Abstract
Acidity is a critical component of the apple fruit consumption experience. In previous biparental family studies, two large-effect acidity QTLs were reported using freshly harvested fruit. Objectives of this study were to determine the number and location of QTLs for acidity variation in a large apple breeding program and ascertain the quantitative effects and breeding relevance of QTL allelic combinations at harvest and after commercially relevant periods of cold storage. Pedigree-connected germplasm of 16 full-sib families representing nine important breeding parents, genotyped for the 8K SNP array, was assessed for titratable acidity at harvest and after 10- and 20-week storage treatments, for three successive seasons. Using pedigree-based QTL mapping software, FlexQTL™, evidence was found for only two QTLs, on linkage groups 16 (the reported Ma locus) and LG 8 (here called Ma3) that jointly explained 66 ± 5% of the phenotypic variation. An additive allele dosage model for the two QTLs effectively explained most acidity variation, with an average of + 1.8 mg/L at harvest per high-acidity allele. The more high-acidity alleles, the faster the depletion with storage, with all combinations appearing to eventually converge to a common baseline. All parent cultivars and selections had one or two of the four possible high-acidity alleles. Each QTL had a rare second high-acidity allele with stronger or reduced effect. Diagnostic SNP markers were identified for QTL alleles derived from distinct sources. Combined QTL effects highlighted utility of the DNA-based information in new cultivar development for targeting desired fruit acidity levels before or after storage.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acidity is a critical component of the apple fruit consumption experience. Apple fruit are popular worldwide, consumed as either fresh or processed products and contributing to human health and well-being (Iwanami et al. 2012). Fruit acidity affects overall apple flavor and the perception of other flavor traits such as sweetness and aroma, therefore influencing consumer satisfaction (Harker et al. 2003; Iwanami et al. 2012). Malic acid is the major organic acid in mature apple fruit (Zhang et al. 2010), and can be measured sensorially by taste or instrumentally by titration or as organic acid content of fruit juice. Commercially, most apple fruit are stored then eaten months to a year after harvest because apples are harvested for only 3 to 4 months yet available to consumers all year round (Watkins 2017). During storage, malic acid content declines (Haynes 1925; Kouassi et al. 2009) and can reach low levels associated with lower satisfaction for some segments of consumers (Harker et al. 2008). However, there is genetic variation for acidity levels before and after storage among common cultivars (Rouchaud et al. 1985; Iwanami et al. 2008) and among germplasm in breeding programs (Ma et al. 2015b; Cliff et al. 2016). An understanding of the components of that genetic variation for fruit acidity would aid development of new apple cultivars that produce fruit with target levels of acidity.
Genetic studies of apple fruit acidity have revealed the presence of major genetic factors. Wellington (1924) described “sweet flavor” (essentially the absence of strong acidity) as a recessive character. By evaluating apple seedlings on a five-point sensory scale from sour to sweet, Crane and Lawrence (1934) ascribed homozygosity or heterozygosity to more than 20 apple cultivars and almost 600 seedlings. Nybom (1959) and Visser and Verhaegh (1978) using independent segregation ratio analysis and analysis of variances, respectively, confirmed the observation that low acidity behaves as a recessive phenotype although it is rarely observed among apple cultivars and selections. Dove and Murphy (1936) and Nybom (1959) suggested that early seedling selection for fruit acidity levels could be conducted using the indirect measure of leaf pH. Visser and Verhaegh (1978) introduced the Malic acid (Ma) locus to explain these observations of relatively simple inheritance.
Quantitative trait loci (QTLs) for apple fruit acidity have been located in the apple genome, especially at the Ma locus. Maliepaard et al. (1998) were the first to map the Ma locus as a morphological marker to the top of linkage group (LG) 16 using a biparental F1 mapping family for which both parents were heterozygous. Using QTL interval mapping with common cultivars as parents, numerous studies reported incomplete dominance of the high-acidity “Ma” allele over the low-acid “ma” allele (King et al. 2000, 2001; Liebhard et al. 2003; Xu et al. 2012; Khan et al. 2013). Liebhard et al. (2003) described a second significant QTL for acidity on LG 8 that explained 46% of the phenotypic variation in a cross between ‘Fiesta’ and ‘Discovery’ that also segregated for Ma (42% of the phenotypic variation). Similarly, Kenis et al. (2008) using the biparental family ‘Telamon’ × ‘Braeburn’ reported significant QTLs for acidity at both loci: in the vicinity of the Ma locus in two consecutive years as well as on LG 8, although the latter was detected in only one of the years. Zhang et al. (2012), using the biparental family ‘Jonathan’ × ‘Golden Delicious,’ reported a LG 8 QTL for malic acid that explained 13.5% of the phenotypic variation (total organic acid content assessed by gas chromatography). Ma et al. (2016) reported QTL at both loci that explained 12% and 14% of the phenotypic variation. Kumar et al. (2012) also reported QTLs for acidity on LG 8 and LG 16 in a genomic selection study with a training population of seven full-sib families. King et al. (2000), using the same mapping population as Maliepaard et al. (1998), reported a QTL for acidity in the vicinity of the Ma locus, with significant influences also on firmness and sensory traits of crispness, juiciness, sponginess, and overall liking. Minor QTLs for acidity were reported on LGs 2, 10, 13, 15, and 17 in the ‘Telamon’ × ‘Braeburn’ family (Kenis et al. 2008). In a cross between ‘Royal Gala’ and the Malus sieversii accession PI 613988, Xu et al. (2012) detected a major QTL at the Ma locus and minor QTLs for acidity on LGs 1 and 6.
Allelic variation in genes of the biochemical pathways underlying fruit acidity biosynthesis and degradation or in genes that affect the transport, storage, and use of acids might underlie the detected QTLs. Several biochemical pathways are involved in the synthesis and metabolism of malate, the precursor of malic acid. In the glycolysis pathway, carboxylation of phosphoenol pyruvate occurs in the cytoplasm, producing oxaloacetate, which is reduced to the volatile compound malate by NAD-dependent malate dehydrogenase (Etienne et al. 2013). Other pathways that could also influence cytoplasmic malate content are as follows: the conversion of malate from pyruvate by NADP-linked malic enzyme in the chloroplast; the conversion to malate of glyoxylate and oxaloacetic acid by malate synthase and NAD-linked malate dehydrogenase, respectively, in the glyoxysome; the conversion of malate from fumarate by fumarase in mitochondria; and the efflux of malate from the vacuole into the cytoplasm (Sweetman et al. 2009). Once fruit are harvested, continued respiration depletes available cellular malate, underlying the typical reduction in acidity throughout the postharvest storage period (Etienne et al. 2013). Bai et al. (2012) provided strong evidence that underlying the Ma locus is an aluminum-activated malate transporter gene (ALMT1) coding for the protein responsible for transporting malic acid molecules from the cytosol to the vacuole where ~ 90% of cell sap is stored. The transport of malate over the vacuole is dependent on the acidity level of the vacuole and membrane potential across the tonoplast (Martinoia et al. 2007). More acidic conditions within the vacuole can lead to a change in the malate’s protonation state, which can further support accumulation, especially if the transporter involved is specific for one form of malate (Martinoia et al. 2007). In plants, the proton pumps V-ATPase and V-PPase are involved in vacuolar acidification (Martinoia et al. 2007). In apple, such genes have been identified, and their involvement in vacuolar acidification and malate accumulation has been confirmed (Hu et al. 2016, 2017; Jia et al. 2018). Coordination of these proton pumps and malate transporters is under research and seems to involve a complex network of regulating and signaling genes that include the MYB genes MdMYB1, MdMYB44, and MdMYB73 (Hu et al. 2016, 2017; Jia et al. 2018) of which MdMYB73 is influenced by, among others, the MdCIbHLH1 gene involved in cold tolerance (Hu et al. 2017).
Knowledge gained from genetic dissection of apple acidity is not yet sufficient for widespread exploitation in breeding. For the most consistently detected trait loci on LG 16 (Ma) and LG 8 (hereby termed Ma3), several unknowns remain. It is unclear whether the QTL effects are maintained across genomic backgrounds of typical breeding germplasm. The high- or low-acidity alleles reported across the various families might be identical-by-descent or represent distinct sources that might differ in quantitative effects. How allelic combinations at the two loci interact to produce genetic potential for fruit acidity is unexplored. Other QTLs might also significantly contribute to acidity genetic potential in wider germplasm than that represented by the biparental family parents. Reports described influences of the loci on freshly harvested fruit, but the more commercially pertinent effects on fruit after storage are not known. Determining the frequency of acidity-associated alleles and their distributions across breeding germplasm would help determine the breeding relevance of significant loci.
Key experimental resources are now in place to characterize the major genetic factors influencing apple fruit acidity at harvest and after storage for diverse breeding germplasm. Genetic improvement in fruit quality via superior new scion cultivars that can satisfy both consumer and industry needs is a primary target of the multi-institutional, US-wide, and international RosBREED project (Iezzoni et al. 2010; www.rosbreed.org) and the participating Washington State University (WSU) Apple Breeding Program (Evans 2013). The pedigree-based analysis (PBA) approach (van de Weg et al. 2004; Bink et al. 2012, 2014) has been established for the aforementioned breeding program in the RosBREED project, enabling efficient detection and validation of segregating alleles in multiple families (Peace et al. 2014). The US apple Crop Reference Set (CR Set), representing alleles of important parents of US apple breeding germplasm, has been established (Peace et al. 2014) and phenotyped for numerous traits, including fruit quality after storage, using standardized protocols (Evans et al. 2012).
The objectives of this study were to determine the number and location of QTLs for acidity variation in a large apple breeding program and ascertain the quantitative effects and breeding relevance of QTL allelic combinations at harvest and after commercially relevant periods of cold storage.
