Abstract
Key message
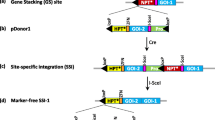
A selectable marker free, highly expressed single copy locus flanked by insulators was created as landing pad for transgene stacking in sugarcane. These events displayed superior transgene expression compared to single-copy transgenic lines lacking insulators. Excision of the selectable marker gene from transgenic sugarcane lines was supported by FLPe/FRT site-specific recombination.
Abstract
Sugarcane, a tropical C4 grass in the genus Saccharum (Poaceae), accounts for nearly 80% of sugar produced worldwide and is also an important feedstock for biofuel production. Generating transgenic sugarcane with predictable and stable transgene expression is critical for crop improvement. In this study, we generated a highly expressed single copy locus as landing pad for transgene stacking. Transgenic sugarcane lines with stable integration of a single copy nptII expression cassette flanked by insulators supported higher transgene expression along with reduced line to line variation when compared to single copy events without insulators by NPTII ELISA analysis. Subsequently, the nptII selectable marker gene was efficiently excised from the sugarcane genome by the FLPe/FRT site-specific recombination system to create selectable marker free plants. This study provides valuable resources for future gene stacking using site-specific recombination or genome editing tools.







Similar content being viewed by others
References
Agapito-Tenfen SZ, Vilperte V, Benevenuto RF, Rover CM, Traavik TI, Nodari RO (2014) Effect of stacking insecticidal cry and herbicide tolerance epsps transgenes on transgenic maize proteome. BMC Plant Biol 14:346
Akbudak MA, Srivastava V (2011) Improved FLP recombinase, FLPe, efficiently removes marker gene from transgene locus developed by Cre-lox mediated site-specific gene integration in rice. Mol Biotechnol 49:82–89
Albert H, Dale EC, Lee E, Ow DW (1995) Site-specific integration of DNA into wild-type and mutant lox sites placed in the plant genome. Plant J 7:649–659
Altpeter F, Oraby H (2010) Sugarcane. In: Kempken F, Jung C (eds) Genetic modification of plants. Springer, Berlin, pp 453–472
Altpeter F, Sandhu S, Davey MR, Anthony P (2010) Genetic transformation—biolistics. In: Davey MR, Anthony P (eds) Plant cell culture. Wiley, Chichester, pp 217–239
Altpeter F, Springer NM, Bartley LE, Blechl A, Brutnell TP, Citovsky V, Conrad L et al (2016) Advancing crop transformation in the era of genome editing. Plant Cell 28:1510–1520
Angell SM, Baulcombe DC (1997) Consistent gene silencing in transgenic plants expressing a replicating potato virus × RNA. EMBO J 16:3675–3684
Assaad FF, Tucker KL, Signer ER (1993) Epigenetic repeat-induced gene silencing (RIGS) in Arabidopsis. Plant Mol Biol 22:067–1085
Bartlett JG, Alves SC, Smedley M, Snape JW, Harwood WA (2008) High-throughput Agrobacterium-mediated barley transformation. Plant Methods 4:22
Bestor TH (2000) Gene silencing as a threat to the success of gene therapy. J Clin Investig 105:409–411
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Buchholz F, Angrand PO, Stewart AF (1998) Improved properties of FLP recombinase evolved by cycling mutagenesis. Nat Biotechnol 16:657–662
Cermak T, Baltes NJ, Cegan R, Zhang Y, Voytas DF (2015) High-frequency, precise modification of the tomato genome. Genome Biol 16:232
Cerutti H, Johnson AM, Gillham NW, Boynton JE (1997) Epigenetic silencing of a foreign gene in nuclear transformants of Chlamydomonas. Plant Cell 9:925–945
Chawla R, Ariza-Nieto M, Wilson AJ, Moore SK, Srivastava V (2006) Transgene expression produced by biolistic-mediated, site-specific gene integration is consistently inherited by the subsequent generations. Plant Biotechnol J 4:209–218
