Abstract
Rice black-streaked dwarf disease (RBSDD) is one of the most serious crop diseases in Asia, causing serious damage to rice production. Therefore, reducing the harmful effects of RBSDD is vital to the food security of China and other Asian countries. In this study, 248 rice varieties from different countries were screened for resistance to RBSDD, and 19 varieties with high resistance to RBSDD were found. Among them, H185, an indica variety, showed stable and high resistance to RBSDD. Using an F2:3 population of H185 and Wuyujing 3 (WYJ3, a highly susceptible japonica rice variety), three QTL conferring resistance to RBSDD, namely qRBSDD2, qRBSDD7, and qRBSDD11 were identified, and they explained 53.6% of the total phenotypic variation. Among them, qRBSDD2 and qRBSDD7, with LOD scores of 4.26 and 4.25, respectively, were repeatedly detected in artificial inoculation conditions, accounting for 28.0% and 29.8% of the total phenotypic variation, respectively. Resistant alleles of the two QTL were all derived from H185, and several BC5F2 lines possessing single or two QTL of qRBSDD2 and qRBSDD7 exhibited higher resistance for RBSDD. The QTL detected in our study open new possibilities for breeding rice cultivars with RBSDD resistance through resistance gene pyramiding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice black-streaked dwarf disease (RBSDD) is a plant disease caused by rice black-streaked dwarf virus (RBSDV), which is transmitted by small brown planthopper (SBPH, Laodelphax striatellus Fallén) in a persistent propagative manner (Van et al. 2000). Rice black-streaked dwarf virus is a double-stranded RNA (dsRNA) virus belonging to the second group of the genus Fijivirus, and the disease it causes is easily transmitted and can lead to epidemics (Lee et al. 2005). In addition to rice, it can also infect wheat, barley, corn, and other gramineous crops, and causes serious damage to crop production (Bai et al. 2002). Typical symptoms of plants infected with RBSDV include stunted growth, dark green leaves, white waxy or black-streaked enations on the abaxial surface of leaves, and poor heads or no heads at all. In rice production, spraying chemical pesticides is still the main method of controlling SBPH and RBSDD, but this has proven ineffective with recent increases in SBPH migration and outbreak frequency. In addition, pesticides are expensive and harmful to the environment, and can also enable SBPH to readily develop resistance (Liu et al. 2020). Therefore, exploring new rice genes from diverse sources and developing new resistant cultivars are considered to be the most economical and effective strategies for SBPH and RBSDD management (Wang et al. 2014; Vicheka et al. 2017).
The Zhejiang Province of China suffered an outbreak of RBSDD in the 1960s, but since then, RBSDD had entered a period of dormancy, therefore failing to arouse the attention of breeders and farmers. Thus, screening of germplasms conferring RBSDD resistance has begun only recently, few varieties and QTL with high RBSDD resistance level have been identified to date, and the molecular mechanism underlying RBSDD resistance remains poorly understood. Using a set of chromosome segment substitution lines (CSSLs) by backcross between Zhenshan 97 and Minghui 63, a major QTL, qRBSDV-6MH, was fine-mapped at a physical distance of 627.6 kb on chromosome 6 (Li et al. 2013). In our previous study, three QTL for RBSDD resistance located on chromosomes 6, 9, and 11, were repeatedly detected by using an F2:3 population derived from a cross between 9194 and Suyunuo (Sun et al. 2017). Xu et al. (2018) identified five QTL for RBSDD resistance, which were distributed on chromosomes 3, 6, 7, 9, and 11, through nature infection in the field of RBSDD resistance of an F2:3 population derived from WR24 and Suyunuo. In addition to bi-parental QTL analysis, genome-wide association study (GWAS) has become a popular strategy for mining QTL in plants. Xiao et al. (2019) identified a significant RBSDD-associated locus, named as qRBSDVD6, by GWAS analysis, and the QTL was confirmed by a recombinant inbred line (RIL) population derived from W5686 and Suyunuo. In another study conducted by Feng et al. (2019), GWAS on RBSDD resistance was conducted in the field using the rice diversity panel 1 (RDP1) cultivars, two QTL, qRBSDV-4.2 and qRBSDV-6.3, were detected in two environment conditions, furthermore, introgression of qRBSDV-6.3 into two susceptible rice cultivars by marker-assisted selection (MAS) demonstrated the effectiveness of qRBSDV-6.3 in enhancing RBSDV resistance. Wang et al. (2022) identified a highly resistant variety W44 through extensive evaluation of a diverse international rice panel, using genome-wide association study, a significant associated locus with largest phenotypic effect, namely, qRBSDV6-1, was finely mapped to a 0.8–1.2 Mb region on the short arm of chromosome 6, and the candidate gene of qRBSDV6-1, aspartic proteinase 47 (OsAP47), was identified and functionally confirmed as the first cloned gene for its resistance to RBSDD in rice, recently. Liu et al. (2021) identified a significant RBSDD-associated locus, named as qRBSDV-6.1, using a different population by multi-locus GWAS, haplotype analyses of the candidate genes LOC_Os06g03150 suggested that the gene is mainly associated with the differentiation of resistance within the xian subgroup, meaning this gene is very important for underlying RBSDD resistance to xian accessions. In addition, the interaction between SBPH, rice plants, and RBSDV is complex and diverse, so there is an urgent need to map more QTL in order to breed resistant cultivars to reduce the damage caused by RBSDD (Zhou et al. 2018).
