Abstract
Triticale is a cereal of high economic importance; however, along with the increase in the area of this cereal, it is more often infected by the fungal pathogen Blumeria graminis, which causes powdery mildew. The rapid development of molecular biology techniques, in particular methods based on molecular markers may be an important tool used in modern plant breeding. Development of genetic maps, location of the QTLs defining the region of the genome associated with resistance and selection of markers linked to particular trait can be used to select resistant genotypes as well as to pyramidize several resistance genes in one variety. In this paper, we present a new, high-density genetic map of triticale doubled haploids (DH) population “Grenado” × “Zorro” composed of DArT, silicoDArT, and SNP markers. Composite interval mapping method was used to detect eight QTL regions associated with the area under disease progress curve (AUDPC) and 15 regions with the average value of powdery mildew infection (avPM) based on observation conducted in 3-year period in three different locations across the Poland. Two regions on rye chromosome 4R, and single loci on 5R and 6R were reported for the first time as regions associated with powdery mildew resistance. Among all QTLs, 14 candidate genes were identified coded cyclin-dependent kinase, serine/threonine-protein kinase-like protein as well as AMEIOTIC 1 homolog DYAD-like protein, DETOXIFICATION 16-like protein, and putative disease resistance protein RGA3. Three of identified candidate genes were found among newly described QTL regions associated with powdery mildew resistance in triticale.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Triticale (xTriticosecale Wittm.) is a human-made wheat-rye hybrid commercialized in the late 1960s (Ammar et al. 2004). Currently cultivated, hexaploid triticale (2n = 6x = 42, AABBRR) accumulates important traits determined by wheat (A and B) and rye (R) genomes (Walker et al. 2011; Klocke et al. 2013). In the last years, triticale has raised its economic importance mainly in Europe. Poland with triticale cultivation area of 1.3 million ha contribute to 1/3 of world production and remains the top producer of this crop (Faostat 2020). Simultaneously, risk of infection by the biotrophic fungal pathogen Blumeria graminis (DC.) Speer which causes powdery mildew has recently increased.
The epidemic appearance of powdery mildew on triticale has been observed in several European countries, including Belgium, France, Germany, and Poland as well (Walker et al. 2011). An epidemics of powdery mildew causes yield drop and requires preventive use of fungicides. The cultivation of triticale varieties resistant to pathogenic fungi offers the most economical and environmentally friendly alternative to chemical protection. So far, 50 loci with more than 78 genes/alleles associated with powdery mildew resistance have been identified on 18 chromosomes of bread wheat and its relatives (Yang et al. 2017) and only 8 resistance genes have been identified in rye (Tyrka and Chelkowski 2004; Schlegel and Korzun, 2021). Many of these resistance genes were broken down by the new races of B. graminis (Menardo et al. 2016), and triticale can benefit both from genes present in rye and introduced into wheat from alien species (Tyrka and Chelkowski 2004; Alam et al. 2013; Schlegel and Korzun, 2021).
Techniques based on DNA molecular markers has become an indispensable tool in modern plant breeding used to monitor introgression and for accumulation of desired genes in breeding materials (Yang et al. 2015). A number of methods based on DNA hybridization (Jaccoud et al. 2001; Cavanagh et al. 2013; Jordan et al. 2015) and next generation sequencing (Vikram et al. 2016; Riaz et al. 2016; Baloch et al. 2017) have been developed and used for wheat or triticale genotyping. Recently, sequencing efforts resulted in assembling of wheat and rye genome (IWGSC 2014, 2018; Bauer et al. 2017; Rabanus-Wallaceet al. 2021). However, in species with sequenced genomes, genetic maps are useful for detecting chromosomal rearrangements (Wingen et al. 2017) and necessary for quantitative trait loci (QTLs) localization (Vinod 2009; Holtz et al. 2016). Therefore, a number of genetic maps have already been developed for wheat (Somers et al. 2004; Mantovani et al. 2008), rye (Korzun et al. 2001; Milczarski et al. 2011), and triticale (Alheit et al. 2011; Tyrka et al. 2011, 2015; Karbarz et al. 2020; Wąsek et al. 2021).
The aims of this study were to (1) develop a high-density genetic map for hexaploid winter triticale composed of diversity arrays technology (DArT), silicoDArT, and DArT-based single nucleotide polymorphism (SNP) markers using DH population of lines derived from two triticale cultivars and (2) identify QTL regions and candidate genes responsible for an adult-plant resistance of triticale (xTriticosecale Wittm.) to powdery mildew infection in natural field conditions.
Materials and methods
Experimental population
The mapping population used in this study consisted 168 doubled haploid (DH) lines derived from F1 hybrid “Grenado” × “Zorro.” “Grenado” was resistant parent and “Zorro” was highly susceptible to infection of B. graminis. These cultivars were registered by Strzelce Plant Breeders Ltd (Plant Breeding and Acclimatization Institute Group, Poland) and Danko Plant Breeders Ltd, respectively. The DH lines were obtained at the Department of Cell Biology of Institute of Plant Physiology Polish Academy of Science (IPP PAS) in Kraków by the anther culture method according to Wędzony (2003).
Plant growth conditions and phenotyping
For the first year of field experiment, lines were reproduced in greenhouse and healthy leaves were sampled for DNA isolation. Seeds of parental lines and each DH line were germinated in plastic pots (3.7 dm3; nine seeds per pot), previously filled with a homogeneous mixture of sand and soil (3:1; v/v). The pots were placed for 8 weeks in a cool chamber at 4 °C (± 1 °C), photoperiod 10-h light/14-h dark. Next, the plants were transferred into a greenhouse chambers with air temperature 26–28/18 °C (± 2 °C) day/night and relative air humidity 40%. All plants were irrigated once a week with a Hoagland’s solution (Hoagland 1948). The seeds were obtained from individual DH lines and their parents from bagged spikes in the greenhouse in the IPP PAS in Kraków. Seed material for the second and the third year of experiment was obtained in field conditions in Danko Plant Breeders Ltd by isolation of five spikes per each DH line before flowering.
Powdery mildew (PM) resistance was assessed in field conditions for three years (2013–2015) in three localizations spread across Poland: Choryń (52° 2′ 26″ N 16° 46′ 59″ E; all three seasons), Laski (51° 47′ N 21° 12′ E; season 2012/2013 and 2013/2014) and Modzurów (50° 9′ 20″ N 18° 7′ 52″ E; season 2014/2015). The lines were sown in two 1-m long rows at the 20 × 2.5 cm spacing. Susceptible cultivar “Zorro” was sewed as spreader every 20 plots. The chemical protection was not applied during plant growth and powdery mildew infection was measured under natural infection. Disease was assessed on a whole plot basis using a 0–9 scale (McNeal et al. 1971), where 0 is immune and 9 is very susceptible (Ziems et al. 2014). Observations were made in periods of heading, flowering, and seed formation. Depending on the weather conditions during field experiments (high temperature and drought) which led to death of some plants, field observations of the PM degree were conducted in one, two, or three stages. Data which were recorded 3 times during one vegetative season in Choryń were used to calculate area under disease progress curve (AUDPC) (Shaner and Finney 1977; Finckh et al. 1999; Jeger and Viljanen-Rollinson 2001), whereas data recorded once or 2 times were used to determine the average value of powdery mildew infection (avPM) according to the 9-grade scale.
