Abstract
Powdery mildew (Pm) caused by Blumeria graminis f. sp. tritici (Bgt) is one of the world’s major wheat diseases and results in large grain yield losses. Discovery and utilization of Pm resistance genes constitute the most common strategy for wheat Pm control. Hongyoumai, a wheat landrace from Henan Province in China, has excellent resistance to infection by Bgt. In order to identify the basis of such Pm resistance, a segregating population was submitted to genetic analysis, which showed that Pm resistance in Hongyoumai was conferred by a single recessive resistance gene. This gene was temporarily named pmHYM. Molecular marker analysis, chromosomal location, resistance spectrum analysis, and an allelism test showed that pmHYM was located on the long arm of chromosome 7B (7BL), most likely representing a new recessive resistance gene allelic with Pm5e and mlXBD. By using 90-kb single-nucleotide polymorphism sequences (SNP) in the BLASTn analysis against the wheat 7BL genome sequence, 12 new simple sequence repeat (SSR) markers linked with pmHYM were developed to map pmHYM co-segregating with the marker Xmp1207 and between markers Xmp925 and Xmp1158, at genetic distances of 2.8 and 2.7 cM, respectively. In addition, physical mapping of the markers linked with pmHYM using Chinese Spring deletion lines indicated a location in the 0.86–1.00 bin of 7BL.



Similar content being viewed by others
References
Brenchley R, Spannagl M, Pfeifer M et al (2012) Analysis of the bread wheat genome using whole-genome shotgun sequencing. Nature 491:705–710
Clarkson J (2000) Virulence survey report for wheat powdery mildew in Europe. http://www.crpmborg/2000/1204clarkson 1996–1998
Costanzo A, Bàrberi P (2014) Functional agrobiodiversity and agroecosystem services in sustainable wheat production. A review. Agron Sustain Dev 34:327–348
Duan X (1994) Monitoring the virulence of wheat powdery mildew and utlization of resistance genes in Europe (in Chinese). Plant Prot 20:36–38
Endo TR, Gill BS (1996) The deletion stocks of common wheat. J Hered 87:295–307
Frye CA, Tang D, Innes RW (2001) Negative regulation of defense responses in plants by a conserved MAPKK kinase. Proc Natl Acad Sci U S A 98:373–378
Fu B, Chen Y, Li N, Ma H, Kong Z, Zhang L, Jia H, Ma Z (2013) pmX: a recessive powdery mildew resistance gene at the Pm4 locus identified in wheat landrace Xiaohongpi. Theor Appl Genet 126:913–921
Hao Y, Parks R, Cowger C, Chen Z, Wang Y, Bland D, Murphy P, Guedira M, Brown-Guedira G, Johnson J (2015) Molecular characterization of a new powdery mildew resistance gene Pm54 in soft red winter wheat. Theor Appl Genet 128:465–476
Hsam SLK, Huang X, Zeller FJ (2001) Chromosomal location of genes for resistance to powdery mildew in common wheat (Triticum aestivum L. em Thell.) 6. Alleles at the Pm5 locus. Theor Appl Genet 102:127–133
Huang X, Hsam SLK, Zeller FJ, Wenzel G, Mohler V (2000) Molecular mapping of the wheat powdery mildew resistance gene Pm24 and marker validation for molecular breeding. Theor Appl Genet 101:407–414
Huang X, Wang LX, Xu M, Röder MS (2003) Microsatellite mapping of the powdery mildew resistance gene Pm5e in common wheat (Triticum aestivum L.) Theor Appl Genet 106:858–865
Jia J, Zhao S, Kong X, Li Y, Zhao G et al (2013) Aegilops tauschii draft genome sequence reveals a gene repertoire for wheat adaptation. Nature 496:91–95
Johnson J, Baenziger P, Yamazaki W, Smith R (1979) Effects of powdery mildew on yield and quality of isogenic lines of 'Chancellor' wheat. Crop Sci 19:349–352
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugenics 12:172–175
Lander E, Green P, Abrahamson J, Barlow A, Daley M, Lincoln S, Newburg L (1987) Mapmaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Law C, Wolfe M (1966) Location of genetic factors for mildew resistance and ear emergence time on chromosome 7B of wheat. Can J Genet Cytol 8:462–470