In order to identify reliable and highly resistant germplasms and QTL for RBSDD resistance, 248 germplasm resources collected from different countries and regions were identified by field identification in our study. Among them, 19 rice varieties (including H185) were highly resistant to RBSDD. An F2:3 population of H185/WYJ3 was then constructed to detect QTL with RBSDD resistance, and two QTL, qRBSDD2 and qRBSDD7, were repeatedly detected in artificial inoculation conditions, accounting for 57.8% of the total phenotypic variation. Furthermore, introgression of qRBSDD2 and qRBSDD7 into a susceptible rice variety by MAS demonstrated that those two QTL are reliable and can act additively. The results of this study will provide useful genetic resources and molecular markers for breeding new cultivars conferring resistance to RBSDD.
Materials and methods
Plant materials
A total of 248 rice varieties collected from different rice-growing regions were selected to evaluate RBSDD resistance by natural infection in the field in 2015 and were provided by the Institute of Germplasm Resources and Biotechnology, Jiangsu Academy of Agricultural Sciences, China. An F2 population containing 188 individuals was constructed by crossing H185 (indica) and WYJ3 (japonica), and then, each F2 plant was self-crossed to generate the corresponding F2:3 line. 188 F2:3 individuals and the two parental varieties were used for field and artificial identification in 2016 and 2017, respectively. Among them, 133 and 83 individuals with good repeatability were selected to detect QTL related to RBSDD. The indica cultivar 9194 with high resistance to RBSDD was used as a resistant control, while the japonica rice cultivar Suyunuo (SYN) was used as a susceptible control, as described in Sun et al. (2017).
Evaluation of RBSDD resistance
In 2015–2016, experimental fields in Kaifeng City in Henan Province were chosen for field natural evaluation, due to the prevalence of RBSDD in this region. Field and artificial inoculation identification of RBSDD were carried out as described in our previous study (Sun et al. 2017). Briefly, about 15 days before harvesting winter wheat, 60 seeds per cultivar or F2:3 individual were sown in a rice seedling bed encircled by wheat plants infected with RBSDV. At 4 weeks after sowing seeds, 40 seedlings per cultivar or F2:3 individual were transplanted into an experimental area at Kaifeng, and the plants were cultured with normal field management, except that pesticides were not applied during the rice-growing period. The plants with the typical symptoms of RBSDD were considered to be susceptible plants. The incidence of RBSDD was recorded at the peak tillering stage as the infected plants showed obvious and typical disease symptoms at this stage. Each cultivar or F2:3 individual was replicated three times, and the average disease incidence (DI, the number of RBSDV-infected plants/the total number of plants counted × 100%) was used for QTL analysis.
