Abstract
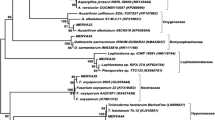
Sponges are well documented to harbor large amounts of microbes. Though it is known that sponge-derived fungi are important sources for marine natural products, the phylogenetic diversity and biological function of sponge-associated fungi remain largely unknown. In this study, the diversity of culturable endozoic fungi in sponges from the South China Sea was revealed based on the ITS phylogenetic analysis. Meanwhile the fungal potential for producing bioactive natural products was estimated according to the detection of Beta-ketosynthase in the polyketide synthase (PKS) gene cluster and cytotoxic activity bioassay. As a result, diverse fungi including 14 genera (Aspergillus, Penicillium, Scolecobasidium, Eurotium, Alternaria, Fusarium, Hypocreales, Yarrowia, Candida, Hypoxylon, Sporidiobolus, Schizophyllum, Bjerkandera, and Trichosporon) in ten orders (Xylariales, Moniliales, Pleosporales, Saccharomycetales, Hypocreales, Eurotiales, Sporidiobolales, Agaricales, Aphyllophorales and Tremellales) of phyla Ascomycota and Basidiomycota were isolated with Aspergillus as the predominant component in the culturable fungal community. Particularly, genera Schizophyllum, Sporidiobolus, and Bjerkandera in phylum Basidiomycota and genus Yarrowia in phylum Ascomycota were isolated from marine sponges for the first time. PKS genes were detected in 12 isolates suggesting their potential for synthesizing PKS compounds. Among the 12 isolates with PKS genes, 9 isolates displayed strong in vitro cytotoxic activity (e.g. IC50 < 50 μg/ml) against human cancer cell lines A-549, Bel-7402, A-375 and MRC-5. This study demonstrates the phylogenetically diverse endozoic fungi in South China Sea sponges, and highlights the potential of sponge-associated fungi in producing biologically active natural products.


Similar content being viewed by others
References
Abdel-Lateffa A, Fischa K, Wrightd AD (2009) Trichopyrone and other constituents from the marine sponge-derived fungus Trichoderma sp. Z Naturforsch B 64:186–192
Abrell LM, Borgeson B, Crews P (1996) Chloro polyketides from the cultured fungus (Aspergillus) separated from a marine sponge. Tetrahedron Lett 37:2331–2334
Amagata T, Morinaka BI, Amagata A, Tenney K, Valeriote FA, Lobkovsky E, Clardy J, Crews PA (2006a) Chemical study of cyclic depsipeptides produced by a sponge-derived fungus. J Nat Prod 69:1560–1565
Amagata T, Minoura K, Numata A (2006b) Gymnastatins F − H, cytostatic metabolites from the sponge-derived fungus Gymnascella dankaliensis. J Nat Prod 69:1384–1388
Amagata T, Amagata A, Tenney K, Valeriote FA, Lobkovsky E, Clardy J, Crews P (2003) Unusual C25 steroids produced by a sponge-derived Penicillium citrinum. Org Lett 5:4393–4396
Amnuaykanjanasin A, Punya J, Paungmoung P, Rungrod A, Tachaleat A, Pongpattanakitshote S, Cheevadhanarak S, Tanti-charoen M (2005) Diversity of type I polyketide synthase genes in the wood-decay fungus Xylaria sp. BCC 1067. FEMS Microbiol Lett 251:125–136
Baker PW, Kennedy J, Dobson ADW, Marchesi JR (2009) Phylogenetic diversity and antimicrobial activities of fungi associated with Haliclona simulans isolated from Irish coastal waters. Mar Biotechnol 11:540–547
Barghoorn ES, Linder DH (1944) Marine fungi: their taxonomy and biology. Farlowia 1:395–467
Bass D, Howe A, Brown N, Barton H, Demidova M, Harlan Michelle H, Li L, Sanders H, Watkinson SC, Willcock S, Richards TA (2007) Yeast forms dominate fungal diversity in the deep oceans. Proc Biol Sci 274:3069–3077
Bhadury P, Mohammad BT, Wright PC (2006) The current status of natural products from marine fungi and their potential as anti infective agents. J Ind Microbiol Biotech 33:325–337
