Abstract
Key message
A powdery mildew resistance gene was introgressed from Aegilops speltoides into winter wheat and mapped to chromosome 5BL. Closely linked markers will permit marker-assisted selection for the resistance gene.
Abstract
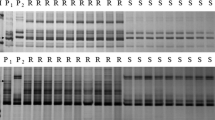
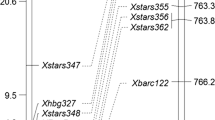
Powdery mildew of wheat (Triticum aestivum L.) is a major fungal disease in many areas of the world, caused by Blumeria graminis f. sp. tritici (Bgt). Host plant resistance is the preferred form of disease prevention because it is both economical and environmentally sound. Identification of new resistance sources and closely linked markers enable breeders to utilize these new sources in marker-assisted selection as well as in gene pyramiding. Aegilops speltoides (2n = 2x = 14, genome SS), has been a valuable disease resistance donor. The powdery mildew resistant wheat germplasm line NC09BGTS16 (NC-S16) was developed by backcrossing an Ae. speltoides accession, TAU829, to the susceptible soft red winter wheat cultivar ‘Saluda’. NC-S16 was crossed to the susceptible cultivar ‘Coker 68-15’ to develop F2:3 families for gene mapping. Greenhouse and field evaluations of these F2:3 families indicated that a single gene, designated Pm53, conferred resistance to powdery mildew. Bulked segregant analysis showed that multiple simple sequence repeat (SSR) and single nucleotide polymorphism (SNP) markers specific to chromosome 5BL segregated with the resistance gene. The gene was flanked by markers Xgwm499, Xwmc759, IWA6024 (0.7 cM proximal) and IWA2454 (1.8 cM distal). Pm36, derived from a different wild wheat relative (T. turgidum var. dicoccoides), had previously been mapped to chromosome 5BL in a durum wheat line. Detached leaf tests revealed that NC-S16 and a genotype carrying Pm36 differed in their responses to each of three Bgt isolates. Pm53 therefore appears to be a new source of powdery mildew resistance.


Similar content being viewed by others
References
Anikster Y, Manisterski J, Leonard KJ (2005) Resistance to leaf rust, stripe rust, and stem rust in Aegilops spp. in Israel. Plant Dis 89:303–308
Blanco A, Gadaleta A, Cenci A, Carluccio AV, Abdelbacki AMM, Simeone R (2008) Molecular mapping of the novel powdery mildew resistance gene Pm36 introgressed from Triticum turgidum var. dicoccoides in durum wheat. Theor Appl Genet 117:135–142
Bowen KL, Everts KL, Leath S (1991) Reduction in yield of winter wheat in North Carolina due to powdery mildew and leaf rust. Phytopathology 81:503–511
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, Forrest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D, Bai G, Pumphrey M, Tomar L, Wong D, Kong S, Reynolds M, da Silva ML, Bockelman H, Talbert L, Anderson JA, Dreisigacker S, Baenziger S, Carter A, Korzun V, Morrell PL, Dubcovsky J, Morell MK, Sorrells ME, Hayden MJ, Akhunov E (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Natl Acad Sci USA 110:8057–8062
Chen XM, Luo YH, Xia XC, Xia LQ, Chen X, Ren ZL, He ZH, Jia JZ (2005) Chromosomal location of powdery mildew resistance gene Pm16 in wheat using SSR marker analysis. Plant Breeding 124:225–228
Chen W, Mingus J, Mammadov J, Backlund JE, Green T, Thompson S, Kumpatla S (2010) KASPar: a simple and cost-effective system for SNP genotyping. In: Proc Plant and Animal Genomes XVIII Conference, San Diego, CA, p 94
Cowger C, Miranda L, Griffey C, Hall M, Murphy JP, Maxwell J (2012) Wheat powdery mildew. In: Sharma I (ed) Disease resistance in wheat. CAB International, Oxfordshire, pp 84–119
Dubcovsky J, Lukaszewski AJ, Echaide M, Antonelli EF, Porter DR (1998) Molecular characterization of two Triticum speltoides interstitial translocations carrying leaf rust and greenbug resistance genes. Crop Sci 38:1655–1660
Dvorak J (1972) Genetic variability in Aegilops speltoides affecting homoeologous pairing in wheat. Can J Genet Cytol 14:371–380
Dvorak J (1977) Transfer of leaf rust resistance from Aegilops speltoides to Triticum aestivum. Can J Genet Cytol 19:133–141
Dvorak J, Knott DR (1980) Chromosome location of two leaf rust resistance genes transferred from Triticum speltoides to T Tatusova. Can J Genet Cytol 22:381–389
Dvorak J, Knott DR (1990) Location of a Triticum speltoides chromosome segment conferring resistance to leaf rust in Triticum aestivum. Genome 33:892–897
Dvorak J, Deal KR, Luo MC (2006) Discovery and mapping of wheat Ph1 suppressors. Genet 174:17–27