Linkage mapping and QTL analysis
The polymorphisms of H185 and WYJ3 were detected by 412 pairs of simple sequence repeats (SSR) and 124 pairs of insertion/deletion (InDel) markers evenly distributed across 12 rice chromosomes. Then, 133 F2:3 individuals of H185/WYJ3 were analyzed by 142 pairs of markers with polymorphic differences among the parent plants. The precise names of SSR marker and sequences of InDel marker are shown in the Supplementary materials. Linkage groups were assembled using Mapmaker/EXP 3.0 (MapMaker3.0b for Windows http://www.softpedia.com/get/Science-CAD/MapMaker.shtml) (Lander et al. 1987). The total length of the map was 1426.3 cM, and the average distance between markers was 10.04 cM, which was suitable for QTL analysis. Following this, QTL analyses of F2:3 population containing 133 or 83 plants was carried out by composite interval mapping using QTL Cartographer, version 2.5 (2010 N.C. State University, Bioinformatics Research Center, Raleigh, NC, USA). The percentage of phenotypic variation explained (PVE) for each QTL and the additive effects were also obtained using the software.
Introgressing and pyramiding qRBSDD2, qRBSDD7 through marker-assisted selection (MAS)
To introgress and pyramid RBSDD resistance QTL to WYJ3, we firstly crossed H185 with WYJ3 and then resultant hybrid F1s were continuously backcrossed with WYJ3 five times. In each generation, two flanked markers of each QTL were deployed to select target recombinants. Finally, eight none, single or two QTL introgressed homozygous plants selected from the BC5F2 population were identified by artificial inoculation, each line had five replicates, and each replicate had 40 plants. SNP genotyping was performed by Huazhi Biotechnology Company using multiplex PCR with next-generation sequencing on the high-throughput genotyping platform, lllumina X-10.
Results
Screening of RBSDD-resistant germplasm resources
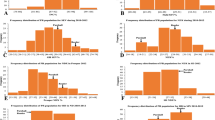
In this study, 248 germplasm resources from different countries and regions were screened for resistance to RBSDD (Fig. 1A). The results showed that the disease incidence (DI) of the 9194 (resistant control) was 9.2%, indicating high resistance to RBSDD, and the DI of the Suyunuo (SYN) (susceptible control) was 82.6%. The overall phenotype of all the tested accessions showed a left bias normal distribution, 19 varieties were highly resistant to RBSDD (DI < 10%), and 45 varieties were susceptible to RBSDD with DI of more than 50% (Fig. 1B). Among the 19 varieties with high resistance to RBSDD, 42.1% were japonica accessions and 57.9% were indica accessions. Among the 45 highly susceptible varieties, japonica and indica rice cultivars accounted for 64.4% and 35.6%, respectively. Among these, the indica cultivar H185 was highly resistant, while the japonica cultivar WYJ3 was highly susceptible (Fig. 1B). According to the analysis of the indica and japonica subsets, the DI of indica varieties was significantly lower than those of japonica varieties (Fig. 1C). Based on the comparison of the DI of different germplasm resources in different countries, the average DI of indica varieties was lower than that of japonica varieties in all countries except Thailand (Fig. S1A), and the same result was found across the different provinces of China (Fig. S1B).
Geographical distribution and evaluation for rice black-streaked dwarf disease (RBSDD) resistance of all the tested rice accessions. A Geographical distribution of 248 rice species in various countries and provinces of China; B frequency distribution of 248 germplasm resources for resistance to RBSDD; C distribution of RBSDD incidence rate of germplasm resources with indica and japonica subgroups
Resistance of H185 to RBSDD
In order to detect QTL conferring resistance to RBSDD, H185 and WYJ3 plants were identified by field identification and artificial inoculation in 2016 and 2017, respectively. Using field identification, the average DI values of H185 and WYJ3 were 8.9% and 58.3%, respectively (Fig. 2A, B, D). Using artificial inoculation, the average DI values of H185 and WYJ3 were 12.4% and 75.1%, respectively, (Fig. 2C, D). There was a significant difference between H185 and WYJ3, meaning that these two cultivars were ideal for RBSDD research.
