Abstract
Apple replant disease (ARD) negatively affects growth and yield of apple plants worldwide. Fungi belonging to the Nectriaceae have often been isolated from roots grown in replant soils and thus are proposed among others as one biotic cause of the disease complex. Microscopic analyses of ARD-affected roots revealed characteristic symptoms associated with fungal infection sites. Here, two extraction methods of such tissue sites were applied to directly identify an unknown fungus that forms typical cauliflower-like structures in diseased root cortex cells. Punching small tissue samples of about 0.5 mm3 volume with the Harris Uni-Core is a quick and easy method to harvest symptomatic material. Secondly, a laser microdissection (LMD) protocol for apple roots was established. This technique allows the extraction of defined cell or tissue fractions from thin cryo-sections. Tissue harvesting was followed by the identification of fungi via PCR amplification of two gene fragments and Sanger sequencing. For Harris samples, Chelex was used for DNA stabilization, while LMD samples were directly submitted to PCR. In Harris samples, mainly the Nectriaceae species Dactylonectria torresensis, Ilyonectria robusta and Rugonectria rugulosa were identified. In addition to these, in LMD samples Cylindrocladiella sp. and Ilyonectria europaea were detected. Thus, the intracellular CF structures contained different species of Nectriaceae in the ARD-affected cortex cells. These results contribute considerably to the etiology of the ARD. Both protocols offer the possibility to identify fungi from selected symptomatic small root sections by molecular tools avoiding isolation and subsequent axenic pure cultures of single fungal isolates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The replant disease of apple negatively affects the plant growth and fruit yield in apple nurseries and orchards worldwide (Geldart 1994; Mazzola and Manici 2012; Winkelmann et al. 2019). Often plant growth-associated measurements like shoot length and plant weight are used to assess the severity of ARD (Mahnkopp et al. 2018; Reim et al. 2019; Yim et al. 2013). In addition, the root system, which is in direct contact with the diseased soil, shows reductions in the root biomass, root tip necrosis and discolorations (Caruso et al. 1989; Grunewaldt-Stöcker et al. 2019; Hoestra 1968). Biotic factors are known to be involved in the etiology since the plant growth can be restored by soil disinfection treatments (Mai and Abawi 1981; Mazzola 1998; Yim et al. 2013). Apple plants can produce phytoalexins to deal with biotic stress. These secondary metabolites were nearly absent in healthy plants, but were produced in high amounts in roots affected by ARD (Weiß et al. 2017a). Nevertheless, it was reported that an effective defense response was missing albeit genes encoding enzymes of phytoalexin biosynthesis were upregulated and the phytoalexins were produced (Weiß et al. 2017a, b).
A detailed microscopic analysis of ARD-affected apple fine roots described early symptoms like necrosis and blackening that often co-localized with fungal infection sites in cortex tissue (Grunewaldt-Stöcker et al. 2019). They could be detected already after two weeks of growth in ARD-affected soil in susceptible plant genotypes such as ‘M26’ and ‘Bittenfelder’. In our studies on early ARD root symptoms, the occurrence of nematodes, Pythium spp. and Rhizoctonia spp. was very rarely observed although these organisms were presented in many reports as candidate inducers of ARD (Braun 1991; Jaffee et al. 1982; Kelderer et al. 2012; Manici et al. 2003; Mazzola 1998; Mazzola and Manici 2012; Tewoldemedhin et al. 2011; Utkhede et al. 1992). In contrast, we found strong evidence for Nectriaceae fungi rapidly infecting young root tissue of the tested susceptible genotypes. The fungi contribute to necrosis and blackening in diseased roots segments by spreading in the tissue and the development of special fungal structures (Grunewaldt-Stöcker et al. 2019). Similarly, also for rose replant disease, a close co-occurrence of blackening and necroses and Nectriaceae was confirmed in detailed histological studies on roots of Rosa corymbifera (Grunewaldt-Stöcker et al. submitted).
In previous experiments, more than 150 endophytic fungal isolates from ARD-affected roots were obtained by microbiological approaches and the pure cultures were identified. Nectriaceae accounted for a large proportion of 37.7% of these ARD-associated fungal root endophytes (Popp et al. 2019). This classical culture-dependent approach required a lot of time, but was indispensable for later inoculation tests and essentially contributed to unravel biotic causes of ARD.
The repeated observation of intracellular fungal structures with a cauliflower-like appearance (CF) in our previously microscopic analyses of apple fine roots (Grunewaldt-Stöcker et al. 2019, Grunewaldt-Stöcker et al. submitted) led to the motivation of this study. Herein, our objective was to collect these fungal structures from the affected cortex cells for an immediate identification by molecular tools. In this study, we established the protocols for two sampling methods applied to fresh or fixed fine roots, namely Harris Uni-Core punching and laser microdissection (LMD) of cryo-sections, and tailored them to the structural properties of apple root tissue. These methods offer the possibility to identify fungi harvested from selected tiny root sections down to selected cells. Moreover, the identification of fungi associated with CF structures presented in this study will help to unravel the biotic complex associated with ARD.
Materials and methods
Plant and soil material
In all experiments, the susceptible apple rootstock ‘M26’ (hereinafter given as M26) was used as the indicator plant for early detection of ARD. For the greenhouse biotests, in vitro propagated plantlets were potted four weeks after acclimatization into soil from three German ARD sites, i.e., Ellerhoop (E), Heidgraben (H) and Ruthe (R), according to Mahnkopp et al. (2018) (n = 10 plants each). For comparison, 10 plants were grown in γ-irradiated (> 10 kGy) Heidgraben soil. Three to six plants per trial and treatment were harvested after 2–4 weeks for tissue sampling. The biotest was repeated three times: 1. in November 2018 (Harris and LMD sampling), 2. February 2019 (only LMD sampling) and 3. May 2019 (only Harris sampling).