Materials and methods
Breeding germplasm
The germplasm set used was a subset of the US apple Crop Reference Set supplemented with a Breeding Pedigree Set specific to the WSU program and chosen to efficiently represent genome-wide alleles of important breeding parents of the WSU breeding program as described by Peace et al. (2014). Parentage records were verified first using three SSR markers, Md-Exp7SSR (Costa et al. 2008), CH05c06, and Hi04e04 (Silfverberg-Dilworth et al. 2006; Velasco et al. 2010) and then with SNP array markers using procedures described in Pikunova et al. (2013) and van de Weg et al. (2018). The final pedigree-connected germplasm consisted of 16 F1 full-sib families with 200 offspring (Online Resource 1) representing nine important breeding parents (‘Arlet,’ ‘Aurora Golden Gala,’ ‘Cripps Pink,’ ‘Delicious,’ ‘Enterprise,’ ‘Honeycrisp,’ ‘Splendour,’ ‘WA 5,’ and selection W1), 25 of their ancestors, and 18 genetically related cultivars and selections supporting the phasing of marker data in ancestral generations.
Phenotypic data
Crop load management and fruit maturity assessment were conducted according to Evans et al. (2012). Titratable acidity (TA, in mg/L malic acid equivalents) using an Auto-Titrator (Metrohm 815 Robotic USB Sample Processor XL, Metrohm USA, Inc., Riverview, FL, USA) was determined for fruit of the germplasm set over three harvest years with three storage treatments within each year: at harvest with no storage (H); after 10 weeks of cold storage followed by 1 week at room temperature (10wk); and after 20 weeks of cold storage followed by 1 week at room temperature (20wk), as described by Evans et al. (2012). These storage periods were chosen for their commercial relevance. For each assessment, the juice of equally sized portions of preferably five but at least four fruit were pooled. Phenotypic data were checked for errors and statistical outliers by plotting phenotypic correlations among pairs of years within a storage treatment and plotting TA levels over storage within each year. To identify erroneous data within expected bounds for an individual but outside the overriding trend of decreasing TA levels over storage, triplets of data points over the three storage periods for each individual within each year were examined for TA increases of more than 1.5 mg/L from H to 10wk or from 10wk to 20wk. Twenty-one such single deviating data points were removed (Online Resources 2 and 3). Inspection of each individual triplet identified one more individual with an unusually low acidity at 20wk in 2012, for which a data point was then made missing (Online Resource 4). To identify erroneous data across years, triplets across years for each individual at H were examined for cases of one data point being at least 2 mg/L higher or lower in TA than the other two data points, and were subsequently made missing (Online Resource 5). To avoid biases from prior breeding selection for superior performance (Peace et al. 2014), only the phenotypic data of offspring within the full-sib families were used for QTL analyses. The original phenotypic data set prior to the above-described curation is provided in Online Resource 6, together with the curated data set.
Heritability
A linear mixed model (individual or animal model) was fitted to the titratable acidity data using univariate analyses in ASReml software 4.1:
where μ is the overall mean; Si is a fixed year-replicate effect; indj is the random additive effect of individual, with \( \mathrm{ind}\sim \mathrm{NID}\left(0,{\sigma}_a^2A\right) \) where A is the matrix numerator relationship based on genome-wide inbreeding coefficients calculated from pedigree information and where \( \mathrm{NID}\Big(0,{\sigma}_a^2 \)) means normally and independently distributed with mean 0 and variance σ2; fam is the family random effect, with \( \mathrm{fam}\sim \mathrm{NID}\left(0,{\sigma}_f^2\right) \); c(fam) is the random effect of individuals within families, with \( c\sim \mathrm{NID}\left(0,{\sigma}_o^2\right) \); and e is the residual term, with \( e\sim \mathrm{NID}\left(0,{\sigma}_e^2\right) \). Narrow-sense heritability (h2) and broad-sense heritability (H2) were calculated from the following expressions:
where \( {\sigma}_a^2 \) is the additive variance, \( {\sigma}_f^2 \) is the family variance, \( {\sigma}_o^2 \) is the variance component associated with individuals within family replicated over years, \( {\sigma}_e^2 \) is the total residual variance, and D2 and I2are dominance and epistasis effects, respectively, expressed as proportions of total phenotypic variance (Costa e Silva et al. 2004). Total phenotypic variance was calculated from the observed phenotypic data and set equal to the above mentioned variance components as \( {\sigma}_p^2={\sigma}_a^2+{\sigma}_f^2+{\sigma}_o^2+{\sigma}_e^2 \), and the residual variance was calculated as \( {\sigma}_e^2={\sigma}_p^2-{\sigma}_a^2-{\sigma}_f^2-{\sigma}_o^2 \).
Genotypic data
Progenies and available progenitors were genotyped with the International RosBREED SNP Consortium apple 8K SNP array v1, an Infinium™ array (Chagné et al. 2012a). GenomeStudio™ genotyping software was used for marker calling. Next, ASSIsT software (Di Guardo et al. 2015) was used for filtering of SNP markers, whereby markers with recognized null alleles were removed, and to prepare an input file in FlexQTL™ format via software FlexQTL DataPrepper (www.flexqtl.nl). Marker data were curated using marker consistency, double recombination reports, graphical genotyping plots as generated by FlexQTL™ and VisualFlexQTL (Bink et al. 2014, www.flexqtl.nl), and problematic markers were eliminated and double-recombinant singletons were made missing through FlexQTL™ parameter settings. Genetic positions of the filtered 1344 SNP markers for QTL analysis were obtained from an early version of Howard et al. (2017), with an average position difference of 0.33 cM and with any additonal SNPs (such as ss475879082, ss475882553, and ss475883359) genetically positioned by relative physical location. SNP data on ancestors that had not been genotyped within the RosBREED project were complemented where feasible with 20K Infinium™ SNP array (Bianco et al. 2014) data from the FruitBreedomics project (Laurens et al. 2010, 2018) (data not shown). The reported causal SNP for the major phenotypic difference at the Ma locus, Ma1-SNP1455 (Bai et al. 2012), was also used to genotype the germplasm set using Taqman® assay AHCTENE (Chagné et al. 2019); this SNP was used to confirm results from the QTL analyses. Positions of markers on the GDDH13v1.1 reference genome were determined from the Genome Database for Rosaceae (Jung et al. 2014) using BLASTn or, in cases of no hits, through manual examination of this genome sequence through the genome browser at https://iris.angers.inra.fr/gddh13/ “visit the apple genome.”
QTL detection and characterization
FlexQTL™ software, which implements pedigree-based QTL analysis via Markov Chain Monte Carlo (MCMC) simulation (Bink et al. 2002, 2014; www.flexqtl.nl), was used for QTL analyses, where an initial genome-wide analysis was followed by an analysis targeting only the set of LGs that harbored significant and repeatable QTLs. The model parameter settings (Online Resource 7) enabled convergence to be reached. The Bayes factor parameter (2lnBF10) was interpreted as non-significant (0–2), positive (2–5), strong (5–10), or decisive (> 10) evidence for presence of QTLs (Kass and Raftery 1995). Chromosome-wide evidence was collected for NQTL models 1/0 and 2/1 to n ∕ n − 1. QTLs were considered significant and repeatable where the 2lnBF10 for QTLs in the same genomic region for at least most of the nine evaluation points were equal to or greater than 5. For data of each storage period, replicated runs were performed until two runs were obtained that met the effective chain size criteria for convergence. QTL regions were recorded as a series of successive 2 cM bins with 2*ln Bayes factors that were greater than 5. MapChart (Voorrips 2002) and VisualFlexQTL™ (www.flexqtl.nl) software were used to visualize FlexQTL™ output on traces of convergence of QTL models, positions, and genotypes. The mode of a QTL was used as its most probable position. Delineation of QTL regions, QTL genotypes (QQ, Qq, or qq), breeding values, and additive variances associated with the QTL regions were estimated with FlexQTL™, where alleles Q and q refer to alleles associated with high and low phenotypic values, respectively.
The QTL analyses of FlexQTL™ aggregates segregation information across SNPs. It translate this information to IBD probabilities of 2 cM bins (chromosome segments), whereby founder alleles are used as statistical factors. This aggregation of segregation information makes the analyses less sensitive for individual markers having (some) missing data. To facilitate downstream analyses by other software, we imputed missing data for the QTL-associated SNP markers where possible. Original and missing-data-curated genotypic data are provided in Online Resource 6.
The proportion of phenotypic variance explained by each QTL was estimated from FlexQTL™ output of an additive genetic model by dividing the variance explained by the QTL region (AVt1) by the total phenotypic variance for each run replicate (similar to Mangandi et al. (2017) but in that paper the QTL variances were incorrectly weighted by the probability, the proportion of sampled QTL models that upheld the QTL; as the QTL reported there had a probability close to 1, the numerical difference is minimal). These estimates were conducted only where the QTL genotypes of full-sib family parents were identical to consensus QTL genotypes across all years and run replicates determined for the at-harvest data.
The SNPs chosen for haplotyping each QTL were those within a region representing a consensus of the highest posterior densities determined by FlexQTL™ for each storage period-year-run replicate QTL analysis. At the Ma3 QTL, three SNPs (ss475876134, ss475882878, and ss475879095) were removed that had a redundant segregation pattern across all germplasm to other SNPs (ss475882875, ss475878886, and ss475879097, respectively). Haplotypes were constructed across the germplasm for Ma and Ma3 using PediHaplotyper (Voorrips et al. 2016), including automated imputation of missing data, with marker phasing by FlexQTL™.
