Abstract
The phyllosphere, or plant leaf surface, represents a microbial ecosystem of considerable size, holding extraordinary biodiversity and enormous potential for the discovery of new products, tools, and applications in biotechnology, agriculture, medicine, and elsewhere. This mini-review highlights the applied microbiology of the phyllosphere as an original field of study concerning itself with the genes, gene products, natural compounds, and traits that underlie phyllosphere-specific adaptations and services that have commercial and economic value for current or future innovation. Examples include plant-growth-promoting and disease-suppressive phyllobacteria, probiotics and fermented foods that support human health, as well as microbials that remedy foliar contamination with airborne pollutants, residual pesticides, or plastics. Phyllosphere microbes promote plant biomass conversion into compost, renewable energy, animal feed, or fiber. They produce foodstuffs such as thickening agents and sugar substitutes, industrial-grade biosurfactants, novel antibiotics and cancer drugs, as well as enzymes used as food additives or freezing agents. Furthermore, new developments in DNA sequence-based profiling of leaf-associated microbial communities allow for surveillance approaches in the context of food safety and security, for example, to detect enteric human pathogens on leafy greens, predict plant disease outbreaks, and intercept plant pathogens and pests on internationally traded goods.
Key points
• Applied phyllosphere microbiology concerns leaf-specific adaptations for economic value
• Phyllobioprospecting searches the phyllosphere microbiome for product development
• Phyllobiomonitoring tracks phyllosphere microbial profiles for early risk detection
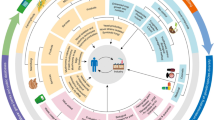
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant foliage is one of the largest and most important resources sustaining life on our planet. Terrestrial vegetation across both hemispheres represents an estimated leaf mass of 30 Gt and a surface area that is twice that of the land mass surface (Bar-On and Milo 2019). Plant leaves provide many services to humans and other life on Earth. They are a source of oxygen, pharmaceuticals (e.g., artemisinin or baccatins, which are precursors to the cancer drug paclitaxel), and recreational drugs (e.g., nicotine, cannabis); they offer shade and landscape aesthetics; serve as nourishment (e.g., leafy greens), drink (e.g., tea), or animal feed (e.g., grass-fed cows); and support through their photosynthetic properties the production of plant-based foods (e.g., fruits, nuts, and vegetables) and other products (e.g., fiber, fuel, lumber, and paper). An underappreciated but crucial source of many other services associated with plant foliage is the rich and diverse microbial communities that plants carry on their leaf surfaces (Bashir et al. 2022). In this review, we will explore these communities in detail, with a particular focus on how the properties, genes, and gene products of these leaf surface microbiota may be applied and exploited for improving plant, human, and environmental health. The novelty of this review lies in the focused and specialized synthesis of the current literature to highlight an active and exciting field of study known as applied microbiology of the phyllosphere, which is presented here in terms of bioprospecting, biosupplementing, bioprocessing, and biomonitoring.
Microbial traits and adaptations to survive and thrive in the leaf surface environment
As a habitat for microorganisms, plant leaves are referred to as the phyllosphere (Koskella 2020; Leveau et al. 2023a; Sohrabi et al. 2023). Microorganisms inhabiting aerial leaf surfaces include bacteria, fungi, yeasts, protists, archaea, and viruses. State-of-the-art DNA-based community profiling techniques have revealed an enormous diversity in the types and functions of leaf surface colonizers (Fadiji and Babalola 2020). Bacteria and filamentous fungi are among the most researched members of the phyllosphere, but the number of studies related to community composition of other microbial groups, for example, yeasts (Gouka et al. 2022), protists (Taerum et al. 2023) and viruses (ter Horst et al. 2023), is on the rise. DNA-based approaches have also revealed that only a fraction of the microbes that can be recovered from leaf surfaces can be cultured in the lab (Müller and Ruppel 2014). In terms of numbers, bacteria are by far the most abundant colonizers of leaf surfaces and can typically be found at densities between 106 and 108 cells/cm2 (Andrews and Harris 2000; Hirano and Upper 2000). Microorganisms that survive and multiply on aerial plant surfaces are called residual (as opposed to transient) epiphytes and are often labeled as pathogens, saprophytes, beneficials, or commensals, depending on the outcome of the interactions that they have with their plant host. Relative abundances of these bacterial epiphytes in the phyllosphere are impacted by a large number of factors such as plant genotype and health, leaf age, environmental conditions, plant neighborhood, geographical location, and the interactions that the epiphytes have among themselves (Bao et al. 2020; Beilsmith et al. 2019; Kumar et al. 2019; Lajoie and Kembel 2021; Meyer et al. 2022; Mina et al. 2020; Schlechter et al. 2019; Shakir et al. 2021; Zhang et al. 2019). The use of gnotobiotic environments for plant growth has allowed a better understanding of the factors shaping phyllosphere microbial communities, including leaf genotype (Schäfer et al. 2022), phytopathogenesis (Vogel et al. 2021), and abiotic stresses (Molina et al. 2020; Yuan et al. 2016).
The leaf surface is considered a harsh environment for microorganisms, as it poses numerous challenges to survival and reproductive success. Many members of the phyllosphere-associated microbiota have acquired and evolved specific adaptations to deal with stresses that are common on the leaf surface, including low nutrient availability, exposure to ultraviolet (UV) light, desiccation, and fluctuating temperatures (Leveau 2019). Such adaptations often represent strategies of “tolerance” (Beattie and Lindow 1995) such as the synthesis of pigments to reduce the effects of harmful ultraviolet radiation (Jacobs et al. 2005; Kumar et al. 2016) and the production of protective biofilms to prevent desiccation (Arun et al. 2020). The latter adaptation is an example of “niche construction” (Manching et al. 2017) or “ecosystem engineering” where microbes change their local environment to increase their chances of survival (Baquero et al. 2021). Other examples involve the secretion of enzymes (Rocky-Salimi et al. 2016) or plant hormones such as indole 3-acetic acid or IAA (Vanderhoef and Dute 1981) to access nutrients, biosurfactants to enhance surface mobility (Nogales et al. 2015), or ice nucleation proteins which not only raise the temperature of freezing thereby breaking plant cells to release nutrients (Avalos-Ruiz et al. 2022) but also contribute to microbial airborne movement between plants (Maki et al. 2023).
Microbial survival in the phyllosphere not only relies on coping with the harsh conditions of a leaf surface but also requires interaction with other cohabitating microorganisms. The multitude of interactions that members of the leaf microbiota have with each other are key to shaping microbial community structure in the phyllosphere (Chaudhry et al. 2021; Rangel et al. 2021). Examples of such interactions are competition for nutrients and space (Schlechter et al. 2023), antibiosis through the production of lytic compounds, enzymes, secondary metabolites, and toxins (Qi et al. 2021; Rangel and Bolton 2022), or signal interference (e.g., quorum quenching) (Ma et al. 2013; Theodora et al. 2019). Each one of these interactions can by itself or in combination with others influence the growth, suppression, or even death of microorganisms inhabiting the same leaf or section of the leaf (Remus-Emsermann and Schlechter 2018). It has been demonstrated that early arrivers to the leaf surface can exert a so-called priority effect and outcompete immigrating microbes (Carlström et al. 2019), thus impacting ecological succession through niche pre-emption (Maignien et al. 2014). However, the survival of these keystone species is also affected by the growth stage of the host, with the persistence of core members contingent on core functions that are adapted for success in the leaf environment (Bell et al. 2021; Müller et al. 2016b). Another type of interaction in the phyllosphere is facilitation: examples include the promotion of fitness of bacterial foliar pathogens through intra-species communication (Li et al. 2019a) and the increased survival of bacterial species through inter-species signaling (He et al. 2022; Li and Tian 2012).
A powerful demonstration of the uniqueness of the leaf surface as a microbial habitat and of the specific microbial adaptations that allow life on the leaf surface comes from the comparative genomics analysis of two closely related foliar bacterial pathogens (Feil et al. 2005): Pseudomonas syringae pv. syringae B728a, which is a highly competent epiphyte and Pseudomonas syringae pv. tomato DC3000, which does not survive the leaf surface very well and prefers an endophytic lifestyle (i.e., inside the leaf tissue). It was shown that many of the genes and gene products unique to B728a are known to contribute to epiphytic fitness or “epiphytness” (Leveau et al. 2023b), such as the ice nucleation protein, an enzymatic pathway for the production of IAA, and genes for antibiosis and repair of UV-damaged DNA (Feil et al. 2005).
Seeing that the phyllosphere is a unique microbial habitat that requires a specific set of microbial adaptations and traits to increase chances of survival in the face of a range of biotic and abiotic challenges, we can expect that foliar microbial communities harbor species, strains, genes, and gene functions that have potential to be exploited for the discovery, development, and/or commercialization of novel products (Thapa and Prasanna 2018). Here, we provide an overview of functions that phyllosphere inhabitants are known to possess and that may be harnessed into products, processes, and technologies (Table 1). We will cover how the approach of bioprospecting (Müller et al. 2016a) contributes to the discovery of phyllosphere-derived microorganisms, enzymes, and metabolites with useful properties. We will also touch upon the use of biomonitoring, as it has been applied to other microbial habitats (Meyer et al. 2023; Michán et al. 2021), defined here as the practice of extracting information from microbial communities on leaf surfaces for the purpose of assessment and decision-making in the face of pathogen threats or other foliar risks.
Biosupplementing for plant health
“Plant probiotics” is an umbrella term for microorganisms that offer some defined benefit to their plant host (Berlec 2012). Well-known examples of plant probiotics are the so-called plant-growth-promoting rhizobacteria (PGPRs), of which formulations are sold commercially as single or mixed strain products (Kumari et al. 2019). The concept of plant-growth-promoting phyllobacteria (PGPPs) is not as developed as that of PGPRs (Orozco-Mosqueda et al. 2021), but there are many ways in which phyllosphere microbes have been shown to contribute to plant health (Stone et al. 2018). The ability to fix atmospheric nitrogen on the leaf surface is one of them (Abadi et al. 2021; Li et al. 2019b). Under natural conditions, this ability is particularly relevant in tropical rainforest systems where the lack of seasonal leaf senescence and poor soil conditions rely heavily on bacterial-leaf N2 fixation (Goncalves et al. 2014; Stanton et al. 2019). The foliar application of nitrogen-fixing bacteria onto agriculturally significant crops has shown increased nitrogen uptake and a reduced need for nitrogen fertilizer input (Abadi et al. 2021; Madhaiyan et al. 2015). Other proposed mechanisms of plant growth stimulation by phyllobacteria that occur naturally but have potential biotechnological utilization are phosphorus solubilization, siderophore production, and the secretion of hormones such as IAA (Abadi et al. 2020; Arun et al. 2020; Berg and Koskella 2018; Cernava et al. 2019; Chacón et al. 2022). Phyllosphere bacteria and fungi also have been shown to alleviate drought stress, for example in rice seedlings (Abadi et al. 2020; Arun et al. 2020; Cernava et al. 2019; Chacón et al. 2022) and panic grass (Aimone et al. 2023).