Cheng M, Frey JE, Pang S, Zhou H, Hironaka CM, Duncan DR, Conner TW, Wan Y (1997) Genetic transformation of wheat mediated by Agrobacterium tumefaciens. Plant Physiol 115:971–980
Chengalrayan K, Gallo-Meagher M (2001) Effect of various growth regulators on shoot regeneration of sugarcane. In Vitro Cell Dev Biol Plant 37:434–439
Chung JH, Whiteley M, Felsenfeld G (1993) A 5′ element of the chicken beta-globin domain serves as an insulator in human erythroid cells and protects against position effect in Drosophila. Cell 74:505–514
Clark AJ, Bissinger P, Bullock DW, Damak S, Wallace R, Whitelaw CB, Yull F (1994) Chromosomal position effects and the modulation of transgene expression. Reprod Fertil Dev 6:589–598
Cordeiro GM, Amouyal O, Eloitt F, Henry RJ (2007) Sugarcane. In: Kole C (ed) Genome mapping and molecular breeding in plants, vol 3. Pulses, sugar and tuber crops. Springer, Berlin, pp 175–204
Dai S, Zheng P, Marmey P, Zhang S, Tian W, Chen S, Beachy RN, Fauquet C (2001) Comparative analysis of transgenic rice plants obtained by Agrobacterium-mediated transformation and particle bombardment. Mol Breed 7:25–33
Davies GJ, Kilby NJ, Riou-Khamlichi C, Murray JAH (1999) Somatic and germinal inheritance of an FLP-mediated deletion in transgenic tobacco. J Exp Bot 50:1447–1456
De Buck S, De Wilde C, Van Montagu M, Depicker A (2000) T-DNA vector backbone sequences are frequently integrated into the genome of transgenic plants obtained by Agrobacterium-mediated transformation. Mol Breed 6:459–468
De Buck S, Peck I, De Wilde C, Marjanac G, Nolf J, De Paepe A, Depicker A (2007) Generation of single-copy T-DNA transformants in Arabidopsis by the CRE/loxP recombination-mediated resolution system. Plant Physiol 145:1171–1182
De Paepe A, De Buck S, Nolf J, Van Lerberge E, Depicker A (2013) Site-specific T–DNA integration in Arabidopsis thaliana mediated by the combined action of CRE recombinase and ϕC31 integrase. Plant J 75:172–184
Depicher A, Montagu MV (1997) Post-transcriptional gene silencing in plants. Curr Opin Cell Biol 9:373–382
Dobie KW, Lee M, Fantes JA, Graham E, Clark AJ, Springbett A, Lathe R, McClenaghan M (1996) Variegated transgene expression in mouse mammary gland is determined by the transgene integration locus. Proc Natl Acad Sci USA 93:6659–6664
Fiering S, Whitelaw E, Martin DI (2000) To be or not to be active: the stochastic nature of enhancer action. Bioessays 22:381–387
Fladung M, Becker D (2010) Targeted integration and removal of transgenes in hybrid aspen (Populus tremula L. X P. tremuloides Michx.) using site-specific recombination systems. Plant Biol 12:334–340
Forsyth A, Weeks T, Richael C, Duan H (2016) Transcription Activator-Like Effector Nucleases (TALEN)-mediated targeted DNA insertion in potato plants. Front Plant Sci 7:1572. eCollection
Garsmeur O, Droc G, Antonise R, Grimwood J, Potier B, Aitken K, Jenkins J, Martin G, Charron C, Hervouet C, Costet L (2018) A mosaic monoploid reference sequence for the highly complex genome of sugarcane. Nat Commun 9:2638
Gatehouse JA (2008) Biotechnological prospects for engineering insect-resistant plants. Plant Physiol 146:881–887
Gidoni D, Bar M, Gilboa N (2001) FLP/FRT-mediated restoration of normal phenotypes and clonal sectors formation in rolC transgenic tobacco. Transgenic Res 10:317–328
Gidoni D, Srivastava V, Carmi N (2008) Site-specific excisional recombination strategies for elimination of undesirable transgenes from crop plants. In Vitro Cell Dev Biol Plant 44:457–467
Giraldo P, Rival-Gervier S, Houdebine LM, Montoliu L (2003) The potential benefits of insulators on heterologous constructs in transgenic animals. Transgenic Res 12:751–755