Sample preparation
After two, three and four weeks of cultivation, apple root systems were rinsed several times in sterile tap water, fine roots were excised and cleaned of adhering soil residues with tweezers, and whole mounts were immediately analyzed microscopically for early symptoms and fungal infection sites according to Grunewaldt-Stöcker et al. (2019). Selected tissue areas, either symptom-free (sf) or symptomatically showing intracellular fungal CF structures, were collected: 1. by punching with Harris Uni-Core (a coring tool designed for sample collection from thin and soft substrates) and direct transfer into Chelex solution (a chelating polymer containing iminodiacetic acid groups used to prevent DNA degradation from degradative enzymes and from impurities that inhibit the PCR) for DNA stabilization and 2. by cutting off infected root segments (3–6 mm length) and fixation in 5% acetic acid for later dissection with Harris Uni-Core or for cryo-sectioning followed by laser-based microdissection. A total of 73 samples were punched with Harris Uni-Core and analyzed by PCR (following "Protocol for apple root tissue harvesting with Harris Uni-Core 0.50 mm and molecular identification of included fungi" section). Using LMD, 53 samples from thin sections were tested (see "Protocol for apple root tissue harvesting with laser microdissection and molecular identification of included fungi" section).
Protocol for apple root tissue harvesting with Harris Uni-Core 0.50 mm and molecular identification of included fungi
-
1.
Microscopy for the selection of infection sites in native, unstained whole mount fine roots (or fixed in 5 % acetic acid for 7–10 h, rinsed in distilled sterile water with 10 % DMSO (dimethylsulfoxid, (CH3)2SO, Merck Supelco, Darmstadt, Germany) and stored at 4 °C): Axio Imager.A2, 400–1000x, bright field with differential interference contrast (DIC) (Zeiss, Göttingen, Germany); cleaned slides, surface-sterilized preparation set.
-
2.
Punching out infection sites including CF structures with Harris Uni-Core 0.5 mm punch (Thermo Fisher, Waltham, USA) on the slide.
-
3.
Transfer of one punched sample each into sterile 0.2 mL PCR reagent tubes with 175 µl 10 % Chelex solution (w/w) (Bio-Rad, Hercules, USA, 200–400 mesh size) for DNA stabilization.
-
4.
After each sample collection, cleaning the punch tip by rinsing in 70% ethanol or 2% NaOCl.
-
5.
Microscopic verification of the cutout position at the root segment.
-
6.
Verification of the tissue sample in the PCR tube by stereo microscopy (Leica MZ 75; 6.3–50x, Wetzlar, Germany).
-
7.
Incubation 20 min at 95 °C.
-
8.
Storage at − 20 °C until PCR.
-
9.
PCR for identification of Nectriaceae fungi: Using primer pairs CYLH3F and CYLH3R (Crous et al. 2004) for amplifying of the histone H3 (HIS) gene or CylEF-1 (5′- ATG GGT AAG GAV GAV AAG AC-3′; J.Z. Groenewald, unpublished) and CylEF-R2 (Crous et al. 2004) for amplification of the translation elongation factor 1-α gene (TEF). A 10 µl PCR mix consisted of 5 µl Phusion Flash High Fidelity PCR Master Mix (Thermo Fisher, Waltham, USA), 1 µl Chelex sample and 1 µl of each primer for HIS or TEF fragment amplification (10 µM). Cycling conditions (C100 thermal cycler, Bio-Rad, Hercules, USA): 98°C for 10 s, followed by 36 cycles at 98°C for 1 s, 56°C for TEF and 63°C for HIS for 5 s and 72°C for 30 s, followed by a final elongation at 72°C for 5 min.
-
10.
Cleaning PCR fragments with SureClean Plus (Bioline, London, UK) according to the manufacturer’s instructions or excising fragments from the agarose gel (in case of additional unspecific amplification) and purification by applying Nucleospin Gel and PCR Clean-up Kit (Macherey & Nagel, Düren, Germany).
-
11.
Cloning with CloneJET PCR Cloning Kit (Thermo Fisher, Waltham, USA): 5 µl reaction buffer, 0.5 µl vector, 3.5 µl purified PCR fragment, 1 µl T4-Ligase (5 U µl−1).
-
12.
Picking of three well-separated single colonies in 75 µl LB.
-
13.
Colony-PCR: 10 µl PCR mix with 5 µl Phusion Flash High Fidelity PCR Master Mix (Thermo Fisher, Waltham, USA), 1 µl colony in LB, 1 µl of each primer pJS (5′-CAC CAT ATC CAT CCG GCG TAA TAC-3′) and pJas (5′-CCT GAT GAG GTG GTT AGC ATA GTT C-3′, E. Maiss, unpublished) (10 µM). Cycling conditions like step 9, but using 62°C for annealing and 34 cycles.
-
14.
Purification of PCR fragments with SureClean Plus according to the manufacturer’s instructions.
-
15.
Sanger sequencing of purified PCR fragments with the sense primer of the amplification (pJS) (Microsynth Seqlab, Göttingen, Germany).
-
16.
NCBI BLASTn analysis (National Center for Biotechnology Information, Bethesda, USA), first hit sorted by max. score (standard settings).
As positive control 1 µl of Dactylonectria torresensis, DNA extract was added to the Chelex mixture. For the negative (no template) control, only Chelex solution was added to the PCR mix.
Protocol for apple root tissue harvesting with laser microdissection and molecular identification of included fungi
-
1.
Microscopy for the selection of infection sites in native, unstained whole mount fine roots: Axio Imager.A2, 400–1000x, bright field with DIC (Zeiss, Göttingen, Germany).
-
2.
Preparation of sufficiently infected root segments of < 2 mm in diameter, maximum length 0.6 cm, preferably straight roots.
-
3.
Fixation of the segments in 5% acetic acid (7–10 h), rinse in distilled sterile water with 10% DMSO, approx. 20 mL and store in rolled rim glasses at 4 °C.
-
4.
Embedding of 3–5 root segments in a plastic mold (Tissue-Tek Cryomold Intermediate, square, 15×15×5mm, Science Services, München, Germany) with Tissue-Tek O.C.T. Compound (Sakura, Alphen aan den Rijn, the Netherlands); align samples parallel to each other.
-
5.
Shock freezing of the mold plus contents in liquid N: with large tweezers only immerse, but do not submerge, so that no liquid N gets onto the Tissue-Tek until the bubbling stops.
-
6.
Transfer the sample to the cryostat, − 20 °C chamber temperature.
-
7.
Block out and freeze on holder with an additional drop of Tissue-Tek; cool in the chamber.
-
8.
Cutting at − 20 °C, 10 µm thickness.