DNA isolation and genotyping
Genomic DNA was isolated from a 90- to100-mg sample of two leaves per each DH line and both parents. The samples were frozen in liquid nitrogen and stored at − 60 °C until the isolation was made. Total genomic DNA isolation for each sample was carried out using the GeneJET Plant Genomic DNA Purification Mini Kit (Thermo Scientific, Waltham, USA). The concentration and purity of the DNA was evaluated using a UV–Vis Q500 (Quawell, San Jose, USA) spectrophotometer. DNA was sent to Diversity Arrays Technology (Yarralumla, Australia) both for profiling using triticale high -resolution array (DArT) with probes representing markers from rye, wheat, and triticale (rPt, wPt, and tPt, respectively) and for DArTseq analysis.
Construction of the genetic map
De novo mapping approach was used to construct genetic map for “Grenado” × “Zorro” DH population. Markers of unknown parental origin and present the frequency < 5% and > 95% were removed from the dataset. All types of DArT markers were binned with QTL IciMapping (Wang et al. 2016). Segregation data were analyzed using JoinMap4 (Van Ooijen 2006) to group all markers using the logarithm of odds (LOD) > 3. Markers within these groups were recurrently ordered using the maximum likelihood option of JoinMap and the RECORD program (Van Os et al. 2005). To establish the marker order, all linkage groups identified for “Grenado” × “Zorro” DH population were compared to reference genetic maps of triticale (Tyrka et al. 2015), reference genome of wheat at URGI (https://urgi.versailles.inra.fr) and partial rye genome (Bauer et al. 2017).
Statistical, QTL, and candidate genes analysis
Mean values from all observations were used to calculate the Pearson’s correlations. The Shapiro–Wilk test was performed to assess deviations from a normal distribution as well as skewness and kurtosis were calculated using Statistica version 12.0 (StatSoft, Inc. USA). High-density genetic map and complete phenotyping data of the degree of powdery mildew infection intriticale were exploited in QTL analysis using WinQTLCartographer2.5 software (Wang et al. 2012). Composite interval mapping (CIM) analysis with a 1000-permutation test and walk speed of 1.0 cM were performed to declare a significant QTL. The LOD threshold was between 2.1 and 8.3 depending on the trait. The percentage of the phenotypic variation covered by QTL was calculated with a single factor regression (R2) and the favorable alleles in each QTL region were selected, based on the additive (Add) effect (negative additive effect refers to cv. “Zorro” while positive to cv. “Grenado”). Candidate genes analysis was performed according to method detailed described by Wąsek et al. (2021).
Results
Phenotypic analysis
Phenotypic variation in powdery mildew infection was assessed for all lines of the “Grenado” × “Zorro” DH population and for both parental lines in Choryń, Laski, and Modzurów during all three vegetative seasons (Table 1, Fig. S1). According to Shapiro–Wilk test, distributions of AUDPC and avPM values over locations and seasons not deviated significantly from a normal distribution. Skewness and kurtosis values also confirmed the proper distribution of observations for the experiments (Table 1). AUDPC values varied significantly depending on the year of experiment. Although, maximum values of AUDPC between years were similar and amounted to 2675.6 and 2530.6, different dynamics of disease development was observed and minimum AUDPC values ranged from 65.2 and 1678.3 in 2015 and 2013, respectively. Average avPM values ranged from 3.3 to 5.7 (Table 1). Besides, statistically significant highly positive correlations between different powdery mildew scores were found within locations that reflect disease progression. Powdery mildew distribution for Choryń in 2015 was significantly, positively correlated also with observations in Modzurów and Laski (Table 2).
The “Grenado” × “Zorro” linkage map
A total of 1891 unique markers (1443 silicoDArT, 326 DArT, and 122 SNP) were assigned to 21 linkage groups corresponding to all triticale chromosomes (Table S1). However, for chromosomes 7A and 1B, additional separate linkage groups were discerned (7A.1 and 1B.1, respectively). These groups were left separate because combining them into a single linkage group was connected with the insertion of large gaps (above 30 cM). The genetic linkage map spanned 5249.9 cM with average marker density of 2.8 cM (Table 3). The A, B, and R genomes covered total distances of 1556.0, 1906.9, and 1787.0 cM, respectively. The A genome had the fewest markers assigned (538) and the highest markers saturation (3.0) comparing to the other triticale genomes. The total number of markers assigned the B and R genomes was 691 and 662, respectively with the corresponding maps saturation of 2.7 and 2.8 (Table 3).
Detection of QTLs for powdery mildew resistance in triticale in all seasons and localizations
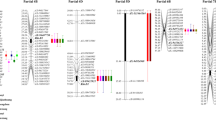
QTLs were calculated from the mean values of data obtained for each experiment separately. Identification of QTL associated with powdery mildew infection was carried out based on the genetic map created de novo for the “Grenado” × “Zorro” DH population. Composite interval mapping (CIM) identified total of 23 QTLs with LOD values ≥ 2.0 on 6 wheat (A and B) chromosomes: 4A, 7A, 7A.1, 2B, 3B, and 7B and 10 on rye (R) chromosomes: 1R, 4R, 5R, and 6R (Table 4, Fig. S2, Fig. S3).
Loci associated with AUDPC evaluated in Choryń in 2013 and 2015 were located on chromosomes 7A, 7A.1, 4R, and 6R (Table 4, Fig. S2, Fig. S3). Those loci explained up to 15.2% and 16.2% of phenotypic variation for Qpm.gz.4R.1 and Qpm.gz.7A1.1 respectively. The highest LOD values were observed for Qpm.gz.4R.1 (6.8), Qpm.gz.7A1.1 (6.8 and 5.7), and Qpm.gz.7B.2 (6.5, Table 4). Also, common QTL regions for both AUDPC measured in 2013 and 2015 were found on chromosomes 4R and 7A.1. Locus Qpm.gz.4R.1 was co-located with Qpm.gz.4R.6 on chromosome 4R between 60.5 cM and 68.9 cM as well as Qpm.gz.7A1.1 with Qpm.gz.7A1.4 on chromosome 7A.1 between 5.9 and 16.3 cM (Table 4, Fig. S2, Fig. S3).