Phenotypes of H185 and Wuyujing 3 (WYJ3) for rice black-streaked dwarf disease (RBSDD) resistance under natural and artificial inoculation conditions. A and B Representative image of H185 and WYJ3 inoculated with RBSDV in the natural field condition in 2016. C Representative image of H185 and WYJ3 artificially inoculated with RBSDV in 2017. D The disease incidence of H185 and WYJ3 by the field identification and artificial inoculation. n = 3, **p < 0.01 by Student’s t-test
Genetic analysis and identification of QTL for resistance to RBSDD
To detect QTL for RBSDD resistance, an F2 population including 188 F2:3 individuals derived from H185 and WYJ3 was used in natural infection experiments in 2016. Of the 188 individuals, 133 with good repeatability (less than 5% variation) were selected to further detect QTL for RBSDD resistance. The DI frequency distribution of the 133 F2:3 individuals varied continuously, ranging from 0 to 61.9%, and there was a peak at 20%, which exhibited characteristics of quantitative traits (Fig. 3A). Those results suggested that resistance to RBSDD was controlled by multiple QTL in this population.
Based on the field phenotypic data of the 133 F2:3 individuals, the RBSDD resistance QTL in H185 were detected by complex interval mapping using the Windows QTL Cartographer V2.5 software. As shown in Table 1 and Fig. 4, three QTL, namely qRBSDD2, qRBSDD7, and qRBSDD11, were detected on chromosomes 2, 7, and 11, respectively. These QTL were located between the markers RM425-RM208, RM3484-RM6776, and RM26105-Indel11-3, respectively. They explained 20.8%, 21.4%, and 11.4% of the phenotypic variance, respectively, with LOD scores of 5.35, 4.81, and 2.65, respectively (Table 1). Jointly, they explained 53.6% of the total phenotypic variance in this population. The additive effect analysis showed that the above-mentioned three QTL were all inherited from the resistant cultivar H185.
Verification of QTL resistant to RBSDD
In order to verify the three QTL resistant to RBSDD in the present study, the two parental cultivars and 83 F2:3 individuals were identified by artificial inoculation in 2017 (Fig. 3B). Using a QTL mapping strategy, two QTL, qRBSDD2 and qRBSDD7, could be re-detected and stably expressed, the LOD scores were both greater than 4, and the phenotypic variance were more than 25%, which is thought to indicate major QTL (Table 1 and Fig. 4). Taken together, these data further confirmed that these QTL were reliable.
Pyramiding effect of QTL on RBSDD resistance
The pyramiding effect of two stable QTL, qRBSDD2 and qRBSDD7, was analyzed by comparing the mean DI of different lines from a BC5F2 population by the backcross between H185 and WYJ3, with WYJ3 as the recurrent parent. The genotype of eight BC5F2 individuals was assessed using a PCR-based genotyping protocol, indicating the proportion of WYJ3 genotype was more than 95% (95.2–97.7%), and none of the BC5F2 lines carried the resistant allele of qRBSDD11 detected in our study. As shown in Fig. 5, two lines carrying none of the QTL showed a susceptible phenotype with an average DI of 67.5% and 63.8%, respectively, while those introgressed lines possessing any single QTL of qRBSDD2 or qRBSDD7 exhibited moderate resistance, with DI ranging from 36.2% to 43.7%. Remarkably, two QTL pyramided lines showed higher resistance for RBSDD than other lines, with an average DI of 21.4% and 22.9%, respectively (Fig. 5). Taken together, these data further confirmed that these two QTL are reliable and can act additively.
Pyramiding effect of two stable QTL associated with resistance to rice black-streaked dwarf disease (RBSDD). The qRBSDD2, qRBSDD7, or two loci pyramided lines, selected from a BC5F2 population by the backcross between donor parent H185 and recurrent parent WYJ3, were identified by artificial inoculation, n = 5. Bars denoted by different letters are significantly different at p < 0.05
Discussion
With global warming and changes to the cultivation system, the outbreak of RBSDD has posed a serious threat to rice production in Asia regions, especially in eastern China. There have been many reports on the symptoms of the disease, but due to complex epidemiological factors, cumbersome field experiments, the need for large-scale investigations, and a longer research cycle, the systematic and in-depth study of RBSDD has become extremely difficult (Li et al. 2013). Therefore, little research has been conducted on the inheritance of or breeding strategies that could be used to enhance RBSDD resistance, and no rice variety resistant to RBSDD has been widely used in rice production. It is worth noting that the average DI of japonica varieties in the Philippines and Indonesia was greater than 50% (Fig. S1A). In China, the resistance of indica varieties in Hubei Province was very weak, with an average DI of more than 60% (Fig. S1B). These countries or regions seriously lack RBSDD-resistant germplasm resources, and the consequences may be very serious once disease outbreaks occur. Therefore, it is urgent to accelerate the screening of rice germplasm resources in order to map major QTL to breed new cultivars with high resistance to RBSDD.