Grønlund JT, Stemmer C, Lichota J, Merkle T, Grasser KD (2007) Functionality of the beta ⁄ six site-specific recombination system in tobacco and Arabidopsis: a novel tool for genetic engineering of plant genomes. Plant Mol Biol 63:545–556
Grosveld F, van Assendelft GB, Greaves DR, Kollias G (1987) Position-independent, high-level expression of the human beta-globin gene in transgenic mice. Cell 51:975–985
Halpin C (2005) Gene stacking in transgenic plants—the challenge for 21st century plant biotechnology. Plant Biotechnol J 3:141–155
Heap I (2018) International survey of herbicide resistant weeds (Online). http://www.weedscience.org/Summary/SOASummary.aspx
Heinz N, Broll S, Schleef M, Baum C, Bode J (2012) Filling a gap: S/MAR-based replicating minicircles. CliniBook— Nonviral Platform; Clinigene Network,pp 271–277
Henry R, Cole C (2010) Genetics, genomics and breeding of sugarcane. CRC Press, Boca Raton
Hily JM, Singer SD, Yang Y, Liu Z (2009) A transformation booster sequence (TBS) from Petunia hybrid functions as an enhancer-blocking insulator in Arabidopsis thaliana. Plant Cell Rep 28:1095–1104
Hobbs SL, Warketin TD, Delong CM (1993) Transgene copy number can be positively or negatively associated with transgene expression. Plant Mol Biol 21:17–26
Hou L, Yau YY, Wei J, Han Z, Dong Z, Ow DW (2014) An open-source system for in planta gene stacking by Bxb1 and Cre recombinases. Mol Plant 7:1756–1765
Hu Q, Nelson K, Luo H (2006) FLP-mediated site-specific recombination for genome modification in turfgrass. Biotechnol Lett 28:1793–1804
Hu Q, Kononowicz-Hodges H, Nelson-Vasilchik K, Viola D, Zeng P, Liu H, Kausch AP, Chandlee JM, Hodges TK, Luo H (2008) FLP recombinase-mediated site-specific recombination in rice. Plant Biotechnol J 6:176–188
Iglesias VA, Moscone EA, Papp I, Neuhuber F, Michalowski S, Phelan T, Spiker S, Matzke M, Matzke AJ (1997) Molecular and cytogenetic analyses of stably and unstably expressed transgene loci in tobacco. Plant Cell 9:1251–1264
Ingelbrecht IL, Van HH, Van MM, Depicker A (1994) Post-transcriptional silencing of reporter transgenes in tobacco correlates with DNA methylation. Proc Natl Acad Sci USA 91:10502–10506
Ingelbrecht IL, Irvine JE, Mirkov E (1999) Posttranscriptional gene silencing in transgenic sugarcane. Dissection of homology-dependent virus resistance in a monocot that has a complex polyploidy genome. Plant Physiol 119:1187–1197
Ingham DJ, Beer S, Money S, Hansen G (2001) Quantitative real-time PCR assay for determining transgene copy number in transformed plants. Biotechniques 31:132–141
Jackson MA, Anderson DJ, Birch RG (2013) Comparison of Agrobacterium and particle bombardment using whole plasmid or minimal cassette for production of high-expressing, low-copy transgenic plants. Transgenic Res 22:143–151
Jeon JS, Lee S, Jung KH, Jun SH, Jeong DH, Lee J, Kim C, Jang S, Yang K, Nam J, An K, Han MJ, Sung RJ, Choi HS, Yu JH, Choi JH, Cho SY, Cha SS, Kim SI, An G (2000) T-DNA insertional mutagenesis for functional genomics in rice. Plant J 22:561–570
Jiang F, Zhang T, Bai S, Wang Z, He K (2016) Evaluation of Bt corn with pyramided genes on efficacy and insect resistance management for the Asian corn borer in China. PLoS ONE 11:e0168442
Jorgensen R, Snyder C, Jones JDG (1987) T-DNA is organized predominantly in inverted repeat structures in plants transformed with Agrobacterium tumefaciens C58 derivatives. Mol Gen Genet 207:471–477
Jorgensen RA, Cluster PD, English J, Que Q, Napoli CA (1996) Chalcone synthase co suppression phenotypes in petunia flowers: comparison of sense vs. antisense constructs and single-copy vs. complex T-DNA sequences. Plant Mol Biol 31:957–973
Jung JH, Altpeter F (2016) TALEN mediated targeted mutagenesis of the caffeic acid O-methyltransferase in highly polyploid sugarcane improves cell wall composition for production of bioethanol. Plant Mol Biol 92:131–142