The avPM which was measured within 2-year time period in three different locations revealed total of 15 loci associated with that trait on chromosomes 4A, 7A.1, 2B, 3B, 7B, 1R, 4R, 5R, and 6R (Table 4, Fig. S2, Fig. S3). Among of all 15 loci, the most significant QTLs are those stable over years and locations. On chromosome 7A.1, loci Qpm.gz.7A1.2 and Qpm.gz.7A1.5 were detected for avPM measured in Choryń location in 2014 and Laski in 2015 (Table 4, Fig. S2). These QTLs covered the same region on 7A.1 chromosome (5.9–29.8 cM) and explained 15.2% of phenotypic variation for Qpm.gz.7A1.5 (Table 4). On chromosome 7B loci, Qpm.gz.7B.1 and Qpm.gz.7B.3 were detected between 174.3 and 190.2 cM in Choryń 2014 and Modzurów 2015 (Table 4, Fig. S2). It explained up to 17.3% of phenotypic variation and also, the same markers have peaked to the maximum LOD position (4,344,428 and 3,623,588). Additionally, on rye chromosome 5R one QTL Qpm.gz.5R.1 was identified. This locus was composed of three regions—between 0.0 and 34.6 cM, 45.7–60.5 cM and 95.2–109.7 cM but all of them have a very similar additive effects and phenotypic variation (Table 4). Therefore, Qpm.gz.5R.1 can be considered as one locus with effect split into three parts.
Candidate genes for adult-plant resistance
Fourteen candidate genes were detected within 11 QTL regions identified in this study on chromosomes: 7A (3), 2B (1), 3B (2), 7B (2), 1R (1), 4R (1), 5R (3), and 6R (1) (Table 5). Among them, two gene records were repeated in different experiments. The first gene encoding GDSL esterase/lipase At4g28780-like (LOC119328445) was identified within Qpm.gz.7A1.1, Qpm.gz.7A1.2, and Qpm.gz.7A1.5 found for AUDCP Choryń 2013, avPM Choryń 2014, and avPM Laski 2015 experiments. The second gene encoding CLAVATA3/ESR (CLE)-related protein 3-like (LOC119335261) was common for Qpm.gz.4R.2 and Qpm.gz.4R.5. The four other candidate genes from QTLs Qpm.gz.6R.1, Qpm.gz.1R.1, Qpm.gz.3B.2, and Qpm.gz.7B.3 coded different kinases like cyclin-dependent kinase A-2-like (LOC119314733), G-typelectin S-receptor-likeserine/threonine-protein kinase At2g19130 (LOC119294828), receptor-like protein kinase At3g47110 (LOC119266893) as well as serine/threonine-protein kinase-like protein ACR4 (LOC119325260), respectively. The remaining genes encoded: protein AMEIOTIC 1 homolog DYAD-like protein (LOC119308950), protein DETOXIFICATION 16-like (LOC119339835), putative disease resistance protein RGA3 (LOC119347815), sodium transporter HKT7-A1, uncharacterized F-box family protein (LOC109735658), uncharacterized ATP-dependent protease ATP asa subunit HslU (LOC119311530) as well as two uncharacterized proteins LOC109764755 and LOC113333611 (Table 5).
Discussion
Based on de novo mapping using unique silicoDArT, DArT, and SNP set of markers, the genetic map for triticale was constructed. This map was used to locate quantitative trait loci (QTL) associated with powdery mildew infection which was measured in a field conditions during 3-year period in three different locations across the Poland.
The genetic map created for “Grenado” × “Zorro” DH population was composed of 1891 markers assigned to 21 chromosomes which corresponds to triticale genome. The majority of this map was constructed of unique 1443 silicoDArT markers with 326 DArT and 122 SSR markers. DArT technique which is quick and highly reproducible can produce thousands of polymorphic loci in a single assay (Wenzl et al. 2004; Alam et al. 2018) that is why is widely used in genetic map construction for multiple crop species (Nsabiyera et al. 2020). However, DArT markers differ in intensity which may have an impact in some applications (Bolibok-Brągoszewska et al. 2009) that is why, a new genotyping technique, SNP chips has been developed and designed for a large number of SNPs (Nsabiyera et al. 2020; von Thaden et al. 2020). SNP chip method enables identification of quantitative trait loci (QTL) for different traits in various plant species (Ballesta et al. 2020; von Thaden et al. 2020). The total length of genetic map described in this paper was 5249.9 cM with the mean markers saturation 2.8 (3.0 for A, 2.7 for B, and 2.8 for R genome). Up to date, not many genetic maps were constructed and described for triticale (González et al. 2005; Alheit et al. 2011; Tyrka et al. 2011, 2015, 2018; Karbarz et al. 2020; Wąsek et al. 2021). The results of total marker number and mean map density are very similar to the genetic map of “Saka3006” × “Modus” DH mapping population described by Tyrka et al. (2011). From all markers, the highest number of them was assigned to the B genome (691) which is not corresponding to other described triticale genetic maps in contrast to the A genome with the lowest total number of markers (538). The A genome was previously described by Tyrka et al. (2011, 2015), Karbarz et al. (2020) and Wąsek et al. (2021) as the one with the lowest number of markers assigned, regardless of marker type used in map construction.
Based on the genetic map, detection of quantitative trait loci (QTL) associated with many important traits can be performed. Studies on localization of genomic regions in crops associated with resistance to fungal pathogens most often focused on fusarium head blight (Buerstmayr et al. 2002, 2003; Giancaspro et al. 2016; Clinesmith et al. 2019) and rusts (Melichar et al. 2008; Prins et al. 2011; Rosewarne et al. 2012; Li et al. 2020) especially in wheat. Regarding to powdery mildew resistance, identification of QTL was widely reported in wheat (Lan et al. 2010; Ren et al. 2017; Liu et al. 2020; Xu et al. 2020) in contrast to triticale (Karbarz et al. 2020). In this paper, detection of QTL regions linked to B. graminis resistance was tested in natural field conditions. Based on field results of triticale resistance, the area under disease progress curve (AUDPC) and the average value of powdery mildew infection (avPM) were calculated to obtain genomic regions associated with these traits.
On chromosome 4A, one locus Qpm.gz.4A.1 was detected in observations conducted in Modzurów in 2015 that explained 13.7% of phenotypic variation (Table 4). On this chromosome, regions with high importance for wheat health were previously described (Chantret et al. 2001; Mingeot et al. 2002; Jakobson et al. 2012). Chromosome 4A has been reported a source of resistance genes not only to powdery mildew (Pm16) but also to leaf stripe and rust resistance (Reader and Miller 1991; Marone et al. 2012, 2013).
Six QTL regions were detected for both AUDPC and avPM in almost all experiments (except Modzurów location in 2015). Wheat chromosome 7A is known as a source of multiple Pm resistance genes (Yang et al. 2017;Nordestgaard et al. 2020) as well as QTL regions associated with powdery mildew resistance. Three of them, Qpm.gz.7A1.1, Qpm.gz.7A1.2, and Qpm.gz.7A1.5 were found for AUDPC and avPM on the same position in a distance between 5.9 and 29.8 cM (Table 4, Fig. S2). Additionally, locus Qpm.gz.7A1.4 was located between 0.0 and 16.3 cM for AUDPC with maximum LOD at the position of 6.4 cM (Table 4). Karbarz et al. (2020) reported locus QPm-7A in triticale associated with AUDPC of B. graminis infection in a distance between 0.0 and 23.3 cM which is very similar to results obtained in this study. Also, Chantret et al. (2001) described loci involved in adult-plant resistance (APR) on 7A in wheat F2:3 population which position of one of them coincides with locus Qpm.gz.7A1.4. Furthermore, the Pm1 gene associated with the stem and leaf rust resistance genes Sr15 and Lr20 as well as gene Pm37 are already reported on chromosome 7A (Neu et al. 2002; Marone et al. 2013). Additionally, genes associated with cellular hydrolase activity, acting on ester bonds, lipid catabolic process, ADP, ATP nucleic acid binding, and zinc ion binding were localized within QTLs on chromosome 7A (Table 5).
