight of 13.89 mg/female (Fig. 3b), and the resistance control PTB33, B5, IR65482–7–216-1-2 and IR71033–121-15 produced of 3.86, 2.89, 7.12 and 3.36 mg/female. Taken together, the order of the gene effect in BPH honeydew weight was Bph14/Bph15 > Bph15 ≥ Bph14 > Bph21 > Bph20 > Bph3 > Bph18 > none (Fig. 3b).
BPH survival rates
To know the influence of antibiosis in the BPH resistance Bph3, Bph14, Bph15, Bph18, Bph20 and Bph21, the BPH survival rate was measured on the pyramided line and introgression lines in every two days interval for 12 days. As shown in Fig. 6, the average BPH survival rate on Jin 23B (14/15) was the lowest among all genotypes, which was 21.11% at 12 DAI. The average numbers of BPH surviving on Jin 23B (15), Jin 23B (14) and Jin 23B (21) decreased gradually and showed a significant difference in number compared with other genotypes at 8 DAI, and the average survival rates were 34.44%, 38.89% and 51.11%, respectively at 12 DAI. Comparatively, the average survival rates on Jin 23B (20), Jin 23B (3) and Jin 23B (18) were 55.56%, 58.89% and 64.45%, respectively at 12 DAI. For Jin 23B (none) and PTB33, the average survival rate was 81.11% and 42.53% at 12 DAI (Fig. 6).
Generally, these findings indicated that the BPHs probably had stronger effect on the antibiotic factors in the pyramiding line and introgression lines. And in the seven genotypes, the order of the gene effect on BPH survival at 8 DAI was: Bph14/Bph15 > Bph15 ≥ Bph14 ≥ Bph21 > Bph20 ≥ Bph3 ≥ Bph18 > none.
The maintainer ability of improved lines
In order to evaluate the fertility of the introgression lines as maintainers we assessed the pollen viabilities of hybrids with Jin 23A by I2-KI staining. Of the nine improved lines with Bph3, one line showed complete fertility (81.7%). Among the eight Bph14 introgressed lines, two lines were completely fertile (94.0%) or partially fertile (53.0%). Among the eight Bph15 introgressed lines, two lines showed completely fertile (70.7%) or partially sterile (15.0%). Among nine Bph18 lines, two lines showed complete fertility (100% and 60.4%) and one line showed partially fertile (56.7%). Among ten Bph21 lines, one line showed partial sterility (16.7%) and one line showed complete fertility (100%). Among eight Bph14/Bph15 lines, one line showed partially fertile (48.3%). For Bph20 introgressed lines, 6 lines were showed complete sterility (0%). The lines with fertile rate more than 0% in the F1 population were excluded, and all the improved lines were showed complete sterility (0%) in the BC1F1 population. Conclusion was that more than five lines of each genotype showed maintaining ability in the Jin 23B background.
Agronomic traits of improved lines of Jin 23B
We compared the agronomic traits of one improved line from each of the 7 backcross populations with Jin 23B at Wuhan under stress-free conditions in 2013 (Table 2). Improved lines of Jin 23B (14), Jin 23B (18) and Jin 23B (14/15) were significantly lower in PH; and Jin 23B (20) had a significant increase in PN (Table 2). In addition, two improved lines, Jin 23B (3) and Jin 23B (20) had significantly decrease in NGP and SF; One improved line, Jin 23B (3), had significantly increase in GW. Furthermore, the yields of improved lines were found similar to the control (Jin 23B) under natural field conditions (Table 2). Generally, these results showed that two improved lines Jin 23B (15) and Jin 23B (21) had similar agronomic trait relative to the control maintaining the elite agronomic traits.