To examine differences in effect among Q alleles of different origin, offspring were assigned to compound Ma and Ma3 QTL genotypes such that comparisons could be based on individuals with the same genotype for the second acidity QTL. Cumulative frequency distributions for TA content at harvest, using averages for each of the 3 years, were generated and year-pairs of distributions were statistically tested for differences with the two-sample Kolmogorov-Smirnov test (p < 0.05). Families derived from selection W1 were excluded from such quantitative analyses of Ma alleles because of functional diversity detected between its Ma3 alleles. To examine differences in effect among QTL alleles at Ma3 for W1, W1 offspring were categorized according to their SNP haplotype at Ma3. Each of the two sets was further subdivided into three classes according to their FlexQTL™-estimated number of “background” q alleles at Ma and Ma3. Next, TA values were normalized by dividing an individual’s observed TA by the average TA of the offspring with the same number of background Q alleles. Effects of the two W1 haplotypes were then compared across each background using the two-sample Kolmogorov-Smirnov test (p < 0.05). Representation of marker haplotypes in the breeding families was calculated based on FlexQTL™-derived identity-by-descent (IBD) estimates (reported in FlexQTL’s output file “pedimap.ped”).
Dominance levels of each QTL were calculated from the difference between TA content of Qq heterozygotes and the mean of the qq and QQ homozygotes, expressed as its proportion of half the difference in TA content between the two homozygotes. The level of dominance of each QTL was estimated for each storage period and year on subsets of offspring that had the same QTL genotype for the other, non-targeted QTL and that also had at least three individuals representing each of the three QTL genotype classes. In case of inconsistencies, the most prevalent genotype was used. These obtained consensus QTL genotypes were used in each of the nine storage period-year combinations.
Results
Acidity changes over storage and stability over years
In the original phenotypic data set, differences between years in TA at harvest were as high as 4.2 mg/L within the same individuals. After conversion of at-harvest data to missing for 22 offspring deviating by at least 2.0 mg/L for 1 year from the other two, higher correlations between years were achieved with r2 increasing from 0.54–0.66 to 0.65–0.78 (Online Resource 8). After this phenotypic data curation, across all progenies and years, TA decreased by an average of 2.5 mg/L over the entire storage and ripening period (Fig. 1). For most offspring, about half this acidity loss had occurred after the mid-duration storage treatment of 10wk (Fig. 1), such that for each year the three data points for the three storage periods were tight to their regression line (Online Resource 2). Within each family, TA distributions tended to be continuous at harvest, and the prevalence of discontinuous distributions increased with storage period (Online Resource 9).
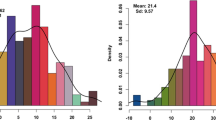
Boxplot distributions of titratable acidity measured for each of 3 years (2010–2012) at three storage durations (H = at harvest with no storage; 10wk = after 10 weeks of cold storage followed by 1 week at room temperature; 20wk = after 20 weeks of cold storage followed by 1 week at room temperature) on 243 apple germplasm individuals representing nine important breeding parents (listed in Table 2)
Broad- and narrow-sense heritability estimates decreased with storage duration, from 0.87 and 0.84 respectively at harvest to 0.63 and 0.22 for 20wk (Table 1). The calculated variance components of the broad-sense heritability associated with family and individuals within families were minor for H and 10wk, thus supporting use of an additive genetic model in QTL analyses. However, for 20wk, these two variance components and additive variance had similar contributions and each had large standard errors.
QTLs detected: Ma and Ma3
For each storage treatment and year combination, two to six simulations with different seeds were needed to obtain full convergence in two of those simulations that were then designated as two replicate runs. At harvest, two QTLs for fruit acidity with decisive statistical evidence (2lnBF10 > 30) were detected at LG 8 and LG 16 in each year and with each replicative run, and no further significant QTLs were found (Online Resource 10). Genetic positions of the mode of each of these two QTLs varied by approximately 5 cM among years (Fig. 2) and were considered to represent the same QTL. The position mode of the LG 8 QTL was at 28 cM for two of the years (2010 and 2012), and the frequency by which this position occurred in the sampled QTL models dominated that of any other position. In 2011, the QTL interval was more dispersed but included the consensus position. For the LG 16 QTL in 2010 and 2011, FlexQTL™ results suggested it was positioned in the first 4 cM of the chromosome, whereas for 2012 it tended to be detected in the 6–8 cM bin. These LG 16 QTL positions coincided with the genetic position of the previously reported Ma (Maliepaard et al. 1998; Liebhard et al. 2003; Khan et al. 2013). Furthermore, two SNPs in the 6–8 cM bin, ss475881682 and ss475876558, flanked the aluminum-activated malate transporter gene ALMT1 (MDP0000252114, Online Resource 11), previously determined to likely represent the gene underlying Ma (Bai et al. 2012; Khan et al. 2013; Ma et al. 2015b). Hereafter, the LG 16 QTL is considered to be Ma. The LG 8 QTL is hereafter referred to as Ma3 (“Ma2” was previously assigned by Bai et al. (2012) to a candidate gene in the vicinity of Ma).
Posterior intensities of acidity QTLs at Ma and Ma3 QTLs along their corresponding linkage groups (LGs) for three storage treatments. Panel a = H treatment, at harvest. Panel b = 10wk treatment (10 weeks of cold storage followed by 1 week at room temperature). Panel c = 20wk treatment (20 weeks of cold storage followed by 1 week at room temperature). The x-axis represents combined genetic locations of LG 16 (blue lines, Ma) and LG 8 (pink lines, Ma3) in centimorgans (cM). The y-axis represents posterior intensities based on an additive genetic model executed by FlexQTL™ software. Increasing color grades correspond to successive years. Solid and dotted lines represent the two run replicates within each year’s runs
QTL analyses for fruit acidity after storage detected QTLs at the same genomic regions as for at harvest, i.e., Ma and Ma3 (Fig. 2), and no additional significant QTLs were found (Online Resource 12). Evidence for the presence of these after-storage QTLs was more variable than for acidity at harvest although the posterior probability remained strong to decisive, except in one case (20wk in 2011) where the evidence was just below the threshold for both run replicates (Online Resource 12).
QTL genotypes of important breeding parents
Genotype estimates revealed that the Q allele representing relatively high-acidity levels was common among breeding parents for Ma and rare for Ma3 (Table 2). For Ma, QTL genotypes for TA at harvest were mostly consistent across years and run replicates (Online Resource 13); two parents were classified as QQ (‘Honeycrisp’ and W1), while the other seven were classified as Qq. For Ma3, QTL genotype estimates for TA at harvest were also mostly consistent across years and run replicates; three were classified as Qq (‘Cripps Pink,’ ‘Enterprise,’ and ‘WA 5’) and the other six were qq (Online Resource 13). Ma and Ma3 genotype estimates for ‘Enterprise’ and W1 were sometimes inconclusive at 10wk or 20wk. Consistent after-storage QTL genotype estimates for those two parents could not be identified from alternative parameter settings nor additional data curation considerations. The final interpretation of QTL genotypes for each of the nine parents (Table 2) was that determined from the at-harvest results. Although zero to four Q alleles were possible within any individual across the two loci, all parent cultivars and selections had only one (‘Delicious,’ ‘Arlet,’ ‘Aurora Golden Gala,’ and ‘Splendour’) or two (all others) Q alleles.
Sources, associated SNPs, and effects of QTL alleles
Ancestral origins of the parental Q and q alleles were successfully traced via SNP haplotypes spanning the QTLs (Table 3). For Ma, each parent of the full-sib families had one or two Q alleles that were traced through the pedigree to eight or nine ancestors: ‘Winesap’ ‘Frostbite,’ ‘Duchess of Oldenburg,’ NJ 27, NJ 136055, perhaps F2-26829-2-2, and the unknown paternal parents of ‘Golden Delicious,’ ‘Jonathan,’ and ‘Lady Williams.’ Ancestors ‘Grimes Golden,’ ‘McIntosh,’ and an unknown parent of ‘Delicious’ were the sources of q alleles for the Ma QTL. For Ma3, the Q alleles for high acidity in important breeding parents ‘Cripps Pink,’ ‘Enterprise,’ and ‘WA 5’ were inherited from ancestors ‘Granny Smith,’ ‘McIntosh,’ and F2-26829-2-2, respectively.
Specific SNP markers were indicative of the high-acidity Ma allele. The B-allele of SNP marker ss475882553 was almost always associated with Q-Ma among the parental haplotypes. Only one Q-allele source (‘Duchess of Oldenburg’) lacked this SNP allele, but a unique B-allele at another SNP (ss475881815) was source-specific for this Q allele. The presence of two distinct Q-allele sources for Ma was thus indicated, a common one characterized by ss475882553-B (Q) and a rare one by ss475881815-B (Q2). The B-alleles for those two SNPs were therefore always associated with a Q allele of Ma. For the reported causal SNP of the Ma locus, Ma1-SNP1455 via the Taqman® assay AHCTENE, the G allele was always associated with Ma, both Q and Q2, and correspondingly the A allele was always associated with q-ma.
Some cultivars shared the same SNP haplotype for Ma3 while these haplotypes came from different lineages and were associated with different-effect QTL alleles (Table 3). Haplotypes BMc-Ma3 of ‘Enterprise’ and BGG+DO-ma3 of ‘Honeycrisp’ were identical-by-state (IBS) but not IBD according to available pedigree information, as ‘Enterprise’ inherited its haplotype from ‘McIntosh’ while that of ‘Honeycrisp’ appeared to have arisen from a recombination event in its paternal parent MN 1627 (data not shown). The other case was haplotype CF2-Ma3 of ‘WA 5’ (source: F2-26829-2-2) and haplotype CES-ma3 of ‘Arlet’ (source: ‘Esopus Spitzenburg’). In this case, recent recombination within the haploblock appeared unlikely because haplotype-sharing extended for 36 cM around this region (Online Resource 14). Lack of discrimination by the SNPs in the region also appeared unlikely because the phenomenon remained when using a higher resolution of 58 SNP markers from the 20K Infinium array (Bianco et al. 2014) within the same interval (data not shown). A comparison of harmonized TA data from subsets of offspring that had otherwise identical QTL genotypes (while distinguishing between Q and Q2 at both loci, results not shown) confirmed the statistical significance of the Q/q contrast for the CF2-Ma3 and CGG+DO-ma3 haplotypes (p = 0.048).