On chromosome arm 2BL, six powdery mildew resistance genes: Pm6, Pm26, MlZec1, Pm33, MlLX9, and Pm51 were previously located (Zhan et al. 2014). In presented study, locus Qpm.gz.2B.1 with LOD value 3.0 was found for avPM (Table 4) with a candidate gene TraesCS2B01G868700LC encoded cation transmembrane transporter activity (Table 5). Marone et al. (2013) also localized QTL region on this chromosome with marker Xcdo244 corresponding to a NBS-LRR gene. Also, Asad et al. (2014) identified QTL for maximum disease severities (MDS) on this chromosome. Locus QPm.caas-2BS.2 was mapped in a position which has a pleiotropic effect on both powdery mildew and stripe rust responses (Guo et al. 2008; Carter et al. 2009).
Two regions on chromosome 3B, Qpm.gz.3B.1 and Qpm.gz.3B.2 were found for avPM measured in 2015 in two different locations with a common chromosome region between 104.0 and 115.3 cM (Table 4, Fig. S2). The highest LOD value (3.0) and phenotypic variation (10.1%) were for Qpm.gz.3B.2 with maximum LOD marker wPt-1159 peak at 115.3 cM. Also, putative receptor-like protein kinase At3g47110 (LOC119266893) gene was located between 104.0 and 133.0 cM on this locus (Table 5). Two loci on a short and long arm of chromosome 3B were described by Asad et al. (2014) explained 9.1% and 18.1% of phenotypic variation. Both of those regions were in close location to Pm13 and Pm41 genes. Another locus on chromosome 3B was reported by Marone et al. (2013) with the marker F103 peak on 3.9 cM position. Although, regions reported so far differ in a genetic position on 3B chromosome from QTL regions described in this paper, comparison of physical regions is necessary to suggest that both loci with high phenotypic variation effect can be a new source of powdery mildew resistance.
Three regions for both, AUDPC and avPM values from 2 years and two different locations were found on chromosome 7B. Those QTL have a common region in a distance from 174.3 to 185.1 cM with the highest LOD value (6.5) and phenotypic variation (14.9%) for Qpm.gz.7B.2 (Table 4, Fig. S2). Genes in this region were involved in the transmembrane antiporter activity, xenobiotic transmembrane transporter activity, and plant epidermal cell differentiation (Table 5). Keller et al. (1999) identified locus on the position 134 cM to 158 cM in four out of the five environments. It was located on a long arm of this chromosome and linked to Pm5 gene. Region described by Marone et al. (2013) was flanked by wPt-8938 and PmTm4 in a position of 137.7 cM on 7B. That locus can be confirmed by Qpm.gz.7B.1 as this region starts from marker 4,339,655 in a position 162.5 cM which is in a close position to wPt-8938 at 159.7 cM of “Grenado” × “Zorro” map (Tab. S1). Additionally, Chantret et al. (2001) and Mingeot et al. (2002) described locus on this chromosome associated with the resistance. These regions on 7B may correspond to Qpm.gz.7B.1 to Qpm.gz.7B.3.
Localization of QTL regions and genes associated with powdery mildew resistance in rye is poorly described so far, comparing to wheat. But close relationship between wheat and rye allows the introduction of desirable agronomic traits from rye to wheat, such as tolerance to various abiotic factors, resistance to pests and fungal diseases, including resistance to powdery mildew (Crespo-Herrera et al. 2017). Long arm of 1R rye chromosome is widely used to obtain a new varieties of wheat using chromosomal translocation of 1BL.1RS or 1AL.1RS and transferring Pm8 and Pm17 genes into the wheat (Duan et al. 2017; Schlegel and Korzun 2021). Remaining rye chromosomes also contain genes which can be used to improve wheat cultivars (Landjeva et al. 2006). Genes Pm7 and Pm20, from rye chromosomes 2RL and 6RL have been already transferred to many wheat cultivars causing powdery mildew resistance (Huang and Röder 2004; An et al. 2013, 2015; Guo et al. 2017; Schlegel and Korzun 2021).
In presented study, QTL regions for AUDCP and avPM have been identified on rye chromosomes 1R, 4R, 5R, and 6R (Table 4, Fig. S3). Locus Qpm.gz.1R.1 on chromosome 1R, covered by markers in a distance 67.3 cM to 91.2 cM was detected for avPM in Laski in 2014. It explained up to 7.4% of phenotypic variation with the LOD value 4.1. The short arm of this chromosome is an important source of genes carrying resistance to leaf and stem rust, yellow rust, and powdery mildew (Schlegel and Meinel 1994; Landjeva et al. 2006) that may correspond to QTL region associated with powdery mildew resistance.
Total of six loci for both, AUDCP and avPM were found on chromosome 4R with the highest LOD value 6.8 and 15.2% of phenotypic variation for Qpm.gz.4R.1. For those, two common regions were identified on a distance 60.5–68.9 cM and 161.9–189.7 cM (Table 4, Fig. S3). Within all identified loci on 4R, CLAVATA3/ESR (CLE)-related protein 3-like protein was found in SECCE4Rv1G0263150 candidate gene (Table 5). It has been reported that rye chromosome 4R contains the elite pool of genes which are applicable for wheat cultivar improvement (Duan et al. 2017). Up to date, five Pm genes derived from rye have been identified and transferred into the wheat genome, especially Pm8 which is one of the most effective and has made a contribution to control wheat powdery mildew (Huang and Röder 2004; Ma et al. 2020). Additionally, Karbarz et al. (2020) described a locus on 4R triticale chromosome, detected for AUDPC which flanking marker rPt-505620 in a position of 175.2 cM is in a close distance to flanking marker rPt-401230 of Qpm.gz.4R.3 at 116.4 cM. We can infer that two new resistance loci to powdery mildew corresponding to three QTLs common with Qpm.gz.4R.1 and two QTLs from region of Qpm.gz.4R.2 were identified.
Qpm.gz.5R.1 region, identified for avPM in Laski in 2014 consisted of three regions separated from each other by 11.1 cM and 34.7 cM (Table 4, Tab. S1). But due to very similar phenotypic and additive effects, it has been considered as one locus on 5R chromosome. Most of the genes located within QTL on chromosome 5R were involved in building the proper chromosome structure at the beginning of meiosis, transition from leptotene to zygotene and homologous chromosome pairing (Table 5). These genes can potentially be important for maintaining the proper functioning of the plant genome despite the ongoing stress associated with powdery mildew infection and defense processes. No QTL for powdery mildew has been detected on the 5R rye chromosome to date so it might be reported as a new source of resistance. To make this effect stronger, the existence of Pm4 gene on this chromosome was confirmed as well as a genes controlling resistance to leaf rust (Baranova et al. 2002; Tyrka and Chelkowski 2004).