A major line of research on how to protect plants from foliar diseases involves the isolation and characterization of so-called biological control agents (BCAs). These BCAs antagonize pathogens or pests through a variety of mechanisms including direct antagonism (parasitism and predation), mixed-path antagonism (production of antibiotics, lytic enzymes, waste products, and chemical interference), or indirect antagonism (competition for limited resources and induction of host plant resistance) (He et al. 2021). Commercialized formulations of both fungal and bacterial antagonists exist (Keswani et al. 2016; Lahlali et al. 2022). One of the classical success stories of phyllosphere-derived biocontrol of plant disease is a product that is registered under the name BlightBan A506® that contains, as the active ingredient, Pseudomonas fluorescens A506, a bacterial isolate from a pear tree in Healdsburg, CA (Wilson and Lindow 1993). A506 has been shown to protect plants from pathogen-induced frost damage (Lindow et al. 1996) and fruit russeting (Lindow et al. 1998) and from symptoms associated with bacterial fire blight caused by Erwinia amylovora (NuFarm 2024). The mechanisms underlying these activities include preemptive exclusion and antibiotic production (Anderson et al. 2004; Temple et al. 2004). Although registration has expired for its use, BlightBan C9-1® was another commercial product for use against fire blight and is derived from the phyllosphere isolate Pantoea vagans C9-1. Interestingly, mixtures of the two BlightBan strains did not provide greater protection against fire blight, because P. fluorescens A506 produces an extracellular protease that inactivates the antibiotic production in P. vagans C9-1 (Anderson et al. 2004).
In the field of biocontrol research, there is a growing interest in using mixtures of microorganisms with plant-protective properties. For example, a cocktail of endophytic bacterial wheat isolates prevented fungal spore germination and protected wheat seedlings from wheat stripe rust caused by Puccinia striiformis f. sp. tritici, both in greenhouse and semi-field conditions (Kiani et al. 2021). Another study found that microbial consortia consisting of bacteria and fungi displayed broad-spectrum hindrance of foliar phytopathogens as a result of their combined broad gene functionality (Minchev et al. 2021). Although it is challenging to register products containing more than one microbial strain (Keswani et al. 2016), ongoing research will likely continue to focus on the compatibility and synergy of BCAs and their modes of action in pursuit of improved control of foliar pathogens (Hawkes et al. 2021; Li et al. 2022). An exciting recent development is the demonstration that protection against foliar disease may be achieved as an emergent property of whole microbial communities (Berg and Koskella 2018) and those disease-suppressive microbial communities may be developed through passaging of leaf microbiota from one generation of plant leaves to the next, in the presence of a foliar pathogen and by selection for low disease phenotype after each passage (Ehau-Taumaunu and Hockett 2023).
Phyllosphere microbiology and human health
Phyllosphere-associated microbial communities play a role not only in the health of plants but also in that of humans who consume plant vegetation, for example, as leafy greens. Unfortunately, consumption of fresh produce carries an increased risk of foodborne diseases due to incomplete removal or inactivation of microorganisms that are pathogenic to humans, for example, norovirus, E. coli, Salmonella, and Cyclospora (Machado-Moreira et al. 2019). However, not all microorganisms on plant foliage are harmful to human consumers. In fact, the concept of the “edible microbiome” is based on the idea that with the ingestion of raw plant foods; we load our gut with microbes that associate with these foods that might have probiotic qualities (Berg et al. 2014a; Soto-Giron et al. 2021). Depending on the type of vegetables consumed, and where and how they are grown, humans are exposed to a variety of different microorganisms on and in their food (Leff and Fierer 2013; Marco et al. 2022; Rook 2013). Of special interest are the lactic acid bacteria (LABs) that are commonly found on leaf surfaces (Yu et al. 2020). Many of these LABs confer health benefits to humans. For example, the plant-associated bacterium Lactiplantibacillus plantarum (formerly Lactobacillus plantarum) can survive gastrointestinal-like conditions, break down food compounds that its human host cannot, antagonize human pathogens, and persist via adherence to epithelial cells (Vitali et al. 2012). Further study into the edible microbiome may uncover additional probiotic strains that could change how we choose or grow our foods, or that serve as the basis for new probiotic products deriving from the phyllosphere which could have an impact on human health and even thwart off human pathogens (Ercolini and Fogliano 2018; Wicaksono et al. 2023b).
Another way in which phyllosphere microbes may benefit human health is by stimulating the immune system. This has been referred to as “natural vaccination” (Berg et al. 2014b) which works by eliciting prophylactic and therapeutic responses in the host. In one study, the microbiota of raw Brassica vegetables were sequenced and screened for isolates that were able to produce an enzyme linked to cancer prevention (Wassermann et al. 2017). Enriched among these were members of the Enterobacteriaceae family. Interestingly, this family represents a major group in the loading of the gut microbiome through the consumption of fresh produce (Erlacher et al. 2015; Rastogi et al. 2012). Arguably, these bacteria represent a long-established part of the human diet and studies have shown that foodborne microbes that accompany plant-based diets can indeed seed the human gut (David et al. 2014; Wicaksono et al. 2023a).
Fermented foods represent another health benefit related to phyllosphere microorganisms (Marco et al. 2017). A growing body of knowledge suggests that the consumption of live microbes associated with fermented foods can alter the gut microbiome (Roselli et al. 2021). Fermentation into foods such as sauerkraut and kimchi typically depends on the microbes that naturally occur on plant leaf surfaces. Lactobacillus species play an important role in the final stages of the fermentation process, and these strains have been shown to have effects on insulin sensitivity (Zhong et al. 2021), Alzheimer’s disease (Kumar et al. 2022) and have anti-cancer (Eweys et al. 2022), anti-mutagenic (Mazanko et al. 2022), immune function (Rastogi and Singh 2022) and anti-obesity (Zhu et al. 2019) properties. We are just beginning to understand the importance of the edible leaf microbiome (Leeming et al. 2021; Tomás-Barberán and Rodríguez 2022), whether raw or fermented and further research will be necessary to unravel how diet and health are related to the incorporation of phyllosphere microbes.
Bioprocessing for environmental health
Phylloremediation refers to the ability of leaf-surface bacteria to degrade airborne or leaf-surface pollutants (Wei et al. 2017). Examples of these are polycyclic aromatic hydrocarbons that build up from deposition of urban area smog (Ali et al. 2015; Gandolfi et al. 2017), residual pesticides that persist as a result of overuse (Katsoula et al. 2020; Kucharska et al. 2020; Scheublin and Leveau 2013), micro- and nano-plastics (Adomako and Yu 2023), or climate-active gases (Bringel and Couee 2015; Crombie et al. 2018). Several studies suggest that the process of phylloremediation may be harnessed by inoculating leaves with plant-associated pollutant-degrading microorganisms (Weyens et al. 2015). Bacteria that degrade phenanthrene (Dharmasiri et al. 2023) or xylene (Sangthong et al. 2016) and fungi that break down aromatic hydrocarbons (Imperato et al. 2021) have successfully been isolated from leaf surfaces. Levels of hydrocarbon pyrene deposits were significantly reduced on jungle geranium leaves after artificially inflating native populations of phyllosphere isolate Kocuria sp. IC3, while the application of a biosurfactant reduced these levels even further (Siriratruengsuk et al. 2017). Similarly, the application of native bacterial degraders of aromatic organic pollutants was shown to reduce aerial concentrations of these compounds (Jindachot et al. 2018). The observation that phenol degradation genes were induced in bacteria from the common phyllosphere genus Arthrobacter during leaf colonization, even in the absence of phenol (Scheublin et al. 2014), suggests that phyllosphere microbiota harbor an untapped and primed capacity to destroy unwanted pollutants and/or chemicals. The future challenge lies in finding out how to expand these efforts of “phylloremediation” to achieve the same effects on a larger scale, e.g., near cities where industrial and vehicle emissions are high (Molina et al. 2021). Phyllosphere bacteria have been shown to degrade phthalates, a common plasticizer in agricultural films. When a phthalate-degrading Rhodococcus isolate from activated sludge was applied onto vegetables grown in plastic filmed greenhouses, the bacterium readily established and foliar accumulation of phthalates from the air was reduced (Zeng et al. 2022). Bacterial degradation of phthalates has also been documented in rice crops where a native Bacillus species was not only capable of degradation, but following its inoculation also shifted the endosphere community towards increased phthalate biodegradation (Liu et al. 2023). The real-world implication of using these types of plastic-degrading isolates or communities is far-reaching, from environmental cleanup to minimizing the consumption of microplastics in our food.
To relieve our dependence on oil and to reduce atmospheric emissions of air pollutants, the search for renewable alternative fuel sources focuses on microbes that can transform plant lignocellulose waste into useful biofuels (Rai et al. 2022). Many epiphytes harbor enzymes for the production of ethanol and butanol (Amadi et al. 2020; Minty et al. 2013), biodiesel or lipid-based fuels (Miao et al. 2020), or methane and hydrogen (Miftah et al. 2022). Hydrogen fuel is of interest because it is carbon-free and produces minimal pollution (Sen et al. 2016). Among the naturally occurring microbes of wheat straw that produce hydrogen, Lactobacillus plays a major role in lignocellulose conversion (Ayala-Campos et al. 2022). Wheat straw microbes outperformed cow manure-, ruminal fluid-, and anaerobic sludge-associated microbes in producing hydrogen, revealing plant materials to be an excellent substrate for manufacturing bioenergy (Pérez-Rangel et al. 2015; Valdez-Vazquez et al. 2017). Additionally, the phyllosphere microbiota of crops used specifically for bioenergy has been well-studied for its capacity to benefit host plants (i.e., aid in nutrient acquisition), defend against pathogens, and mitigate drought or salt stresses (Zhalnina et al. 2021). The microbiome of bioenergy crops such as switchgrass and miscanthus has been tracked over seasons to reveal useful functions and core microbes to supply a foundation for future manipulation to maximize yields (Grady et al. 2019; Howe et al. 2023).
Bioprospecting the leaf microbiome
Bioprospecting is the search for genes, natural compounds, and whole organisms in nature that have commercial economic value (Müller et al. 2016a). Perhaps the best-documented example of bioprospecting in the context of phyllosphere microbiology is the isolation and application of microorganisms for the management of foliar diseases and pests, for instance, the aforementioned BlightBan A506. Many other examples exist of industrial products or applications that have their origin within the leaf-associated microbiota and are produced commercially and at-scale for use in markets as diverse as cosmetics, food, pharmaceutics, and textiles (Müller et al. 2016a). One is xanthan gum, which is sold under the name Keltro® or Xantural®, and is used as a thickening agent for stabilizing emulsified suspensions (CPKelco 2024). The bacterium that produces this compound naturally is the leaf-colonizing pathogen Xanthomonas campestris, a causative agent of bacterial leaf spots on brassicas worldwide. Xanthan is a virulence factor in this organism, as it plays a significant role in biofilm formation, producing a hydrated, protected environment during leaf colonization (Bianco et al. 2016). For the production of this economically important polymer, which is used in foods, household cleaners, pesticides, paints, cosmetics, pharmaceuticals, textile dyes, and petrol recovery and production, cultures of X. campestris are grown in large bioreactors (Elella et al. 2021). Another phyllosphere-based product is erythritol which is widely used as a zero-calorie sugar substitute but is also an additive in some pharmaceuticals (Rzechonek et al. 2018). The biosynthetic pathway for producing erythritol is widespread among osmophilic phyllosphere bacteria, yeast, and fungi, including members of Pseudozyma sp., Aureobasidium sp., Aspergillus nidulans, Ustilaginomycetes, and the lactic acid bacteria Leuconostoc oenos, L. plantarum, and Acetobacter xylinum (Monedero et al. 2010; Moon et al. 2010). Often, the erythritol biosynthetic pathway genes from these microbes are cloned into organisms that are more easily managed for the production of this sweetener (Regnat et al. 2018).