-
9.
Placing max. three cuts on special slides with a polyethylene naphthalate membrane field (MembraneSlide 1.0 PEN (D), Zeiss, Göttingen, Germany). These were irradiated with UV light (15 W, 245 nm) for 30 min prior to use in order to reduce electrostatic charge.
-
10.
Laser microdissection with the LMD 7000 instrument (Leica, Wetzlar, Germany): General Instructions according to Leica regulations.
UV-A laser adjustment for apple root tissue: power 30–40; aperture 2; speed 3; specimen balance 25; pulse frequency 4018 at 40× magnification, 5000 at 63× magnification. Specimen were laser dissected in single cuts and in combined cuts of two to five areas of interest, each of 300 µm2 up to 1400 µm2 depending on the size and assembly of the CF structures. The areas were selected on the screen and marked with a free hand marker line.
-
11.
Collection of tissue samples in caps of sterilized PCR reagent tubes (0.2 mL) in 12.5 µl 2x Phire Plant Tissue PCR buffer template (Thermo Fisher, Waltham, USA) and subsequent microscopic control of specimens in the cap (according to Leica instructions) to ensure a precise landing.
-
12.
Filling the caps with additional 12.5 µl buffer, closing, removing the caps from the holder and keeping them cold (− 20°C).
-
13.
PCR of the samples by adding 1 µl Phire Hot Start II DNA polymerase (Thermo Fisher, Waltham, USA), 0.5 µM of each sense and antisense primer (either HIS or TEF primers, see previous protocol) and sterile bi-distilled water to the PCR mix with a final volume of 50 µl (according to manufacturer's instructions). Cycling conditions: 98°C for 5 min, followed by 40 cycles at 98°C for 5 s, 56°C for TEF and 63°C for HIS for 5 s and 72°C for 20 s, and a final elongation at 72°C for 1 min.
-
14.
After successful amplification, excising fragments from the agarose gel and purification by applying Nucleospin Gel and PCR Clean-up Kit (Macherey & Nagel, Düren, Germany).
-
15.
Cloning, colony PCR and analyses of sequences, see previous protocol step 11–16.
Results
ARD symptoms in apple roots and sampling
The induction of ARD symptoms in M26 plantlets was successful as visually assessed in shoot and root performance already after two and four weeks of cultivation in ARD-affected soils of the three locations. All plantlets had a disturbed development in shoot growth and showed only few, blackened or necrotic brown roots. In contrast, control plants from irradiated soil of the same site Heidgraben (H γ) showed undisturbed shoot and root growth and developed new light brown roots (Fig. 1a, e). Fine roots were analyzed microscopically to select fungal infection sites in tissue parts with cortex cell alterations like necrosis and blackening. In particular, small clusters of light brown or clearly necrotic cells with CF structures were selected for further analyses (Fig. 1b–d).
Procedure of sample selection: a Exemplary basic plant material of M26 apple plants, harvested after four weeks of cultivation in untreated ARD-affected soil, site Heidgraben (H ARD ut), with root necroses and reduction in growth (a, left), in gamma-irradiated soil (H ARD-γ) with bright newly formed roots and healthy growth (a, right), b close-up of diseased roots, c and d preselection of root segments for necroses and blackening detected by bright-field microscopy. Infected brown cells scattered in the rhizodermis (c, Ruthe ARD ut, after 28 days of cultivation), close-up of necrotic cell clusters, best suited for punching (d, Ellerhoop ARD ut) and e healthy appearing part of a fine root (Ruthe ARD ut, after 25 days of cultivation). f Segment with cauliflower-like (CF) structures in early developmental stages and g segment with mycelium (arrows) and CF structures
Applying the punching method to obtain tissue samples from whole mounts, entirely necrotic areas with many CF structures were omitted to avoid the contamination by secondary colonizers. Instead, sites with early stages of infection were preferred (Fig. 1f, g). On the other hand, cryo-sections of infected tissue enabled to clearly select and dissect the CF structures, which were cut off as a series of several adjacent nodules with LMD (Fig. 2). Healthy non-infected samples served as control (Fig. 3).
Harris Uni-Core tissue harvest
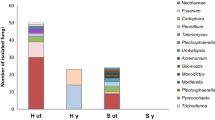
A total of 73 tissue samples taken with the Harris Uni-Core were analyzed by PCR. Of these, 44 were successful for at least one of the two primer pairs (14 in biotest 1, 30 in biotest 3) and 5 samples from γ-irradiated soil (biotest 3) gave no PCR fragment (Table 1 and Table ESM 1). The amplification of HIS gene yielded much more and more diverse results than that of the TEF gene (42 vs. 14). The hits in the BLAST analysis of sequences generated with TEF primers mostly showed Hypocreales sp. (ICMP 13358) with two exceptions (Ilyonectria venezuelensis in sample 319, R-ARD-ut and D. torresensis in sample 335, H-ARD_sf; Table ESM 1). According to HIS primers, Rugonectria rugulosa was identified in 17 samples, D. torresensis in 15 samples and Ilyonectria species in 6 samples (Table 1). They originated from all three ARD soils. In contrast, Fusarium graminearum, Leptosphaeria sp. and Conocephalum sp. were only marginally present in biotest 3 (Table ESM 1). Punching samples from symptom-free roots grown in γ-irradiated ARD soil from the Heidgraben soil gave no PCR fragments with both primer pairs.
Regarding the details of sample collection (Table ESM 1), in 10 of 12 plants which came from untreated soils and were tested with more than one root sample, different fungi could be identified with HIS gene primers. In particular, R. rugulosa and D. torresensis were involved in co-infections. Rugonectria rugulosa was the most widespread as it was detected in both biotests on at least one plant cultivated in untreated ARD soil from each location. Nectriaceae species were mainly found in root segments selected for ARD symptoms, i.e., necrotic and brown areas with CF structures. However, obviously symptom-free root sections of plants from untreated ARD Heidgraben samples also contained D. torresensis, Ilyonectria europaea and R. rugulosa (Table ESM 1, samples 334, 335 and 306), and two symptom-free marked samples from untreated ARD Ellerhoop and Ruthe soil both harbored D. torresensis (Table ESM 1, samples 394 and 396). Four plants grown in γ-irradiated soil from the site Heidgraben had roots showing light browning. The PCR from these samples (Table ESM 1, H-ARD-γs, samples 400, 403, 405 and 406) revealed the presence of D. torresensis, I. robusta, R. rugulosa and F. graminearum.