Bingle LEH, Simpson TJ, Lazarus CM (1999) Ketosynthase domain probes identify two subclasses of fungal polyketide synthase genes. Fungal Gen Bio 26:209–223
Blunt JW, Copp BR, Hu WP, Munro MHG, Northcote PT, Prinsep MR (2009) Marine natural products. Nat Prod Rep 26:170–244
Boot CM, Tenney K, Valeriote FA, Crews P (2006) Highly N-methylated linear peptides produced by an atypical sponge-derived Acremonium sp. J Nat Prod 69:83–92
Brauers G, Ebel R, Edrada RA, Wray V, Berg A, Gräfe U, Proksch P (2001) Hortein, a new natural product from the fungus Hortaea werneckii associated with the sponge Aplysina aerophoba. J Nat Prod 64:651–652
Brauers G, Edrada RA, Ebel R, Proksch P, Vr W, Berg A, Gräfe U, Schächtele C, Totzke F, Finkenzeller G, Marme D, Kraus J, Münchbach M, Michel M, Bringmann G, Schaumann K (2000) Anthraquinones and betaenone derivatives from the sponge-associated fungus Microsphaeropsis species: novel inhibitors of protein kinases. J Nat Prod 63:739–745
Bringmann G, Lang G, Steffens S, Schaumann K (2004) Petrosifungins A and B, novel cyclodepsipeptides from a sponge-derived strain of Penicillium brevicompactum. J Nat Prod 67:311–315
Bringmann G, Lang G, Steffens S, Günther E, Schaumann K (2003) Evariquinone, isoemericellin, and stromemycin from a sponge derived strain of the fungus Emericella variecolor. Phytochem 63:4437–4443
Bugni TS, Ireland CM (2004) Marine-derived fungi: a chemically and biologically diverse group of microorganisms. Nat Prod Rep 21:143–163
Debbab A, Aly AH, Proksch P (2011) Bioactive secondary metabolites from endophytes and associated marine derived fungi. Fungal Divers 49:1–12
Ding B, Yin Y, Zhang F, Li Z (2011) Recovery and phylogenetic diversity of culturable fungi associated with marine sponges Clathrina luteoc ulcitella and Holoxea sp. in the South China Sea. Mar Biotechnol 13:713–721
Edgcomb VP, Beaudoin D, Gast R, Biddle JF, Teske A (2011) Marine subsurface eukaryotes: the fungal majority. Environ Microbiol 13:172–183
Edrada RA, Heubes M, Brauers G, Wray V, Berg A, Gräfe U, Wohlfarth M, Mühlbacher J, Schaumann K, Sudarsono BG, Proksch P (2002) Online analysis of xestodecalactones A − C, novel bioactive metabolites from the fungus Penicillium cf. montanense and their subsequent isolation from the sponge Xestospongia exigua. J Nat Prod 65:1598–1604
Ein-Gil N, Ilan M, Carmel S, Smith GW, Pawlik JR, Yarden O (2009) Presence of Aspergillus sydowii, a pathogen of gorgonian sea fans in the marine sponge Spongia obscura. ISME J 3:752–755
Fieseler L, Horn M, Wagner M, Hentschel U (2004) Discovery of the novel candidate phylum “Poribacteria” in marine sponges. Appl Environ Microbiol 70:3724–3732
Gao Z, Li B, Zheng C, Wang G (2008) Molecular detection of fungal communities in the Hawaiian marine sponges Suberites zeteki and Mycale armata. Appl Environ Microbiol 74:6091–6101
Gesner S, Cohen N, Ilan M, Yarden O, Carmeli S (2005) Pandangolide 1a, a metabolite of the sponge-associated fungus Cladosporium sp., and the absolute stereochemistry of pandangolide 1 and iso-Cladospolide B. J Nat Prod 68:1350–1353
Hentschel U, Usher KM, Taylor MW (2006) Marine sponges as microbial fermenters. FEMS Microbiol Ecol 55:167–177
Hendrickson L, Davis CR, Roach C, Nguyen DK, Aldrich T, McAda PC, Reeves CD (1999) Lovastatin biosynthesis in Aspergillus terreus: Characterization of blocked mutants, enzyme activities and a multifunctional polyketide synthase gene. Chem Biol 6:429–439
Hiort J, Maksimenka K, Perović-Ottstadt S, Lin W, Wray V, Steube K, Schaumann K, Weber H, Proksch P, Ebel R, Müller WEG, Bringmann G (2004) New natural products from the sponge-derived fungus Aspergillus niger. J Nat Prod 67:1532–1543