Li N, Wen Z, Wang J, Fu B, Liu J, Xu H, Kong Z, Zhang L, Jia H, Ma Z (2014) Transfer and mapping of a gene conferring later-growth-stage powdery mildew resistance in a tetraploid wheat accession. Mol Breed 33:669–677
Ling H, Zhao S, Liu D, Wang J, Sun H et al (2013) Draft genome of the wheat A-genome progenitor Triticum urartu. Nature 496:87–90
Liu R, Meng J (2003) MapDraw: a microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Hereditas (Beijing) 25:317–321
Ma H, Kong Z, Fu B, Li N, Zhang L, Jia H, Ma Z (2011) Identification and mapping of a new powdery mildew resistance gene on chromosome 6D of common wheat. Theor Appl Genet 123:1099–1106
Ma Z, Sorrells ME, Tanksley SD (1994) RFLP markers linked to powdery mildew resistance genes Pm1, Pm2, Pm3, and Pm4 in wheat. Genome 37:871–875
Ma P, Xu H, Xu Y, Li L, Qie Y, Luo Q, Zhang X, Li X, Zhou Y, An D (2015) Molecular mapping of a new powdery mildew resistance gene Pm2b in Chinese breeding line KM2939. Theor Appl Genet 128:613–622
McDonald B, Linde C (2002) The population genetics of plant pathogens and breeding strategies for durable resistance. Euphytica 124:163–180
McIntosh R, Dubcovsky J, Rogers W, Morris C, Appels R, Xia X (2014) Catalogue of gene symbols for wheat: 2013–2014 supplement. Ann Wheat Newslett 60:153–175 http://www.shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp
Michelmore R, Paran I, Kesseli R (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci U S A 88:9828–9832
Mohler V, Bauer C, Schweizer G, Kempf H, Hartl L (2013) Pm50: a new powdery mildew resistance gene in common wheat derived from cultivated emmer. J Appl Genet 54:259–263
Nematollahi G, Mohler V, Wenzel G, Zeller FJ, Hsam SLK (2008) Microsatellite mapping of powdery mildew resistance allele Pm5d from common wheat line IGV1-455. Euphytica 159:307–313
Niewoehner A, Leath S (1998) Virulence of Blumeria graminis f. Sp. tritici on winter wheat in the eastern United States. Plant Dis 82:64–68
Nishimura MT, Stein M, Hou B-H, Vogel JP, Edwards H, Somerville SC (2003) Loss of a callose synthase results in salicylic aciddependent disease resistance. Science 301:969–972
Paillard S, Schnurbusch T, Winzeler M, Messmer M, Sourdille P, Abderhalden O, Keller B, Schachermayr G (2003) An integrative genetic linkage map of winter wheat (Triticum aestivum L.) Theor Appl Genet 107:1235–1242
Peng J, Ronin Y, Fahima T, Röder MS, Li Y, Nevo E, Korol A (2003) Domestication quantitative trait loci in Triticum Dicoccoides, the progenitor of wheat. Proc Natl Acad Sci U S A 100:2489–2494
Peng F, Song N, Shen H, Wu H, Dong H, Zhang J, Li Y, Peng H, Ni Z, Liu Z, Yang T, Li B, Xie C, Sun Q (2014) Molecular mapping of a recessive powdery mildew resistancegene in spelt wheat cultivar Hubel. Mol Breed 34:491–500
Petersen S, Lyerly JH, Worthington ML, Parks WR, Cowger C, Marshall DS, Brown-Guedira G, Murphy PJ (2015) Mapping of powdery mildew resistance gene Pm53 introgressed from Aegilops speltoides into soft red winter wheat. Theor Appl Genet 128:303–312
Röder M, Korzun V, Wendehake K, Plaschke J, Tixier M, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Šafář J, Simkova H, Kubalakova M, Cihalikova J, Suchankova P, Bartos J, Dolezel J (2010) Development of chromosome-specific BAC resources for genomics of bread wheat. Cytogenet Genome Res 129:211–223
Santos F, Pena SDJ, Epplen JT (1993) Genetic and population study of a Y-linked tetranucleotide repeat DNA polymorphism with a simple non-isotopic technique. Hum Genet 90:655–656
Si Q, Zhang X, Duan X, Sheng B, Zhou Y (1992) On gene analysis and classification of powdery mildew (Erysiphe graminis f. Sp. tritici) resistant wheat varieties. Acta Phytopathol Sin 22:349–355
Somers D, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.) Theor Appl Genet 109:1105–1114
Song Q, Shi J, Singh S, Fickus EW, Costa JM, Lewis J, Gill BS, Ward R, Cregan PB (2005) Development and mapping of microsatellite (SSR) markers in wheat. Theor Appl Genet 110:550–560