Biosurfactants are biological molecules that have both hydrophobic and hydrophilic qualities that allow for reduced surface tension. This property is useful for solubilizing compounds and creating emulsifications. Many types of biosurfactants are produced by microorganisms including those that are members of the microbial communities that associate with plant leaf surfaces. Bacterial biosurfactants are cheaper and easier to produce than their synthetic chemical counterparts. Pathways from leaf-dwelling bacteria such as Arthrobacter sp., Pseudomonas sp., Acinetobacter sp., and Bacillus sp. are utilized for petrol recovery, foods, pharmaceuticals, and cosmetics (Campos et al. 2013; Varvaresou and Iakovou 2015). Dozens of companies market biosurfactants for various uses in what is a multibillion-dollar industry (Sarubbo et al. 2022).
Other phyllosphere-derived industrial products involve enzymes and other proteins. Phytases break down phytate, which is the main form of phosphate storage in plants. These enzymes are commonly found to be produced by plant-associated microbes including Aspergillus sp., Pseudomonas sp., Serratia sp., and Bacillus sp. (Singh et al. 2020). They are marketed under the names Finase®, Natuphos®, and Allzyme® and sold commercially to ranchers worldwide as an amendment to feed for monogastric animals to better utilize phosphorus (Handa et al. 2020). Another enzyme of biotech interest is asparaginase which is used as an anti-cancer treatment by denying asparagine to growing tumor cells. Asparaginase from E. coli is marketed under different trade names, but some patients are intolerant to this enzyme, which is why, instead, wider use of the one from the phyllosphere colonizer (and foliar pathogen) Erwinia chrysanthemi is employed (Emadi et al. 2018; Salzer et al. 2014), which is marketed under the name Erwinaze (Keating 2013). Another therapeutic drug derived from a phyllosphere microorganism is the antibiotic mupirocin, produced by P. fluorescens (Khoshnood et al. 2019). It is used for topical treatment against methicillin-resistant Staphylococcus aureus and is prescribed under the name Bactroban® (GlaxoSmithKline 2024). The ice-nucleating activity of P. syringae bacteria has been harnessed and turned into technologies that allow altering weather by cloud seeding, snow-making at ski resorts, and freezing foods (Baloh et al. 2019; Pokharel et al. 2021; Wex et al. 2015). SnoMax® has been one of the most successful products based on P. syringae’s ice-nucleating protein and has been in use since 1987 for the production of artificial snow (Telemet 2024). Phyllosphere yeasts have been shown to contribute to the degradation of biodegradable plastics used in compost bags, mulches, and other agricultural supplies (Watanabe et al. 2014). These plastics are made from molecules similar to the cutin monomer found in plant cell walls which are the target of degradation by enzymes produced by certain phyllosphere inhabitants (Kitamoto et al. 2011; Saika et al. 2019). Enzymes and microbes with such activities represent an integral part of so-called life-cycle-engineered plastics (García-Depraect et al. 2021).
Phyllosphere microorganisms have long been known to participate in carbon cycling by contributing to the decomposition of leaf litter (Fanin et al. 2021). The process of decomposition can be harnessed with relatively little human intervention to produce compost and silage. Composting is the controlled biodegradation of organic materials, including leaves and other plant waste, whereas animal silage is preserved forage that is used as animal feed during the dry season. For both compost and silage, the required fermentation is carried out by members of the microbial community that is naturally associated with the plant material (Kraut-Cohen et al. 2016). Foliar microbes also play a role during the curing of tobacco and the retting of hemp for fiber production (Law et al. 2016, 2020; Zhou et al. 2021, 2020). Microbial succession is important for these processes, and the enzymatic activity that facilitates this degradation largely derives from leaf-associated microbes (Alper and Stephanopoulos 2009). Based on these examples, the phyllosphere presents an ideal environment from which to isolate microorganisms with applicable potential for biomass conversion of raw plant materials or agricultural waste into useful substances.
In recent years, the development of culture-independent, DNA-based methods has allowed (meta)genomic mining of diverse microbial environments, including the phyllosphere, for novel functions (Khoiri et al. 2021; Methe et al. 2020; Mishra et al. 2021). To give just one example of the power of such an approach is the study by Helfrich et al. (2018) who sequenced the genomes of 224 bacterial strains isolated from Arabidopsis leaves to reveal more than 1000 putative natural product biosynthetic gene clusters, many predicted to code for completely novel types of compounds. In mining the phyllosphere microbiota by metagenomics, it benefits to cast the net as wide as possible and take full advantage of the variation that is naturally present in leaf microbial communities due to geography (i.e., location, growing conditions, and season) and plant genotype (Abdelfattah et al. 2021; Miura et al. 2019). Such an approach also may lead to new strategies that rely on engineered plant genotypes and/or artificially imposed environments to select microorganisms with useful functions (Hale et al. 2014; Singh et al. 2021).
Biomonitoring the leaf microbiota
Phyllobiomonitoring is defined here as the interrogation of leaf surface microbiota for information that helps with the assessment of plant leaf-associated risks. This idea may be applied to several different fields, including agriculture, food safety, and biosecurity. Typically, biomonitoring involves metagenomics to establish the normal operating range of leaf-associated communities in a greenhouse, field, or grove and to interpret deviations from the normal operating range as a potential signal for a change in the environment or physiology of the plant (Burke et al. 2011; Chandran et al. 2020). These analyses may uncover different states of what constitutes a healthy plant leaf microbiota (Müller et al. 2016c; Vorholt et al. 2017) or whether plants are experiencing biotic or abiotic stresses (Chen et al. 2020; Saleem et al. 2017; Yin et al. 2018). There may be good reason to consider phyllosphere communities as sentinels for changes on a global scale. For example, climate disruptions have been shown to cause perturbations in phyllosphere microbial communities, in turn influencing host plant survival, fitness, ecology, and susceptibility to foliar disease (Perreault and Laforest-Lapointe 2021; Rosado et al. 2018; Trivedi et al. 2022; Zhu et al. 2021). Global warming has a demonstrable effect on the structure of bacterial (Aydogan et al. 2020) and fungal (Faticov et al. 2021) phyllosphere community structure, as does drought (Debray et al. 2021), suggesting a role for phyllobiomonitoring in collecting and analyzing data that can be fed into predictive models for climate warning systems.
Similar surveillance approaches may be applied to the field of food safety to alert growers or processors on changes in the microbiota of their crops which correlate with a higher probability of plant or human pathogens (Williams and Marco 2014). The need to monitor for human pathogen outbreaks has increased dramatically; as diets have shifted, demand for raw produce has gone up, and the number of reported cases of enteric disease resulting from the ingestion of contaminated fresh produce has risen, in particular, leafy greens since the 1990s (Marshall et al. 2020). Whereas sources of contamination and duration of persistence of major enteric human pathogens such as Salmonella enterica and E. coli O157:H7 on leafy greens have been studied extensively, it appears that these human pathogens, which are actually poor leaf colonizers, can benefit from the presence and activity of naturally present microorganisms on the leaf surface, both pre-harvest (Brandl et al. 2023; Cooley et al. 2006; Potnis et al. 2014) and post-harvest (Williams et al. 2013). Culture-independent community profiling to monitor for such facilitators of pathogen establishment would be a great asset to the produce industry.
Microbial forensics utilizes microbiological knowledge and methods to provide evidence in unlawful matters (Fletcher et al. 2020; Schmedes et al. 2016). Generally, it concerns the abuse of pathogenic organisms and/or their metabolites to cause social or economic damage. This field has become more and more valuable as increased travel and trade within or between countries have led to greater opportunities for the transport and introduction of non-native pathogenic microorganisms (Ochoa-Corona 2011). Current biosecurity tools for monitoring agricultural products have addressable gaps in standardization and validation during all precautionary steps from initial assessment and containment to control of the pathogen (Schmedes and Budowle 2019). This threat to food security can occur at any point in the field-to-table chain, from field to postharvest on domestic and imported goods. Phyllobiomonitoring may directly target one or more pathogens at any point in this series. This has already been implicated in tracking sources of food-borne outbreaks of human pathogens (Marine et al. 2015; Ottesen et al. 2019). Tracing bacterial community profiles at each step of the food processing process could enhance agricultural biosecurity and give concrete evidence at what step contamination occurred (Doyle et al. 2017). Snapshots of community profiles could be uploaded to databases that include what a “normal” microbiome looks like from crops grown in specific locations and seasons throughout their processing. Technologies regarding new detection or tracking of phyllosphere microbes will offer safety checks in the chain of food handling and agricultural trade.
Concluding remarks
Applied microbiology of the phyllosphere is an exciting area of research and development. Its potential is not bound by source material: for example, the total number of bacterial cells estimated to be associated with all of Earth’s combined terrestrial plant foliage is 1026 (Lindow and Brandl 2003). This number dwarfs by many orders of magnitude the number of bacterial (and other microbial) cells that have ever been cultured, assayed, and/or sequenced in labs around the world to produce the cumulative body of knowledge that currently exists on the topic of phyllosphere microbiology, of which only the tiniest fraction is covered in this review. The 1026 number is a powerful reminder of the vast functional diversity that is contained in the phyllosphere microbiota that we still know very little about. As the examples in this review make clear, the diversity, and the many agricultural, biotechnological, and other products, applications, and services that emerge from it, touch on many facets of life on this planet. Applied phyllosphere microbiology has a proven impact and future promise to enhance the health and productivity of plants, humans, and the environment.