Laser microdissection cell harvest
The implementation of the same molecular procedure for LMD samples as for Harris samples failed. A set of > 50 samples (including all single-cut dissections) was lost when Chelex was implemented for a DNA stabilization to enable multiple tests as with Harris samples: All PCR products with the HIS primers (also negative controls without fungal material) led to unspecific amplifications which did not contain the expected histone 3 gene fragment after sequencing. Therefore, we used the Phire Plant Direct PCR Kit to identify Nectriaceae fungi in LMD samples. The optimization of the PCR was difficult, since only one attempt per sample was possible and an appropriate positive control was missing. Furthermore, it was not clear whether samples were still suitable for PCR after the cryo-sectioning procedure. Consequently, often a DNA smear was observed on the gel instead of a clear band of the expected size. A total of 53 samples were taken by LMD, but only 20 led to reliable results in fungal identification. The amplification of the fungal genes of interest frequently resulted in only very weak bands. These were excised from the agarose gel and cloned for sequencing (Table 2). All fungi investigated after LMD harvesting were members of the Nectriaceae, namely I. europaea and I. robusta, D. torresensis, Cylindrocladiella sp. and R. rugulosa. Cloning of PCR fragments showed that more than one species could be detected in one sample (17, 18, 19; Table ESM 2). Samples tested with TEF primers (131, 136; Table ESM 2) again gave best hits to Hypocreales sp. (ICMP 13358) in the BLAST analysis.
Discussion
Harris Uni-Core tissue harvest
The easy direct removal of tiny small tissue cubes with the Harris Uni-Core device from selected infection sites enables a quick identification of endophytes, which colonize the respective tissue section. In comparison, the microbiological method of isolating endophytic fungi from surface-sterilized root samples is much more laborious and time-consuming. Sampling with the Harris Uni-Core gains cubes of about 0.5 mm3 volume, which contain rhizodermal and cortical cells of differing size and may harbor various fungi. Therefore, it is recommended to select individual infection sites at an early stage or cluster sites with only few symptomatically altered cortex cells to avoid the detection of secondary invaders. Next, the use of a specific PCR with appropriate primers for the target fungi is essential. This approach has not yet been reported to be used in ARD studies. Moreover, the punching method allows the identification of fungal endophytes that are difficult to cultivate.
Laser microdissection cell harvest
LMD has been applied for some time to different specimens in order to gain insight into particular cell components or single-cell reactions in animal and human tissues (Emmert-Buck et al. 1996). In plant science, this method is gaining more and more attention with increasing fields of investigations, especially combined with molecular genetic tools. Today, LMD is one of the important methods for harvesting selected cells or cell contents to analyze DNA and RNA, e.g., for studies of gene expression on single-cell level or for profiling proteins and metabolites (Hölscher and Schneider 2008; Kerk et al. 2003; Nelson et al. 2006). It has been applied to study stem, inflorescence and leaf tissue (e.g., Abbott et al. 2010; Agustí et al. 2009; Becker et al. 2017; Fang and Schneider 2014; Harrop et al. 2016), as well as to characterize host–pathogen interactions in Arabidopsis thaliana leaves with powdery mildew infections (Chandran et al. 2010). In studies on mycorrhizal colonization root tissue sections of a defined size were cut off laser-based. However, root samples are still mostly prepared as paraffin sections (Balestrini et al. 2007, 2014; Belmondo et al. 2014; Berruti et al. 2013; Campos-Soriano et al. 2011; Corradi and Bonfante 2012; Fiorilli et al. 2016; Giovannetti et al. 2014), a very elaborate preparation method, or—just as cumbersome—after embedding fixed roots in Steedman’s wax (Gomez and Harrison 2009; Huisman et al. 2016). Only Gambetta et al. (2013) used both paraffin sections and unfixed thin cryo-sections for mRNA quantification in LMD-treated grapevine fine root tissue samples when studying mechanisms of water uptake. Ishimaru et al. (2015) substituted the paraffin method by embedding ice-cold 3:1 ethanol:acetic acid fixed samples in 2% carboxymethylcellulose before cryo-sectioning. Since the cryo-sectioning of unfixed fine root segments of apple (shock-frozen in liquid nitrogen, being the standard for animal tissue, e.g., Bölling 2012) failed to produce acceptable thin sections for microscopic evaluation, it was necessary to test other fixation methods. Embedding DMSO-stabilized or acetic acid–ethanol fixed root samples in Tissue-Tek O.C.T. Compound were unsuccessful. After several attempts, the problematic mixed cell structures of the complex root tissue (with cortex and stele) could be preserved in cryo-sections after fixing the samples in a series of acetic acid and DMSO.
In addition, the idea of a laser-based dissection of the CF structures in fine roots of replant-diseased apple was more difficult to realize than expected. As the CF structures are mainly bound to the plant cell wall at the fungal penetration site, it was necessary to cut the plant cell wall as well. The extremely solid plant cell walls of mature cortex tissue even in 5 µm sections were an enormous obstacle for the laser cutting with pulse frequencies of < 50 Hz, which were commonly used in homogeneous animal tissue, or even with a pulse frequency of 964 Hz, which has been adapted for root zone analyses (Gotté et al. 2016). Only by extending the pulse frequency to outstandingly high dimensions up to 5000 Hz, it was possible to separate the area of interest from the surrounding tissue, sometimes only by a second laser-cut. Ultimately, we achieved our LMD goal of obtaining fungal structures from a minute amount of cortex cells, using thin cryo-sections of apple roots with ARD symptoms for further identification with molecular tools (see "Protocol for apple root tissue harvesting with laser microdissection"). To our knowledge, this LMD approach has been successfully applied for the first time to apple roots as well as in the context of ARD.
PCR as the basis for the identification of fungi
Using Chelex for DNA stabilization is cheap and easy (Liu et al. 2015). It allowed the amplification of fragments of both genes of interest (HIS and TEF) for the Harris Uni-Core samples; however, the amplification of the HIS gene fragment failed for two samples, while the TEF primers delivered an amplification. Singh et al. (2018) also reported difficulties of PCR after Chelex application due to inhibitory substances.