Discussion
Mapping and cloning of functional genes in rice have facilitated MAS of genes for resistance to biotic stress in rice genetic improvement in recent years. Marker-assisted selection combined with backcross breeding is a powerful method of coping with defects in otherwise very susceptible rice cultivars which had application in bacterial blight, sheath blight, blast, stem borer, and gall midge resistance (Huang et al. 1997; Katiyar et al. 2001; Datta et al. 2002; Maruthasalam et al. 2007; Koide et al. 2010; Jiang et al. 2012). For BPH resistance, recent studies have focused on incorporating BPH resistance genes such as Bph14, Bph15, and Bph18 into the elite rice cultivars Minhui 63, 9311 and their hybrids (Hu et al. 2012, 2013).
In the present study, six dominant BPH-resistance genes, Bph3, Bph14, Bph15, Bph18, Bph20 and Bph21 were introgressed or pyramided into an important hybrid parent Jin 23B using MAS-based backcrossing, and improved lines containing single or two BPH-resistance genes were obtained. BPH bioassays showed that each of the six BPH resistance genes conferred different levels of resistance at both seedling and tillering stage of plants. The line with the combined Bph14 and Bph15 genes was more resistant than lines with the respective single genes and was the most resistant line overall. And the resistance level of single gene introgression line with Bph15 was significant higher than that with Bph3 or Bph18 at seedling stage. Pyramiding line or introgression lines carrying gene Bph14/Bph15, Bph15 or Bph14 had a moderately resistant level at 25 DAI at tillering stage. And single introgression lines carrying Bph20, Bph21, Bph3 or Bph18 had a moderately susceptible level at 25 DAI at tillering stage. In honeydew weight and BPH survival rate, the pyramiding line with two genes had a significant difference from single introgression lines. And the single introgression lines had different among themselves (Figs. 3b, 6). The resistance levels of donor parent PTB33 at both seedling and tillering stages were significantly higher and the honeydew weight of PTB33 was significantly lower than Jin 23B (3), indicating that PTB33 may have more than one BPH resistance gene. In fact, Lakshminarayana and Khush (1976) reported that PTB33 carried two BPH resistance genes, namely bph2 and Bph3. From the resistance level of Jin 23B (3), we hypothesis that Jin 23B (3) did not get the bph2 gene from PTB33.
Gene effect of the BPH resistance conferred by Bph3, Bph14, Bph15, Bph18, Bph20, and Bph21 was almost the same in honeydew weight, BPH survival rate, and the tests in seedlings stage. The pyramiding line carrying Bph14 and Bph15 has higher resistance level than the lines carrying single resistance gene. The results indicated an additive effect of pyramiding genes and showed that pyramiding two resistance genes have larger effects to BPH under Jin 23B background. In addition, single gene introgression line carrying Bph15 had a largest effect than other single gene introgression lines. And single gene introgression line carrying Bph18 had a lowest effect than other single gene introgression lines.
With the development of high-throughput and resolution genotyping platform in whole genome scale, whole genome SNP array-based background selection has become a feasible strategy with higher precision and efficiency for crop genetic improvement. The breeding chip RICE6K was employed in background examination of the BPH resistance improved lines. After three times backcross, all the improved lines had the segment of the target gene from donors. The genetic background recovery rate of Jin 23B (18), Jin 23B (21) and Jin 23B (14/15) were less than 90%, and the other segments from the donor parents may have had resistance QTL with positive effects on BPH resistance. Further backcrosses with Jin 23B would eliminate those segments. In addition, Jairin et al. (2007b) reported that IR71033–121-15 carried a QTL on short arm of chromosome 6. It would appear from the RICE6K, results that lines Jin 23B (20) and Jin 23B (21) did not get the chromosome 6 resistance QTL from the IR71033–121-15 donor.
Eight BPH-resistance genes (Bph1, Bph2, Bph7, Bph9, Bph10, Bph18, Bph21, Bph26) that are clustered on the long arm of chromosome 12, and they may be allelic. Among them Bph26, Bph9 and Bph18 have been cloned. Zhao et al. (2016) reported that Bph9 shares 91.32% and 96.05% nucleotide sequence identity with Bph26 and Bph18, respectively, and these allelotypes confer varying levels of resistance to different biotypes of BPH. The coding region of BPH1/9 shows a high level of diversity in rice germplasm (Zhao et al. 2016). In our study the allele Bph18 and Bph21 were introduced into Jin 23B and the resistance level conferred by Bph21 was higher than Bph18 at seedling stage.