References
Abadi VAJM, Sepehri M, Rahmani HA, Zarei M, Ronaghi A, Taghavi SM, Shamshiripour M (2020) Role of dominant phyllosphere bacteria with plant growth–promoting characteristics on growth and nutrition of maize (Zea mays L.). J Soil Sci Plant Nutr 20(4):2348–2363. https://doi.org/10.1007/s42729-020-00302-1
Abadi V, Sepehri M, Rahmani H, Dolatabad H, Shamshiripour M, Khatabi B (2021) Diversity and abundance of culturable nitrogen-fixing bacteria in the phyllosphere of maize. J Appl Microbiol 131(2):898–912. https://doi.org/10.1111/jam.14975
Abdelfattah A, Freilich S, Bartuv R, Zhimo VY, Kumar A, Biasi A, Salim S, Feygenberg O, Burchard E, Dardick C (2021) Global analysis of the apple fruit microbiome: are all apples the same? Environ Microbiol 23(10):6038–6055. https://doi.org/10.1111/1462-2920.15469
Adomako MO, Yu F-H (2023) Potential effects of micro-and nanoplastics on phyllosphere microorganisms and their evolutionary and ecological responses. Sci Total Environ 884:163760. https://doi.org/10.1016/j.scitotenv.2023.163760
Aimone CD, Giauque H, Hawkes C (2023) Fungal symbionts generate water-saver and water-spender plant drought strategies via diverse effects on host gene expression. Phytobiomes J 7(2):172–183. https://doi.org/10.1094/PBIOMES-01-22-0006-FI
Ali N, Al-Awadhi H, Dashti N, Khanafer M, El-Nemr I, Sorkhoh N, Radwan SS (2015) Bioremediation of atmospheric hydrocarbons via bacteria naturally associated with leaves of higher plants. Int J Phytoremediat 17(12):1160–1170. https://doi.org/10.1080/15226514.2015.1045125
Alper H, Stephanopoulos G (2009) Engineering for biofuels: exploiting innate microbial capacity or importing biosynthetic potential? Nat Rev Microbiol 7(10):715–723. https://doi.org/10.1038/nrmicro2186
Amadi OC, Onyenma NC, Nwagu TN, Nnamchi CI, Ndubuisi IA, Akachukwu SO, Moneke AN, Okolo BN, Agu RC (2020) Total utilization of different parts of wild cocoyam in production of sugar feedstock for bioethanol production: an integrated approach. Bioresour Technol Rep 12:100550. https://doi.org/10.1016/j.biteb.2020.100550
Anderson LM, Stockwell VO, Loper JE (2004) An extracellular protease of Pseudomonas fluorescens inactivates antibiotics of Pantoea agglomerans. Phytopathology 94(11):1228–1234. https://doi.org/10.1094/PHYTO.2004.94.11.1228
Andrews JH, Harris RF (2000) The ecology and biogeography of microorganisms of plant surfaces. Annu Rev Phytopathol 38:145–180. https://doi.org/10.1146/annurev.phyto.38.1.145
Arun KD, Sabarinathan KG, Gomathy M, Kannan R, Balachandar D (2020) Mitigation of drought stress in rice crop with plant growth-promoting abiotic stress-tolerant rice phyllosphere bacteria. J Basic Microb 60(9):768–786. https://doi.org/10.1002/jobm.202000011
Avalos-Ruiz D, Ten LN, Kim C-K, Lee S-Y, Jung H-Y (2022) Isolation and identification of ice nucleation active Fusarium strains from rapid apple declined trees in Korea. Plant Pathol J 38(4):403. https://doi.org/10.5423/PPJ.NT.04.2022.0051
Ayala-Campos OR, Sanchez A, Rebollar EA, Valdez-Vazquez I (2022) Plant-associated microbial communities converge in fermentative hydrogen production and form a core microbiome. Int J Hydrogen Energy 47(46):20049–20063. https://doi.org/10.1016/j.ijhydene.2022.04.155
Aydogan EL, Budich O, Hardt M, Choi YH, Jansen-Willems AB, Moser G, Müller C, Kämpfer P, Glaeser SP (2020) Global warming shifts the composition of the abundant bacterial phyllosphere microbiota as indicated by a cultivation-dependent and -independent study of the grassland phyllosphere of a long-term warming field experiment. FEMS Microbiol Ecol 96(8):fiaa087. https://doi.org/10.1093/femsec/fiaa087
Baloh P, Els N, David RO, Larose C, Whitmore K, Sattler B, Grothe H (2019) Assessment of artificial and natural transport mechanisms of ice nucleating particles in an Alpine Ski Resort in Obergurgl. Austria Front Microbiol 10:2278. https://doi.org/10.3389/fmicb.2019.02278
Bao L, Gu L, Sun B, Cai W, Zhang S, Zhuang G, Bai Z, Zhuang X (2020) Seasonal variation of epiphytic bacteria in the phyllosphere of Gingko biloba, Pinus bungeana and Sabina chinensis. FEMS Microbiol Ecol 96(3):fiaa017. https://doi.org/10.1093/femsec/fiaa017
Baquero F, Coque TM, Galán JC, Martinez JL (2021) The origin of niches and species in the bacterial world. Front Microbiol 12:657986. https://doi.org/10.3389/fmicb.2021.657986
Bar-On YM, Milo R (2019) The global mass and average rate of rubisco. P Natl Acad Sci 116(10):4738–4743. https://doi.org/10.1073/pnas.1816654116
Bashir I, War AF, Rafiq I, Reshi ZA, Rashid I, Shouche YS (2022) Phyllosphere microbiome: diversity and functions. Microbiol Res 254:126888. https://doi.org/10.1016/j.micres.2021.126888
Beattie GA, Lindow SE (1995) The secret life of foliar bacterial pathogens on leaves. Annu Rev Phytopathol 33(1):145–172. https://doi.org/10.1146/annurev.py.33.090195.001045
Beilsmith K, Thoen MP, Brachi B, Gloss AD, Khan MH, Bergelson J (2019) Genome-wide association studies on the phyllosphere microbiome: embracing complexity in host–microbe interactions. Plant J 97(1):164–181. https://doi.org/10.1111/tpj.14170
Bell JK, Helgason B, Siciliano SD (2021) Brassica napus phyllosphere bacterial composition changes with growth stage. Plant Soil 464(1):501–516. https://doi.org/10.1007/s11104-021-04965-2
Berg M, Koskella B (2018) Nutrient- and dose-dependent microbiome-mediated protection against a plant pathogen. Curr Bio 28(15):2487–2492. https://doi.org/10.1016/j.cub.2018.05.085
Berg G, Erlacher A, Grube M (2014a) The edible plant microbiome: importance and health issues. In: Lugtenberg B (ed) Principles of plant-microbe interactions. Springer International Publishing, Cham, Switzerland, pp 419–426
Berg G, Erlacher A, Smalla K, Krause R (2014b) Vegetable microbiomes: is there a connection among opportunistic infections, human health and our ‘gut feeling’? Microb Biotechnol 7(6):487–495. https://doi.org/10.1111/1751-7915.12159
Berlec A (2012) Novel techniques and findings in the study of plant microbiota: search for plant probiotics. Plant Sci 193:96–102. https://doi.org/10.1016/j.plantsci.2012.05.010
Bianco MI, Toum L, Yaryura PM, Mielnichuk N, Gudesblat GE, Roeschlin R, Marano MR, Ielpi L, Vojnov AA (2016) Xanthan pyruvilation is essential for the virulence of Xanthomonas campestris pv. campestris. Mol Plant Microbe Interact 29(9):688–699. https://doi.org/10.1094/MPMI-06-16-0106-R
Brandl MT, Mammel MK, Simko I, Richter TK, Gebru ST, Leonard SR (2023) Weather factors, soil microbiome, and bacteria-fungi interactions as drivers of the epiphytic phyllosphere communities of romaine lettuce. Food Microbiol 113:104260. https://doi.org/10.1016/j.fm.2023.104260
Bringel F, Couee I (2015) Pivotal roles of phyllosphere microorganisms at the interface between plant functioning and atmospheric trace gas dynamics. Front Microbiol 6:486. https://doi.org/10.3389/fmicb.2015.00486
Burke C, Steinberg P, Rusch D, Kjelleberg S, Thomas T (2011) Bacterial community assembly based on functional genes rather than species. P Natl Acad Sci 108(34):14288–14293. https://doi.org/10.1073/pnas.1101591108
Campos JM, Montenegro Stamford TL, Sarubbo LA, de Luna JM, Rufino RD, Banat IM (2013) Microbial biosurfactants as additives for food industries. Biotechnol Prog 29(5):1097–1108. https://doi.org/10.1002/btpr.1796
Carlström CI, Field CM, Bortfeld-Miller M, Müller B, Sunagawa S, Vorholt JA (2019) Synthetic microbiota reveal priority effects and keystone strains in the Arabidopsis phyllosphere. Nat Ecol Evol 3(10):1445–1454. https://doi.org/10.1038/s41559-019-0994-z
Cernava T, Erlacher A, Soh J, Sensen CW, Grube M, Berg G (2019) Enterobacteriaceae dominate the core microbiome and contribute to the resistome of arugula (Eruca sativa Mill.). Microbiome 7(1):1–12. https://doi.org/10.1186/s40168-019-0624-7
Chacón FI, Sineli PE, Mansilla FI, Pereyra MM, Diaz MA, Volentini SI, Poehlein A, Meinhardt F, Daniel R, Dib JR (2022) Native cultivable bacteria from the blueberry microbiome as novel potential biocontrol agents. Microorganisms 10(5):969. https://doi.org/10.3390/microorganisms10050969
Chandran H, Meena M, Sharma K (2020) Microbial biodiversity and bioremediation assessment through -omics approaches. Front Environ Chem 1:570326. https://doi.org/10.3389/fenvc.2020.570326
Chaudhry V, Runge P, Sengupta P, Doehlemann G, Parker JE, Kemen E (2021) Shaping the leaf microbiota: plant–microbe–microbe interactions. J Exp Bot 72(1):36–56. https://doi.org/10.1093/jxb/eraa417
Chen T, Nomura K, Wang X, Sohrabi R, Xu J, Yao L, Paasch BC, Ma L, Kremer J, Cheng Y (2020) A plant genetic network for preventing dysbiosis in the phyllosphere. Nature 580(7805):653–657. https://doi.org/10.1038/s41586-020-2185-0
Cooley MB, Chao D, Mandrell RE (2006) Escherichia coli O157: H7 survival and growth on lettuce is altered by the presence of epiphytic bacteria. J Food Protect 69(10):2329–2335. https://doi.org/10.4315/0362-028X-69.10.2329
CPKelco (2024) Xanthan gum products. Accessed January 16, 2024. https://www.cpkelco.com/products/xanthan-gum/
Crombie AT, Larke-Mejia NL, Emery H, Dawson R, Pratscher J, Murphy GP, McGenity TJ, Murrell JC (2018) Poplar phyllosphere harbors disparate isoprene-degrading bacteria. P Natl Acad Sci 115(51):13081–13086. https://doi.org/10.1073/pnas.1812668115
David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE, Ling AV, Devlin AS, Varma Y, Fischbach MA (2014) Diet rapidly and reproducibly alters the human gut microbiome. Nature 505(7484):559–563. https://doi.org/10.1038/nature12820
Debray R, Socolar Y, Kaulbach G, Guzman A, Hernandez CA, Curley R, Dhond A, Bowles T, Koskella B (2021) Water stress and disruption of mycorrhizas induce parallel shifts in phyllosphere microbiome composition. New Phytol 234(6):2018–2031. https://doi.org/10.1111/nph.17817
Dharmasiri RBN, Undugoda LJS, Nilmini AHL, Nugara NNRN, Manage PM, Udayanga D (2023) Phylloremediation approach to green air: phenanthrene degrading potential of Bacillus spp. inhabit the phyllosphere of ornamental plants in urban polluted areas. Int J Environ Sci Technol 20:13359–13372. https://doi.org/10.1007/s13762-023-04883-z