The average effect of each Q-dose for each QTL across the 3 years was + 1.8 mg/L (H), + 1.2 mg/L (10wk), and + 0.9 mg/L (20wk). Loss of acidity over storage was greater for genotypes associated with higher acidity at harvest (i.e., acidity loss 4× Q > 3× Q > 2× Q > 1× Q > 0 Q). Extrapolated linear regression lines indicated that, with a hypothetical storage treatment of “40wk” (40 weeks cold storage and 1 week at room temperature), fruit acidity levels converged to a common baseline level that depended on the year (Fig. 3). Loss of acidity over storage was greatest in 2011 for all genotypes, and this year had the lowest extrapolated baseline (Fig. 3c). Averaged over the three observed years, the extrapolated baseline was approximately 1.0 mg/L (Fig. 3d).
Average fruit acidity content of Ma-Ma3 compound genotypes over three storage periods observed for all three fruiting seasons (years) together (a) and each year individually (b–d). Regression lines are extrapolated beyond the experimentally observed storage periods. Compound genotypes are specified by two digits indicating the number of Q alleles for Ma and Ma3 respectively. For example, “12” indicates the presence of one and two doses of the Q allele for Ma and Ma3 respectively. QTL genotypes of offspring estimated with FlexQTL™ were assigned from the at-harvest results and not considering the subsequently determined Q2 alleles (thus, ‘Honeycrisp’ = QQ for Ma and W1 = qq for Ma3 here)
To determine whether the two Ma-Q-allele types, common Q and ‘Duchess of Oldenburg’–derived “Q2,” were associated with consistently significant differences in effect size, comparisons were made among offspring that had these different Q-allele sources but the same genetic background at Ma3. These comparisons were among Ma-Qq, Ma-Q2q, and Ma-qq, with Ma3 always being homozygous qq. A statistically significant difference in TA content was observed (p < 0.005), whereby the rare ‘Duchess of Oldenburg’ Q2 allele was associated with a higher TA content, + 0.5 mg/L at harvest across 3 years, compared to the common, multi-ancestor Q allele (Fig. 4).
Evidence was obtained for the presence in the selection W1 of a Ma3 allele of higher acidity (Q2) with an effect in between that of the common q and Q alleles. Two families had W1 (QQ/qq or QQ/Q2q) as a parent: ‘Cripps Pink’ (Qq/Qq) × W1 and W1 × ‘WA 5’ (Qq/Qq). W1 segregated for its Ma3 q alleles, D-Ma3 and F-ma3, which were respectively inherited without recombination by 13 and 29 offspring. Each of the two sets was further subdivided into three classes according to their number of Ma-Q and Ma3-Q alleles. The set of offspring having the D-Ma3 allele had a higher average TA content at H and 10wk with each number of “background” Q alleles and in each year (Online Resource 15). The distributions of the normalized TA values were statistically significantly different at H and 10wk for each year (0.003 < p < 0.047), except for H in 2011 (p = 0.38). The effect averaged + 0.8 mg/L (Online Resource 15), which was less than the effects of + 1.8 mg/L (H) and + 1.2 mg/L (10wk) for the common Ma3-allele. Hence, W1 appeared to segregate for a Q2 allele with an intermediate effect between q and Q.
Combined QTL effects
Phenotypic variance explained by Ma, Ma3, and Ma+Ma3 was 26.9 ± 2.4%, 39.1 ± 0.6%, and 66.0 ± 1.9%, respectively, across the six storage-year-replication combinations for which the estimated parental QTL genotypes were consistent with the consensus genotypes (Online Resources 13 and 16). The explained variance decreased with increasing storage duration for Ma (from 30.2 ± 2.1 for H to 20.5 ± 0.3% for 10wk) and remained stable for Ma3 (38.3 ± 0.3% for H and 40.7 ± 0.5% for 10wk). Estimated dominance levels of Ma and Ma3 for TA content across all years, run replications, and QTL genotypes were 5.8 ± 1.1% and 1.5 ±1.9% respectively.
Considering both QTLs at once, fruit acidity increased with Q-allele dose at each storage period (Fig. 3). A single Q-dose at Ma was associated with a higher TA content than was a single Q-dose at Ma3, adding 2.0 and 0.7 mg/L, respectively, at harvest (across all 3 years) compared to qq genotypes. In the presence of a total of two Q alleles across both loci, offspring heterozygous at both loci had slightly higher or equal (but never lower) TA content than offspring having both Q alleles at Ma (Fig. 3). Comparisons to offspring with both Q alleles at Ma3 could not be made as such germplasm was not present in the dataset. In the presence of a total of three Q alleles, offspring homozygous at Ma3 tended to have a higher TA content than those homozygous at Ma. The effect of both loci being QQ-homozygous could not be estimated due to too few such offspring.
Discussion
Two large-effect QTLs influencing apple fruit acidity levels before and after commercially relevant fruit storage treatment were detected in US breeding germplasm. These QTLs were characterized across breeding families derived from nine parents with common ancestors but also distinct founders. The two QTLs that were consistently detected, Ma on LG 16 and Ma3 on LG 8 (the latter locus being newly named here), represented loci detected previously in biparental family studies. Combined effects of alleles from the two loci highlighted the utility of this DNA-based information in new cultivar development for targeting desired fruit acidity levels before or after storage.
General genetic influences on fruit acidity from harvest through storage
The high heritability estimates for apple fruit acidity at harvest (H2 = 0.87 ± 0.02) and for the 10wk treatment (H2 = 0.73 ± 0.05), largely due to additive variance, across the 16 full-sib families from nine parents and 3 years of evaluation was as expected from previous studies (Baojiang et al. 1995; Maliepaard et al. 1998; Liebhard et al. 2003; Kouassi et al. 2009). These high estimates signified a large opportunity to identify the underlying genetic factors segregating in this germplasm. While it is difficult to harvest apple fruit in a standardized manner from genetically variable populations (Evans et al. 2012; Howard et al. 2018), the high heritability estimates obtained indicate that any discrepancies in determining standardized maturity across trees within and among years were minimal for fruit acidity compared to genetic influences. Kouassi et al. (2009) reported narrow-sense heritabilities for four storage treatments of 0.79 ± 0.01 to 0.81 ± 0.01 for TA on 2207 pedigreed individuals in 29 families from breeding programs of six European countries. Those heritability estimates were therefore stable over storage and were also the highest among numerous fruit quality traits evaluated in that study, in contrast with our study’s decrease in this general genetic signal with storage. The difference between our results and those of the previously mentioned report might be due to differences in phenotyping or statistical methods or storage treatments. More likely, there is a fundamental difference in QTL genotypes and alleles present—for example, several ancestors of Kouassi et al. (2009) were associated with extreme breeding values for acidity—perhaps homozygous QQ for Ma3 and Ma, or having “Q3” alleles associated with extremely high acidity—and germplasm of these types were not represented in the present study. The low additive genetic variance calculated for the 20wk treatment (Table 1) indicated limitations for an additive genetic model to adequately describe the phenotypic data. Reanalyzing the 20wk dataset with a genetic model in FlexQTL™ that allowed for the presence of both additive and dominance effects did not find dominance to be a significant effect for Ma and Ma3 (results not shown). The difference between the univariate analysis results, which indicated significant dominance effects for 20wk, and the FlexQTL™ evidence showing otherwise might be due to dominance effects contributed by many small-effect trait loci other than Ma and Ma3 and thus were not detected by FlexQTL™.
Identities and effects of detected QTLs
The two detected QTLs influencing apple fruit acidity for all storage treatments were the only ones that had statistical evidence. The detection of these two QTLs was consistent with previous reports on biparental families using fruit evaluated only at harvest. Our detected “Ma” locus was very likely that controlled by the aluminum-activated malate transporter ALMT1 gene on LG 16 described by Bai et al. (2012) and Khan et al. (2013), as evidenced by genomic positioning. Our “Ma3” might be identical to the acidity QTL reported on LG 8 by Liebhard et al. (2003), Kenis et al. (2008), Zhang et al. (2012), and Ma et al. (2015b) based on co-localization on aligned genetic linkage maps and/or the GDDH13 reference genome sequence (Online Resource 17).
Effects of the two QTLs detected were maintained across many genomic backgrounds typical of breeding germplasm, providing confidence in breeding consideration of genotypes for these QTLs. Our 9-parent, 16-family germplasm set had a wider range of genetic backgrounds than the biparental families of Liebhard et al. (2003), Zhang et al. (2012), and Ma et al. (2016) (‘Fiesta’ × ‘Discovery,’ ‘Jonathan’ × ‘Golden Delicious,’ and ‘Jiguan’ × ‘Wangshanhong,’ respectively), allowing for more allelic combinations of the two large-effect QTLs and other influencing loci. Our large and similar effects calculated for the Ma and Ma3 QTLs were in agreement with those previous biparental studies. The phenotypic variance explained for each (Ma, 27 ± 2% and Ma3, 39 ± 1%, with a combined effect of 66 ± 2%) was slightly less than Liebhard et al. (2003) reported (42% and 46%, respectively), higher than reported by Ma et al. (2016) (12% and 14%, respectively) and Zhang et al. (2012) (13.5% for Ma3) for their single Ma3. It is noteworthy that QTL effects were calculated to be so high in the present study, indicating that the two QTLs indeed account for the bulk of genetic variation for fruit acidity across a wide range of breeding germplasm. Kenis et al. (2008), using another biparental family (‘Telamon’ × ‘Braeburn’), reported 30–34% of the phenotypic variation being explained by a LG 16 QTL (Ma), similar to above, but only 8% by a LG 8 QTL (Ma3). That lower influence of Ma3 can be now explained by the segregation of a Q2 allele associated with acidity levels not as high as the common Q allele.