Because resistance mechanisms against insect-borne viruses are very complex, phenotypic identification of viral diseases has always been an issue (Mandadi and Scholthof 2013; Souza and Carvalho 2019). Rice black-streaked dwarf disease is transmitted by SBPH in a persistent manner, but RBSDV cannot be transmitted to the next generation via eggs, which makes artificial inoculation and identification of RBSDD extremely difficult (Xiao et al. 2019). In addition, field natural identification is convenient and easy to implement. Large-scale identification of RBSDD relies mainly on field natural identification in severely diseased areas. However, owing to the uneven distribution of vector insects, the accuracy of field natural identification needs to be further improved (Zheng et al. 2012). In order to improve the reliability of field natural evaluation, the following measures were taken in our study: (A) Selecting regions with serious disease outbreaks as identification nurseries on the premise that the DI of susceptible varieties was more than 50% in the past year, and eventually, experimental fields in Kaifeng City in Henan Province were chosen for field natural evaluations due to the prevalence of RBSDD in this region. (B) Increasing the SBPH population: Experimental fields were surrounded by wheat fields, after wheat harvest, a large population of SBPH moved to the experimental field to ensure the quantity of vector insects. (C) Improved uniformity of insect exposure: After the wheat harvest, the SBPH were expelled several times a day in the seedling field. (D) Evaluation of RBSDD resistance by artificial inoculation: RBSDV-carrying SBPH individuals were used to inoculate the F2:3 population, which ensured the uniformity and adequacy of inoculation intensity. The above measures ensured the reliability of the phenotypic evaluation.
The present study found that the resistant parents, such as Minghui63, Tetep, WR24, and, W5686, were all indica varieties in the previous QTL mapping of RBSDD resistance (Li et al. 2013; Zhou et al. 2015; Xu et al. 2018; Xiao et al. 2019). Similarly, H185, the resistant parent in our study, was also an indica variety, which indicated that indica rice varieties were more resistant to RBSDD than japonica rice in the evolutionary process. This hypothesis was further confirmed by comparing the DI of indica and japonica subsets in different countries and regions.
Previous studies have shown that resistance in the F2 population obtained by crossing resistant and susceptible germplasms shows the characteristics of a normal distribution (Zhou et al. 2015; Zhang et al. 2016; Xu et al. 2018), meaning that resistance of RBSDD is controlled by multiple QTL. These genetic characteristics have been confirmed in maize rough dwarf disease (MRDD), which was also caused by RBSDV (Luan et al. 2012; Shi et al. 2012; Tao et al. 2013). In this study, three RBSDD-resistant QTL were also detected in the F2:3 population of H185/WYJ3, which supports the results of previous studies. These results provide useful information for enhancing the breeding of new varieties resistant to RBSDD, and lay a foundation for exploring the molecular mechanism behind RBSDD resistance.
Among the three QTL detected in our study, qRBSDD2 and qRBSDD7 explained 20.8% and 21.4% of the phenotypic variance, respectively, with LOD scores of 5.35 and 4.81, respectively, suggesting that they are major QTL involved in resistance to RBSDD. In order to further confirm the existence of these two QTL, 83 F2:3 individuals of H185/WYJ3 were re-identified by artificial inoculation, the presence of these two QTL was re-detected at the same location, which indicated that qRBSDD2 and qRBSDD7 were stably expressed major QTL for RBSDD resistance. Marker-assisted pyramiding of QTL is an effective way to improve quantitative traits. In this study, we determined the pyramiding effect of these two QTL in BC5F2 population derived from the backcross between donor parent H185 and recurrent parent WYJ3, and the pyramiding lines of the two QTL exhibited higher resistance for RBSDD than lines with no or a single QTL. Therefore, qRBSDD2 and qRBSDD7 have great potential for enhancing the breeding of new rice varieties resistant to RBSDD. Two tightly linked markers with the two QTL (SR2-4 and SR7-3, Supplementary materials) were designed and can be used in breeding cultivars with RBSDD resistance by marker-assisted selection (MAS).