Most of all, to pursue a durable and broad-spectrum resistance to BPH we should not only make use of gene pyramiding, but also exploit genetic diversity as an ecological approach to pest control, which can be highly effective over a large area and contribute to the sustainability of crop production (Zhu et al. 2000). It is more important that diversity represents insurance of buffering from failure of a single widely used gene. Because different BPH resistance genes are identified from different indica and wild rice accessions. Using of improved lines containing one or both BPH resistance genes planted in the different or same fields should restrict rapid increases in insect population and therefore reduce crop damage. Fortunately, Bph14, Bph15 and Bph18 have been pyramided into several rice hybrids in different genetic backgrounds, indicating that planting resistant pyramided hybrids in conjunction with conventional susceptible hybrids could effectively decreases the BPH population with the consequence of less serious BPH outbreaks, reduced costs of labor and pesticides, and increased rice production (Hu et al. 2011). The improved lines generated in this study can be used to develop multiple gene pyramided lines in the same genetic background for BPH control in hybrids.
Data from filed trails demonstrated that most of the required traits were well recovered in the improved lines (Table 2). The Bph3 and Bph20 introgression lines had a significantly lower NGP and SF relative to the control, but showed significantly increase in GW or PN. The yields of improved lines were similar to the control. Two improved lines Jin 23B (15) and Jin 23B (21) showed no difference in agronomic traits compared to Jin 23B. Jin 23A is an elite indica CMS line widely used in hybrid rice production in China. Through hybrid with Jin 23A, these BPH resistance gene can be transferred into Jin 23A, which can be used for development of BPH resistance hybrids in future. In addition, I2-KI stain of the F1 and BC1F1 hybrids show that the improved Jin 23B lines have good maintainer ability. These results show that these BPH resistance genes in Jin 23B have considerable commercial value for use in sustainable rice production.
From a global viewpoint, the increasingly severe occurrence of insects and diseases and indiscriminate pesticide is a challenge for sustainable rice production. A combination of approaches based on recent advances in genomic research has been formulated to address these challenges, with the long-term goal to develop rice cultivars referred to as Green Super Rice (Zhang 2007). Green Super Rice should possess resistance to multiple insect pests and diseases. In our previous work we have got the improved line Jin 23B (Pi1/Pi2/D12) containing three blast resistance gene (Jiang et al. 2012). This line can be hybridized with the present BPH resistance lines. The objective of the new improved lines is to address the problems of both blast and BPH.
Conclusions
Overall, six dominant BPH-resistance genes Bph3, Bph14, Bph15, Bph18, Bph20 and Bph21 have an effect on BPH growth and development and antibiotic factors, resulting both seedling and tillering resistance. Bioassay test showed that the order of the gene effect being Bph14/Bph15 > Bph15 ≥ Bph14 ≥ Bph20 ≥ Bph21 ≥ Bph3 > Bph18 > none at seedling stage of rice. Furthermore, field trial data demonstrated that yields of the improved lines were similar to the Jin 23B control under field conditions. These improved versions can be used in hybrid production in China.