Ma is reported to have additive gene action in papers where titratable acidity was used as the measure of fruit acidity (Liebhard et al. 2003), and dominant gene action was reported where pH was used (Maliepaard et al. 1998; Xu et al. 2012; Khan et al. 2013). These conclusions appear to be conflicting, but might be explained by differences in the scales of observation, TA being linear and pH being logarithmic. In QTL discovery approaches that assume additive gene action, such as in our current analysis, the use of TA would therefore be the appropriate measure of acidity. The appropriate QTL analytical model for sensory evaluations of acidity would depend on whether the scale was more aligned with TA or pH. In the study of Jia et al. (2018) where malic acid content was assessed by HPLC, genetic action of the Ma allele varied between full-sib families, being dominant in ‘Jonathan’ × ‘Golden Delicious’ and additive in Malus asiatica ‘Zisai Pearl’ × M. domestica ‘Red Fuji.’ The cause for this variable genetic action is not clear.
Two-allele model and beyond
In QTL modeling, FlexQTL™ assumes the presence of just two functional QTL alleles (Q and q), associated with high and low phenotypic values respectively. These associations are based on phenotypic contrasts among Q and q alleles determined to be different from IBD analysis across the pedigree structure. Observations of extended shared haplotypes across and around the QTL interval and other chromosomal regions (results not shown) indicate a recent shared, still unknown, ancestor, such as for BF2/Jt-Ma from F2-26829-2-2 or the unknown father of ‘Jonathan’ and BWs-Ma from ‘Winesap’ (Table 3). However, as QTL alleles calculated to be functionally equivalent arose from numerous ancestral sources (founders) (Table 3), some of these sources might actually have a different QTL allele. Indeed, two extended shared haplotypes at Ma3 and beyond (CF2-Ma3 and CES-ma3, Online Resource 14) had contrasting Q/q designations. As the contrasting Q/q designations were confirmed by comparing offspring with otherwise identical QTL genotypes (p = 0.01), a mutation from Q to q or vice versa is likely to have occurred and might be of relatively recent origin. In cases such as both C F2-Ma3 and CES-ma3 segregating, diagnostic DNA tests will need to ascertain lineage-specific effects or IBD estimates prior to use in marker-assisted selection.
It appears likely that each of the two QTLs have more than two-allele types, as represented by the Q2 alleles. The higher-than-high-acidity Ma-Q2 allele of ‘Honeycrisp,’ inherited from ‘Duchess of Oldenburg,’ had a distinctive SNP haplotype including a unique allele for the second SNP. The reported causal SNP for the Ma locus, Ma1-SNP1455 (Bai et al. 2012; Chagné et al. 2019) was not able to distinguish the Ma-Q2 from Ma-Q. However, a combination of any two of the three SNPs of Ma1-SNP1455, ss475881815, and ss475882553 would be effective to do so (and to establish a germplasm individual’s Q-allele dosage). ‘Duchess of Oldenburg’ is an old Russian cultivar with origins distinct from the bulk of European and US apple germplasm (Beach et al. 1905), so the unique effect of its allele is not surprising. Assuming that the high-acidity Q allele is ancestral to low-acid q allele contributing to increased animal palatability (Ma et al. 2015a; Duan et al. 2017), the seniority, genesis, and DNA sequence differences of Ma-Q2 and the common Ma-Q are yet to be clarified.
Ma3-Q2 had low representation as it was present in just two full-sib families that also showed distorted segregation. Therefore, its discovery was at risk of being false, being due to the coincidental more than usual presence of phenotypic outliers. Confirmation on it being true comes from its co-localization with a QTL of relatively small effect reported by Kenis et al. (2008). That reported result can now be explained by both parents in that study being Q2q rather than qq, thus having a smaller phenotypic contrast between the QTL’s segregating alleles than would Qq parents. Like W1 in the present study that inherited the Ma3-Q2 allele from ‘McIntosh,’ the mapping family’s parent ‘Telamon’ in Kenis et al. (2008) is also a (direct) ‘McIntosh’ descendant and the allele it inherited from ‘McIntosh’ was determined by us, using FruitBreedomics SNP data (results not shown), to be IBD with the W1 allele. Similarly, the second allele of ‘Telamon’ inherited from ‘Crimson Golden’ through ‘Golden Delicious’ was part of haplotype E-ma3 (Table 3). The existence of multiple degrees of Q-allele effects that were well represented in the present study’s germplasm could explain why FlexQTL™ was not able to reach convergence in its QTL genotype estimates for W1 in some runs for 2010.
Use of the IBD approach (Bink et al. 2014) might be useful to infer QTL genotypes of other breeding parents, ancestors, and founders that are genetically related to our currently examined important breeding parents. Such inferences would help extend QTL genotyping results on specific germplasm to wider germplasm. Comparisons among QTL studies could provide insights into the robustness of QTL effects across genetic backgrounds and environments. For example, the presence of a Ma3-Q2 allele in our study was consistent with another QTL mapping study (Kenis et al. 2008) on germplasm that had a common founder origin of the favorable QTL allele. In addition to confirmations, we also noticed some inconsistencies.
Our data allowed complete QTL genotyping of ‘Jonathan’ and ‘Golden Delicious,’ two major founders in apple breeding worldwide. These founders were predicted to be heterozygous for Ma and homozygous qq for Ma3. In contrast, Ma seemed not to segregate in studies of a ‘Jonathan’ × ‘Golden Delicious’ full-sib family with acidity measured as malate content via HPLC, whereas a QTL was found for Ma3 segregating from ‘Jonathan’ (Zhang et al. 2012; Sun et al. 2015; Jia et al. 2018). Assuming the ‘Jonathan’ used in these studies was true-to-type, and if titratable acidity and HPLC-assessed malate content are equivalent for apple fruit, then the effect of Ma3 might be sensitive to environmental factors and/or genetic background.
Diagnostic markers and causal genes
Two or more of the available SNP markers for each locus would be required to diagnose the alleles present in any individual. For the Ma locus, the SNP marker ss475882553 (GDsnp01588) was predictive for Q/q from ten of the eleven ancestral sources. Its predictiveness is further confirmed by consistency of its QTL genotypes in parents of reported QTL studies on acidity. For example, ‘Braeburn,’ ‘Discovery,’ ‘Fiesta,’ ‘Prima,’ and ‘Telamon’ were reported to be heterozygous at Ma (Maliepaard et al. 1998; Liebhard et al. 2003; Kenis et al. 2008) and were all heterozygous for this marker (in the present dataset or that of FruitBreedomics). Only for ‘Duchess of Oldenburg’ and derived germplasm carrying the Q2 allele from this source was ss475882553 not predictive. However, the nearby SNP marker ss475881815 alone discriminated this rare Q-allele source. Statements on its predictiveness in segregating families require insights on its distance from the actual gene to which this Q2 allele belongs. If this would indeed be the ALMT1 gene of Ma, the physical distance is more than 3 Mb, suggesting a genetic distance of approximately 7 cM. Also, further information on the strength of the currently found association would be required prior to its application on wider germplasm.
The reported causal polymorphism for Ma/ma is based on an intragenic SNP creating a truncated, ineffective protein for the ma allele, providing the basis for a functional diagnostic DNA marker (Ma et al. 2015b; Jia et al. 2018). However, it does not appear that ‘Duchess of Oldenburg’ or its descendants were included in the study that identified that causal mutation, allowing for the possibility that this cultivar carries a different mutation at the Ma locus or in a nearby gene such as the Ma2 gene described by Bai et al. (2012). Because none of the current single SNP markers at the Ma3 locus were predictive for any Q/q source across the germplasm (they were only predictive within families), extrapolating the Q/q status of Ma locus alleles across germplasm requires consideration of multiple SNPs at once, i.e., haplotypes.
Sun identified 194 genes on the GD 1.0 draft genome sequence corresponding to the LG 8 QTL found in ‘Jonathan’ × ‘Golden Delicious,’ and performed some initial candidate gene expression studies. Results from qPCR, complementation, transient expression, and protein studies reported by Jia et al. (2018), continuing the work of Sun et al. (2015), gave insight into the gene networks and signaling pathways involved in vacuolar malic acid content: MdPP2CH inactivated three vacuolar H+-ATPases that serve as proton pumps across the tonoplast (MdVHA-A3, MdVHA-B2, and MdVHA-D2) as the aluminum-activated malate transporter MdALMTII. Also, MdPP2CH was determined to be suppressed by the early auxin response gene, MdSAUR37. Jia et al. (2018) focused on QTL LG 8 co-localizing genes that showed the highest differential segregation between two bulks of high- and low-malate offspring and prioritized genes that were segregating in both the parents ‘Jonathan’ and ‘Golden Delicious,’ despite the LG 8 QTL segregating in ‘Jonathan’ only. These prioritized genes showed a distorted ratio of zygotic genotypes among the 246 randomly chosen offspring of ‘Jonathan’ × ‘Golden Delicious.’ For example, their MdPP2CH marker showed a AA:AG:GG segregation of 82:97:67, a significant deviation from the expected 1:2:1 ratio (p = 0.002), and showed a significant lack of heterozygotes (p = 0.005). The presence of distorted segregation for the LG 8 QTL region in the randomly chosen offspring indicates the presence of selection pressure against certain alleles or zygotes. Depending on how this selection-sensitive trait co-segregated with malate content, the differential segregation between the bulks might have arisen not only from artificial selection for contrasting malate content. This feature might make the bulked segregant analysis approach that was used ineffective for the intended purpose. An alternative, efficient way to identify candidate genes and develop predictive markers for Ma3 might be selective phenotyping and intense genotyping of offspring with recombination in the QTL interval. Segregation distortions in the randomly chosen offspring might indicate the presence of lethal alleles (Gao and van de Weg, 2006). If true, the observed zygotic genotype ratio was consistent with biparental segregation of a single, recessive, lethal allele for a locus located approximately 10 cM from the LG 8 QTL, for which the lethal allele of ‘Jonathan’ was in coupling phase with the Q-allele for malate content (data not shown).