At present, approximately 20 QTL conferring resistance to RBSDD have been detected on different rice chromosomes. In previous studies, Zheng et al. (2012) detected a RBSDD-resistant QTL named qRBSDV2 on the short arm of chromosome 2, while the qRBSDD2 detected in the present study is located on the long arm of chromosome 2. Xu et al. (2018) detected a RBSDD-resistant QTL named qRBSDV7WR24 on chromosome 7, while according to the physical locations in the rice genomes, qRBSDV7WR24 and qRBSDD7 detected in this study are quite far apart, hence, qRBSDD2 and qRBSDD7 may be novel QTL involved in RBSDD resistance. Generally, chromosome 11 is considered to be a "hotspot" of rice resistance genes to many diseases, such as rice blast, bacterial blight, and stripe blight (Chauhan et al. 2002; Sallaud et al. 2003; Xiang et al. 2006; Zhang et al. 2011). In our previous study, a QTL conferring resistance to RBSDD, named as qRBSDV11, was detected on chromosome 11 using the F2:3 segregation population of 9194/Suyunuo, which explained 31.1% of the phenotypic variance with a LOD score of 7.16 (Sun et al. 2017). The chromosomal location of this QTL overlaps with that of the qRBSDD11 identified in this study, and these two QTL are likely controlled by the same gene or allele. The results indicate that qRBSDD11 is stable in different backgrounds and is valuable for MAS in breeding cultivars resistant to RBSDD.
References
Bai FW, Jian Y, Qu ZC, Zhang HW, Jia X, Ye MM, Shen DL (2002) Phylogenetic analysis reveals that a dwarfing disease on different cereal crops in china is due to rice black streaked dwarf virus (RBSDV). Virus Genes 25(2):201–206. https://doi.org/10.1023/a:1020170020581
Chauhan RS, Farman ML, Zhang HB, Leong SA (2002) Genetic and physical mapping of a new blast resistance locus, Pi-CO39(t), that corresponds to the avirulence gene AVR1-CO39 of Magnaporthe grisea. Mol Genet Genomics 267:603–612. https://doi.org/10.1007/s00438-002-0691-4
Feng ZM, Kang HX, Li MY, Zou LH, Wang XQ, Zhao JH, Wei L, Zhou NN, Li QQ, Lan Y, Zhang YF, Chen ZX, Liu WD, Pan XB, Wang GL, Zuo SM (2019) Identification of new rice cultivars and resistance loci against rice black-streaked dwarf virus disease through genome-wide association study. Rice 12:49. https://doi.org/10.1186/s12284-019-0310-1
Lander ES, Philip G, Jeff A, Aaron B, Mark JD, Stephen EL, Lee N (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181. https://doi.org/10.1016/j.ygeno.2008.12.003
Lee BC, Hong YK, Hong SJ, Park ST, Lee KW (2005) Occurrence and detection of rice black-streaked dwarf virus in Korea. J Plant Pathol 21(2):172–173. https://doi.org/10.5423/PPJ.2005.21.2.172
Li AH, Pan CH, Wu LB, Dai ZY, Zuo SM, Xiao N, Ling Y, Li YH, Zhang XX, Xue WX, Zhang HX, Pan XB (2013) Identification and fine mapping of qRBSDV-6MH, a major QTL for resistance to rice black-streaked dwarf virus disease. Mol Breeding 32(1):1–13. https://doi.org/10.1007/s11032-012-9807-7
Liu QC, Deng SN, Liu BS, Tao YF, Ai HY, Liu JJ, Zhang YZ, Zhao Y, Xu ML (2020) A helitron-induced RabGDIα variant causes quantitative recessive resistance to maize rough dwarf disease. Nat Commun 11:495. https://doi.org/10.1038/s41467-020-14372-3
Liu Q, Lan GF, Zhu YJ, Chen K, Shen CC, Zhao XQ, Zhang F, Xu JL, Li ZK (2021) Genome-wide association study on resistance to rice black-streaked dwarf disease caused by rice black-streaked dwarf virus. Plant Dis 105(3):607–615. https://doi.org/10.1094/PDIS-10-19-2263-RE