Everts KL, Leath S, Finney PL (2001) Impact of powdery mildew and leaf rust on milling and baking quality of soft red winter wheat. Plant Dis 85:423–429
Faris JD, Xu SS, Cai X, Friesen TL, Jin Y (2008) Molecular and cytogenetic characterization of a durum wheat-Aegilops speltoides chromosome translocation conferring resistance to stem rust. Chromosome Res 16:1097–1105
Friebe B, Jiang J, Raupp WJ, McIntosh RA, Gill BS (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87
Gill BS, Friebe BR, White FF (2011) Alien introgressions represent a rich source of genes for crop improvement. Proc Natl Acad Sci USA 108:7657–7658
Gupta PK, Rustgi S, Sharma S, Singh R, Kumar N, Balyan HS (2003) Transferable EST-SSR markers for the study of polymorphism and genetic diversity in bread wheat. Mol Genet Genomics 270:315–323
Helguera M, Vanzetti L, Soria M, Khan IA, Kolmer J, Dubcovsky J (2005) PCR markers for Triticum speltoides leaf rust resistance gene Lr51 and their use to develop isogenic hard red spring wheat lines. Crop Sci 45:728–734
Hsam SLK, Lapochkina IF, Zeller FJ (2003) Chromosomal location of genes for resistance to powdery mildew in common wheat (Triticum aestivum L. em Thell.). 8. Gene Pm32 in a wheat-Aegilops speltoides translocation line. Euphytica 133:367–370
Huang XQ, Roder MS (2004) Molecular mapping of powdery mildew resistance genes in wheat: a review. Euphytica 137:203–223
Huang S, Sirikhachornkit A, Su X, Faris J, Gill B, Haselkorn R, Gornicki P (2002) Genes encoding plastid acetyl-CoA carboxylase and 3-phosphoglycerate kinase of the Triticum/Aegilops complex and the evolutionary history of polyploidy wheat. Proc Natl Acad Sci USA 99:8133–8138
Jauhar PP (2007) Meiotic restitution in wheat polyhaploids (amphihaploids): a potent evolutionary force. J Hered 98:188–193
Jia J, Devos KM, Chao S, Miller TE, Reader SM, Gale MD (1996) RFLP-based maps of the homoeologous group-6 chromosomes of wheat and their application in the tagging of Pm12, a powdery mildew resistance gene transferred from Aegilops speltoides to wheat. Theor Appl Genet 92:559–565
Johnson JW, Baenziger PS, Yamazaki WT, Smith RT (1979) Effects of powdery mildew on yield and quality of isogenic lines of Chancellor wheat. Crop Sci 19:349–352
Kerber ER, Dyck PL (1990) Transfer to hexaploid wheat of linked genes for adult-plant leaf rust and seedling stem rust resistance from an amphiploid of Aegilops-speltoides × Triticum monocuccum. Genome 33:530–537
Klindworth DL, Niu Z, Chao S, Friesen TL, Jin Y, Faris JD, Cai X, Xu SS (2012) Introgression and characterization of a Goatgrass gene for a high level of resistance to Ug99 stem rust in tetraploid wheat. G3-Genes Genomes Genet 2:665–673
Komsambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Lapochkina IF, Solomatin DA, Serezhkina GV, Grishina EE, Vishnyakova KS, Pukhalskii VA (1996) Common wheat lines with genetic material from Aegilops speltoides Tausch. Genetika 32:1651–1656
Leath S, Bowen KL (1989) Effects of powdery mildew, triadimenol seed treatment, and triadimefon foliar sprays on yield of winter wheat in North Carolina. Phytopathol 79:152–155
Leath S, Heun M (1990) Identification of powdery mildew resistance genes in cultivars of soft red winter wheat. Plant Dis 74:747–752
Lincoln SE, Daly MJ, Lander ES (1993) Constructing linkage maps with MAPMAKER/Exp version 3.0: a tutorial reference manual 3rd edn Whitehead Inst. For Medical Research, Cambridge
Liu ZY, Sun QX, Ni ZF, Nevo E, Yang TM (2002) Molecular characterization of a novel powdery mildew resistance gene Pm30 in wheat originating from wild emmer. Euphytica 123:21–29
Luo MC, Deal K, Yang ZL, Dvorak J (2005) Comparative genetic maps reveal extreme crossover localization in the Aegilops speltoides chromosomes. Theor Appl Genet 111:1098–1106
McDonald BA, Linde C (2002) The population genetics of plant pathogens and breeding strategies for durable resistance. Euphytica 124:163–180
McIntosh RA (1983) Genetic and cytogenetic studies involving Lr18 for resistance to Puccinia recondita. In: Sakamoto S (ed) Proc sixth Int Wheat Genet Symp. Maruzen, Kyoto, pp 777–783
McIntosh RA, Miller TE, Chapman V (1982) Cytogenetical studies in wheat. XII. Lr28 for resistance to Puccinia recondita and Sr34 for resistance to P. graminis tritici. Z Pflanzenzuchtg 89:295–306
McIntosh RA, Hart GE, Gale MD (1995a) Catalogue of gene symbols for wheat. In: Li ZS and Xin ZY(eds) Proc 8th Intl Wheat Genet Symposium, Beijing, China, 1993, pp 1333–1500