Kannan B, Jung JH, Moxley GW, Lee SM, Altpeter F (2018) TALEN-mediated targeted mutagenesis of more than 100 COMT copies/alleles in highly polyploid sugarcane improves saccharification efficiency without compromising biomass yield. Plant Biotechnol J 16:856–866
Kanno T, Naito S, Shimamoto G (2000) Post-transcriptional gene silencing in cultured rice cells. Plant Cell Physiol 41:321–326
Karunanandaa B, Qi Q, Hao M, Baszis SR, Jensen PK, Wong YH, Jiang J, Venkatramesh M, Gruys KJ, Moshiri F, Post-Beittenmiller D, Weiss JD, Valentin HE (2005) Metabolically engineered oilseed crops with enhanced seed tocopherol. Metab Eng 7:384–400
Kohli A, Gahakwa D, Vain P, Laurie DA, Christou P (1999) Transgene expression in rice engineered through particle bombardment: Molecular factors controlling stable expression and transgene silencing. Planta 208:88–97
Kohli A, Twyman RM, Abranches R, Wegel E, Stoger E, Christou P (2003) Transgene integration, organization and interaction in plants. Plant Mol Biol 52:247–258
Kumar S, Thompson WF (2009) Simultaneous excision of two transgene flanking sequences and resolution of complex integration loci. Plant Mol Biol 69:23–32
Li Z, Xing A, Moon BP, McCardell RP, Mills K, Falco SC (2009) Site-specific integration of transgenes in soybean via recombinase-mediated DNA cassette exchange. Plant Physiol 151:1087–1095
Li R, Han Z, Hou L, Kaur G, Qian Y, Ow DW (2016) Method for biolistic site-specific integration in plants catalyzed by Bxb1 integrase. Methods Mol Biol 1469:15–30
Lowe BA, Prakash NS, Way M, Mann MT, Spencer TM, Boddupalli RS (2009) Enhanced single copy integration events in corn via particle bombardment using low quantities of DNA. Transgenic Res 18:831–840
Lutz KA, Corneille S, Azhagiri AK, Svab Z, Maliga P (2004) A novel approach to plastid transformation utilizes the phiC31 phage integrase. Plant J 37:906–913
Lyznik L, Gordon-Kamm W, Tao Y (2003) Site-specific recombination for genetic engineering in plants. Plant cell Rep 21:925–932
Meyer P, Saedler H (1996) Homology-dependent gene silencing in plants. Annu Rev Plant Physiol Plant Mol Biol 47:23–48
Moellenbeck DJ, Peters ML, Bing JW, Rouse JR, Higgins LS, Sims L, Nevshemal T, Marshall L, Ellis RT, Bystrak PG, Lang BA, Stewart JL, Kouba K, Sondag V, Gustafson V, Nour K, Xu D, Swenson J, Zhang J, Czapla T, Schwab G, Jayne S, Stockhoff BA, Narva K, Schnepf HE, Stelman SJ, Poutre C, Koziel M, Duck N (2001) Insecticidal proteins from Bacillus thuringiensis protect corn from corn rootworms. Nat Biotechnol 19:668–672
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Muskens MW, Vissers AP, Mol JN, Kooter JM (2000) Role of inverted DNA repeats in transcriptional and post-transcriptional gene silencing. Plant Mol Biol 43:243–260
Nandy S, Srivastava V (2011) Site-specific gene integration in rice genome mediated by the FLP-FRT recombination system. Plant Biotechnol J 9:713–721
Naqvi S, Farre G, Sanahuja G, Capell T, Zhu C, Christou P (2009) When more is better: multigene engineering in plants. Trends Plant Sci 15:48–56
Ng HH, Bird AP (1999) DNA Methylation and chromatin modification. Curr Opin Genet Dev 9:158–163
Nguyen LD, Underwood JL, Nandy S, Akbudak MA, Srivastava V (2014) Strong activity of FLPe recombinase in rice plants does not correlate with the transmission of the recombined locus to the progeny. Plant Biotechnol Rep 8:455–462
Oz T, Karan R, Merotto A, Altpeter F (2019) CRISPR/Cas9 and template-mediated multi-allelic homologous recombination of the acetolactate synthase gene in sugarcane confers herbicide resistance (submitted for publication)
Peterson KR, Clegg CH, Huxley C, Josephson BM, Haugen HS, Furukawa T (1993) Transgenic mice containing a 248-kb yeast artificial chromosome carrying the human beta-globin locus display proper developmental control of human globin genes. Proc Natl Acad Sci USA 90:7593–7597