Also, for the LMD sections, the optimization of PCR conditions was difficult having only one attempt per sample for testing by direct PCR. Unfortunately, the implementation of Chelex to enable multiple testing failed because this led always to artifacts and thus to a loss of many samples (data not shown). Compared to the tissue cubes taken with the Harris Uni-Core, the very small laser-cut samples posed further challenges to the process. The amplification of fungal genes of interest was often very weak. The fungal chitin rich cell wall is extremely strong and has to be disrupted without harming the genomic DNA (Bir et al. 1995; Moťková and Vytřasová 2012; Yamamoto et al. 2010). With this background, the laser strength might have affected the fungal DNA, and during the microdissection process, the cut plant cells also might release compounds inhibiting the amplification. Nucleases may modify or degrade template DNA (Schrader et al. 2012). Samples gained from apple tissue are known to have a high content of polyphenols and polysaccharides (Green et al. 1999). These substances can impair the PCR as well (Peist et al. 2001; Schrader et al. 2012; Wei et al. 2008). It cannot be excluded that even the high laser temperature could have had a modifying effect on plant tissue ingredients with negative consequences for the analysis of fungal DNA. Despite all encountered difficulties, we were finally able to amplify fungal DNA from 20 LMD samples. This allowed a reliable assignment of the identified fungi to the isolated CF structures.
Identification of Nectriaceae in symptomatic tissue
For the identification of fungi, the HIS primers CYLH3F and CYLH3R (Crous et al. 2004) were reported to be best suited to distinguish between Nectriaceae species (Cabral et al. 2012a; Lawrence et al. 2019). With both collection methods, Harris Uni-Core and LMD, a group of Nectriaceae species was verified in the root tissue areas of interest. These were especially R. rugulosa, D. torresensis and I. robusta, dominating the symptomatic samples irrespective of the site from which the ARD soils were derived. In the LMD samples, focused on fungi in CF structures, Cylindrocladiella sp. and I. europaea were additionally discovered. These results correspond well with isolation tests on roots of apple (Manici et al. 2018; Popp et al. 2019) and roses (Grunewaldt-Stöcker et al. submitted). A similar spectrum of Nectriaceae species has also been reported to cause black foot disease in grapevine (Cabral et al. 2012b; Carlucci et al. 2017; Reis et al. 2013) and black root rot on strawberry and raspberry plants (Weber and Entrop 2017). Since the fungal endophytes causing the CF structures can infect root cells directly, other fungal species such as Conocephalum sp., F. graminearum and Leptosphaeria sp., also detected in our study, are more likely secondary invaders. The markedly larger proportion of Nectriaceae species further supports the assumption of their essential pathogenic role within the ARD complex.
In Harris samples, Nectriaceae species were also detected in symptom-free tissue from untreated ARD soils (Table 1). This could be explained by microscopically undetected small amounts of mycelium in the tissue cube, which might be sufficient to cause amplification. This, in turn, underlines the particular sensitivity of the analysis. Hits on Nectriaceae in root samples with slight browning from γ-irradiated ARD soil raise questions about the effectiveness of soil sterilization. They might be attributed to a survival of thick-walled chlamydospores or CF structures of Nectriaceae followed by re-colonization. Grunewaldt-Stöcker et al. (2019) have already reported on less frequent but existing ARD symptoms of necrosis and blackening in roots grown in γ-irradiated ARD soils. McNamara et al. (2007) described a dose of 10 kGy as a sub-sterilizing dose killing most of the fungi in communities of forest soils; there, it led to a reduction in the number of fungal species detectable by plate counting, but nevertheless some fungi survived. We used gamma irradiation at a minimum dose of 10 kGy to eliminate most of the harmful pathogens in ARD soils. This was largely successful as the plants showed stronger growth compared to those in untreated soil. Same effects on growth were reported for earlier in similar biotests conducted by Mahnkopp et al. (2018) and Yim et al. (2015). Finally, the symptom-free samples from roots grown in irradiated soil proved to be fungus-free (Table ESM1).
A basic question about the precision of molecular analyses arises when considering the different results of fungal identity after colony sequencing: In both, Harris samples (Table ESM 1, Sample 391, R ARD-ut) and LMD samples (Table ESM 2, Samples 17, R ARD-ut; 18, 19, H ARD-ut), even fungi of different genera were detected. This result was most likely due to the fact that for LMD 3–5 dissected cell areas each from one individual root segment had to be combined in one PCR tube to ensure amplification, but this also might indicate co-infections occurring on a small scale. The spectrum of Nectriaceae found in our samples raises several questions about fungal infections in ARD-affected tissue: How can co-infections of diverse species in tiny necrotic areas be explained? Are differing results after colony sequencing due to taxonomic uncertainties in comparisons with gene bank information? Phytopathologists and taxonomists are asked to further work on this important group of pathogens causing ARD symptoms.
Conclusion
The Harris Uni-Core punching method was advantageous for the quick and easy root tissue harvest to determine the fungi contained. The LMD method, on the other hand, is based on individual tissue layers produced by cryo-sectioning and therefore enables the direct collection of defined cell contents or tissue parts down to analyses of selected cells. In the latter case, the processing requires special instruments and a more laborious sample selection, but offers highly specific results for a localized identification of fungi in individual cortex cells of apple roots. Both protocols offer the possibility to identify fungi from selected tiny root areas of interest and can be used for the detection of other microorganisms of apple with other specific primers. This avoids the time-consuming procedures required for the isolation and cultivation of root endophytes and yet uncultivable fungi can be identified. Both methods were used for the first time in ARD studies and were adjusted to the properties of mature apple root tissue. As a result, different Nectriaceae were identified, which form intracellular CF structures during the infection process, resulting in necrosis and blackening of the affected cortex cell area. This confirms their contribution to the induction of ARD. The function of fungal CF structures for the pathogen survival and/or their effect on the host cell should be further investigated in detail.