Hochmuth T, Piel J (2009) Polyketide synthases of bacterial symbionts in sponges-evolution-based applications in natural products research. Phytochem 70:1841–1849
Höller U, Wright AD, Matthee GF, König GM, Draeger S, Aust HJ, Schulz B (2000) Fungi from marine sponges: diversity, biological activity and secondary metabolites. Mycol Res 104:1354–1365
Ingavat N, Dobereiner J, Wiyakrutta S, Mahidol C, Ruchirawat S, Kittakoop P (2009) Aspergillusol A, an α-glucosidase inhibitor from the marine-derived fungus Aspergillus aculeatus. J Nat Prod 72:2049–2052
Jadulco R, Edrada RA, Ebel R, Berg A, Schaumann K, Wray V, Steube K, Proksch P (2004) New communesin derivatives from the fungus Penicillium sp. derived from the Mediterranean sponge Axinella verrucosa. J Nat Prod 67:78–81
Jadulco R, Brauers G, Edrada RA, Ebel R, Wray V, Sudarsono PP (2002) New metabolites from sponge-derived fungi Curvularia lunata and Cladosporium herbarum. J Nat Prod 65:730–733
Jadulco R, Proksch P, Wray V, Sudarsono BA, Gräfe U (2001) New macrolides and furan carboxylic acid derivative from the sponge-derived fungus Cladosporium herbarum. J Nat Prod 64:527–530
Jiang S, Sun W, Chen M, Dai S, Zhang L, Liu Y, Lee K, Li X (2007) Diversity of culturable actinobacteria isolated from marine sponge Haliclona sp. Antonie Leeuwenhoek 92:405–416
Jones EBG (2011) Fifty years of marine mycology. Fungal Divers 50:73–112
Kim T, Fuerst J (2006) Diversity of polyketide synthase genes from bacteria associated with the marine sponge Pseudoceratina clavata: culture-dependent and culture-independent approaches. Environ Microbiol 8:1460–1470
Kobayashi M, Uehara H, Matsunami K, Aoki S, Kitagawa I (1993) Trichoharzin, a new polyketide produced by the imperfect fungus Trichoderma harzianum separated from the marine spongeMicale Cecilia. Tetrahedron Lett 49:7925–7928
König GM, Kehraus S, Seiber SF, Abdel-Lateff A, Müller D (2006) Natural products from marine organisms and their associated microbes. Chem Biochem 7:229–238
Lang G, Wiese J, Schmaljohann R, Imhoff JF (2007) New pentaenes from the sponge-derived marine fungus Penicillium rugulosum: structure determination and biosynthetic studies. Tetrahedron 63:11844–11849
Lee OO, Wang Y, Yang JK, Lafi F, Al-Suwailem A, Qian PY (2011) Pyrosequencing reveals highly diverse and species-specific microbial communities in sponges from the Red Sea. ISME J 5:650–664
Lee YM, Li H, Hong J, Cho HY, Bae KS, Kim MA, Kim D, Jung JH (2010) Bioactive metabolites from the sponge-derived Fungus AspergillusVersicolor. Arch Pharm Res 33:231–235
Li ZY, Liu Y (2006) Marine sponge Craniella austrialiensis-associated bacterial diversity revelation based on 16 S rDNA library and biologically active actinomycetes screening, phylogenetic analysis. Lett Appl Microbiol 43:410–416
Li Q, Wang G (2009) Diversity of fungal isolates from three Hawaiian marine sponges. Microbiol Res 164:233–241
Lin W, Brauers G, Ebel R, Wray V, Berg A, Sudarsono PP (2003) Novel chromone derivatives from the fungus Aspergillus versicolor isolated from the marine sponge Xestospongia exigua. J Nat Prod 66:57–61
Liu H, Edrada-Ebel R, Ebel R, Wang Y, Schulz B, Draeger S, Müller WEG, Wray V, Lin W, Proksch P (2009) Drimane sesquiterpenoids from the fungus Aspergillus ustus isolated from the marine sponge Suberites domuncula. J Nat Prod 72:1585–1588
Liu WC, Li CQ, Zhu P, Yang JL, Cheng KD (2010) Phylogenetic diversity of culturable fungi associated with two marine sponges: Haliclona simulans and Gelliodes carnosa, collected from the Hainan Island coastal waters of the South China Sea. Fungal Divers 42:1–15
