Abstract
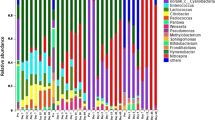
Ensiling is a feed preservation method of moist forage crops that generally depends on naturally developing lactic acid bacteria to convert water-soluble carbohydrates into organic acids. While bacterial community dynamics have been previously assessed in bench-scale and pilot ensiling facilities, almost no studies have assessed the microbiomes of large-scale silage facilities. This study analyzed bacterial community composition in mature silage from bunker silos in three commercial production centers as related to pH, organic matter, volatile fatty acid composition, and spatial distribution within the ensiling bunker. It revealed significant physicochemical differences between “preserved” regions situated in the center and along the walls of the silage bunkers that were characterized by high concentrations of lactic acid and other volatiles and pH values below 5, and “spoiled” regions in the corners (shoulders) of the bunkers that had low lactic acid concentrations and high pH values. Preserved silage was dominated (>90 %) by lactic acid bacteria and characterized by high similarity and low taxonomic diversity, whereas spoiled silage had highly diverse microbiomes with low abundances of lactic acid bacteria (<5 %) that were sometimes characterized by high levels of Enterobacteriaceae. Spatial position had a much stronger impact on the microbial community composition than feedstock type, sampling date, or production center location supporting previous studies demonstrating that ecology and not geography is a major driver of environmental microbiomes.





Similar content being viewed by others
References
Atamna-Ismaeel N, Finkel OM, Glaser F, Sharon I, Schneider R, Post AF, Spudich JL, von Mering C, Vorholt JA, Iluz D, Béjà O, Belkin S (2012) Microbial rhodopsins on leaf surfaces of terrestrial plants. Environ Microbiol 14(1):140–146
Baran MR, Negri LC, Chineze PHN, Bronkhorst DE, Pereira CES, Bogado ALG, da Silva LC, Marcasso RA, Okano W (2015) Proteus mirabilis causes of renal affection associated with pathological changes in Nellore bull—case report. Brazilian J Hyg Anim Sanity 9(1):78–90. doi:10.5935/1981-2965.20150008
Barker SB, Summerson WH (1941) The colorimetric determination of lactic acid in biological material. J Biol Chem 138(2):535–554
Bodenhausen N, Horton MW, Bergelson J (2013) Bacterial communities associated with the leaves and the roots of Arabidopsis thaliana. PLoS One 8(2):e56329
Bolsen K, Ashbell G, Weinberg Z (1996) Silage fermentation and silage additives—review. Asian-Australasian J Anim Sci 9(5):483–494
Buxton DR, Muck RE, Harisson JH (2003) Silage science and technology agronomy monograph 42. American Society of Agronomy, Crop Science Society of America, Soil Science Society of America, Madison
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336
Coker C, Poore CA, Li X, Mobley HL (2000) Pathogenesis of Proteus mirabilis urinary tract infection. Microbes Infect 2(12):1497–1505
Dini-Andreote F, Stegen JC, van Elsas JD, Salles JF (2015) Disentangling mechanisms that mediate the balance between stochastic and deterministic processes in microbial succession. Proc Natl Acad Sci 112(11):E1326–E1332
Driehuis F, Elferink SO (2000) The impact of the quality of silage on animal health and food safety: a review. Vet Q 22(4):212–216
Duniere L, Sindou J, Chaucheyras-Durand F, Chevallier I, Thevenot-Sergentet D (2013) Silage processing and strategies to prevent persistence of undesirable microorganisms. Anim Feed Sci Technol 182(1–4):1–15. doi:10.1016/j.anifeedsci.2013.04.006
Eikmeyer FG, Köfinger P, Poschenel A, Jünemann S, Zakrzewski M, Heinl S, Mayrhuber E, Grabherr R, Pühler A, Schwab H (2013) Metagenome analyses reveal the influence of the inoculant Lactobacillus buchneri CD034 on the microbial community involved in grass ensiling. J Biotechnol 167(3):334–343
Li Y, Nishino N (2011) Monitoring the bacterial community of maize silage stored in a bunker silo inoculated with Enterococcus faecium, Lactobacillus plantarum and Lactobacillus buchneri. J Appl Microbiol 110(6):1561–1570