Change history
26 August 2021
A Correction to this paper has been published: https://doi.org/10.1007/s41348-021-00506-2
References
Abbott E, Hall D, Hamberger B, Bohlmann J (2010) Laser microdissection of conifer stem tissues: isolation and analysis of high quality RNA, terpene synthase enzyme activity and terpenoid metabolites from resin ducts and cambial zone tissue of white spruce (Picea glauca). BMC Plant Biol 10:106. https://doi.org/10.1186/1471-2229-10-106
Agustí J, Merelo P, Cercós M, Tadeo FR, Talón M (2009) Comparative transcriptional survey between laser-microdissected cells from laminar abscission zone and petiolar cortical tissue during ethylene-promoted abscission in citrus leaves. BMC Plant Biol 9:127. https://doi.org/10.1186/1471-2229-9-127
Balestrini R, Gómez-Ariza J, Lanfranco L, Bonfante P (2007) Laser microdissection reveals that transcripts for five plant and one fungal phosphate transporter genes are contemporaneously present in arbusculated cells. Mol Plant Microbe Interact 20:1055–1062. https://doi.org/10.1094/MPMI-20-9-1055
Balestrini R, Nerva L, Sillo F, Girlanda M, Perotto S (2014) Plant and fungal gene expression in mycorrhizal protocorms of the orchid Serapias vomeracea colonized by Tulasnella calospora. Plant Signal Behav 9:e977707. https://doi.org/10.4161/15592324.2014.977707
Becker MG, Zhang X, Walker PL, Wan JC, Millar JL, Khan D, Granger MJ, Cavers JD, Chan AC, Fernando DWG, Belmonte MF (2017) Transcriptome analysis of the Brassica napus-Leptosphaeria maculans pathosystem identifies receptor, signaling and structural genes underlying plant resistance. Plant J 90:573–586. https://doi.org/10.1111/tpj.13514
Belmondo S, Fiorilli V, Pérez-Tienda J, Ferrol N, Marmeisse R, Lanfranco L (2014) A dipeptide transporter from the arbuscular mycorrhizal fungus Rhizophagus irregularis is upregulated in the intraradical phase. Front Plant Sci 5:436. https://doi.org/10.3389/fpls.2014.00436
Berruti A, Borriello R, Lumini E, Scariot V, Bianciotto V, Balestrini R (2013) Application of laser microdissection to identify the mycorrhizal fungi that establish arbuscules inside root cells. Front Plant Sci 4:135. https://doi.org/10.3389/fpls.2013.00135
Bir N, Paliwal A, Muralidhar K, Reddy P, Sarma PU (1995) A rapid method for the isolation of genomic DNA from Aspergillus fumigatus. Prep Biochem 25:171–181. https://doi.org/10.1080/10826069508010119
Bölling A (2012) Kurzzeiteffekte einer intrauterinen Seminalplasmaapplikation auf die Genexpression immun- und follikelreifungsassoziierter Transkripte im weiblichen Genitaltrakt des Schweins. Dissertation, Tierärztliche Hochschule Hannover, pp 33–34
Braun PG (1991) The combination of Cylindrocarpon lucidum and Pythium irregulare as a possible cause of apple replant disease in Nova Scotia. Can J Plant Pathol 13:291–297. https://doi.org/10.1080/07060669109500914
Cabral A, Groenewald JZ, Rego C, Oliveira H, Crous PW (2012a) Cylindrocarpon root rot: multi-gene analysis reveals novel species within the Ilyonectria radicicola species complex. Mycol Progress 11:655–688. https://doi.org/10.1007/s11557-011-0777-7
Cabral A, Rego C, Crous PW, Oliveira H (2012b) Virulence and cross-infection potential of Ilyonectria spp. to grapevine. Phytopathol Mediterr 51:340–354
Campos-Soriano L, Gómez-Ariza J, Bonfante P, San Segundo B (2011) A rice calcium-dependent protein kinase is expressed in cortical root cells during the presymbiotic phase of the arbuscular mycorrhizal symbiosis. BMC Plant Biol 11:90. https://doi.org/10.1186/1471-2229-11-90
Carlucci A, Lops F, Mostert L, Halleen F, Raimondo ML (2017) Occurrence of fungi causing black foot on young grapevines and nursery rootstock plants in Italy. Phytopathol Mediterr 56:10–39. https://doi.org/10.14601/Phytopathol_Mediterr-18769
Caruso FL, Neubauer BF, Begin MD (1989) A histological study of apple roots affected by replant disease. Can J Bot 67:742–749. https://doi.org/10.1139/b89-100
Chandran D, Inada N, Hather G, Kleindt CK, Wildermuth MC (2010) Laser microdissection of Arabidopsis cells at the powdery mildew infection site reveals site-specific processes and regulators. Proc Natl Acad Sci U S A 107:460–465. https://doi.org/10.1073/pnas.0912492107
Corradi N, Bonfante P (2012) The arbuscular mycorrhizal symbiosis: origin and evolution of a beneficial plant infection. PLoS Pathog 8:e1002600. https://doi.org/10.1371/journal.ppat.1002600
Crous PW, Groenewald JZ, Risède J-M, Simoneau P, Hywel-Jones NL (2004) Calonectria species and their Cylindrocladium anamorphs: species with sphaeropedunculate vesicles. Stud Mycol:415–430
Emmert-Buck MR, Bonner RF, Smith PD, Chuaqui RF, Zhuang Z, Goldstein SR, Weiss RA, Liotta LA (1996) Laser capture microdissection. Science 274:998–1001. https://doi.org/10.1126/science.274.5289.998
Fang J, Schneider B (2014) Laser microdissection: a sample preparation technique for plant micrometabolic profiling. Phytochem Anal 25:307–313. https://doi.org/10.1002/pca.2477
Fiorilli V, Belmondo S, Khouja HR, Abbà S, Faccio A, Daghino S, Lanfranco L (2016) RiPEIP1, a gene from the arbuscular mycorrhizal fungus Rhizophagus irregularis, is preferentially expressed in planta and may be involved in root colonization. Mycorrhiza 26:609–621. https://doi.org/10.1007/s00572-016-0697-0
Gambetta GA, Fei J, Rost TL, Knipfer T, Matthews MA, Shackel KA, Walker MA, McElrone AJ (2013) Water uptake along the length of grapevine fine roots: developmental anatomy, tissue-specific aquaporin expression, and pathways of water transport. Plant Physiol 163:1254–1265. https://doi.org/10.1104/pp.113.221283