Maldonado M, Cortadellas N, Trillas MI, Ruetzler K (2005) Endosymbiotic yeast maternally transmitted in a marine sponge. Biol Bull 209:94–106
Menezesa CBA, Bonugli-Santosa RC, Miquelettoa PB, Passarinia MRZ, Silvaa CHD, Justoa MR, Leala MR, Fantinatti-Garbogginia F, Oliveiraa VM, Berlinckb RGS, Sette LD (2010) Microbial diversityassociated with algae, ascidians and sponges from the north coast of São Paulostate, Brazil. Microbiol Res 165:466–482
Mohapatra BR, Banerjee UC, Bapuji M (1998) Characterization of a fungal amylase from Mucor sp. associated with the marine sponge Spirastrella sp. J Biotechnol 60:113–117
Müller WEG, Brümmer F, Batel R, Müller IM, Schröder HC (2003) Molecular biodiversity. Case study: Porifera (sponges). Naturwissenschaften 90:103–120
Paz Z, Komon-Zelazowska M, Druzhinina IS, Aveskamp MM, Shnaiderman A, Aluma Y, Carmeli S, Ilan M, Yarden O (2010) Diversity and potential antifungal properties of fungi associated with a Mediterranean sponge. Fungal Divers 42:17–26
Piel J (2004) Metabolites from symbiotic bacteria. Nat Prod Rep 21:519–538
Piel J, Hui D, Fusetani N, Matsunaga S (2004) Targeting modular polyketide synthases with iteratively acting acyltransferases from metagenomes of uncultured bacterial consortia. Environ Microbiol 6:921–927
Proksch P, Edrada-Ebel R-A, Ebel R (2003) Drugs from the sea: opportunities and obstacles. Mar Drugs 1:5–17
Proksch P, Ebel R, Edrada R, Riebe F, Liu H, Diesel A, Bayer M, Li X, Lin WH, Grebenyuk V, Müller WEG, Draeger S, Zuccaro A, Schulz B (2008) Sponge associated fungi and their bioactive compounds - The Suberites case. Bot Mar 51:209–218
Proksch P, Putz A, Ortlepp S, Kjer J, Bayer M (2010) Bioactive natural products from marine sponges and fungal endophytes. Phytochem Rev 9:475–489
Pruesse E, Quast C, Knittel K, Fuchs B, Ludwig W, Peplies J, Glockner F (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequenced at a compatible with ARB. Nucleic Acids Res 35:7188–7196
Raghukumar C (2008) Marine fungal biotechnology: an ecological perspective. Fungal Divers 31:19–35
Rahbæk L, Sperry S, Piper JE, Crews P (1998) Deoxynortrichoharzin, a new polyketide from the saltwater culture of a sponge-derived Paecilomyces Fungus. J Nat Prod 61:1571–1573
Richards TA, Jones M, Leonard G, Bass D (2012) Marine fungi: Their ecology and molecular diversity. Annu Rev Mar Sci 4:495–522
Rot C, Goldfarb I, Ilan M, Huchon D (2006) Putative cross-kingdom horizontal gene transfer in sponge (Porifera) mitochon-dria. BMC Evol Biol 6:71
Saleem M, Ali SM, Hussain S, Jabbar A, Ashraf M, Lee YS (2007) Marine natural products of fungal origin. Nat Prod Rep 24:1142–1152
Schirmer A, Gadkari R, Reeves C, Ibrahim F, Delong E, Hutchinson C (2005) Metagenomic analysis reveals diverse polyketide synthase gene clusters in microorganisms associated with the marine sponge Discodermia dissoluta. Appl Environ Microbiol 71:4840–4849
Schmitt S, Tsai P, Bell J, Fromont J, Ilan M, Lindquist N, Perez T, Rodrigo A, Schupp PJ, Vacelet J, Webster N, Hentschel U, Taylor MW (2012) Assessing the complex sponge microbiota: core, variable and species-specific bacterial communities in marine sponges. ISME J 6:564–576
Siegl A, Hentschel U (2010) PKS and NRPS gene clusters from microbial symbiont cells of marine sponges by whole genome amplification. Environ Microbiol Rep 2:507–513
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Taylor MW, Hill RT, Piel J, Thacker RW, Hentschel U (2007a) Soaking it up: the complex lives of marine sponges and their microbial associates. ISME J 1:187–190
Taylor MW, Radax R, Steger D, Wagner M (2007b) Sponge-associated microorganisms: evolution, ecology, and biotechnological potential. Microbiol Mol Biol Rev 71:295–347