References
Ali MP, Chowdhury TR (2014) Tagging and mapping of genes and QTLs of Nilaparvata lugens resistance in rice. Euphytica 195:1–30
Brar DS, Virk PS, Jena KK, Khush GS (2010) Breeding for resistance to planthoppers in rice. Planthopper: new threats to the sustainability of intensive rice production systems in Asia. IRRI, pp 401–427
Cha YS, Ji H, Yun DW, Ahn BO, Lee MC, Suh SC, Lee CS, Ahn EK, Jeon YH, Jin ID, Sohn JK, Koh HJ, Eun MY (2008) Fine mapping of the rice Bph1 gene, which confers resistance to the brown planthopper (Nilaparvata lugens Stål), and development of STS markers for marker-assisted selection. Mol Cells 26:146–151
Datta K, Baisakh N, Thet KM, Tu J, Datta SK (2002) Pyramiding transgenes for multiple resistance in rice against bacterial blight, yellow stem borer and sheath blight. Theor Appl Genet 106:1–8
Du B, Zhang WL, Liu BF, Hu J, Wei Z, Shi ZY, He RF, Zhu LL, Chen RZ, Han B, He GC (2009) Identification and characterization of Bph14, a gene conferring resistance to brown planthopper in rice. Proc Natl Acad Sci U S A 106:22163–22168
Govinda Raj K, Virmani SS (1988) Genetics of fertility restoration of WA type cytoplasmic male sterility in rice. Crop Sci 28:787–792
Hu J, Cheng MX, Gao GJ, Zhang QL, Xiao JH, He YQ (2013) Pyramiding and evaluation of three dominant brown planthopper resistance genes in the elite indica rice 9311 and its hybrids. Pest Manag Sci 69:802–808
Hu J, Li X, Wu CJ, Yang CJ, Hua HX, Gao GJ, Xiao JH, He YQ (2012) Pyramiding and evaluation of the brown planthopper resistance genes Bph14 and Bph15 in hybrid rice. Mol Breeding 29:61–69
Hu J, Xiao C, He YQ (2016) Recent progress on the genetics and molecular breeding of brown planthopper resistance in rice. Rice 9(1):1–12
Hu J, Yang CJ, Zhang QL, Gao GJ, He YQ (2011) Resistance of pyramided rice hybrids to brown planthoppers. Chin J Appl Entomol 48:1341–1347
Huang N, Angeles ER, Domingo J, Magpantay G, Singh S, Zhang G, Kumaravadivel N, Bennett J, Khush GS (1997) Pyramiding of bacterial blight resistance genes in rice: marker-assisted selection using RFLP and PCR. Theor Appl Genet 95:313–320
Huang Z, Xu CG, Tu JM, Li XH, Zhang QF (2001) Identification and mapping of two brown planthopper resistance genes in rice. Theor Appl Genet 102:929–934
Jairin J, Phengrat K, Teangdeerith S, Vanavichit A, Toojinda T (2007a) Mapping of a broad-spectrum brown planthopper resistance gene, Bph3, on rice chromosome 6. Mol Breeding 19:35–44
Jairin J, Teangdeerith S, Leelagud P, Phengrat K, Vanavichit A, Toojinda T (2007b) Detection of brown planthopper resistance genes from different rice mapping populations in the same genomic location. ScienceAsia 33(3):347–352
Jena KK, Jeung JU, Lee JH, Choi HC, Brar DS (2006) High-resolution mapping of a new brown planthopper (BPH) resistance gene, Bph18(t), and marker-assisted selection for BPH resistance in rice (Oryza sativa L.). Theor Appl Genet 112:288–297
Jena KK, Kim SM (2010) Current status of brown planthopper (BPH) resistance and genetics. Rice 3:161–171
Ji H, Kim SR, Kim YH, Suh JP, Park HM, Sreenivasulu N, Misra G, Kim SM, Hechanova SL, Kim H, Lee GS, Yoon UH, Kim TH, Lim H, Suh SC, Yang J, An G, Jena KK (2016) Map-based cloning and characterization of the BPH18 gene from wild rice conferring resistance to brown planthopper (BPH) insect pest. Sci Rep 6:34376