Limitations in this study
This study observed and managed large experimental variation in the phenotypic data. While TA content generally decreased during storage, some samples showed a large increase of up to 3.7 mg/L. Also, individuals could show large differences between years in TA at harvest. Such phenomena have not been reported in previous QTL mapping studies, which might indicate previous ignorance or the presence here of a more than usual level of experimental error. Curation for extreme outliers improved the between-year correlations, although still only to moderate levels (Online Resources 3, 4, 5, and 8), probably leaving considerable experimental variation in the genetic analyses. This residual variation might explain the variability in the observed mode and interval size of the QTLs and in the parental QTL genotype estimates among storage treatments and years (Fig. 2, Online Resource 13). The presence of outliers might raise uncertainty on the true presence of multiple Q alleles, especially for Ma3 as its discovery was based on a low number of offspring. In that respect, it was useful that our findings on Ma3-Q2 were consistent with previous research of Kenis et al. (2008).
Another limitation was in the bi-allelic nature of the QTL models used in QTL mapping. In the presence of multiple Q alleles of different effect, FlexQTL™ could be wrong in its Q vs. q designations. The presence in the germplasm of more than one kind of high-acidity Q allele for Ma and Ma3, with quantitatively differing effects for each, might have contributed to the software’s inability to determine consistent QTL genotype estimates for certain parents and might have led to incorrectly designated alleles for others. For example, the presence of the Ma-Q2 allele in ‘Honeycrisp’ might have hindered FlexQTL™’s ability to accurately designate Q vs. q alleles for that cultivar for both QTLs. This possible issue had only a minor role in our study. A co-factor analyses with SNP ss475882553 as a co-factor representing Ma-Q2 did not affect the previous consensus QTL genotypes—with one exception (Online Resource 18). Previously, W1 was assigned Ma3-qq, whereas in the co-factor analyses results were inconclusive: estimates were inconsistent between years and even within a single simulation (2012-10wk, R2, Online Resource 18). Accounting for the higher-than-high-acidity Ma-Q2 allele increased awareness of QTL genotype assignment issues with W1, which we can now understand from the presence of a less-strong Ma3-Q2 allele in this selection.
While the previous two limitations related to confidence in the discovered QTL models, a third limitation relates to obtaining practical insight from the implications of our QTL model. Under-representation in some parts of the compound QTL genotypes (Online Resources 19 and 20) hampers strong conclusions on the performance of QTLs across wide germplasm. Under-representation is true for most QTL mapping studies on a multi-locus trait. In our study, classes with three to four Q-doses were mostly under-represented. This outcome is readily expected from binomial probability. Assuming regular segregation and a balanced representation of QTL genotypes across the parent pairs, the class of smallest sample size theoretically holds 1/16 (6.3%) and 1/81 (1.2%) of the total population in the case of a bi-allelic or tri-allelic two-locus trait, respectively. With a population size of around 160 phenotyped offspring per dataset such as in our study, such a class would hold 10 and 2 offspring, respectively. In addition, QTL genotypes are usually not equally distributed over parent pairs and sampling errors occur also, increasing chances for genotype classes with no or very low representation. However, because of obtained insights on effects of trait loci, numbers of alleles, and parental QTL genotypes, directed crosses can now be made for future studies to examine the effects of particular compound QTL genotypes.
Q-allele dosage model of apple fruit acidity
The presence of just two major QTLs and the consideration of just two alleles per locus provided a simplified interpretation of fruit acidity genetic control that might lend itself to easy integration into routine breeding decisions. The largely additive genetic variation, compared to relatively low dominance and epistasis, and the similar-sized effects of the two QTLs, as observed for the H and 10wk evaluations, indicated that the dosage of high-acidity alleles could be a simple way to understand effects of the two loci. In this model, as long as high-acidity alleles can be distinguished and counted for any individual, there is no need to consider the particular allelic combinations. Some interesting features arose from this model, displayed in Fig. 3. First, at harvest were observed the largest phenotypic distinctions among Ma-Ma3 compound genotypes, which went along with the highest broad- and narrow-sense heritability estimates (Table 1). Second, the higher the Q-allele dosage the greater the acidity depletion with storage. This phenomenon indicates physiological limitations of fruit acidity maintenance over long periods of storage, and has breeding implications. As organic acids are considered an energy source for fruit metabolism during storage (Etienne et al. 2013), the greater acidity depletion in fruit of cultivars with higher Q-allele dosage might mean that high levels of titratable acidity are depleted at the expense of other energy reserves. Third, the order of QTL genotypes for TA content remained the same during the examined storage period: no crossing regression lines. Therefore, fruit acidity after storage appears to be a function of acidity at harvest, which itself could be explained as a function of Q-allele dosage.
The apparent baseline of fruit acidity reached after extended storage of fruit that was indicated by extrapolating beyond the experimentally observed results for each compound genotype has important implications, if true. If indeed TA decreases linearly over time in storage, all genotypes were predicted to eventually reach the same level after about 9–12 months, with QTL genotype no longer influencing fruit acidity. Fluctuations over the 3 years in the rate of loss and storage time to reach the baseline might be due to particular pre-harvest conditions experienced each year by the trees, as they were all grown at the same location. These observations imply that TA measured at harvest for a set of cultivars representing several Q-allele dosages could be used to predict acidity loss patterns throughout the subsequent storage season. Further study is warranted to investigate whether fruit acidity decreases linearly with time in storage, and, if so, if the arising predictive relationship that takes advantage of new genetic information on Q-allele dosage holds up in evaluations of fruit stored for durations between 2 and 12 months and in germplasm grown at other locations under other conditions over several years. It would also be valuable to identify pre-harvest factors that influence initial acidity levels and final baselines both for prediction purposes and to inform management practices that could help achieve fruit within target acidity levels for as long as possible to extend market life.
Breeding targets for Q-allele dosage
Our results indicate that breeders can select for particular Q-allele dosages to achieve a particular desired level of high acidity after storage—such as doses of 3× and 4× to achieve fruit with 5 mg/L TA after a 20wk treatment (Fig. 3a). Similarly, the results indicate that, to avoid acidity being too low, particular Q-allele dosages should be avoided—such as 0× or 1× doses to avoid fruit with less than 3.5 mg/L TA after a 20wk treatment (Fig. 3a). As acidity levels of > 5 mg/L are generally considered desirable for apple fruit (Harker et al. 2008) and most apple fruit consumed are subject to several months of storage, a breeding target of 3–4 Q alleles might appear to be feasible. However, there are several arguments against that strategy. First, the acidity associated with such fruit is expected to be unpalatable at harvest. Second, the relatively rapid depletion of acidity in storage of these highest Q-allele dosages would be expected to be associated with an inconsistent perceived flavor over time, which could confuse consumers. Third, the rapid depletion of acidity would be expected to be associated with the shortest time within the desired range, and thus the shortest market life. Indeed, breeders appear to have avoided such a strategy as all nine important breeding parents in the present study carried only one or two Q alleles. And, from our observations over wider germplasm, few if any cultivars and selections, from heritage to modern, appear to have more than two Q alleles. Besides too many Q alleles, zero Q alleles with its associated bland flavor also appear to have been completely avoided in apple cultivar development. Because of the high frequency and multiple sources of Q and q alleles for both QTLs, there are numerous ways that breeders are able to achieve the desired number of Q alleles in new cultivar development. Fine-tuning of the Q-allele dosage model to account for some epistasis between the two loci and effects of alternative (Q2) alleles could further improve the explanatory power of the Q-allele dosage model.
Data archiving statement
Genotypic data (1344 filtered SNP markers) of the multi-family germplasm set investigated has been submitted to the Genome Database for Rosaceae (www.rosaceae.org).