Doyle CJ, O’Toole PW, Cotter PD (2017) Metagenome-based surveillance and diagnostic approaches to studying the microbial ecology of food production and processing environments. Environ Microbiol 19(11):4382–4391. https://doi.org/10.1111/1462-2920.13859
Ehau-Taumaunu H, Hockett KL (2023) Passaging phyllosphere microbial communities develop suppression towards bacterial speck disease in tomato. Phytobiomes J 7(2):233–243. https://doi.org/10.1094/PBIOMES-05-22-0030-FI
Elella MHA, Goda ES, Gab-Allah MA, Hong SE, Pandit B, Lee S, Gamal H, urRehman A, Yoon KR (2021) Xanthan gum-derived materials for applications in environment and eco-friendly materials: a review. J Environ Chem Eng 9(1):104702. https://doi.org/10.1016/j.jece.2020.104702
Emadi A, Law JY, Strovel ET, Lapidus RG, Jeng LJ, Lee M, Blitzer MG, Carter-Cooper BA, Sewell D, Van Der Merwe I (2018) Asparaginase Erwinia chrysanthemi effectively depletes plasma glutamine in adult patients with relapsed/refractory acute myeloid leukemia. Cancer Chemother Pharmacol 81:217–222. https://doi.org/10.1007/s00280-017-3459-6
Ercolini D, Fogliano V (2018) Food design to feed the human gut microbiota. J Agric Food Chem 66(15):3754–3758. https://doi.org/10.1021/acs.jafc.8b00456
Erlacher A, Cardinale M, Grube M, Berg G (2015) Biotic stress shifted structure and abundance of Enterobacteriaceae in the lettuce microbiome. PLoS ONE 10(2):e0118068. https://doi.org/10.1371/journal.pone.0118068
Eweys AS, Zhao Y-S, Darwesh OM (2022) Improving the antioxidant and anticancer potential of Cinnamomum cassia via fermentation with Lactobacillus plantarum. Biotechnol Rep 36:e00768. https://doi.org/10.1016/j.btre.2022.e00768
Fadiji AE, Babalola OO (2020) Metagenomics methods for the study of plant-associated microbial communities: a review. J Microbiol Methods 170:105860. https://doi.org/10.1016/j.mimet.2020.105860
Fanin N, Lin D, Freschet GT, Keiser AD, Augusto L, Wardle DA, Veen G (2021) Home-field advantage of litter decomposition: from the phyllosphere to the soil. New Phytol 231(4):1353–1358. https://doi.org/10.1111/nph.17475
Faticov M, Abdelfattah A, Roslin T, Vacher C, Hambäck P, Blanchet FG, Lindahl BD, Tack AJ (2021) Climate warming dominates over plant genotype in shaping the seasonal trajectory of foliar fungal communities on oak. New Phytol 231(5):1770–1783. https://doi.org/10.1111/nph.17434
Feil H, Feil WS, Chain P, Larimer F, DiBartolo G, Copeland A, Lykidis A, Trong S, Nolan M, Goltsman E (2005) Comparison of the complete genome sequences of Pseudomonas syringae pv. syringae B728a and pv. tomato DC3000. P Natl Acad Sci 102(31):11064–11069. https://doi.org/10.1073/pnas.0504930102
Fletcher J, Barnaby NG, Burans J, Melcher U, Luster DG, Nutter FW Jr, Scherm H, Schmale DG III, Thomas CS, Corona FMO (2020) Forensic plant pathology. In: Budowle B, Schutzer SE, Morse SA (eds) Microbial forensics, 3rd edn. Elsevier, London, UK, pp 49–70
Gandolfi I, Canedoli C, Imperato V, Tagliaferri I, Gkorezis P, Vangronsveld J, Schioppa EP, Papacchini M, Bestetti G, Franzetti A (2017) Diversity and hydrocarbon-degrading potential of epiphytic microbial communities on Platanus x acerifolia leaves in an urban area. Environ Pollut 220:650–658. https://doi.org/10.1016/j.envpol.2016.10.022
García-Depraect O, Bordel S, Lebrero R, Santos-Beneit F, Börner RA, Börner T, Muñoz R (2021) Inspired by nature: microbial production, degradation and valorization of biodegradable bioplastics for life-cycle-engineered products. Biotechnol Adv 53:107772. https://doi.org/10.1016/j.biotechadv.2021.107772
GlaxoSmithKline (2024) Bactroban. Accessed January 16, 2024. https://au.gsk.com/media/6279/bactroban_ointment_cmi_au_004_approved-1.pdf
Goncalves AZ, Hoffmann FL, Mercier H, Mazzafera P, Romero GQ (2014) Phyllosphere bacteria improve animal contribution to plant nutrition. Biotropica 46(2):170–174. https://doi.org/10.1111/btp.12086
Gouka L, Raaijmakers JM, Cordovez V (2022) Ecology and functional potential of phyllosphere yeasts. Trends Plant Sci 27:1109–1123. https://doi.org/10.1016/j.tplants.2022.06.007
Grady KL, Sorensen JW, Stopnisek N, Guittar J, Shade A (2019) Assembly and seasonality of core phyllosphere microbiota on perennial biofuel crops. Nat Commun 10(1):1–10. https://doi.org/10.1038/s41467-019-11974-4
Hale IL, Broders K, Iriarte G (2014) A Vavilovian approach to discovering crop-associated microbes with potential to enhance plant immunity. Front Plant Sci 5:492. https://doi.org/10.3389/fpls.2014.00492
Handa V, Sharma D, Kaur A, Arya SK (2020) Biotechnological applications of microbial phytase and phytic acid in food and feed industries. Biocatal Agric Biotechnol 25:101600. https://doi.org/10.1016/j.bcab.2020.101600
Hawkes CV, Kjøller R, Raaijmakers JM, Riber L, Christensen S, Rasmussen S, Christensen JH, Dahl AB, Westergaard JC, Nielsen M (2021) Extension of plant phenotypes by the foliar microbiome. Annu Rev Plant Biol 72:823–846. https://doi.org/10.1146/annurev-arplant-080620-114342
He D-C, He M-H, Amalin DM, Liu W, Alvindia DG, Zhan J (2021) Biological control of plant diseases: an evolutionary and eco-economic consideration. Pathogens 10(10):1311. https://doi.org/10.3390/pathogens10101311
He Y-W, Deng Y, Miao Y, Chatterjee S, Tran TM, Tian J, Lindow SE (2022) DSF-family quorum sensing signal-mediated intraspecies, interspecies, and inter-kingdom communication. Trends Microbiol. https://doi.org/10.1016/j.tim.2022.07.006
Helfrich EJ, Vogel CM, Ueoka R, Schäfer M, Ryffel F, Müller DB, Probst S, Kreuzer M, Piel J, Vorholt JA (2018) Bipartite interactions, antibiotic production and biosynthetic potential of the Arabidopsis leaf microbiome. Nat Microbiol 3(8):909–919. https://doi.org/10.1038/s41564-018-0200-0
Hirano SS, Upper CD (2000) Bacteria in the leaf ecosystem with emphasis on Pseudomonas syringae - a pathogen, ice nucleus, and epiphyte. Microbiol Mol Biol R 64(3):624–653. https://doi.org/10.1128/Mmbr.64.3.624-653.2000
Howe A, Stopnisek N, Dooley SK, Yang F, Grady KL, Shade A (2023) Seasonal activities of the phyllosphere microbiome of perennial crops. Nat Commun 14(1):1039. https://doi.org/10.1038/s41467-023-36515-y
Imperato V, Portillo-Estrada M, Saran A, Thoonen A, Kowalkowski Ł, Gawronski SW, Rineau F, Vangronsveld J, Thijs S (2021) Exploring the diversity and aromatic hydrocarbon degrading potential of epiphytic fungi on hornbeams from chronically polluted areas. J Fungi 7(11):972. https://doi.org/10.3390/jof7110972
Jacobs JL, Carroll TL, Sundin GW (2005) The role of pigmentation, ultraviolet radiation tolerance, and leaf colonization strategies in the epiphytic survival of phyllosphere bacteria. Microb Ecol 49(1):104–113. https://doi.org/10.1007/s00248-003-1061-4
Jindachot W, Treesubsuntorn C, Thiravetyan P (2018) Effect of individual/co-culture of native phyllosphere organisms to enhance Dracaena sanderiana for benzene phytoremediation. Water Air Soil Pollut 229(3):1–11. https://doi.org/10.1007/s11270-018-3735-z
Katsoula A, Vasileiadis S, Sapountzi M, Karpouzas DG (2020) The response of soil and phyllosphere microbial communities to repeated application of the fungicide iprodione: accelerated biodegradation or toxicity? FEMS Microbiol Ecol 96(6):fiaa056. https://doi.org/10.1093/femsec/fiaa056
Keating GM (2013) Asparaginase Erwinia chrysanthemi (Erwinaze(R)): a guide to its use in acute lymphoblastic leukemia in the USA. BioDrugs 27(4):413–418. https://doi.org/10.1007/s40259-013-0051-4
Keswani C, Bisen K, Singh V, Sarma BK, Singh HB (2016) Formulation technology of biocontrol agents: present status and future prospects. In: Arora N, Mehnaz S, Balestrini R (eds) Bioformulations: for sustainable agriculture. Springer, New Delhi, pp 35–52
Khoiri AN, Cheevadhanarak S, Jirakkakul J, Dulsawat S, Prommeenate P, Tachaleat A, Kusonmano K, Wattanachaisaereekul S, Sutheeworapong S (2021) Comparative metagenomics reveals microbial signatures of sugarcane phyllosphere in organic management. Front Microbiol 12:310. https://doi.org/10.3389/fmicb.2021.623799
Khoshnood S, Heidary M, Asadi A, Soleimani S, Motahar M, Savari M, Saki M, Abdi M (2019) A review on mechanism of action, resistance, synergism, and clinical implications of mupirocin against Staphylococcus aureus. Biomed Pharmacother 109:1809–1818. https://doi.org/10.1016/j.biopha.2018.10.131
Kiani T, Mehboob F, Hyder MZ, Zainy Z, Xu L, Huang L, Farrakh S (2021) Control of stripe rust of wheat using indigenous endophytic bacteria at seedling and adult plant stage. Sci Rep 11(1):14473. https://doi.org/10.1038/s41598-021-93939-6
Kitamoto HK, Shinozaki Y, Cao XH, Morita T, Konishi M, Tago K, Kajiwara H, Koitabashi M, Yoshida S, Watanabe T, Sameshima-Yamashita Y, Nakajima-Kambe T, Tsushima S (2011) Phyllosphere yeasts rapidly break down biodegradable plastics. AMB Express 1:44. https://doi.org/10.1186/2191-0855-1-44
Koskella B (2020) The Phyllosphere. Curr Bio 30(19):R1143–R1146. https://doi.org/10.1016/j.cub.2020.07.037
Kraut-Cohen J, Tripathi V, Chen Y, Gatica J, Volchinski V, Sela S, Weinberg Z, Cytryn E (2016) Temporal and spatial assessment of microbial communities in commercial silages from bunker silos. Appl Microbiol Biotechnol 100(15):6827–6835. https://doi.org/10.1007/s00253-016-7512-x
Kucharska K, Wachowska U, Czaplicki S (2020) Wheat phyllosphere yeasts degrade propiconazole. BMC Microbiol 20(1):1–14. https://doi.org/10.1186/s12866-020-01885-6
Kumar J, Babele PK, Singh D, Kumar A (2016) UV-B radiation stress causes alterations in whole cell protein profile and expression of certain genes in the rice phyllospheric bacterium Enterobacter cloacae. Front Microbiol 7:1440. https://doi.org/10.3389/fmicb.2016.01440
Kumar I, Mondal M, Gurusamy R, Balakrishnan S, Natarajan S (2019) Plant-microbiome interaction and the effects of biotic and abiotic components in agroecosystem. In: Singh D, Gupta V, Prabha R (eds) Microbial interventions in agriculture and environment. Springer, Singapore, pp 517–546