Luan J, Wang F, Li Y, Zhang B, Zhang J (2012) Mapping quantitative trait loci conferring resistance to Rice black-streaked virus in maize (Zea mays L.). Theor Appl Genet 125:781–791. https://doi.org/10.1007/s00122-012-1871-1
Mandadi KK, Scholthof KBG (2013) Plant immune responses against viruses: how does a virus cause disease? Plant Cell 25:1489–1505. https://doi.org/10.1105/tpc.113.111658
Sallaud C, Lorieux M, Roumen E, Tharreau D, Berruyer R, Svestasrani P, Garsmeur O, Ghesquiere A, Notteghem JL (2003) Identification of five new blast resistance genes in the highly blast-resistant rice variety IR64 using a QTL mapping strategy. Theor Appl Genet 106:794–803. https://doi.org/10.1007/s00122-002-1088-9
Shi LY, Hao ZF, Weng JF, Xie CX, Liu CL, Zhang DG, Li MS, Bai L, Li XH, Zhang SH (2012) Identification of a major quantitative trait locus for resistance to maize rough dwarf virus in a chinese maize inbred line X178 using a linkage map based on 514 gene-derived single nucleotide polymorphisms. Mol Breeding 30(2):615–625. https://doi.org/10.1007/s11032-011-9652-0
Souza PFN, Carvalho FEL (2019) Killing two birds with one stone: how do plant viruses break down plant defenses and manipulate cellular processes to replicate themselves? J Plant Biol 62:170–180. https://doi.org/10.1007/s12374-019-0056-8
Sun ZG, Liu YQ, Xiao SZ, Hu JL, Pan G, He J, Xu TT, Huang J, Qiu ZY, Fan DJ, Zhang L, Liu LL, Jiang L, Cheng XN, Zhai HQ, Wan JM (2017) Identification of quantitative trait loci for resistance to rice black-streaked dwarf virus disease and small brown planthopper in rice. Mol Breed 37:72. https://doi.org/10.1007/s11032-017-0669-x
Tao YF, Liu QC, Wang HH, Zhang YJ, Huang XY, Wang BB, Lai JS, Ye JR, Liu BS, Xu ML (2013) Identification and fine-mapping of a QTL, qMRDD1, that confers recessive resistance to maize rough dwarf disease. BMC Plant Biol 13(1):145. https://doi.org/10.1186/1471-2229-13-145
Van RMH, Mayo MA, Fauquet CM, Maniloff J (2000) Virus nomenclature: consensus versus chaos. Arch Virol 145(10):2227–2232. https://doi.org/10.1007/s007050070053
Vicheka T, Yun S, Lindain AF, Sopheareth M, Chung IK, Kim KM (2017) Defense gene expression and phenotypic changes of rice (Oryza sativa L.) at the reproductive stage in response to whitebacked planthopper (Sogatella furcifera Horvath) infestation. Cereal Res Commun 45:456–465. https://doi.org/10.1556/0806.45.2017.025
Wang Q, Liu YQ, He J, Zheng XM, Hu JL, Liu YL, Dai HM, Zhang YX, Wang BX, Wu WX, Gao H, Zhang YH, Tao XR, Deng HF, Yuan DY, Jiang L, Zhang X, Guo XP, Cheng XN, Wu CY, Wang HY, Yuan LP, Wan JM (2014) STV11 encodes a sulphotransferase and confers durable resistance to rice stripe virus. Nat Commun 5:4768. https://doi.org/10.1038/ncomms5768
Wang ZY, Zhou L, Lan Y, Li XJ, Wang J, Dong JF, Guo W, Jing DD, Liu Q, Zhang SH, Liu ZY, Shi WJ, Yang W, Yang TF, Sun F, Du LL, Fu H, Ma YM, Shao YD, Chen L, Li S, Fan YJ, Wang YY, Leung H, Liu B, Zhou YJ, Zhao JL, Zhou T (2022) An aspartic protease 47 causes quantitative recessive resistance to rice black-streaked dwarf virus disease and southern rice black-streaked dwarf virus disease. New Phytol 233:2520–2533. https://doi.org/10.1111/nph.17961
Xiang Y, Cao YL, Xu CG, Li XH, Wang SP (2006) Xa3, conferring resistance for rice bacterial blight and encoding a receptor kinase-like protein, is the same as Xa26. Theor Appl Genet 113:1347–1355. https://doi.org/10.1007/s00122-006-0388-x