Geldart GH (1994) The impact of replant problem on the economics of high density apple plantings. Acta Hortic.:11–18. https://doi.org/10.17660/ActaHortic.1994.363.2
Giovannetti M, Tolosano M, Volpe V, Kopriva S, Bonfante P (2014) Identification and functional characterization of a sulfate transporter induced by both sulfur starvation and mycorrhiza formation in Lotus japonicus. New Phytol 204:609–619. https://doi.org/10.1111/nph.12949
Gomez SK, Harrison MJ (2009) Laser microdissection and its application to analyze gene expression in arbuscular mycorrhizal symbiosis. Pest Manag Sci 65:504–511. https://doi.org/10.1002/ps.1715
Gotté M, Bénard M, Kiefer-Meyer M-C, Jaber R, Moore JP, Vicré-Gibouin M, Driouich A (2016) endoplasmic reticulum body-related gene expression in different root zones of Arabidopsis isolated by laser-assisted microdissection. Plant Genome. https://doi.org/10.3835/plantgenome2015.08.0076
Green MJ, Thompson DA, MacKenzie DJ (1999) Easy and efficient DNA extraction from woody plants for the detection of phytoplasmas by polymerase chain reaction. Plant Dis 83:482–485. https://doi.org/10.1094/PDIS.1999.83.5.482
Grunewaldt-Stöcker G, Mahnkopp F, Popp C, Maiss E, Winkelmann T (2019) Diagnosis of apple replant disease (ARD): Microscopic evidence of early symptoms in fine roots of different apple rootstock genotypes. Sci Hort 243:583–594. https://doi.org/10.1016/j.scienta.2018.09.014
Harrop TW, Ud Din I, Gregis V, Osnato M, Jouannic S, Adam H, Kater MM (2016) Gene expression profiling of reproductive meristem types in early rice inflorescences by laser microdissection. Plant J 86:75–88. https://doi.org/10.1111/tpj.13147
Hoestra H (1968) Replant diseases of apple in the Netherlands: Ph.D. thesis. Meded. Landbouwhogesch., Wageningen, the Netherlands, pp 20–24
Hölscher D, Schneider B (2008) Application of laser-assisted microdissection for tissue and cell-specific analysis of RNA, proteins, and metabolites. In: Lüttge U, Beyschlag W, Murata J (eds) Progress in botany, vol 69. Springer, Berlin, pp 141–167
Huisman R, Hontelez J, Mysore KS, Wen J, Bisseling T, Limpens E (2016) A symbiosis-dedicated SYNTAXIN OF PLANTS 13II isoform controls the formation of a stable host-microbe interface in symbiosis. New Phytol 211:1338–1351. https://doi.org/10.1111/nph.13973
Ishimaru T, Ida M, Hirose S, Shimamura S, Masumura T, Nishizawa NK, Nakazono M, Kondo M (2015) Laser microdissection-based gene expression analysis in the aleurone layer and starchy endosperm of developing rice caryopses in the early storage phase. Rice (N Y) 8:57. https://doi.org/10.1186/s12284-015-0057-2
Jaffee BA, Abawi GS, Mai WF (1982) Role of soil microflora and Pratylenchus penetrans in apple replant disease. Phytopathology 72:247. https://doi.org/10.1094/Phyto-72-247
Kelderer M, Manici LM, Caputo F, Thalheimer M (2012) Planting in the ‘inter-row’ to overcome replant disease in apple orchards: a study on the effectiveness of the practice based on microbial indicators. Plant Soil 357:381–393. https://doi.org/10.1007/s11104-012-1172-0
Kerk NM, Ceserani T, Tausta SL, Sussex IM, Nelson TM (2003) Laser capture microdissection of cells from plant tissues. Plant Physiol 132:27–35. https://doi.org/10.1104/pp.102.018127
Lawrence DP, Nouri MT, Trouillas FP (2019) Taxonomy and multi-locus phylogeny of cylindrocarpon-like species associated with diseased roots of grapevine and other fruit and nut crops in California. FUSE 4:59–75. https://doi.org/10.3114/fuse.2019.04.06
Liu L, Wang CL, Peng WY, Yang J, Lan MQ, Zhang B, Li JB, Zhu YY, Li CY (2015) Direct DNA extraction method of an obligate parasitic fungus from infected plant tissue. Genet Mol Res 14:18546–18551. https://doi.org/10.4238/2015.December.28.1
Mahnkopp F, Simon M, Lehndorff E, Pätzold S, Wrede A, Winkelmann T (2018) Induction and diagnosis of apple replant disease (ARD): a matter of heterogeneous soil properties? Sci Hort 241:167–177. https://doi.org/10.1016/j.scienta.2018.06.076
Mai WF, Abawi GS (1981) Controlling replant disease of pome and stone fruit in northeastern United States by preplant fumigation. Plant Dis 65:859–864
Manici LM, Ciavatta C, Kelderer M, Erschbaumer G (2003) Replant problems in South Tyrol: role of fungal pathogens and microbial population in conventional and organic apple orchards. Plant Soil 256:315–324
Manici LM, Kelderer M, Caputo F, Saccà ML, Nicoletti F, Topp AR, Mazzola M (2018) Involvement of Dactylonectria and Ilyonectria spp. in tree decline affecting multi-generation apple orchards. Plant Soil 425:217–230. https://doi.org/10.1007/s11104-018-3571-3
Mazzola M (1998) Elucidation of the microbial complex having a causal role in the development of apple replant disease in Washington. Phytopathology 88:930–938. https://doi.org/10.1094/PHYTO.1998.88.9.930
Mazzola M, Manici LM (2012) Apple replant disease: role of microbial ecology in cause and control. Annu Rev Phytopathol 50:45–65. https://doi.org/10.1146/annurev-phyto-081211-173005
McNamara NP, Griffiths RI, Tabouret A, Beresford NA, Bailey MJ, Whiteley AS (2007) The sensitivity of a forest soil microbial community to acute gamma-irradiation. Appl Soil Ecol 37:1–9. https://doi.org/10.1016/j.apsoil.2007.03.011
Moťková P, Vytřasová J (2012) Comparison of methods for isolating fungal DNA. Czech J. Food Sci. 29:S76-S85. https://doi.org/10.17221/266/2011-CJFS
Nelson T, Tausta SL, Gandotra N, Liu T (2006) Laser microdissection of plant tissue: what you see is what you get. Annu Rev Plant Biol 57:181–201. https://doi.org/10.1146/annurev.arplant.56.032604.144138
Peist R, Honsel D, Twieling G, Löffert D (2001) PCR inhibitors in plant DNA preparations. QIAGEN News 3:7–9