Kumar MR, Azizi NF, Yeap SK, Abdullah JO, Khalid M, Omar AR, Osman M, Leow ATC, Mortadza SAS, Alitheen NB (2022) Clinical and preclinical studies of fermented foods and their effects on Alzheimer’s disease. Antioxidants 11(5):883. https://doi.org/10.3390/antiox11050883
Kumari B, Mallick M, Solanki MK, Solanki AC, Hora A, Guo W (2019) Plant growth promoting rhizobacteria (PGPR): modern prospects for sustainable agriculture. In: Ansari R, Mahmood I (eds) Plant health under biotic stress. Springer, Singapore, pp 109–127
Lahlali R, Ezrari S, Radouane N, Kenfaoui J, Esmaeel Q, El Hamss H, Belabess Z, Barka EA (2022) Biological control of plant pathogens: a global perspective. Microorganisms 10(3):596. https://doi.org/10.3390/microorganisms10030596
Lajoie G, Kembel SW (2021) Plant-bacteria associations are phylogenetically structured in the phyllosphere. Mol Ecol 30(21):5572–5587. https://doi.org/10.1111/mec.16131
Law AD, Fisher C, Jack A, Moe LA (2016) Tobacco, microbes, and carcinogens: correlation between tobacco cure conditions, tobacco-specific nitrosamine content, and cured leaf microbial community. Microb Ecol 72(1):120–129. https://doi.org/10.1007/s00248-016-0754-4
Law AD, McNees CR, Moe LA (2020) The microbiology of hemp retting in a controlled environment: steering the hemp microbiome towards more consistent fiber production. Agronomy 10(4):492. https://doi.org/10.3390/agronomy10040492
Leeming ER, Louca P, Gibson R, Menni C, Spector TD, Le Roy CI (2021) The complexities of the diet-microbiome relationship: advances and perspectives. Genome Med 13(1):1–14. https://doi.org/10.1186/s13073-020-00813-7
Leff JW, Fierer N (2013) Bacterial communities associated with the surfaces of fresh fruits and vegetables. PLoS ONE 8(3):e59310. https://doi.org/10.1371/journal.pone.0059310
Leveau JHJ (2019) A brief from the leaf: latest research to inform our understanding of the phyllosphere microbiome. Curr Opin Microbiol 49:41–49. https://doi.org/10.1016/j.mib.2019.10.002
Leveau JHJ, Beattie GA, Lindow SE, Mahaffee WF (2023a) Phyllosphere, front and center: focus on a formerly ‘ecologically neglected’ microbial milieu. Phytobiomes J 7(2):140–144. https://doi.org/10.1094/PBIOMES-08-23-0088-E
Leveau JHJ, Coaker GL, Marco ML (2023b) Phyllosphere 2022: 11th international symposium on leaf surface microbiology. Phytobiomes J 7(2):151–159. https://doi.org/10.1094/PBIOMES-05-23-0029-MR
Li YH, Tian XL (2012) Quorum sensing and bacterial social interactions in biofilms. Sensors 12(3):2519–2538. https://doi.org/10.3390/s120302519
Li L, Li J, Zhang Y, Wang N (2019a) Diffusible signal factor (DSF)-mediated quorum sensing modulates expression of diverse traits in Xanthomonas citri and responses of citrus plants to promote disease. BMC Genomics 20(1):1–22. https://doi.org/10.1186/s12864-018-5384-4
Li Y, Sun H, Wu Z, Li H, Sun Q (2019b) Urban traffic changes the biodiversity, abundance, and activity of phyllospheric nitrogen-fixing bacteria. Environ Sci Pollut Res 26(16):16097–16104. https://doi.org/10.1007/s11356-019-05008-1
Li P-D, Zhu Z-R, Zhang Y, Xu J, Wang H, Wang Z, Li H (2022) The phyllosphere microbiome shifts toward combating melanose pathogen. Microbiome 10(1):1–17. https://doi.org/10.1186/s40168-022-01234-x
Lindow SE, Brandl MT (2003) Microbiology of the phyllosphere. Appl Environ Microb 69(4):1875–1883. https://doi.org/10.1128/AEM.69.4.1875-1883.2003
Lindow SE, McGourty G, Elkins R (1996) Interactions of antibiotics with Pseudomonas fluorescens strain A506 in the control of fire blight and frost injury to pear. Phytopathology 86(8):841–848. https://doi.org/10.1094/Phyto-86-841
Lindow SE, Desurmont C, Elkins R, McGourty G, Clark E, Brandl MT (1998) Occurrence of indole-3-acetic acid-producing bacteria on pear trees and their association with fruit russet. Phytopathology 88(11):1149–1157. https://doi.org/10.1094/PHYTO.1998.88.11.1149
Liu L-H, Zhang J-Y, Tang G-X, Huang Y-H, Xie X-Q, Geng J, Lü H-X, Li H, Li Y-W, Mo C-H (2023) Endophytic phthalate-degrading Bacillus subtilis N-1-gfp colonizing in soil-crop system shifted indigenous bacterial community to remove di-n-butyl phthalate. J Hazard Mater 449:130993. https://doi.org/10.1016/j.jhazmat.2023.130993
Ma A, Lv D, Zhuang X, Zhuang G (2013) Quorum quenching in culturable phyllosphere bacteria from tobacco. Int J Mol Sci 14(7):14607–14619. https://doi.org/10.3390/ijms140714607
Machado-Moreira B, Richards K, Brennan F, Abram F, Burgess CM (2019) Microbial contamination of fresh produce: what, where, and how? Compr Rev Food Sci Food Saf 18(6):1727–1750. https://doi.org/10.1111/1541-4337.12487
Madhaiyan M, Alex THH, Ngoh ST, Prithiviraj B, Ji L (2015) Leaf-residing Methylobacterium species fix nitrogen and promote biomass and seed production in Jatropha curcas. Biotechnol Biofuels 8(1):1–14. https://doi.org/10.1186/s13068-015-0404-y
Maignien L, DeForce EA, Chafee ME, Eren AM, Simmons SL (2014) Ecological succession and stochastic variation in the assembly of Arabidopsis thaliana phyllosphere communities. mBio 5(1):e00682-13. https://doi.org/10.1128/mbio.00682-13
Maki T, Hosaka K, Lee KC, Kawabata Y, Kajino M, Uto M, Kita K, Igarashi Y (2023) Vertical distribution of airborne microorganisms over forest environments: a potential source of ice-nucleating bioaerosols. Atmos Environ 302:119726. https://doi.org/10.1016/j.atmosenv.2023.119726
Manching HC, Carlson K, Kosowsky S, Smitherman CT, Stapleton AE (2017) Maize phyllosphere microbial community niche development across stages of host leaf growth. F1000Res 6:1698. https://doi.org/10.12688/f1000research.12490.3
Marco ML, Heeney D, Binda S, Cifelli CJ, Cotter PD, Foligne B, Ganzle M, Kort R, Pasin G, Pihlanto A, Smid EJ, Hutkins R (2017) Health benefits of fermented foods: microbiota and beyond. Curr Opin Biotech 44:94–102. https://doi.org/10.1016/j.copbio.2016.11.010
Marco ML, Hutkins R, Hill C, Fulgoni VL III, Cifelli CJ, Gahche J, Slavin JL, Merenstein D, Tancredi DJ, Sanders ME (2022) A classification system for defining and estimating dietary intake of live microbes in US adults and children. J Nutr 152(7):1729–1736. https://doi.org/10.1093/jn/nxac074
Marine SC, Pagadala S, Wang F, Pahl DM, Melendez MV, Kline WL, Oni RA, Walsh CS, Everts KL, Buchanan RL, Micallef SA (2015) The growing season, but not the farming system, is a food safety risk determinant for leafy greens in the mid-Atlantic region of the United States. Appl Environ Microb 81(7):2395–2407. https://doi.org/10.1128/Aem.00051-15
Marshall KE, Hexemer A, Seelman SL, Fatica MK, Blessington T, Hajmeer M, Kisselburgh H, Atkinson R, Hill K, Sharma D (2020) Lessons learned from a decade of investigations of Shiga toxin–producing Escherichia coli outbreaks linked to leafy greens, United States and Canada. Emerg Infect Dis 26(10):2319. https://doi.org/10.3201/eid2610.191418
Mazanko M, Prazdnova E, Kulikov M, Maltseva T, Rudoy D, Chikindas M (2022) Antioxidant and antimutagenic properties of probiotic Lactobacilli determined using LUX-biosensors. Enzyme Microb Technol 155:109980. https://doi.org/10.1016/j.enzmictec.2021.109980
Methe BA, Hiltbrand D, Roach J, Xu W, Gordon SG, Goodner BW, Stapleton AE (2020) Functional gene categories differentiate maize leaf drought-related microbial epiphytic communities. PLoS ONE 15(9):e0237493. https://doi.org/10.1371/journal.pone.0237493
Meyer KM, Porch R, Muscettola IE, Vasconcelos ALS, Sherman JK, Metcalf CJE, Lindow SE, Koskella B (2022) Plant neighborhood shapes diversity and reduces interspecific variation of the phyllosphere microbiome. ISME J 16(5):1376–1387. https://doi.org/10.1038/s41396-021-01184-6
Meyer L, Guyot S, Chalot M, Capelli N (2023) The potential of microorganisms as biomonitoring and bioremediation tools for mercury-contaminated soils. Ecotoxicol Environ Saf 262:115185. https://doi.org/10.1016/j.ecoenv.2023.115185
Miao Z, Tian X, Liang W, He Y, Wang G (2020) Bioconversion of corncob hydrolysate into microbial lipid by an oleaginous yeast Rhodotorula taiwanensis AM2352 for biodiesel production. Renew Energy 161:91–97. https://doi.org/10.1016/j.renene.2020.07.007
Michán C, Blasco J, Alhama J (2021) High-throughput molecular analyses of microbiomes as a tool to monitor the wellbeing of aquatic environments. Microb Biotechnol 14(3):870–885. https://doi.org/10.1111/1751-7915.13763
Miftah AK, Sittijunda S, Imai T, Salakkam A, Reungsang A (2022) Biohydrogen and methane production from sugarcane leaves pretreated by deep eutectic solvents and enzymatic hydrolysis by cellulolytic consortia. Fermentation 8(8):396. https://doi.org/10.3390/fermentation8080396
Mina D, Pereira JA, Lino-Neto T, Baptista P (2020) Epiphytic and endophytic bacteria on olive tree phyllosphere: exploring tissue and cultivar effect. Microb Ecol 80(1):145–157. https://doi.org/10.1007/s00248-020-01488-8
Minchev Z, Kostenko O, Soler R, Pozo MJ (2021) Microbial consortia for effective biocontrol of root and foliar diseases in tomato. Front Plant Sci 12:756368. https://doi.org/10.3389/fpls.2021.756368
Minty JJ, Singer ME, Scholz SA, Bae CH, Ahn JH, Foster CE, Liao JC, Lin XN (2013) Design and characterization of synthetic fungal-bacterial consortia for direct production of isobutanol from cellulosic biomass. P Natl Acad Sci 110(36):14592–14597. https://doi.org/10.1073/pnas.1218447110
Mishra S, Goyal D, Phurailatpam L (2021) Targeted 16S rRNA gene and ITS2 amplicon sequencing of leaf and spike tissues of Piper longum identifies new candidates for bioprospecting of bioactive compounds. Arch Microbiol 203(7):3851–3867. https://doi.org/10.1007/s00203-021-02356-w
Miura T, Sánchez R, Castañeda LE, Godoy K, Barbosa O (2019) Shared and unique features of bacterial communities in native forest and vineyard phyllosphere. Ecol Evol 9(6):3295–3305. https://doi.org/10.1002/ece3.4949
Molina MdC, White JF, García-Salgado S, Quijano MÁ, González-Benítez N (2020) A gnotobiotic model to examine plant and microbiome contributions to survival under arsenic stress. Microorganisms 9(1):45. https://doi.org/10.3390/microorganisms9010045