Two regions on 6R chromosome were detected for AUDCP and avPM in Choryń in a 2-years period (2014 and 2015). Those loci were in a different position on this chromosome and explained up to 11.1% of phenotypic variation for Qpm.gz.6R.2 and LOD value 3.4 for Qpm.gz.6R.1. Also, for Qpm.gz.6R.1, gene encoded cyclin-dependent kinase A-2-like (LOC119314733) protein was identified (Table 5). The Pm20 gene has been identified and derived from 6RL of Prolific rye (Zhuang 2003; An et al. 2015) that may correspond to one QTL region on the 6R rye chromosome associated with powdery mildew resistance, while the second locus on this chromosome is new.
In conclusion, availability of the winter triticale DH population allowed to create a new, high-density genetic map for this crop specie. Based on this map, a total of 23 QTL regions were identified based on a 3-year field experiment on triticale resistance to powdery mildew infection conducted in three different locations across the Poland. Among those regions, two found on rye chromosome 4R and single loci on 5R and 6R were reported for the first time as regions associated with powdery mildew resistance. The information of significant QTL regions associated with powdery mildew resistance together with candidate gene–coded proteins taking part in triticale defense against fungal pathogen can be an important tool used in modern breeding programs. Molecular markers against Blumeria graminis after careful validation in available triticale varieties can be used for pyramiding two or more than two APR genes or QTLs from donor to recipient parent. To assist molecular breeding programs, described in this paper, regions associated with PM resistance can be used in marker-assisted selection (MAS) as well as in marker-assisted recurrent selection (MARS) and genomic selection (GS).
References
Alam A, Xue F, Ali M, Wang C, Ji W (2013) Identification and molecular mapping of powdery mildew resistance gene Pmg25 in common wheat originated from wild emmer (Triticum turgidum var. dicoccoides). Pak J Botany 45(1):203–208
Alam M, Neal J, O’Connor K, Kilian A, Topp B (2018) Ultra-high-throughput DArTseq-based silicoDArT and SNP markers for genomic studies in macadamia. PLoS ONE 13(8):e0203465. https://doi.org/10.1371/journal.pone.0203465
Alheit KV, Reif JC, Maurer HP, Hahn V, Weissmann EA, Miedaner T, Würschum T (2011) Detection of segregation distortion loci in triticale (xTriticosecale Wittmack) based on a high-density DArT marker consensus genetic linkage map. BMC Genom 12:380. https://doi.org/10.1186/1471-2164-12-380
Ammar K, Mergoum M, Rajaram S (2004) The history and evolution in triticale. In: Mergoum M, Gomez-Macpherson H (eds) Triticale improvement and production, 1st edn. FAO, Rome, pp 1–10
An D, Zheng Q, Zhou Y, Ma P, Lv Z, Li L, Li B, Luo Q, Xu H, Xu Y (2013) Molecular cytogenetic characterization of a new wheat–rye 4R chromosome translocation line resistant to powdery mildew. Chrom Res 21(4):419–432. https://doi.org/10.1007/s10577-013-9366-8
An D, Zheng Q, Luo Q, Ma P, Zhang H, Li L, Han F, Xu H, Xu Y, Zhang X, Zhou Y (2015) Molecular cytogenetic identification of a new wheat-rye 6R chromosome disomic addition line with powdery mildew resistance. PLoS ONE 10(8):e0134534. https://doi.org/10.1371/journal.pone.0134534
Asad MA, Bai B, Lan C, Yan J, Xia X, He Z (2014) Identification of QTL for adult-plant resistance to powdery mildew in Chinese wheat landrace Pingyuan 50. Crop J 2(5):308–314. https://doi.org/10.1016/j.cj.2014.04.009
Ballesta P, Bush D, Silva FF, Mora F (2020) Genomic predictions using low-density SNP markers, pedigree and GWAS information: a case study with the non-model species Eucalyptus cladocalyx. Plants 9(1):99. https://doi.org/10.3390/plants9010099
Baloch FS, Alsaleh A, Shahid MQ, Çiftçi V, de Miera LES, Aasim M, Nadeem MA, Aktas H, Özkan H, Hatipoğlu R (2017) A whole genome DArTseq and SNP analysis for genetic diversity assessment in durum wheat from central fertile crescent. PLoS ONE 12(1):e0167821. https://doi.org/10.1371/journal.pone.0167821
Baranova OA, Dmitriev AP, Voylocov AV, Solodukhina OV (2002) Mapping of resistance genes to brown rust in 1R chromosome of rye (Secale cereale L). Plant Protect Sci Prague 38:588–589. https://doi.org/10.17221/10562-PPS
Bauer E, Schmutzer T, Barilar I, Mascher M, Gundlach H, Martis MM, Twardziok SO, Hackauf B, Gordillo A, Wilde P, Schmidt M, Korzun V, Mayer KF, Schmid K, Schön CC, Scholz U (2017) Towards a whole-genome sequence for rye (Secale cereale L.). Plant J 89:853–869. https://doi.org/10.1111/tpj.13436
Bolibok-Brągoszewska H, Heller-Uszyńska K, Wenzl P, Uszyński G, Kilian A, Rakoczy-Trojanowska M (2009) DArT markers for the rye genome-genetic diversity and mapping. BMC Genomics 10(1):1–11. https://doi.org/10.1186/1471-2164-10-578
Carter AH, Chen XM, Garland-Campbell K, Kidwell KK (2009) Identifying QTL for high-temperature adult-plant resistance to stripe rust (Puccinia striiformis f. sp. tritici) in the spring wheat (Triticum aestivum L.) cultivar ‘Louise.’ Theor Appl Genet 119(6):1119–1128. https://doi.org/10.1007/s00122-009-1114-2
Buerstmayr H, Lemmens M, Hartl L, Doldi L, Steiner B, Stierschneider M, Ruckenbauer P (2002) Molecular mapping of QTLs for Fusarium head blight resistance in spring wheat. I. Resistance to fungal spread (Type II resistance). Theor Appl Genet 104(1):84–91. https://doi.org/10.1007/s001220200009
Buerstmayr H, Steiner B, Hartl L, Griesser M, Angerer N, Lengauer D, Miedaner T, Schneider B, Lemmens M (2003) Molecular mapping of QTLs for Fusarium head blight resistance in spring wheat II Resistance to fungal penetration and spread. Theor Appl Genet 107(3):503–508. https://doi.org/10.1007/s00122-003-1272-6
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, Forrest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D, Bai G, Pumphrey M, Tomar L, Wong D, Kong S, Reynolds M, Lopez da Silva M, Bockelman H, Talbert L, Anderson JA, Dreisigacker S, Baenziger S, Carter A, Korzun V, Morrell PL, Dubcovsky J, Morell MK, Sorrells ME, Hayden MJ, Akhunov E (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. PNAS 110(20):8057–8062. https://doi.org/10.1073/pnas.1217133110
Chantret N, Mingeot D, Sourdille P, Bernard M, Jacquemin JM, Doussinault G (2001) A major QTL for powdery mildew resistance is stable over time and at two development stages in winter wheat. Theor Appl Genet 103(6–7):962–971. https://doi.org/10.1007/s001220100645
Clinesmith MA, Fritz AK, Lemes da Silva C, Bockus WW, Poland JA, Dowell FE, Peiris KH (2019) QTL mapping of fusarium head blight resistance in winter wheat cultivars ‘art’ and ‘everest.’ Crop Sci 59(3):911–924. https://doi.org/10.2135/cropsci2018.04.0276
Crespo-Herrera LA, Garkava-Gustavsson L, Åhman I (2017) A systematic review of rye (Secale cereale L.) as a source of resistance to pathogens and pests in wheat (Triticum aestivum L.). Hereditas 154(1):14. https://doi.org/10.1186/s41065-017-0033-5
Faostat (2020) Food and Agriculture Organization of the United Nations. http://www.fao.org/ (28.12.2020).