Popp C, Grunewaldt-Stöcker G, Maiss E (2019) A soil-free method for assessing pathogenicity of fungal isolates from apple roots. J Plant Dis Prot 126:329–341. https://doi.org/10.1007/s41348-019-00236-6
Reim S, Siewert C, Winkelmann T, Wöhner T, Hanke M-V, Flachowsky H (2019) Evaluation of Malus genetic resources for tolerance to apple replant disease (ARD). Sci Hort 256:108517. https://doi.org/10.1016/j.scienta.2019.05.044
Reis P, Cabral A, Nascimento T, Oliveira H, Rego C (2013) Diversity of Ilyonectria species in a young vineyard affected by black foot disease. Phytopathol Mediterr 52:335–346
Schrader C, Schielke A, Ellerbroek L, Johne R (2012) PCR inhibitors - occurrence, properties and removal. J Appl Microbiol 113:1014–1026. https://doi.org/10.1111/j.1365-2672.2012.05384.x
Singh UA, Kumari M, Iyengar S (2018) Method for improving the quality of genomic DNA obtained from minute quantities of tissue and blood samples using Chelex 100 resin. Biol Proced Online 20:12. https://doi.org/10.1186/s12575-018-0077-6
Tewoldemedhin YT, Mazzola M, Labuschagne I, McLeod A (2011) A multi-phasic approach reveals that apple replant disease is caused by multiple biological agents, with some agents acting synergistically. Soil Biol and Biochem 43:1917–1927. https://doi.org/10.1016/j.soilbio.2011.05.014
Utkhede RS, Vrain TC, Yorston JM (1992) Effects of nematodes, fungi and bacteria on the growth of young apple trees grown in apple replant disease soil. Plant Soil 139:1–6. https://doi.org/10.1007/BF00012835
Weber RWS, Entrop A-P (2017) Dactylonectria torresensis as the main component of the black root rot complex of strawberries and raspberries in northern Germany. Erwerbs-Obstbau 59:157–169. https://doi.org/10.1007/s10341-017-0343-9
Wei T, Lu G, Clover G (2008) Novel approaches to mitigate primer interaction and eliminate inhibitors in multiplex PCR, demonstrated using an assay for detection of three strawberry viruses. J Virol Methods 151:132–139. https://doi.org/10.1016/j.jviromet.2008.03.003
Weiß S, Liu B, Reckwell D, Beerhues L, Winkelmann T (2017a) Impaired defense reactions in apple replant disease-affected roots of Malus domestica 'M26'. Tree Physiol 37:1672–1685. https://doi.org/10.1093/treephys/tpx108
Weiß S, Bartsch M, Winkelmann T (2017b) Transcriptomic analysis of molecular responses in Malus domestica 'M26' roots affected by apple replant disease. Plant Mol Biol 94:303–318. https://doi.org/10.1007/s11103-017-0608-6
Winkelmann T, Smalla K, Amelung W, Baab G, Grunewaldt-Stöcker G, Kanfra X, Meyhöfer R, Reim S, Schmitz M, Vetterlein D, Wrede A, Zühlke S, Grunewaldt J, Weiß S, Schloter M (2019) Apple replant disease: Causes and mitigation strategies. Curr Issues Mol Biol 30:89–106. https://doi.org/10.21775/cimb.030.089
Yamamoto N, Kimura M, Matsuki H, Yanagisawa Y (2010) Optimization of a real-time PCR assay to quantitate airborne fungi collected on a gelatin filter. J Biosci Bioeng 109:83–88. https://doi.org/10.1016/j.jbiosc.2009.06.015
Yim B, Smalla K, Winkelmann T (2013) Evaluation of apple replant problems based on different soil disinfection treatments—links to soil microbial community structure? Plant Soil 366:617–631. https://doi.org/10.1007/s11104-012-1454-6
Yim B, Winkelmann T, Ding G-C, Smalla K (2015) Different bacterial communities in heat and gamma irradiation treated replant disease soils revealed by 16S rRNA gene analysis: contribution to improved aboveground apple plant growth? Front Microbiol 6:1224. https://doi.org/10.3389/fmicb.2015.01224
Acknowledgements
Our thanks go to Mrs. Ewa Schneider for providing in vitro propagated and acclimatized M26 plants and Mrs. Birgit Milde for technical support in the cloning of PCR amplificates for sequencing, both members of IGPS Leibniz University Hannover. We also thank Mrs. Marion Langeheinrich for her support in cryo-sectioning and Mrs. Ines Blume for her invaluable help in working with the LMD system, both members of the Anatomical Institute of the University of Veterinary Medicine Hannover (TiHo). The cooperation and hospitality of Prof. Dr. med. vet. Christiane Pfarrer offered to GGS and DW during their work at TiHo are highly appreciated. The German Federal Ministry of Research and Education funded this work in the project ORDIAmur (FKZ 031B0512A) within the framework of the BonaRes program.
Funding
The German Federal Ministry of Research and Education funded this work in the project ORDIAmur (FKZ 031B0512A) within the framework of the BonaRes program.
Author information
Authors and Affiliations
Contributions
CP and GGS conceived and designed the experiments. GGS and DW performed the microscopic and histological analyses. CP and DW performed the molecular analyses. CP, DW and EM analyzed the molecular data. EM contributed reagents, materials and analysis tools. CP, GGS, EM and TW contributed to the writing of the manuscript. The authors declare that there is no conflict of interest.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. The research did involve neither human participants nor animals.
Availability of data and material
Not applicable.
Code availability
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to a retrospective Open Access order.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Popp, C., Wamhoff, D., Winkelmann, T. et al. Molecular identification of Nectriaceae in infections of apple replant disease affected roots collected by Harris Uni-Core punching or laser microdissection. J Plant Dis Prot 127, 571–582 (2020). https://doi.org/10.1007/s41348-020-00333-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41348-020-00333-x