Porcu S, Kitamura M, Witkowska E, Zhang Z, Mutero A, Lin C, Chang J, Gaensler KML (1997) The human beta globin locus introduced by YAC transfer exhibits a specific and reproducible pattern of developmental regulation in transgenic mice. Blood 90:4602–4609
Sandhu S, Altpeter F (2008) Co-integration, co-expression and inheritance of unlinked minimal transgene expression cassettes in an apomictic turf and forage grass (Paspalum notatum Flugge). Plant Cell Rep 27:1755–1765
Shou H, Frame BR, Whitham SA, Wang K (2004) Assessment of transgenic maize events produced by particle bombardment or Agrobacterium-mediated transformation. Mol Breed 13:201–208
Singer T, Burke E (2003) High-throughput TAIL-PCR as a tool to identify DNA flanking insertions. Methods Mol Biol 236:241–272
Singer SD, Hily JM, Liu Z (2010) A 1-kb bacteriophage lamda fragment functions as an insulator to effectively block enhancer-promoter interactions in Arabidopsis thaliana. Plant Mol Biol Rep 28:69–76
Singer SD, Cox KD, Liu Z (2011) Enhancer-promoter interference and its prevention in transgenic plants. Plant Cell Rep 30:723–731
Singer SD, Liu Z, Cox KD (2012) Minimizing the unpredictability of transgene expression in plants: the role of genetic insulators. Plant Cell Rep 31:13–25
Srivastava V, Gidoni D (2010) Site-specific gene integration technologies for crop improvement. In Vitro Cell Dev Biol Plant 46(3):219–232
Srivastava V, Ow DW (2001) Biolistic mediated site-specific integration in rice. Mol Breed 8:345–350
Srivastava V, Thomson J (2016) Gene stacking by recombinases. Plant Biotechnol J 14:471–482
Srivastava V, Ariza-Nieto M, Wilson AJ (2004) Cre-mediated site-specific gene integration for consistent transgene expression in rice. Plant Biotechnol J 2:169–179
Stam M, De Bruin R, Kenter S, Van Der Hoorn RAL, Van Blokland R, Mol JNM, Kooter JM (1997) Post-transcriptional silencing of chalcone synthase in Petunia by inverted transgene repeats. Plant J 1:63–82
Storer NP, Thompson GD, Head GP (2012) Application of pyramided traits against Lepidoptera in insect resistance management for Bt crops. GM Crops Food 3:154–162
Taparia Y, Fouad W, Gallo M, Altpeter F (2012a) Rapid production of transgenic sugarcane with the introduction of simple loci following biolistic transfer of a minimal expression cassette and direct embryogenesis. In Vitro Cell Dev Biol Plant 48:15–22
Taparia Y, Gallo M, Altpeter F (2012b) Comparison of direct and indirect embryogenesis protocols, biolistic gene transfer and selection parameters for efficient genetic transformation of sugarcane. Plant Cell Tissue Organ Cult 111:131–141
Tew TL, Cobill RM (2008) Genetic improvement of sugarcane (Saccharum spp.) as an energy crop. In: Vermerris W (ed) Genetic improvement of bioenergy crops. Springer, New York, pp 273–294
Thomson JG, Yau YY, Blanvillain R, Chiniquy D, Thilmony R, Ow DW (2009) ParA resolvase catalyzes site-specific excision of DNA from the Arabidopsis genome. Transgenic Res 18:237–248
Thomson JG, Chan R, Thilmony R, Yau YY, Ow DW (2010) PhiC31 recombination system demonstrates heritable germinal transmission of site-specific excision from the Arabidopsis genome. BMC Biotechnol 10:17
Tinland B, Schoumacher F, Gloeckler V, Bravo-Angel AM, Hohn B (1995) The Agrobacterium tumefaciens virulence D2 protein is responsible for precise integration of T-DNA into the plant genome. EMBO J 14:3585–3595
Travella S, Ross SM, Harden J, Everett C, Snape JW, Harwood WA (2005) A comparison of transgenic barley lines produced by particle bombardment and Agrobacterium-mediated techniques. Plant Cell Rep 23:780–789