Molina L, Wittich R-M, van Dillewijn P, Segura A (2021) Plant-bacteria interactions for the elimination of atmospheric contaminants in cities. Agronomy 11(3):493. https://doi.org/10.3390/agronomy11030493
Monedero V, Perez-Martinez G, Yebra MJ (2010) Perspectives of engineering lactic acid bacteria for biotechnological polyol production. Appl Microbiol Biotechnol 86(4):1003–1015. https://doi.org/10.1007/s00253-010-2494-6
Moon HJ, Jeya M, Kim IW, Lee JK (2010) Biotechnological production of erythritol and its applications. Appl Microbiol Biotechnol 86(4):1017–1025. https://doi.org/10.1007/s00253-010-2496-4
Müller T, Ruppel S (2014) Progress in cultivation-independent phyllosphere microbiology. FEMS Microbiol Ecol 87(1):2–17. https://doi.org/10.1111/1574-6941.12198
Müller CA, Obermeier MM, Berg G (2016a) Bioprospecting plant-associated microbiomes. J Biotechnol 235:171–180. https://doi.org/10.1016/j.jbiotec.2016.03.033
Müller DB, Schubert OT, Rost H, Aebersold R, Vorholt JA (2016b) Systems-level proteomics of two ubiquitous leaf commensals reveals complementary adaptive traits for phyllosphere colonization. Mol Cell Proteomics 15(10):3256–3269. https://doi.org/10.1074/mcp.M116.058164
Müller DB, Vogel C, Bai Y, Vorholt JA (2016c) The plant microbiota: systems-level insights and perspectives. Annu Rev Genet 50:211–234. https://doi.org/10.1146/annurev-genet-120215-034952
Nogales J, Vargas P, Farias GA, Olmedilla A, Sanjuán J, Gallegos M-T (2015) FleQ coordinates flagellum-dependent and-independent motilities in Pseudomonas syringae pv. tomato DC3000. Appl Environ Microb 81(21):7533–7545. https://doi.org/10.1128/AEM.01798-15
NuFarm (2024) BlightBan A506. Accessed January 16, 2024. https://nufarm.com/uscrop/product/blightbana506/
Ochoa-Corona FM (2011) Biosecurity, microbial forensics and plant pathology: education challenges, overlapping disciplines and research needs. Australasian Plant Pathol 40(4):335–338. https://doi.org/10.1007/s13313-011-0052-z
Orozco-Mosqueda M, Flores A, Rojas-Sánchez B, Urtis-Flores CA, Morales-Cedeño LR, Valencia-Marin MF, Chávez-Avila S, Rojas-Solis D, Santoyo G (2021) Plant growth-promoting bacteria as bioinoculants: attributes and challenges for sustainable crop improvement. Agronomy 11(6):1167. https://doi.org/10.3390/agronomy11061167
Ottesen A, Ramachandran P, Reed E, Gu G, Gorham S, Ducharme D, Newell M, Rideout S, Turini T, Hill T (2019) Metagenome tracking biogeographic agroecology: phytobiota of tomatoes from Virginia, Maryland, North Carolina and California. Food Microbiol 79:132–136. https://doi.org/10.1016/j.fm.2018.12.001
Pérez-Rangel M, Quiroz-Figueroa FR, González-Castañeda J, Valdez-Vazquez I (2015) Microscopic analysis of wheat straw cell wall degradation by microbial consortia for hydrogen production. Int J Hydrogen Energy 40(1):151–160. https://doi.org/10.1016/j.ijhydene.2014.10.050
Perreault R, Laforest-Lapointe I (2021) Plant-microbe interactions in the phyllosphere: facing challenges of the anthropocene. ISME J 16(2):339–345. https://doi.org/10.1038/s41396-021-01109-3
Pokharel B, Wang S-YS, Gu H, LaPlante MD, Serago J, Gillies R, Meyer J, Beall S, Ikeda K (2021) A modeling examination of cloud seeding conditions under the warmer climate in Utah, USA. Atmos Res 248:105239. https://doi.org/10.1016/j.atmosres.2020.105239
Potnis N, Soto-Arias JP, Cowles KN, van Bruggen AHC, Jones JB, Barak JD (2014) Xanthomonas perforans colonization influences Salmonella enterica in the tomato phyllosphere. Appl Environ Microb 80(10):3173–3180. https://doi.org/10.1128/AEM.00345-14
Qi SS, Bogdanov A, Cnockaert M, Acar T, Ranty-Roby S, Coenye T, Vandamme P, König GM, Crüsemann M, Carlier A (2021) Induction of antibiotic specialized metabolism by co-culturing in a collection of phyllosphere bacteria. Environ Microbiol 23(4):2132–2151. https://doi.org/10.1111/1462-2920.15382
Rai AK, Al Makishah NH, Wen Z, Gupta G, Pandit S, Prasad R (2022) Recent developments in lignocellulosic biofuels, a renewable source of bioenergy. Fermentation 8(4):161. https://doi.org/10.3390/fermentation8040161
Rangel LI, Bolton MD (2022) The unsung roles of microbial secondary metabolite effectors in the plant disease cacophony. Curr Opin Plant Biol 68:102233. https://doi.org/10.1016/j.pbi.2022.102233
Rangel LI, Hamilton O, de Jonge R, Bolton MD (2021) Fungal social influencers: secondary metabolites as a platform for shaping the plant-associated community. Plant J 108(3):632–645. https://doi.org/10.1111/tpj.15490
Rastogi S, Singh A (2022) Gut microbiome and human health: exploring how the probiotic genus Lactobacillus modulate immune responses. Front Pharmacol 13:1042189. https://doi.org/10.3389/fphar.2022.1042189
Rastogi G, Sbodio A, Tech JJ, Suslow TV, Coaker GL, Leveau JHJ (2012) Leaf microbiota in an agroecosystem: spatiotemporal variation in bacterial community composition on field-grown lettuce. ISME J 6(10):1812–1822. https://doi.org/10.1038/ismej.2012.32
Regnat K, Mach R, Mach-Aigner A (2018) Erythritol as sweetener—wherefrom and whereto? Appl Microbiol Biotechnol 102:587–595. https://doi.org/10.1007/s00253-017-8654-1
Remus-Emsermann MNP, Schlechter RO (2018) Phyllosphere microbiology: at the interface between microbial individuals and the plant host. New Phytol 218(4):1327–1333. https://doi.org/10.1111/nph.15054
Rocky-Salimi K, Hashemi M, Safari M, Mousivand M (2016) A novel phytase characterized by thermostability and high pH tolerance from rice phyllosphere isolated Bacillus subtilis BS 46. J Adv Res 7(3):381–390. https://doi.org/10.1016/j.jare.2016.02.003
Rook GA (2013) Regulation of the immune system by biodiversity from the natural environment: an ecosystem service essential to health. P Natl Acad Sci 110(46):18360–18367. https://doi.org/10.1073/pnas.1313731110
Rosado BH, Almeida LC, Alves LF, Lambais MR, Oliveira RS (2018) The importance of phyllosphere on plant functional ecology: a phyllo trait manifesto. New Phytol 219(4):1145–1149. https://doi.org/10.1111/nph.15235
Roselli M, Natella F, Zinno P, Guantario B, Canali R, Schifano E, De Angelis M, Nikoloudaki O, Gobbetti M, Perozzi G (2021) Colonization ability and impact on human gut microbiota of foodborne microbes from traditional or probiotic-added fermented foods: a systematic review. Front Nutr 8:689084. https://doi.org/10.3389/fnut.2021.689084
Rzechonek DA, Dobrowolski A, Rymowicz W, Mirończuk AM (2018) Recent advances in biological production of erythritol. Crit Rev Biotechnol 38(4):620–633. https://doi.org/10.1080/07388551.2017.1380598
Saika A, Koike H, Yarimizu T, Watanabe T, Kitamoto H, Morita T (2019) Deficiency of biodegradable plastic-degrading enzyme production in a gene-deletion mutant of phyllosphere yeast, Pseudozyma antarctica defective in mannosylerythritol lipid biosynthesis. AMB Express 9(1):1–11. https://doi.org/10.1186/s13568-019-0825-2
Saleem M, Meckes N, Pervaiz ZH, Traw MB (2017) Microbial interactions in the phyllosphere increase plant performance under herbivore biotic stress. Front Microbiol 8:41. https://doi.org/10.3389/fmicb.2017.00041
Salzer WL, Asselin BL, Plourde PV, Corn T, Hunger SP (2014) Development of asparaginase Erwinia chrysanthemi for the treatment of acute lymphoblastic leukemia. Ann NY Acad Sci 1329(1):81–92. https://doi.org/10.1111/nyas.12496
Sangthong S, Suksabye P, Thiravetyan P (2016) Air-borne xylene degradation by Bougainvillea buttiana and the role of epiphytic bacteria in the degradation. Ecotoxicol Environ Saf 126:273–280. https://doi.org/10.1016/j.ecoenv.2015.12.017
Sarubbo LA, Maria da Gloria CS, Durval IJB, Bezerra KGO, Ribeiro BG, Silva IA, Twigg MS, Banat IM (2022) Biosurfactants: production, properties, applications, trends, and general perspectives. Biochem Eng J 181:108377. https://doi.org/10.1016/j.bej.2022.108377
Schäfer M, Vogel CM, Bortfeld-Miller M, Mittelviefhaus M, Vorholt JA (2022) Mapping phyllosphere microbiota interactions in planta to establish genotype–phenotype relationships. Nat Microbiol 7(6):856–867. https://doi.org/10.1038/s41564-022-01132-w
Scheublin TR, Leveau JHJ (2013) Isolation of Arthrobacter species from the phyllosphere and demonstration of their epiphytic fitness. MicrobiologyOpen 2(1):205–213. https://doi.org/10.1002/mbo3.59
Scheublin TR, Deusch S, Moreno-Forero SK, Muller JA, van der Meer JR, Leveau JHJ (2014) Transcriptional profiling of Gram-positive Arthrobacter in the phyllosphere: induction of pollutant degradation genes by natural plant phenolic compounds. Environ Microbiol 16(7):2212–2225. https://doi.org/10.1111/1462-2920.12375
Schlechter RO, Miebach M, Remus-Emsermann MNP (2019) Driving factors of epiphytic bacterial communities: a review. J Adv Res 19:57–65. https://doi.org/10.1016/j.jare.2019.03.003
Schlechter RO, Kear EJ, Bernach M, Remus DM, Remus-Emsermann MNP (2023) Metabolic resource overlap impacts competition among phyllosphere bacteria. ISME J 17:1445–1454. https://doi.org/10.1038/s41396-023-01459-0
Schmedes SE, Sajantila A, Budowle B (2016) Expansion of microbial forensics. J Clin Microbiol 54(8):1964–1974. https://doi.org/10.1128/JCM.00046-16
Schmedes SE, Budowle B (2019) Microbial forensics. In: Schmidt T (ed) Encyclopedia of microbiology, 4th edn. Elsevier, Amsterdam, pp 134–145. https://doi.org/10.1016/B978-0-12-801238-3.02483-1
Sen B, Aravind J, Kanmani P, Lay C-H (2016) State of the art and future concept of food waste fermentation to bioenergy. Renew Sustain Energy Rev 53:547–557. https://doi.org/10.1016/j.rser.2015.08.065
Shakir S, Zaidi SSEA, de Vries FT, Mansoor S (2021) Plant genetic networks shaping phyllosphere microbial community. Trends Genet 37(4):306–316. https://doi.org/10.1016/j.tig.2020.09.010
Singh B, Boukhris I, Kumar V, Yadav AN, Farhat-Khemakhem A, Kumar A, Singh D, Blibech M, Chouayekh H, Alghamdi OA (2020) Contribution of microbial phytases to the improvement of plant growth and nutrition: a review. Pedosphere 30(3):295–313. https://doi.org/10.1016/S1002-0160(20)60010-8