Xiao SZ, Wang BX, Liu YQ, Miao TH, Zhang HL, Wen PZ, He J, Huang J, Liu DM, Qiu ZY, Liu LL, Liu SJ, Jiang L, Cheng XN, Wang CM, Xu DY, Wan JM (2019) Genome-wide association study and linkage analysis on resistance to rice black-streaked dwarf virus disease. Mol Breed 39:73. https://doi.org/10.1007/s11032-019-0980-9
Xu TT, Liu YQ, Zhang LL, Liu LL, Wang CM, Hu JL, Sun ZG, Pan G, Xiao SZ, He J, Huang J, Qiu ZY, Fan DJ, Jiang L, Cheng XN, Zhai HQ, Wan JM (2018) Mapping of quantitative trait loci associated with rice black-streaked dwarf virus disease and its insect vector in rice (Oryza sativa L.). Plant Breed 137:698–705. https://doi.org/10.1111/pbr.12620
Zhang YX, Wang Q, Jiang L, Liu LL, Wang BX, Shen YY, Wan JM (2011) Fine mapping of qSTV11KAS, a major QTL for rice stripe disease. Theor Appl Genet 122:1591–1604. https://doi.org/10.1007/s00122-011-1557-0
Zhang HG, Ge YS, Wang MY, Liu JN, Si H, Zhang LJ, Liang GH, Gu MH, Tang SZ (2016) Mapping QTLs conferring resistance to rice black-streaked dwarf disease in rice (Oryza sativa L.). Euphytica 212(2):1–8. https://doi.org/10.1007/s10681-016-1782-3
Zheng TQ, Yang J, Zhong WG, Zhai HQ, Zhu LH, Fan FJ, Li ZK (2012) Novel loci for field resistance to black-streaked dwarf and stripe viruses identified in a set of reciprocal introgression lines of rice (Oryza sativa L.). Mol Breed 29:925–938. https://doi.org/10.1007/s11032-011-9691-6
Zhou T, Du L, Wang LJ, Wang Y, Gao CY, Lan Y, Zhou YJ (2015) Genetic analysis and molecular mapping of QTLs for resistance to rice black-streaked dwarf disease in rice. Sci Rep 5:10509. https://doi.org/10.1038/srep10509
Zhou C, Li S, Zhou Y, Fan Y (2018) A simple method for obtaining rice black-streaked dwarf virus-infected small brown planthopper nymphs. J Virol Methods 251:80–82. https://doi.org/10.1016/j.jviromet.2017.10.018
Acknowledgements
We would like to thank Wiley Editing Services for English language editing.
Funding
This study was funded by the China Agriculture Research System (CARS-01–61), Natural Science Foundation of Jiangsu Province of China (BK20201214), The Project for Agricultural Significant New Varieties Breeding of Jiangsu Province (PZCZ201704), Natural Science Foundation of Hunan Province of China (2019JJ50714), and the Financial Grant Support Program of Lianyungang City (QNJJ2001).
Author information
Authors and Affiliations
Contributions
ZS conducted linkage mapping and wrote the paper; GP conducted the SSR and InDel marker genotyping of parental lines and individuals of the F2:3 population; HA conducted the disease phenotyping and analyzed the data; BW and DX designed and supervised the study. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Jizeng Jia.
Supplementary Information
Below is the link to the electronic supplementary material.
42976_2023_423_MOESM1_ESM.pdf
Fig. S1 Evaluation for rice black-streaked dwarf disease (RBSDD) resistance of rice accessions with indica and japonica subgroups. (A) The average disease incidence of indica and japonica subgroups in various countries; (B) The average disease incidence of indica and japonica subgroups in different provinces of China. (PDF 361 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sun, Z., Pan, G., An, H. et al. Molecular mapping of QTL for rice black-streaked dwarf disease resistance in rice (Oryza sativa L.). CEREAL RESEARCH COMMUNICATIONS 52, 413–421 (2024). https://doi.org/10.1007/s42976-023-00423-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42976-023-00423-0