Martiny JBH, Bohannan BJ, Brown JH, Colwell RK, Fuhrman JA, Green JL, Horner-Devine MC, Kane M, Krumins JA, Kuske CR (2006) Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4(2):102–112
Mendez-Garcia C, Pelaez AI, Mesa V, Sanchez J, Golyshina OV, Ferrer M (2015) Microbial diversity and metabolic networks in acid mine drainage habitats. Front Microbiol 6:17. doi:10.3389/fmicb.2015.00475
Minh TTT, Huu VN, Nishino N (2014) A pilot examination of the fermentation products, aerobic stability and bacterial community of total mixed ration silage produced in Vietnam. Grassl Sci 60(1):63–68. doi:10.1111/grs.12041
Moonsamy P, Williams T, Bonella P, Holcomb C, Höglund B, Hillman G, Goodridge D, Turenchalk G, Blake L, Daigle D (2013) High throughput HLA genotyping using 454 sequencing and the Fluidigm Access Array™ system for simplified amplicon library preparation. Tissue Antigens 81(3):141–149
Muck R (2013) Recent advances in silage microbiology. Agricultural Food Sci 22(1):3–15
Naoki N, Yuji T (2008) Variations in bacterial communities in laboratory-scale and big bale silos assessed by fermentation products, colony counts and denaturing gradient gel electrophoresis profiles. Lett Appl Microbiol 46(3):283–288. doi:10.1111/j.1472-765X.2007.02306.x
O’Hara CM, Brenner FW, Miller JM (2000) Classification, identification, and clinical significance of Proteus, Providencia, and Morganella. Clin Microbiol Rev 13(4):534–546
O’Malley MA (2007) The nineteenth century roots of ‘everything is everywhere’. Nat Rev Microbiol 5(8):647–651
Oakley BB, Lillehoj HS, Kogut MH, Kim WK, Maurer JJ, Pedroso A, Lee MD, Collett SR, Johnson TJ, Cox NA (2014) The chicken gastrointestinal microbiome. FEMS Microbiol Lett 360(2):100–112. doi:10.1111/1574-6968.12608
Ogiy S, Chen Y, Pasvolsky R, Weinberg ZG, Shemesh M (2015) High resolution melt analysis to confirm the establishment of Lactobacillus plantarum and Enterococcus faecium from silage inoculants during ensiling of wheat. Grassl Sci 62(1):29–36
Reisman HB (1993) Problems in scale-up of biotechnology production processes. Crit Rev Biotechnol 13(3):195–253
Rózalski A, Sidorczyk Z, Kotełko K (1997) Potential virulence factors of Proteus bacilli. Microbiol Mol Biol Rev 61(1):65–89
Samuni-Blank M, Izhaki I, Laviad S, Bar-Massada A, Gerchman Y, Halpern M (2014) The role of abiotic environmental conditions and herbivory in shaping bacterial community composition in floral nectar. PloS one 9(6):e99107
Stevenson DM, Muck RE, Shinners KJ, Weimer PJ (2006) Use of real time PCR to determine population profiles of individual species of lactic acid bacteria in alfalfa silage and stored corn stover. Appl Microbiol Biotechnol 71(3):329–338
Wang C, Han H, Gu X, Yu Z, Nishino N (2014) A survey of fermentation products and bacterial communities in corn silage produced in a bunker silo in China. Anim Sci J 85(1):32–36
Weinberg ZG, Ashbell G (1994) Changes in gas composition in corn silages in bunker silos during storage and feedout. Can Agric Eng 36(3):155–158
Weinberg ZG, Muck R (1996) New trends and opportunities in the development and use of inoculants for silage. FEMS Microbiol Rev 19(1):53–68
Weinberg ZG, Muck RE, Weimer PJ, Chen Y, Gamburg M (2004) Lactic acid bacteria used in inoculants for silage as probiotics for ruminants. Appl Biochem Biotechnol 118(1–3):1–9
Woolford M (1990) The detrimental effects of air on silage. J Appl Bacteriol 68(2):101–116
Wu JJ, Du RP, Gao M, Sui YQ, Xiu L, Wang X (2014) Naturally occurring lactic acid bacteria isolated from tomato pomace silage. Asian-Australasian J Anim Sci 27(5):648–657. doi:10.5713/ajas.2013.13670
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical statement
ᅟ
Funding
This study was supported by a grant (301-0793-14) from the Israeli Ministry of Agriculture and Rural Development.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 180 kb)
Rights and permissions
About this article
Cite this article
Kraut-Cohen, J., Tripathi, V., Chen, Y. et al. Temporal and spatial assessment of microbial communities in commercial silages from bunker silos. Appl Microbiol Biotechnol 100, 6827–6835 (2016). https://doi.org/10.1007/s00253-016-7512-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7512-x