Sourdille P, Singh S, Cadalen T, Brown-Guedira GL, Gay G, Qi L, Gill BS, Dufour P, Murigneux A, Bernard M (2004) Microsatellite-based deletion bin system for the establishment of genetic physical map relationships in wheat (Triticum aestivum L.) Funct Integr Genomics 4:12–25
Suenaga K, Khairallah M, William HM, Hoisington DA (2005) A new intervarietal linkage map and its application for quantitative trait locus analysis of “gigas” features in bread wheat. Genome 48:65–75
Wang S, Wong D, Forrest K, Allen A et al (2014) Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796
Xiao M, Song F, Jiao J, Wang X, Xu H, Li H (2013) Identification of the gene Pm47 on chromosome 7BS conferring resistance to powdery mildew in the Chinese wheat landrace Hongyanglazi. Theor Appl Genet 126:1397–1403
Xu H, Yi Y, Ma P, Qie Y, Fu X, Xu Y, Zhang X, An D (2015) Molecular tagging of a new broad-spectrum powdery mildew resistance allele Pm2c in Chinese wheat landrace Niaomai. Theor Appl Genet 128:2077–2084
Xue F, Wang C, Li C, Duan X, Zhou Y, Zhao N, Wang Y, Ji W (2012) Molecular mapping of powdery mildew resistance gene in common wheat landrace Baihulu and its allelism with Pm24. Theor Appl Genet 125:1425–1432
Xue F, Zhai W, Duan X, Zhou Y, Ji W (2009) Microsatellite mapping of a powdery mildew resistance gene in wheat landrace Xiaobaidong (in Chinese). Acta Agron Sin 35:1806–1811
Xue S, Zhang Z, Lin KZ, Cao Y, Li C, Yi H, Mei M, Zhu H, Wu J, Xu H, Zhao D, Tian D, Zhang C, Ma Z (2008) A high-density intervarietal map of the wheat genome enriched with markers derived from expressed sequence tags. Theor Appl Genet 117:181–189
Zhang R, Sun B, Chen J, Cao A, Xing L, Feng Y, Lan C, Chen P (2016) Pm55, A developmental-stage and tissue-specific powdery mildew resistance gene introgressed from Dasypyrum villosum, into common wheat. Theor Appl Genet 129:1975–1984
Zhao Z, Sun H, Song W, Lu M, Huang J, Wu L, Wang X, Li H (2014) Genetic analysis and detection of the gene MlLX99 on chromosome 2BL conferring resistance to powdery mildew in the wheat cultivar Liangxing 99. Theor Appl Genet 126:3081–3089
Acknowledgements
This project was supported by the National Key Research and Development Program of China (2016YFD0100102-3), the National Natural Science Foundation of China (No. 31501313), the Natural Science Foundation of Jiangsu Province (Grant no. BK20140757), and the Jiangsu Provincial Foundation of Agricultural Scientific Innovation (Grant no. cx [14]5006) in China. We are grateful to Thomas S. Cox, the Land Institute, USA, for improving the manuscript.
Authorʼs contribution
B. Fu, J. Wu, and S. Cai designed the study; B. Fu collected the plant materials and generated the populations; B. Fu, Z Zhang, Q Zhang, and X Wu performed the experiments; B. Fu and Z. Zhang analyzed the data; B. Fu wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that these experiments comply with the current laws of China and no conflicts of interest.
Additional information
Key message
The new powdery mildew resistance allele pmHYM, very likely at the Pm5 locus, was identified and mapped in the Chinese wheat landrace Hongyoumai.
Electronic supplementary material
Supplementary Figure 1
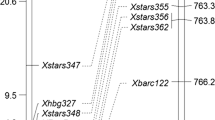
Comparison of previously reported genetic linkage maps of various powdery mildew resistance genes at or near the Pm5 locus on chromosome 7BL. Bold markers are common to the four genes. The vertical arrow indicates the direction toward the telomere. (50 kb).
Supplementary Table 1
(14 kb).
Rights and permissions
About this article
Cite this article
Fu, B., Zhang, Z., Zhang, Q. et al. Identification and mapping of a new powdery mildew resistance allele in the Chinese wheat landrace Hongyoumai. Mol Breeding 37, 133 (2017). https://doi.org/10.1007/s11032-017-0728-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-017-0728-3