Jiang HC, Feng YT, Bao L, Li X, Gao GJ, Zhang QL, Xiao JH, Xu CG, He YQ (2012) Improving blast resistance of Jin 23B and its hybrid rice by marker-assisted gene pyramiding. Mol Breeding 30:1679–1688
Katiyar S, Verulkar S, Chandel G, Zhang Y, Huang BC, Bennett J (2001) Genetic analysis and pyramiding of two gall midge resistance genes (Gm-2 and Gm-6t) in rice (Oryza sativa L.). Euphytica 122:327–334
Khush GS, Brar DS (1991) Genetics of resistance to insects in crop plants. Adv Agron 45:223–274
Koide Y, Kawasaki A, Telebanco-Yanoria MJ, Hairmansis A, Nguyet NTM, Bigirimana J, Fujita D, Kobayashi N, Fukuta Y (2010) Development of pyramided lines with two resistance genes, Pish and Pib, for blast disease (Magnaporthe oryzae B. Couch) in rice (Oryza sativa L.). Plant Breed 129:670–675
Lakshminarayana A, Khush GS (1976) New genes for resistance to brown planthopper in rice. Crop Sci 17:96–100
Liu YQ, Wu H, Chen H, Liu YL, He J, Kang HY, Sun ZG, Pan G, Wang Q, Hu JL, Zhou F, Zhou KN, Zheng XM, Ren YL, Chen LM, Wang YH, Zhao ZG, Lin QB, Wu FQ, Zhang X, Guo XP, Cheng XN, Jiang L, Wu CY, Wang HY, Wan JM (2015) A gene cluster encoding lectin receptor kinases confers broadspectrum and durable insect resistance in rice. Nat Biotechnol 33:301–305
Maruthasalam S, Kalpana K, Kumar KK, Loganathan M, Poovannan K, Raja JAJ, Kokiladevi E, Samiyappan R, Sudhakar D, Balasubramanian P (2007) Pyramiding transgenic resistance in elite indica rice cultivars against the sheath blight and bacterial blight. Plant Cell Rep 26:791–804
McCouch SR, Teytelman L, Xu Y, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, Xing Y, Zhang Q, Kono I, Yano M, Fjellstrom R, DeClerck G, Schneider D, Cartinhour S, Ware D, Stein L (2002) Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res 9:199–207
Pathak MD, Cheng CH, Fortuno ME (1969) Resistance to Nephotettix impicticeps and Nilaparvata lugens in varieties of rice. Nature 223:502–504
Pathak PK, Saxena RC, Heinrichs EA (1982) Parafilm sachet for measuring honeydew excretion by Nilaparvata lugens on rice. J Econ Entomol 75:194–195
Qiu YF, Guo JP, Jing SL, Zhu LL, He GC (2011) Development and characterization of japonica rice lines carrying the brown planthopper-resistance genes BPH12 and BPH6. Theor Appl Genet 124:485–494
Rahman ML, Jiang WZ, Chu SH, Qiao YJ, Ham TH, Woo MO, Lee J, Khanam MS, Chin JH, Jeung JU, Brar DS, Jena KK, Koh HJ (2009) High-resolution mapping of two rice brown planthopper resistance genes, Bph20(t) and Bph21(t), originating from Oryza minuta. Theor Appl Genet 119:1237–1246
Ren JS, Gao FY, Wu XT, Lu XJ, Zeng LH, Lv JQ, Su XW, Luo H, Ren GJ (2016) Bph32, a novel gene encoding an unknown SCR domain-containing protein, confers resistance against the brown planthopper in rice. Sci Rep 6:37645
Sebastian LS, Ikeda R, Huang N (1996) Molecular mapping of resistance to rice tungro spherical virus and green leafhopper. Phytopathology 86:25–30
Sharma PN, Torii A, Takumi S, Mori N, Nakamura C (2004) Marker-assisted pyramiding of brown planthopper (Nilaparvata lugens Stål) resistance genes Bph1 and Bph2 on rice chromosome 12. Hereditas 140:61–69
Shi BH, Zhang JH, Zheng YM, Liu YQ, Vera Cruz CM, Zheng TQ, Zhao MF (2012) Identification of a new resistance gene Pi-Da(t) from Dacca6 against rice blast fungus (Magnaporthe oryzae) in Jin23B background. Mol Breeding 30(2):1089–1096