References
Bai Y, Dougherty L, Li M, Fazio Z, Cheng L, Xu K (2012) A natural mutation-led truncation in one of the two aluminum-activated malate transporter-like genes at the Ma locus is associated with low fruit acidity in apple. Mol Gen Genomics 287:663–678
Baojiang L, Guiyong L, Shixi J (1995) Studies on the inheritance and selection of sweetness and acidity of apples. Acta Hortic 403:233–240
Beach SA, Booth NO, Taylor OM (1905) Apples of New York (Vol II) State of New York – Department of Agriculture. J. B. Lyon Company, New York, pp 150–152
Bianco L, Cestaro A, Sargent DJ, Banchi E, Derdak S, Di Guardo N, Salvi S, Viola R, Gut I, Laurens F, Chagné D, Velasco R, van de Weg E, Troggio M (2014) Development and validation of a 20K Single Nucleotide Polymorphism (SNP) whole genome genotyping array for apple (Malus × domestica Borkh). PLoS ONE 9: e110377
Bink MCAM, Uimari P, Sillanpaa J, Janss LLG, Jansen RC (2002) Multiple QTL mapping in related plant populations via a pedigree-analysis approach. Theor Appl Genet 104:751–762
Bink MCAM, Totir LR, ter Braak CJF, Winkler CR, Boer MP, Smith OS (2012) QTL linkage analysis of connected populations using ancestral marker and pedigree information. Theor Appl Genet 124:1097–1113
Bink MCAM, Jansen J, Madduri M, Voorrips RE, Durel C-E, Kouassi AB, Laurens F, Mathis F, Gessler C, Gobbin D, Rezzonico F, Patocchi A, Kellerhals M, Boudichevskaia A, Dunemann F, Peil A, Nowicka A, Lata B, Stankiewicz-Kosyl M, Jeziorek K, Pitera E, Soska A, Tomala K, Evans KM, Fernández-Fernández F, Guerra W, Korbin M, Keller S, Lewandowski M, Plocharski W, Rutkowski K, Zurawicz E, Costa F, Sansavini S, Tartarini S, Komjanc M, Mott D, Antofie A, Lateur M, Rondia A, Gianfranceschi L, van de Weg WE (2014) Bayesian QTL analyses using pedigreed families of an outcrossing species, with application to fruit firmness in apple. Theor Appl Genet 127:1073–1090
Chagné D, Crowhurst RN, Troggio M, Davey MW, Gilmore B, Lawley C, Vanderzande S, Hellens RP, Kumar S, Cestaro A, Velasco R, Main D, Rees J, Iezzoni A, Mockler T, Wilhelm L, van de Weg E, Gardiner SE, Bassil N, Peace C (2012a) Genome-wide SNP detection, validation, and development of an 8K SNP array for apple. PLoS One 7:e31745
Chagné D, Krieger C, Rassam M, Sullivan M, Fraser J, André C, Pindo M, Troggio M, Gardiner SE, Henry RA, Allan AC, McGhie TK, Laing WA (2012b) QTL and candidate gene mapping for polyphenolic composition in apple fruit. BMC Plant Biol 12:12
Chagné D, Vanderzande S, Kirk C, Profitt N, Weskett R, Gardiner SE, Peace CP, Volz RK, Bassil NV (2019) Validation of SNP markers for fruit quality and disease resistance loci in apple (Malus × domestica Borkh.) using the OpenArray platform. Hortic Res (in press)
Cliff MA, Stanich K, Lu R, Hampson CR (2016) Use of descriptive analysis and preference mapping for early-stage assessment of new and established apples. J Sci Food Agric 96:2170–2183
Costa e Silva J, Borralho NMG, Potts BM (2004) Additive and non-additive genetic parameters from clonally replicated and seedling progenies of Eucalyptus globulus. Theor Appl Genet 108:1113–1119
Costa F, van de Weg WE, Stella S, Dondini L, Pratesi D, Musacchi S, Sansavin S (2008) Map position and functional allelic diversity of Md-Exp7, a new putative expansin gene associated with fruit softening in apple (Malus × domestica Borkh.) and pear (Pyrus communis). Tree Genet Genomes 4:575–586
Crane MB, Lawrence WJC (1934) Genetical studies in cultivated apples. J Genetics 265–296
Daccord N, Celton JM, Linsmith G, Becker C, Choisne N, Schijlen E, van de Geest H, Bianco L, Muranty H, Micheletti D, Di Pierro EA, Gouzy J, Gaillard S, Velasco R, Durel CE, Rees DJG, Laurens F, Lespinasse Y, Quesneville H, Aubourg S, Weigel D, van de Weg E, Troggio M, Bucher E (2017) High-quality de novo assembly of the apple genome and methylome dynamics of early fruit development. Nat Genet 49:1099–1106
Di Guardo M, Micheletti D, Bianco L, Koehorst-van Putten HJ, Longhi S, Costa F, Aranzana MJ, Velasco R, Arús P, Troggio M, van de Weg EW (2015) ASSIsT: an automatic SNP scoring tool for in- and outbreeding species. Bioinformatics 31:3873–3874
Di Pierro EA, Gianfranceschi L, Di Guardo M, Koehorst - van Putten HJJ, Kruisselbrink JW, Longhi S, Troggio M, Bianco L, Muranty H, Pagliarani G, Tartarini S, Letschka T, Lozano-Luis L, Garkava-Gustavsson L, Micheletti D, Bink MCAM, Voorrips RE, Aziz E, Velasco R, Laurens F, van de Weg E (2016) A highdensity, multi-parental SNP genetic map on apple validates a new mapping approach for outcrossing species. Hortic Res 3:16057
Dove WF, Murphy E (1936) The vitamin C-content of apples and its relation to human welfare. Science 83:325–327
Duan N, Bai Y, Sun H, Wang N, Ma Y, Li M, Wang X, Jiao C, Legall N, Mao L, Wan S, Wang K, He T, Feng S, Zhang Z, Mao Z, Shen X, Chen X, Jiang Y, Wu S, Yin C, Ge S, Yang L, Jiang S, Xu H, Liu J, Wang D, Qu C, Wang Y, Zuo W, Xiang L, Liu C, Zhang D, Gao Y, Xu Y, Xu K, Chao T, Fazio G, Shu H, Zhong G-Y, Cheng L, Fei Z, Chen X (2017) Genome re-sequencing reveals the history of apple and supports a two-stage model for fruit enlargement. Nature Communications 8(1):249. https://doi.org/10.1038/s41467-017-00336-7
Etienne A, Génard M, Lobit P, Mbeguié-A-Mbéguié D, Bugaud C (2013) What controls fleshy fruit acidity? A review of malate and citrate accumulation in fruit cells. J Exp Bot 64:1451–1469
Evans K (2013) Apple breeding in the Pacific Northwest. Acta Hort 976:75–78
Evans K, Guan Y, Luby J, Clark M, Schmitz C, Brown S, Orcheski B, Peace C, van de Weg E, Iezzoni A (2012) Large-scale standardized phenotyping of apple in RosBREED. Acta Hortic 945:233–238
Gao Z, van de Weg WE (2006) The Vf gene for scab resistance in apple is linked to sub-lethal genes. Euphytica 151:123–132
Harker FR, Lau K, Gunson FA (2003) Juiciness of fresh fruit: a time-intensity study. Postharvest Biol and Technol 29:55–60
Harker FF, Kupferman EM, Marin AB, Gunson FA, Triggs FM (2008) Eating quality standards for apples based on consumer preferences. Postharvest Biol Technol 50:70–78
Haynes D (1925) Chemical studies in the physiology of apples I. Change in the acid content of stored apples and its physiological significance. Ann Bot 1:77–96
Howard NP, van de Weg E, Bedford DS, Peace C, Vanderzande S, Clark MD, Teh SL, Cai L, Luby JJ (2017) Elucidation of the ‘Honeycrisp’ pedigree through haplotype analysis with a multi-family integrated SNP linkage map and a large apple (Malus × domestica) pedigree-connected SNP data set. Hortic Res 4:17003
Howard NP, van de Weg E, Tillman J, Tong CBS, Silverstein KAT, Luby J (2018) Two QTL characterized for soft scald and soggy breakdown in apple (Malus × domestica) through pedigree-based analysis of a large population of interconnected families. Tree Genet Genomes 14:2
Hu DG, Sun CH, Ma QJ, You CX, Cheng L, Hao YJ (2016) MdMYB1 regulates anthocyanin and malate accumulation by directly facilitating their transport into vacuoles in apples. Plant Physiol 170:1315–1330
Hu DG, Li YY, Zhang QY, Li M, Sun CH, Yu JQ, Hao YJ (2017) R2R3-MYB transcription factor MdMYB73 is involved in malate accumulation and vacuolar acidification in apple. Plant J 91:443–454
Iezzoni A, Weebadde C, Luby J, Chengyan Y, van de Weg E, Fazio G, Main D, Peace C, Bassil N, McFerson J (2010) RosBREED: enabling marker-assisted breeding in Rosaceae. Acta Hort 859:389–394
Iwanami H, Moriya S, Kotoda N, Takahashi S, Abe K (2008) Storability in cold temperatures can be evaluated based on changes in fruit quality in apple genotypes under shelf life conditions. HortScience 43:655–660
Iwanami H, Moriya S, Kotoda N, Mimida N, Takahashi-Sumiyoshi S, Abe K (2012) Mode of inheritance in fruit acidity in apple analysed with a mixed model of a major gene and polygenes using large complex pedigree. Plant Breed 131:322–328
Jia D, Shen F, Wang Y, Wu T, Xu X, Zhang X, Han Z (2018) Apple fruit acidity is genetically diversified by natural variations in three hierarchical epistatic genes: MdSAUR37, MdPP2CH and MdALMTII. Plant J 95:427–443
Jung S, Ficklin SP, Lee T, Cheng C-H, Blenda A, Zheng P, Yu J, Bombarely A, Cho I, Ru S, Evans K, Peace C, Abbott AG, Mueller LA, Olmstead MA, Main D (2014) The Genome Database for Rosaceae (GDR): year 10 update. Nucleic Acids Res 42:D1237–D1244
Kass RE, Raftery AE (1995) Bayes factors. J Am Stat Assoc 90(430):773–795
Kenis K, Keulemans J, Davey M (2008) Identification and stability of QTLs for fruit quality traits in apple. Tree Genet Genomes 4:647–661