Thirunavukkarasu N, Suryanarayanan T, Girivasan KP, Venkatachalam A, Geetha V, Ravishankar JP, Doble M (2012) Fungal symbionts of marine sponges from Rameswaram, southern India: species composition and bioactive metabolites. Fungal Divers 55:37–46
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL-X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Varoglu M, Crews P (2000) Biosynthetically diverse compounds from a saltwater culture of sponge-derived Aspergillus niger. J Nat Prod 63:41–43
Wang G, Li Q, Zhu P (2008) Phylogenetic diversity of culturable fungi associated with the Hawaiian sponges Suberites zeteki and Gelliodes fibrosa. Antonie Leeuwenhoek 93:163–174
Webster NS, Taylor MW (2012) Marine sponges and their microbial symbionts: love and other relationships. Environ Microbiol 14:335–346
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and application. Academic Press Inc, San Diego, pp 315–322
Wiese J, Ohlendorf B, Blümel M, Schmaljohann R, Imhoff JF (2011) Phylogenetic identification of fungi isolated from the marine sponge Tethya aurantium and identification of their secondary metabolites. Mar Drugs 9:561–585
Xie LW, Jiang SM, Zhu HH, Sun W, Ouyang YC, Dai SK, Li X (2008) Potential inhibitors against Sclerotinia sclerotiorum, produced by the fungus Myrothecium sp. associated with the marine sponge Axinella sp. Eur J Plant Pathol 122:4571–4578
Xin Z, Fang Y, Du L, Zhu T, Duan L, Chen J, Gu Q, Zhu W (2007) Aurantiomides A−C, quinazoline alkaloids from the sponge-derived fungus Penicillium aurantiogriseum SP0-19. J Nat Prod 70:853–855
Yang LH, Miao L, Lee OO, Li X, Xiong H, Pang K, Vrijmoed L, Qian P (2007) Effect of culture conditions on antifouling compound production of a sponge-associated fungus. Appl Microbiol Biotech 74:1221–1231
Yin Y, Gao Q, Zhang F, Li Z (2012) Medium optimization for the high yield production of single (+)-terrein by Aspergillus terreus strain PF26 derived from marine sponge Phakellia fusca. Process Biochem 47:887–891
Zhang P, Bao B, Dang HT, Hong J, Lee HJ, Yoo ES, Yoo ES, Bae KS, Jung JH (2009a) Anti-inflammatory sesquiterpenoids from a sponge-derived fungus Acremonium sp. J Nat Prod 72:270–275
Zhang W, Li Z, Miao X, Zhang F (2009b) The screening of antimicrobial bacteria with diverse novel non-ribosomal peptide synthetase (NRPS) genes from South China Sea sponges. Mar Biotechnol 11:346–355
Zhang W, Zhang FL, Li ZY, Miao XL, Meng QP, Zhang XS (2009c) Investigation of sponge-associated cultivable bacteria with polyketide synthase genes and antimicrobial activity in the South China Sea. J Appl Microbiol 107:567–575
Zhou K, Zhang X, Zhang F, Li Z (2011) Phylogenetically diverse cultivable fungal community and polyketide synthase (PKS), non-ribosomal peptide synthase (NRPS) genes associated with the South China Sea sponges. Microb Ecol 62:644–654
Acknowledgement
This work was supported by the National Natural Science Foundation of China (81102417) and the High-Tech Research and Development Program of China (2011AA09070203)
Author information
Authors and Affiliations
Corresponding author
Additional information
Zhisheng Yu and Baohua Zhang contributed equally to this paper.
Rights and permissions
About this article
Cite this article
Yu, Z., Zhang, B., Sun, W. et al. Phylogenetically diverse endozoic fungi in the South China Sea sponges and their potential in synthesizing bioactive natural products suggested by PKS gene and cytotoxic activity analysis. Fungal Diversity 58, 127–141 (2013). https://doi.org/10.1007/s13225-012-0192-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-012-0192-7