Singh E, Schenk PM, Carvalhais LC (2021) Sample preparation for culture-independent profiling and isolation of phyllosphere bacteria to identify potential biopesticides. In: Carvalhais LC, Dennis P (eds) The plant microbiome, vol 2232. Humana, New York, pp 193–208
Siriratruengsuk W, Furuuchi M, Prueksasit T, Luepromchai E (2017) Potential of pyrene removal from urban environments by the activities of bacteria and biosurfactant on ornamental plant leaves. Water Air Soil Pollut 228:1–16. https://doi.org/10.1007/s11270-017-3435-0
Sohrabi R, Paasch BC, Liber JA, He SY (2023) Phyllosphere microbiome. Annu Rev Plant Biol 74:539–568. https://doi.org/10.1146/annurev-arplant-102820-032704
Soto-Giron MJ, Kim J-N, Schott E, Tahmin C, Ishoey T, Mincer TJ, DeWalt J, Toledo G (2021) The edible plant microbiome represents a diverse genetic reservoir with functional potential in the human host. Sci Rep 11(1):1–14. https://doi.org/10.1038/s41598-021-03334-4
Stanton DE, Batterman SA, Von Fischer JC, Hedin LO (2019) Rapid nitrogen fixation by canopy microbiome in tropical forest determined by both phosphorus and molybdenum. Ecology 100(9):e02795. https://doi.org/10.1002/ecy.2795
Stone BW, Weingarten EA, Jackson CR (2018) The role of the phyllosphere microbiome in plant health and function. Annual Plant Reviews 1:1–24. https://doi.org/10.1002/9781119312994.apr0614
Taerum SJ, Steven B, Gage D, Triplett LR (2023) Dominance of ciliophora and chlorophyta among phyllosphere protists of solanaceous plants. Phytobiomes J 7(2):270–280. https://doi.org/10.1094/PBIOMES-04-22-0021-FI
Telemet (2024) SnowMax snow inducer. Accessed January 16, 2024. https://telemet.com/snomax-snow-inducer/
Temple TN, Stockwell VO, Loper JE, Johnson KB (2004) Bioavailability of iron to Pseudomonas fluorescens strain A506 on flowers of pear and apple. Phytopathology 94(12):1286–1294. https://doi.org/10.1094/PHYTO.2004.94.12.1286
ter Horst AM, Fudyma JD, Bak A, Hwang MS, Santos-Medellín C, Stevens KA, Rizzo DM, Al Rwahnih M, Emerson JB (2023) RNA viral communities are structured by host plant phylogeny in oak and conifer leaves. Phytobiomes J 7(2):288–296. https://doi.org/10.1094/PBIOMES-12-21-0080-R
Thapa S, Prasanna R (2018) Prospecting the characteristics and significance of the phyllosphere microbiome. Ann Microbiol 68(5):229–245. https://doi.org/10.1007/s13213-018-1331-5
Theodora NA, Dominika V, Waturangi DE (2019) Screening and quantification of anti-quorum sensing and antibiofilm activities of phyllosphere bacteria against biofilm forming bacteria. BMC Res Notes 12(732):1–5. https://doi.org/10.1186/s13104-019-4775-1
Tomás-Barberán FA, Rodríguez JM (2022) Interactions of food with the microbiota of the digestive tract. In: Glibetic M (ed) Comprehensive gut microbiota, vol 3. Elsevier, Serbia, pp 1–11
Trivedi P, Batista BD, Bazany KE, Singh BK (2022) Plant–microbiome interactions under a changing world: responses, consequences and perspectives. New Phytol. https://doi.org/10.1111/nph.18016
Valdez-Vazquez I, Morales AL, Escalante AE (2017) History of adaptation determines short-term shifts in performance and community structure of hydrogen-producing microbial communities degrading wheat straw. Microb Biotechnol 10(6):1569–1580. https://doi.org/10.1111/1751-7915.12678
Vanderhoef LN, Dute RR (1981) Auxin-regulated wall loosening and sustained growth in elongation. Plant Physiol 67(1):146–149. https://doi.org/10.1104/pp.67.1.146
Varvaresou A, Iakovou K (2015) Biosurfactants in cosmetics and biopharmaceuticals. Lett Appl Microbiol 61(3):214–223. https://doi.org/10.1111/lam.12440
Vitali B, Minervini G, Rizzello CG, Spisni E, Maccaferri S, Brigidi P, Gobbetti M, Di Cagno R (2012) Novel probiotic candidates for humans isolated from raw fruits and vegetables. Food Microbiol 31(1):116–125. https://doi.org/10.1016/j.fm.2011.12.027
Vogel CM, Potthoff DB, Schäfer M, Barandun N, Vorholt JA (2021) Protective role of the Arabidopsis leaf microbiota against a bacterial pathogen. Nat Microbiol 6(12):1537–1548. https://doi.org/10.1038/s41564-021-00997-7
Vorholt JA, Vogel C, Carlström CI, Mueller DB (2017) Establishing causality: opportunities of synthetic communities for plant microbiome research. Cell Host Microbe 22(2):142–155. https://doi.org/10.1016/j.chom.2017.07.004
Wassermann B, Rybakova D, Muller C, Berg G (2017) Harnessing the microbiomes of Brassica vegetables for health issues. Sci Rep 7(1):17649. https://doi.org/10.1038/s41598-017-17949-z
Watanabe T, Shinozaki Y, Yoshida S, Koitabashi M, Sameshima-Yamashita Y, Fujii T, Fukuoka T, Kitamoto HK (2014) Xylose induces the phyllosphere yeast Pseudozyma antarctica to produce a cutinase-like enzyme which efficiently degrades biodegradable plastics. J Biosci Bioeng 117(3):325–329. https://doi.org/10.1016/j.jbiosc.2013.09.002
Wei X, Lyu S, Yu Y, Wang Z, Liu H, Pan D, Chen J (2017) Phylloremediation of air pollutants: exploiting the potential of plant leaves and leaf-associated microbes. Front Plant Sci 8:1318. https://doi.org/10.3389/fpls.2017.01318
Wex H, Augustin-Bauditz S, Boose Y, Budke C, Curtius J, Diehl K, Dreyer A, Frank F, Hartmann S, Hiranuma N, Jantsch E, Kanji ZA, Kiselev A, Koop T, Mohler O, Niedermeier D, Nillius B, Rosch M, Rose D, Schmidt C, Steinke I, Stratmann F (2015) Intercomparing different devices for the investigation of ice nucleating particles using Snomax (R) as test substance. Atmos Chem Phys 15(3):1463–1485. https://doi.org/10.5194/acp-15-1463-2015
Weyens N, Thijs S, Pope R, Witters N, Przybysz A, Espenshade J, Gawronska H, Vangronsveld J, Gawronski S (2015) The role of plant-microbe interactions and their exploitation for phytoremediation of air pollutants. Int J Mol Sci 16(10):25576–25604. https://doi.org/10.3390/ijms161025576
Wicaksono WA, Cernava T, Wassermann B, Abdelfattah A, Soto-Giron MJ, Toledo GV, Virtanen SM, Knip M, Hyöty H, Berg G (2023a) The edible plant microbiome: evidence for the occurrence of fruit and vegetable bacteria in the human gut. Gut Microbes 15(2):2258565. https://doi.org/10.1080/19490976.2023.2258565
Wicaksono WA, Reisenhofer-Graber T, Erschen S, Kusstatscher P, Berg C, Krause R, Cernava T, Berg G (2023b) Phyllosphere-associated microbiota in built environment: do they have the potential to antagonize human pathogens? J Adv Res 43:109–121. https://doi.org/10.1016/j.jare.2022.02.003
Williams TR, Marco ML (2014) Phyllosphere microbiota composition and microbial community transplantation on lettuce plants grown indoors. mBio 5(4):e01564-14. https://doi.org/10.1128/mBio.01564-14
Williams TR, Moyne AL, Harris LJ, Marco ML (2013) Season, irrigation, leaf age, and Escherichia coli inoculation influence the bacterial diversity in the lettuce phyllosphere. PLoS ONE 8(7):e68642. https://doi.org/10.1371/journal.pone.0068642
Wilson M, Lindow SE (1993) Interactions between the biological control agent Pseudomonas fluorescens A506 and Erwinia amylovora in pear blossoms. Phytopathology 83(1):117–123. https://doi.org/10.17660/ActaHortic.1993.338.51
Yin X, Kelly KN, Maharaj NN, Rolshausen P, Leveau JHJ (2018) A microbiota-based approach to citrus tree health: in search of microbial biomarkers to pre-diagnose trees for HLB infection. Citrograph 9(4):58–63. https://citrus-research-board-static.sfo2.digitaloceanspaces.com/citrograph/pdf/CRB-Citrograph-Mag-Q4-Fall2018-Web-1.pdf
Yu AO, Leveau JHJ, Marco ML (2020) Abundance, diversity and plant-specific adaptations of plant-associated lactic acid bacteria. Environ Microbiol Rep 12(1):16–29. https://doi.org/10.1111/1758-2229.12794
Yuan Z, Druzhinina IS, Labbé J, Redman R, Qin Y, Rodriguez R, Zhang C, Tuskan GA, Lin F (2016) Specialized microbiome of a halophyte and its role in helping non-host plants to withstand salinity. Sci Rep 6(1):32467. https://doi.org/10.1038/srep32467
Zeng L-J, Huang Y-H, Lü H, Geng J, Zhao H-M, Xiang L, Li H, Li Y-W, Mo C-H, Cai Q-Y (2022) Uptake pathways of phthalates (PAEs) into Chinese flowering cabbage grown in plastic greenhouses and lowering PAE accumulation by spraying PAE-degrading bacterial strain. Sci Total Environ 815:152854. https://doi.org/10.1016/j.scitotenv.2021.152854
Zhalnina K, Hawkes C, Shade A, Firestone MK, Pett-Ridge J (2021) Managing plant microbiomes for sustainable biofuel production. Phytobiomes J 5(1):3–13. https://doi.org/10.1094/PBIOMES-12-20-0090-E
Zhang Z, Kong X, Jin D, Yu H, Zhu X, Su X, Wang P, Zhang R, Jia M, Deng Y (2019) Euonymus japonicus phyllosphere microbiome is significantly changed by powdery mildew. Arch Microbiol 201(8):1099–1109. https://doi.org/10.1007/s00203-019-01683-3
Zhong H, Wang J, Hafeez MA, Guan R, Feng F (2021) Lactobacillus plantarum ZJUFB2 prevents high fat diet-induced insulin resistance in association with modulation of the gut microbiota. Front Nutr 8:754222. https://doi.org/10.3389/fnut.2021.754222
Zhou J, Yu L, Zhang J, Zhang X, Xue Y, Liu J, Zou X (2020) Characterization of the core microbiome in tobacco leaves during aging. MicrobiologyOpen 9(3):e984. https://doi.org/10.1002/mbo3.984
Zhou J, Yu L, Zhang J, Liu J, Zou X (2021) Dynamic characteristics and co-occurrence patterns of microbial community in tobacco leaves during the 24-month aging process. Ann Microbiol 71(1):1–13. https://doi.org/10.1186/s13213-021-01620-0
Zhu K, Tan F, Mu J, Yi R, Zhou X, Zhao X (2019) Anti-obesity effects of Lactobacillus fermentum CQPC05 isolated from Sichuan pickle in high-fat diet-induced obese mice through PPAR-α signaling pathway. Microorganisms 7(7):194. https://doi.org/10.3390/microorganisms7070194
Zhu YG, Xiong C, Wei Z, Chen QL, Ma B, Zhou SYD, Tan J, Zhang LM, Cui HL, Duan GL (2021) Impacts of global change on phyllosphere microbiome. New Phytol. https://doi.org/10.1111/nph.17928
Author information
Authors and Affiliations
Contributions
LIR: conceptualization, writing original draft, review, and editing. JHJL: conceptualization, supervision, review, and editing. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rangel, L.I., Leveau, J.H.J. Applied microbiology of the phyllosphere. Appl Microbiol Biotechnol 108, 211 (2024). https://doi.org/10.1007/s00253-024-13042-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13042-4