Duan Q, Wang YY, Qiu L, Ren TH, Li Z, Fu SL, Tang ZX (2017) Physical location of new PCR-based markers and powdery mildew resistance gene(s) on rye (Secale cereale L.) chromosome 4 using 4R dissection lines. Frontiers in Plant Science 8:1716. https://doi.org/10.3389/fpls.2017.01716
Finckh MR, Gacek ES, Czembor HJ, Wolfe MS (1999) Host frequency and density effects on powdery mildew and yield in mixtures of barley cultivars. Plant Pathol 48(6):807–816. https://doi.org/10.1046/j.1365-3059.1999.00398.x
Giancaspro A, Giove SL, Zito D, Blanco A, Gadaleta A (2016) Mapping QTLs for Fusarium head blight resistance in an interspecific wheat population. Front Plant Sci 7:1381. https://doi.org/10.3389/fpls.2016.01381
González JM, Muñiz LM, Jouve N (2005) Mapping of QTLs for androgenetic response based on a molecular genetic map of ×Triticosecale Wittmack. Genome 48(6):999–1009. https://doi.org/10.1139/g05-064
Guo Q, Zhang ZJ, Xu YB, Li GH, Feng J, Zhou Y (2008) Quantitative trait loci for high-temperature adult-plant and slow-rusting resistance to Puccinia striiformis f. sp. tritici in wheat cultivars. Phytopathology 98(7):803–809. https://doi.org/10.1094/PHYTO-98-7-0803
Guo J, Zhao Z, Song J, Liu C, Zhai S, Li H, Liu A, Cheng D, Han R, Liu J, Kong L, Zhao Z, Kong L (2017) Molecular and physical mapping of powdery mildew resistance genes and QTLs in wheat: a review. Agricult Sci Technol 18(6):965
Hoagland DR (1948) Lectures on the inorganic nutrition of plants. Chronica Botanica Co., Waltham
Holtz Y, Ardisson M, Ranwez V, Besnard A, Leroy P, Poux G, Roumet P, Viader V, Santoni S, David J (2016) Genotyping by sequencing using specific allelic capture to build a high-density genetic map of Durum wheat. PLoS ONE 11(5):e0154609. https://doi.org/10.1371/journal.pone.0154609
Huang XQ, Röder MS (2004) Molecular mapping of powdery mildew resistance genes in wheat: a review. Euphytica 137(2):203–223. https://doi.org/10.1023/B:EUPH.0000041576.74566.d7
Jaccoud D, Peng K, Feinstein D, Kilian A (2001) Diversity arrays: a solid state technology for sequence information independent genotyping. Nucleic Acids Res 29:e25. https://doi.org/10.1093/nar/29.4.e25
Jakobson I, Reis D, Tiidema A, Peusha H, Timofejeva L, Valárik M, Kladivová M, Šimková H, Doležel J, Järve K (2012) Fine mapping, phenotypic characterization and validation of non-race-specific resistance to powdery mildew in a wheat - Triticum militinae introgression line. Theor Appl Genet 125(3):609–623. https://doi.org/10.1007/s00122-012-1856-0
Jeger MJ, Viljanen-Rollinson SLH (2001) The use of the area under the disease-progress curve (AUDPC) to assess quantitative disease resistance in crop cultivars. Theor Appl Genet 102(1):32–40. https://doi.org/10.1007/s001220051615
Jordan KW, Wang S, Lun Y, Gardiner LJ, MacLachlan R, Hucl P, Wiebe K, Wong D, Forrest KL, Sharpe AG, Sidebottom CHD, Hall N, Toomajian C, Close T, Dubcovsky J, Akhunova A, Talbert L, Bansal UK, Bariana HS, Hayden MJ, Pozniak C, Jeddeloh JA, Hall A (2015) Akhunov E (2015) A haplotype map of allohexaploid wheat reveals distinct patterns of selection on homoeologous genomes. Genome Biol 16:48. https://doi.org/10.1186/s13059-015-0606-4
Karbarz M, Tyrka M, Woźniak-Strzembicka A, Czajowski G, Wardyńska A, Tyrka D, Pojmaj M, Wędzony M (2020) Quantitative trait loci mapping of adult-plant resistance to powdery mildew in triticale. Ann Appl Biol 177(2):223–231. https://doi.org/10.1111/aab.12613
Keller M, Keller B, Schachermayr G, Winzeler M, Schmid JE, Stamp P, Messmer MM (1999) Quantitative trait loci for resistance against powdery mildew in a segregating wheat×spelt population. Theor Appl Genet 98(6–7):903–912. https://doi.org/10.1007/s001220051149
Klocke B, Flath K, Miedaner T (2013) Virulence phenotypes in powdery mildew (Blumeria graminis) population and resistance genes in triticale (xTriticosecale). Eur J Plant Pathol 137(3):463–476. https://doi.org/10.1007/s10658-013-0257-9
Korzun V, Malyshev S, Voylokov AV, Börner A (2001) A genetic map of rye (Secale cereale L.) combining RFLP, isozyme, protein, microsatellite and gene loci. Theor Appl Genet 102(5):709–717. https://doi.org/10.1007/s001220051701
Lan C, Ni X, Yan J, Zhang Y, Xia X, Chen X, He Z (2010) Quantitative trait loci mapping of adult-plant resistance to powdery mildew in Chinese wheat cultivar Lumai 21. Mol Breeding 25(4):615–622. https://doi.org/10.1007/s11032-009-9358-8
Landjeva S, Korzun V, Tsanev V, Vladova R, Ganeva G (2006) Distribution of the wheat-rye translocation 1RS.1BL among bread wheat varieties of Bulgaria. Plant Breeding 125:102–104. https://doi.org/10.1111/j.1439-0523.2006.01142.x
Li H, Bariana H, Singh D, Zhang L, Dillon S, Whan A, Urmil B, Ayliffe M (2020) A durum wheat adult plant stripe rust resistance QTL and its relationship with the bread wheat Yr80 locus. Theor Appl Genet 133(11):3049–3066. https://doi.org/10.1007/s00122-020-03654-5
Liu Z, Wang Q, Wan H, Yang F, Wei H, Xu Z, Ji H, Xia X, Li J, Yang W (2020) QTL mapping for adult-plant resistance to powdery mildew in Chinese elite common wheat Chuanmai104. Cereal Research Communications 49:99–108. https://doi.org/10.1007/s42976-020-00082-5
Ma P, Han G, Zheng Q, Liu S, Han F, Wang J, Luo Q, An D (2020) Development of novel wheat-rye chromosome 4R translocations and assignment of their powdery mildew resistance. Plant Dis 104(1):260–268. https://doi.org/10.1094/PDIS-01-19-0160-RE
Mantovani P, Maccaferri M, Sanguineti MC, Tuberosa R, Catizone I, Wenzl P, Thomson B, Carling J, Huttner E, DeAmbrogio E, Kilian A (2008) An integrated DArT-SSR linkage map of durum wheat. Mol Breeding 22(4):629–648. https://doi.org/10.1007/s11032-008-9205-3
Marone D, Panio G, Ficco DBM, Russo MA, De Vita P, Papa R, Rubiales D, Cattivelli L, Mastrangelo AM (2012) Characterization of wheat DArT markers:genetic and functional features. Mol Genet Genomics 287:741–753. https://doi.org/10.1007/s00438-012-0714-8