McIntosh RA, Wellings CR, Park RF (1995b) Wheat rusts: an atlas of resistance genes. CSIRO Publications, Victoria
McIntosh RA, Yamazaki Y, Dubcovsky J, Rogers J, Morris C, Somers DJ, Appels R, Devos KM (2013) Catalogue of gene symbols for wheat. In: KOMUGI-integrated wheat science database at http://www.shigen.nig.ac.jp/wheat/komugi/genes/download.jsp. Accessed 4 April 2014
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis—a rapid method to detect markers in genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Miller T, Reader S, Ainsworth C et al (1988) The introduction of a major gene for resistance to powdery mildew of wheat, Erysiphe graminis f. sp. tritici, from Aegilops speltoides into wheat, Triticum aestivum. In: Joma ML, Slootmaker LAJ et al (eds) Cereal breeding related to integrated cereal production. Pudoc, The Netherlands, pp 179–183
Miranda LM, Murphy JP, Marshall D, Leath S (2006) Pm34: a new powdery mildew resistance gene transferred from Aegilops tauschii coss to common wheat (Triticum aestivum L.). Theor Appl Gent 113:1497–1504
Niewoehner AS, Leath S (1998) Virulence of Blumeria graminis f. sp. tritici on winter wheat in the eastern United States. Plant Dis 82:64–68
Parks R, Carbone I, Murphy JP, Marshall D, Cowger C (2008) Virulence structure of the Eastern US wheat powdery mildew population. Plant Dis 92:1074–1082
Parry DW (1990) Diseases of small grain cereals. Plant pathology in agriculture. Cambridge University Press, Cambridge, pp 160–224
Qu LJ, Foote TN, Roberts MA, Money TA, Aragon-Alcaide L, Snape JW, Moore G (1998) A simple PCR-based method for scoring the ph1b deletion in wheat. Theor Appl Genet 96:371–375
Reader SM, Miller TE (1991) The introduction into bread wheat of a major gene for resistance to powdery mildew from wild emmer wheat. Euphytica 53:57–60
Riley R, Chapman V, Kimber G (1961) Origin of genetic control of diploid-like behavior of polyploid wheat. J Hered 52:22–25
Roberts MA, Reader SM, Dalgliesh C, Miller TE, Foote TN, Fish LJ, Snape JW, Moore G (1999) Induction and characterization of Ph1 wheat mutants. Genet 153:1909–1918
SAS Institute Inc. (2011) Base SAS® 9.3 Procedures guide. Cary: SAS Institute
Schneider A, Molnar I, Molnar-Lang M (2008) Utilisation of Aegilops (goatgrass) species to widen the genetic diversity of cultivated wheat. Euphytica 163:1–19
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotech 18:233–234
Somers DJ, Kirkpatrick R, Moniwa M, Walsh A (2003) Mining single-nucleotide polymorphisms from hexaploid wheat ESTs. Genome 46:431–437
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Starling TM, Roane CW, Camper HMJ (1986) Registration of cultivar ‘Saluda’ wheat. Crop Sci 26:200
Stein N, Herren G, Keller B (2001) A new DNA extraction method for high-throughput marker analysis in a large-genome species such as Triticum aestivum. Plant Breeding 120:354–356
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang S, Wong D, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Whan A, Stephen S, Barker G, Wieseke R, Plieske J, International Wheat Genome Sequencing Consortium, Lillemo M, Mather D, Appels R, Dolferus R, Brown-Guedira G, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo M-C, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden M, Akhunov E (2014) Characterization of polyploidy wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol J. doi:10.1111/pbi.12183
Weisz R (2013) Small grains production guide. North Carolina Cooperative Extension, Raleigh
Worthington W, Lyerly J, Petersen S, Brown-Guedira G, Marshall D, Cowger C, Parks R, Murphy JP (2014) MlUM15: an Aegilops neglecta-derived powdery mildew resistance gene in common wheat. Crop Sci 54 (in press)
Acknowledgments
The research was supported by the North Carolina Small Grain Growers Association. Thanks to Jon Raupp at Kansas State University Wheat Genetics Resource Center for his help in obtaining seeds of Chinese Spring deletion bin lines. We gratefully acknowledge Prof. Bob McIntosh for his guidance and review of this paper.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Evans Lagudah.
Rights and permissions
About this article
Cite this article
Petersen, S., Lyerly, J.H., Worthington, M.L. et al. Mapping of powdery mildew resistance gene Pm53 introgressed from Aegilops speltoides into soft red winter wheat. Theor Appl Genet 128, 303–312 (2015). https://doi.org/10.1007/s00122-014-2430-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-014-2430-8