Suh JP, Yang SJ, Jeung JU, Pamplona A, Kim JJ, Lee JH, Hong HC, Yang CL, Kim YG, Jena KK (2011) Development of elite breeding lines conferring Bph18 gene-derived resistance to brown planthopper (BPH) by marker-assisted selection and genome-wide background analysis in japonica rice (Oryza sativa L.). Field Crop Res 120:215–222
Tamura Y, Hattori M, Yoshioka H, Yoshioka M, Takahashi A, Wu JZ, Sentoku N, Yasui H (2014) Map-based cloning and characterization of a brown planthopper resistance gene BPH26 from Oryza sativa L. ssp. indica cultivar ADR52. Sci Reports 4:5872
Tanaka K, Endo S, Kazana H (2000) Toxicity of insecticides to predators of rice planthoppers: spiders, the mirid bug and the dryinid wasp. Appl Entomol Zool 35:177–187
Virmani SS, Viraktamath BC, Casa CL, Toledo RS, Lopez MT, Manalo JO (1997) Hybrid rice breeding manual. IRRI, Philippines, p 139
Wang Y, Cao LM, Zhang YX, Cao CX, Liu F, Huang FK, Qiu YF, Li RB, Lou XJ (2015) Map-based cloning and characterization of BPH29, a B3 domain-containing recessive gene conferring brown planthopper resistance in rice. J Exp Bot 66:6035–6045
Watanabe T, Kitagawa H (2000) Photosynthesis and translocation of assimilates in rice plants following phloem feeding by the planthopper Nilaparvata lugens (Homoptera: Delphacidae). J Econ Entomol 93:1192–1198
Yang HY, You AQ, Yang ZF, Zhang F, He RF, Zhu LL, He GC (2004) High-resolution genetic mapping at the Bph15 locus for brown planthopper resistance in rice (Oryza sativa L.). Theor Appl Genet 110:182–191
Yu HH, Xie WB, Li J, Zhou FS, Zhang QF (2014) A whole-genome SNP array (RICE6K) for genomic breeding in RICE. Plant Biotechnol J 12(1):28
Zhang QF (2007) Strategies for developing green super rice. Proc Natl Acad Sci U S A 104:16402–16409
Zhao Y, Huang J, Wang ZZ, Jing SL, Wang Y, Ouyang YD, Cai BD, Xin XF, Liu X, Zhang CX, Pan YF, Ma R, Li QF, Jiang WH, Zeng Y, Shangguan XX, Wang HY, Du B, Zhu LL, Xu X, Feng YQ, He SY, Chen RZ, Zhang QF, He GC (2016) Allelic diversity in an NLR gene BPH9 enables rice to combat planthopper variation. Proc Natl Acad Sci U S A 113:12850–12855
Zhu YY, Chen HR, Fan JH, Wang YY, Li Y, Chen JB, Fan JX, Yang SS, Hu LP, Leung H, Mew TW, Teng PS, Wang ZH, Mundt CC (2000) Genetic diversity and disease control in rice. Nature 406:718–722
Acknowledgements
We are very grateful to Professors DS Brar in IRRI and Guangcun He in Wuhan University for providing seeds of donor parent of BPH-resistance cultivars and the markers.
Funding
This work was supported by grants from the National Program on R&D of Transgenic Plants (2016ZX08001002–002), the National 863 Project (2014AA10A604) and the earmarked fund for the China Agriculture Research System (CARS-01-03) of China and the Bill & Melinda Gates Foundation.
Availability of data and materials
The data sets supporting the results of this article are included within the article.
Author information
Authors and Affiliations
Contributions
YH designed the experiments. HJ, JH, ZL and JL performed the experiments. GG and QZ help to field management. JX assisted in the project development. HJ analyzed data. HJ and YH wrote the manuscript. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Jiang, H., Hu, J., Li, Z. et al. Evaluation and breeding application of six brown planthopper resistance genes in rice maintainer line Jin 23B. Rice 11, 22 (2018). https://doi.org/10.1186/s12284-018-0215-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12284-018-0215-4