Marone D, Russo MA, Laidò G, De Vita P, Papa R, Blanco A, Gadaleta A, Rubiales D, Mastrangelo AM (2013) Genetic basis of qualitative and quantitative resistance to powdery mildew in wheat: from consensus regions to candidate genes. BMC Genomics 14(1):1–17. https://doi.org/10.1186/1471-2164-14-562
McNeal FH, Konzak CF, Smith EP, Täte WS, Russell TS (1971) A uniform system for recording and processing cereal research data. Agricultural research service bulletin. Department of Agriculture, Washington, pp 34–121
Melichar JPE, Berry S, Newell C, MacCormack R, Boyd LA (2008) QTL identification and microphenotype characterisation of the developmentally regulated yellow rust resistance in the UK wheat cultivar Guardian. Theor Appl Genet 117(3):391–399. https://doi.org/10.1007/s00122-008-0783-6
Menardo F, Praz CR, Wyder S, Ben-David R, Bourras S, Matsumae H, McNally KE, Parlange F, Riba A, Roffler S, Schaefer LK, Shimizu KK, Valenti L, Zbinden H, Wicker T, Keller B (2016) Hybridization of powdery mildew strains gives rise to pathogens on novel agricultural crop species. Nat Genet 48(2):201–205. https://doi.org/10.1038/ng.3485
Milczarski P, Bolibok-Bragoszewska H, Myśków B, Stojałowski S, Heller-Uszyńska K, Góralska M, Brągoszewski P, Uszyński G, Kilian A (2011) Rakoczy-Trojanowska M (2011) A high density consensus map of rye (Secale cereale L.) based on DArT markers. PLoS One. 6(12):1–14. https://doi.org/10.1371/journal.pone.0028495
Mingeot D, Chantret N, Baret PV, Dekeyser A, Boukhatem N, Sourdille P, Doussinault G, Jacquemin JM (2002) Mapping QTL involved in adult plant resistance to powdery mildew in the winter wheat line RE714 in two susceptible genetic backgrounds. Plant Breeding 121(2):133–140. https://doi.org/10.1046/j.1439-0523.2002.00679.x
Neu C, Stein N, Keller B (2002) Genetic mapping of theLr20–Pm1resistancelocus reveals suppressed recombination on chromosome arm 7AL in hexaploid wheat. Genome 45:737–744. https://doi.org/10.1139/g02-040
Nordestgaard NV, Thach T, Sarup P, Rodriguez-Algaba J, Andersen JR, Hovmøller MS, Jahoor A, Jørgensen LN, Orabi J (2020) Multi-parental populations suitable for identifying sources of resistance to powdery mildew in winter wheat. Front Plant Sci 11:2103. https://doi.org/10.3389/fpls.2020.570863
Nsabiyera V, Baranwal D, Qureshi N, Kay P, Forrest K, Valárik M, Dolezel J, Hayden MJ, Bariana HS, Bansal UK (2020) Fine mapping of Lr49 using 90K SNP chip array and flow-sorted chromosome sequencing in wheat. Front Plant Sci 10:1787. https://doi.org/10.3389/fpls.2019.01787
Prins R, Pretorius ZA, Bender CM, Lehmensiek A (2011) QTL mapping of stripe, leaf and stem rust resistance genes in a Kariega×Avocet S doubled haploid wheat population. Mol Breeding 27(2):259–270. https://doi.org/10.1007/s11032-010-9428-y
Rabanus-Wallace MT, Hackauf B, Mascher M et al (2021) Chromosome-scale genome assembly provides insights into rye biology, evolution and agronomic potential. Nat Genet 53:564–573. https://doi.org/10.1038/s41588-021-00807-0
Reader SM, Miller TE (1991) The introduction into bread wheat of a major gene for resistance to powdery mildew from wild emmer wheat. Euphytica 53(1):57–60. https://doi.org/10.1007/BF00032033
Ren Y, Hou W, Lan C, Basnet BR, Singh RP, Zhu W, Cheng X, Cui D, Chen F (2017) QTL analysis and nested association mapping for adult plant resistance to powdery mildew in two bread wheat populations. Front Plant Sci 8:1212. https://doi.org/10.3389/fpls.2017.01212
Rosewarne GM, Singh RP, Huerta-Espino J, Herrera-Foessel SA, Forrest KL, Hayden MJ, Rebetzke GJ (2012) Analysis of leaf and stripe rust severities reveals pathotype changes and multiple minor QTLs associated with resistance in an Avocet× Pastor wheat population. Theor Appl Genet 124(7):1283–1294. https://doi.org/10.1007/s00122-012-1786-x
Riaz A, Hathorn A, Dinglasan E, Ziems L, Richard C, Singh D, Mitrofanova O, Afanasenko O, Aitken E, Godwin I, Hickey L (2016) Into the vault of the Vavilov wheats: old diversity for new alleles. Genet Resour Crop Evol 64:531–544. https://doi.org/10.1007/s10722-016-0380-5
Schlegel R, Korzun V (2021) Genes, markers and linkage data of rye (Secale cereale L.), 11th updated inventory, V. 01.21, pp. 1–115. http://www.rye-gene-map.de
Schlegel R, Meinel A (1994) A quantitative trait locus (QTL) on chromosome arm 1RS of rye and its effect on yield performance of hexaploid wheat. Cereal research communications, 7–13. https://www.jstor.org/stable/23783805
Shaner G, Finney RE (1977) The effect of nitrogen fertilization on the expression of slow-mildewing resistance in Knox wheat. Phytopathology 67(8):1051–1056
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L). Theor Appl Genet 109(6):1105–1114. https://doi.org/10.1007/s00122-004-1740-7
The International Wheat Genome Sequencing Consortium (IWGSC) (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
The International Wheat Genome Sequencing Consortium (IWGSC) (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 2018(361):661
Tyrka M, Chełkowski J (2004) Enhancing the resistance of triticale by using genes from wheat and rye. J Appl Genet 45:283–296
Tyrka M, Bednarek PT, Kilian A, Wędzony M, Hura T, Bauer E (2011) Genetic map of triticale compiling DArT. SSR and AFLP Markers Genome 54(5):391–401. https://doi.org/10.1139/g11-009
Tyrka M, Tyrka D, Wędzony M (2015) Genetic map of triticale integrating microsatellite, DArT and SNP Markers. Plos ONE 10(12):1–17. https://doi.org/10.1371/journal.pone.0145714
Tyrka M, Oleszczuk S, Rabiza-Swider J, Woś H, Wędzony M, Zimny J, Ponitka A, Ślusarkiewicz-Jarzina A, Metzger RJ, Baenziger PS, Lukaszewski AJ (2018) Populations of doubled haploids for genetic mapping in hexaploid winter triticale. Mol Breeding 38(4):1–15. https://doi.org/10.1007/s11032-018-0804-3
Van Ooijen JW (2006) JoinMap ® 4. Software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wageningen, Netherlands 2006.