Wang MB, Waterhouse PM (2000) High-efficiency silencing of a beta-glucuronidase gene in rice is correlated with repetitive transgene structure but is independent of DNA methylation. Plant Mol Biol 43:67–82
West AG, Gaszner M, Felsenfeld G (2002) Insulators: many functions, many mechanisms. Gene Dev 16:271–288
Windels P, De Buck S, Depicker A (2008) Agrobacterium tumefaciens-mediated transformation: patterns of T-DNA integration into the host genome. In: Agrobacterium: from biology to biotechnology. Springer, New York, pp 442–483
Woo H, Cho H, Lim S, Shin K, Lee S, Lee K, Kim D, Cho Y (2009) Auto-excision of selectable marker genes from transgenic tobacco via a stress inducible FLP/FRT site-specific recombination system. Transgenic Res 18(3):455–465
Wu H, Altpeter F (2015a) Sugarcane (Saccharum spp. hybrids). In: Wang K (ed) Agrobacterium protocols. Springer, Heidelberg, pp 307–316
Wu H, Sparks CA, Jones HD (2006) Characterisation of T-DNA loci and vector backbone sequences in transgenic wheat produced by Agrobacterium-mediated transformation. Mol Breed 18:195–208
Wu H, Awan FS, Vilarinho A, Zeng Q, Kannan B, Phipps T, McCuiston J, Wang W, Caffall K, Altpeter F (2015b) Transgene integration complexity and expression stability following biolistic or Agrobacterium-mediated transformation of sugarcane. In Vitro Cell Dev Biol –Plant. https://doi.org/10.1007/s11627-015-9710-0
Yang Y, Singer SD, Liu Z (2011) Evaluation and comparison of the insulation efficiency of three enhancer-blocking insulators in plants. Plant Cell Tiss Org 105:405–414
Yoder JI, Goldsbrough AP (1994) Transformation systems for generating marker-free transgenic plants. Bio Technology 12:263–267
Zale J, Jung JH, Kim JY, Pathak B, Karan R, Liu H, Chen X, Wu H, Candreva J, Zhai Z, Shanklin J, Altpeter F (2016) Metabolic engineering of sugarcane to accumulate energy-dense triacylglycerols in vegetative biomass. Plant Biotechnol J 14:661–669
Zhang Y, Zheng Y, Xiao N, Wang L, Zhang Z, Fang R, Chen X (2012) Functional analysis of the HS185 regulatory element in the rice HSP70 promotor. Mol Biol Rep 39:1649–1657
Zhu C, Naqvi S, Breitenbach J, Sandmann G, Christou P, Capell T (2008) Combinatorial genetic transformation generates a library of metabolic phenotypes for the carotenoid pathway in maize. Proc Natl Acad Sci USA 105:18232–18237
Acknowledgements
The authors are thankful to Syngenta Crop Protection, LLC. and CPBR for financial support. This work was also co-funded by the DOE Center for Advanced Bioenergy and Bioproducts Innovation (U.S. Department of Energy, Office of Science, Office of Biological and Environmental Research under Award Number DE-SC0018420). Any opinions, findings, and conclusions or recommendations expressed in this publication are those of the authors and do not necessarily reflect the views of the U.S. Department of Energy. The authors would like to thank Dr. Hardev Sandhu (Everglades Research and Educational Center, UF-IFAS, Belle Glade, FL) for providing tops of sugarcane cultivar CP 88-1762 and Sun Gro Horticulture, Apopka, FL, for donation of the Fafard #2 potting mix.
Author information
Authors and Affiliations
Contributions
F.A. and V.S. conceived and designed the experiments; Y.Z., J.Y.K. and J.H.J. constructed the plasmids; Y.Z., B.P. and B.W. generated the transgenic plants; Y.Z., B.K. and D.W. analyzed the transgenic plants; C.F., W.Y. and S.D. performed the TaqMan® qPCR assay; R.K. and Y.Z. carried out Southern Blot hybridization; Y.Z. and F.A. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, Y., Kim, J.Y., Karan, R. et al. Generation of a selectable marker free, highly expressed single copy locus as landing pad for transgene stacking in sugarcane. Plant Mol Biol 100, 247–263 (2019). https://doi.org/10.1007/s11103-019-00856-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-019-00856-4