Van Os H, Stam P, Visser RGF, Van Eck HJ (2005) RECORD: a novel method for ordering loci on a genetic linkage map. Theor Appl Genet 112:30–40. https://doi.org/10.1007/s00122-005-0097-x
Vinod KK (2009) Genetic mapping of quantitative trait loci and marker assisted selection in plantation crops. Vitro Techniques in Plantation Crops. pp 111–132
Vikram P, Franco J, Burgueño-Ferreira J, Li H, Sehgal D, Saint Pierre C, Ortiz C, Sneller C, Tattaris M, Guzman C, Sansaloni CP, Fuentes-Davila G, Reynolds M, Sonders K, Singh P, Payne T, Wenzl P, Sharma A, Bains NS, Singh GP, Crossa J, Singh S (2016) Unlocking the genetic diversity of Creole wheats. Sci Rep 6:1–13. https://doi.org/10.1038/srep23092
Von Thaden A, Nowak C, Tiesmeyer A, Reiners TE, Alves PC, Lyons LA, Mattucci F, Randi E, Cragnolini M, Galian J, Hrgyeli Z, Kitchener AC, Lambinet C, Lucas JM, Molich T, Ramos L, Schockert V, Cocchiararo B (2020) Applying genomic data in wildlife monitoring: development guidelines for genotyping degraded samples with reduced single nucleotide polymorphism panels. Mol Ecol Resour 20(3):662–680. https://doi.org/10.1111/1755-0998.13136
Walker AS, Bouguennec A, Confais J, Morgant G, Leroux P (2011) Evidence of host-range expansion from new powdery mildew (Blumeria graminis) infection of triticale (xTriticosecale) in France. Plant Pathol 60(2):207–220. https://doi.org/10.1111/j.1365-3059.2010.02379.x
Wang J, Li H, Zhang L, Meng L (2016) Users’ manual of QTL IciMapping. The Quantitative Genetics Group, Institute of Crop Science, Chinese Academy of Agricultural Sciences (CAAS), Beijing 100081, China, and Genetic Resources Program, International Maize and Wheat Improvement Center (CIMMYT), Apdo. Postal 6–641, 06600 Mexico, D.F., Mexico.
Wang S, Basten CJ, Zeng BZ (2012) Windows QTL Cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh
Wąsek I, Dyda M, Gołębiowska G, Tyrka M, Rapacz M, Szechyńska-Hebda M, Wędzony M (2021) Quantitative trait loci and candidate genes associated with freezing tolerance of winter triticale (× Triticosecale Wittmack). J Appl Genet. https://doi.org/10.1007/s13353-021-00660-1
Wenzl P, Carling J, Kudrna D, Jaccoud D, Huttner E, Kleinhofs A, Kilian A (2004) Diversity Arrays Technology (DArT) for whole-genome profiling of barley. Proc Natl Acad Sci 101(26):9915–9920. https://doi.org/10.1073/pnas.0401076101
Wędzony M (2003) Protocol for anther culture in hexaploid triticale (xTriticosecale Wittm.). In: Maluszynski M, Kasha KJ, Forster BP, Szarejko I (eds) Doubled haploid production in crop plants a manual. Kluwer Academic Publishers, Dordrecht/Boston/London, pp 123–128
Wingen LU, West C, Waite ML, Collier S, Orford S, Goram R, Yang CY, King J, Allen AM, Burridge A, Edwards KJ, Griffiths S (2017) Wheat landrace genome diversity. Genetics 205:1657–1676. https://doi.org/10.1534/genetics.116.194688
Xu X, Zhu Z, Jia A, Wang F, Wang J, Zhang Y, Fu C, Fu L, Bai G, Xia X, Hao Y, He Z (2020) Mapping of QTL for partial resistance to powdery mildew in two Chinese common wheat cultivars. Euphytica 216(1):1–12. https://doi.org/10.1007/s10681-019-2537-8
Yang H, Li C, Lam HM, Clements J, Yan G, Zhao S (2015) Sequencing consolidates molecular markers with plant breeding practice. Theor Appl Genet 128(5):779–795. https://doi.org/10.1007/s00122-015-2499-8
Yang L, Zhang X, Zhang X, Wang J, Luo M, Yang M, Wang H, Xiang L, Zeng F, Yu D, Fu D, Rosewarne M (2017) Identification and evaluation of resistance to powdery mildew and yellow rust in a wheat mapping population. PLoS ONE 12(5):e0177905. https://doi.org/10.1371/journal.pone.0177905
Zhan H, Li G, Zhang X, Li X, Guo H (2014) Chromosomal location and comparative genomics analysis of powdery mildew resistance gene Pm51 in a putative wheat-Thinopyrum pontificum introgression line. PLoS ONE 9(11):e113455. https://doi.org/10.1371/journal.pone.0113455
Zhuang QS (2003) Chinese wheat improvement and pedigree analysis. China Agriculture Press, Beijing
Ziems LA, Hickey LT, Hunt CH, Mace ES, Platz GJ, Franckowiak JD, Jordan DR (2014) Association mapping of resistance to Puccinia hordei in Australian barley breeding germplasm. Theor Appl Genet 127:1199–1212. https://doi.org/10.1007/s00122-014-2291-1
Acknowledgements
We thank Małgorzata Niewińska from DANKO Plant Breeders Ltd for taking care of triticale phenotyping and Elżbieta Golemiec for “Grenado” × “Zorro” population preparation.
Funding
This research was supported by the National Centre for Research and Development, Project GENMARK: PBS1/A8/1/2012.
Author information
Authors and Affiliations
Contributions
MD—writing the manuscript, QTL and statistical analysis and performing the experiments; MT—genetic map construction and QTL analysis, writing the manuscript; GG—statistical and candidate genes analysis, writing the manuscript; MR—planning the experiments, obtaining the funding, text consulting; MW—planning and supervision on the experiments, obtaining the funding and text consulting. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by: Izabela Pawłowicz
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dyda, M., Tyrka, M., Gołębiowska, G. et al. Genetic mapping of adult-plant resistance genes to powdery mildew in triticale. J Appl Genetics 63, 73–86 (2022). https://doi.org/10.1007/s13353-021-00664-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-021-00664-x