Khan SA, Beekwilder J, Schaart JG, Mumm R, Soriano JM, Jacobsen E, Schouten HJ (2013) Differences in acidity of apples are probably mainly caused by a malic acid transporter gene on LG16. Tree Genet Genomes 9:475–487
King GJ, Maliepaard C, Lynn JR, Alston FH, Durel C-E, Evans KM, Griffon B, Laurens F, Manganaris AG, Schrevens E, Tartarini S, Verhaegh J (2000) Quantitative genetic analysis and comparison of physical and sensory descriptors relating to fruit flesh firmness in apple (Malus pumila Mill.). Theor Appl Genet 100:1074–1084
King GJ, Lynn JR, Dover CJ, Evans KM, Seymour GB (2001) Resolution of quantitative trait loci for mechanical measures accounting for genetic variation in fruit texture of apple (Malus pumila Mill.). Theor Appl Genet 102:1227–1235
Kouassi AB, Durel CE, Fabrizio C, Tartarini S, Van de Weg E, Evans K, Fernandez-Fernandez F, Govan C, Boudichevskaja A, Dunemann F, Antofie A, Lateur M, Stankiewicz-kosyl M, Soska A, Tomala K, Lewandowski M, Rutkovski K, Zurawicz E, Walter G, Laurens F (2009) Estimation of genetic parameters and prediction of breeding values for apple (Malus x domestica) fruit quality traits using pedigreed plant material in Europe. Tree Gen Genomes 5:659–672
Kumar S, Chagne D, Bink MC, Volz RK, Whitworth C, Carlisle C (2012) Genomic selection for fruit quality traits in apple (Malus x domestica Borkh.). PLoS One 7:e36674
Laurens F, Durel CE, Patocchi A, Peil A, Salvi S, Tartarini S, Velasco R, van de Weg E (2010) Review on apple genetics and breeding programmes and presentation of a new European initiative to increase fruit breeding efficiency. J Fruit Sci 27:102–107
Laurens F, Aranzana MJ, Arus P, Bassi D, Bink M, Bonany J, Caprera A, Corelli-Grappadelli L, Costes E, Durel CE, Mauroux JB, Muranty H, Nazzicari N, Pascal T, Patocchi A, Peil A, Quilot-Turion B, Rossini L, Stella A, Troggio M, Velasco R, van de Weg E (2018) An integrated approach for increasing breeding efficiency in apple and peach in Europe. Hortic Res 5:11
Liebhard R, Gianfranceschi L, Koller B, Ryder CD, Tarchini R, van de Weg WE, Gessler (2002) Development and characterisation of 140 new microsatellites in apple (Malus x domestica Borkh). Mol Breed 10:217–241
Liebhard R, Koller B, Gianfranceschi L, Gessler C (2003) Creating a saturated reference map for the apple (Malus × domestica Borkh.) genome. Theor Appl Genet 106:1497–1508
Ma B, Chen J, Zheng H, Fang T, Ogutu C, Li S, Han Y, Wu B (2015a) Comparative assessment of sugar and malic acid composition in cultivated and wild apples. Food Chem 172:86–91
Ma B, Liao L, Zheng H, Chen J, Wu B, Ogutu C, Li S, Korban SS, Han Y (2015b) Genes encoding aluminum-activated malate transporter II and their association with fruit acidity in apple. Plant Genome 8:0
Ma BQ, Zhao S, Wu BH, Wang D, Peng Q, Owiti A, Fang T, Liao L, Ogutu C, Korban SS, Li S, Han Y (2016) Construction of a high density linkage map and its application in the identification of QTLs for soluble sugar and organic acid components in apple. Tree Genet Genomes 12:1
Maliepaard C, Alston FH, van Arkel G, Brown LM, Chevreau E, Dunemann F, Evans KM, Gardiner S, Guilford P, van Heusden AW, Janse J, Laurens F, Lynn JR, Manganaris AG, den Nijs APM, Periam N, Rikkerink E, Roche P, Ryder C, Sansavini S, Schmidt H, Tartarini S, Verhaegh JJ, Vrielink-van Ginkel M, King GJ (1998) Aligning male and female linkage maps of apple (Malus pumila Mill.) using multi-allelic markers. Theor Appl Genet 97:60–73
Mangandi J, Verma S, Osorio L, Peres NA, van de Weg E, Whitaker VM (2017) Pedigree-based analysis in a multiparental population of octoploid strawberry reveals QTL alleles conferring resistance to Phytophthora cactorum. G3: Genes Genom Genet 7:1707–1719
Martinoia E, Maeshima M, Neuhaus HE (2007) Vacuolar transporters and their essential role in plant metabolism. J Exp Bot 58:83–102
Nybom N (1959) On the inheritance of acidity in cultivated apples. Hereditas 45:332–350
Peace CP, Luby J, van de Weg E, Bink M, Iezzoni AF (2014) A strategy for developing representative germplasm sets for systematic QTL validation, demonstrated for apple, peach and sweet cherry. Tree Genet Genomes 10:1679–1694
Pikunova A, Madduri M, Sedov E, Noordijk Y, Peil A, Troggio M, Bus VGM, van de Weg E (2013) ‘Schmidt’s Antonovka’ is identical to ‘Common Antonovka’, an apple cultivar widely used in Russia in breeding for biotic and abiotic stresses. Tree Genet Genomes 10:261–271
Rouchaud L, Moons C, Meyer JA (1985) Cultivar differences in the influence of harvest date and cold storage on the free sugar and acids contents, and the eating quality of apples. J Hortic Sci Biotech 60:29l–296
Silfverberg-Dilworth E, Matasci CL, van de Weg WE, Van Kaauwen MPW, Walser M, Kodde LP, Soglio V, Gianfranceschi L, Durel CE, Costa F, Yamamoto T, Koller B, Gessler C, Patochhi A (2006) Microsatellite markers spanning the apple (Malus x domestica Borkh.) genome. Tree Genet Genomes 2:202–224
Sun R, Chang YS, Yang FQ, Wang Y, Li H, Zhao YB, Chen DM, Wu T, Zhang XZ, Han ZH (2015) A dense SNP genetic map constructed using restriction site-associated DNA sequencing enables detection of QTLs controlling apple fruit quality. BMC Genomics 16:747
Sweetman C, Deluc LG, Cramer GR, Ford CM, Soole KL (2009) Regulation of malate metabolism in grape berry and other developing fruits. Phytochemistry 70:1329–1344
van de Weg WE, Voorrips RE, Finkers HJ, Kodde LP, Jansen J, Bink MCAM (2004) Pedigree genotyping: a new pedigree-based approach of QTL identification and allele mining. Acta Hortic 663:45–50
van de Weg E, Di Guardo M, Jänsch N, Socquet-Juglard D, Costa F, Baumgartner I, Broggini GAL, Kellerhals M, Troggio M, Laurens F, Durel CE, Patocchi A (2018) Epistatic fire blight resistance QTL alleles in the apple cultivar ‘Enterprise’ and selection X-6398 discovered and characterized through pedigree-informed analysis. Mol Breed 38:5
Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A, Kalyanaraman A, Fontana P, Bhatnagar SK, Troggio M, Pruss D, Salvi S, Pindo M, Baldi P, Castelletti S, Cavaiuolo M, Coppola G, Costa F, Cova V, Dal Ri A, Goremykin V, Komjanc M, Longhi S, Magnago P, Malacarne G, Malnoy M, Micheletti D, Moretto M, Perazzolli M, Si-Ammour A, Vezzulli S, Zini E, Eldredge G, Fitzgerald LM, Gutin N, Lanchbury J, Macalma T, Mitchell JT, Reid J, Wardell B, Kodira C, Chen Z, Desany B, Niazi F, Palmer M, Koepke T, Jiwan D, Schaeffer S, Krishnan V, Wu C, Chu VT, King ST, Vick J, Tao Q, Mraz A, Stormo A, Stormo K, Bogden R, Ederle D, Stella A, Vecchietti A, Kater MM, Masiero S, Lasserre P, Lespinasse Y, Allan AC, Bus V, Chagné D, Crowhurst RN, Gleave AP, Lavezzo E, Fawcett JA, Proost S, Rouzé P, Sterck L, Toppo S, Lazzari B, Hellens RP, Durel CE, Gutin A, Bumgarner RE, Gardiner SE, Skolnick M, Egholm M, van de Peer Y, Salamini F, Viola R (2010) The genome of the domesticated apple (Malus × domestica Borkh.). Nat Genet 42:833–839
Visser T, Verhaegh JJ (1978) Inheritance and selection of some fruit characters of apple. I. Inheritance of low and high acidity. Euphytica 27:753–760
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Voorrips RE, Bink MCAM, Kruisselbrink JW, Koehorst-van Putten HJJ, van de Weg WE (2016) PediHaplotyper: software for consistent assignment of marker haplotypes in pedigrees. Mol Breed 36:119
Watkins CB (2017) Advances in post-harvest storage and handling of apples. In: Evans K (ed) Achieving sustainable cultivation of apples. Burleigh Dodds, Cambridge, pp 337–370
Wellington R (1924) An experiment in breeding apples. II. - New York State Agr. Exp. Sta. Techn. Bull. No. 106. 149 pp
Xu K, Wang A, Brown S (2012) Genetic characterization of the Ma locus with pH and titratable acidity in apple. Mol Breed 30:899–912
Zhang YZ, Li PM, Cheng LL (2010) Developmental changes of carbohydrates, organic acids, amino acids, and phenolic compounds in ‘Honeycrisp’ apple flesh. Food Chem 123:1013–1018
Zhang Q, Ma BQ, Li H, Chang Y, Han Y, Li J, Wei G, Zhao S, Khan MA, Zhou Y, Gu C, Zhang X, Han Z, Korban SS, Li S, Han Y (2012) Identification, characterization, and utilization of genome-wide simple sequence repeats to identify a QTL for acidity in apple. BMC Genomics 13:537
Acknowledgements
Dr. Luis F. Osorio provided helpful guidance with ASReml statistical analysis, and Lichun Cai and Stijn Vanderzande helped curate raw SNP data. Jack Klipfel conducted laboratory analyses for Ma1-SNP1455. Some genetic data and technical expertise came from the FruitBreedomics project no 265582: “Integrated approach for increasing breeding efficiency in fruit tree crops” (www.fruitbreedomics.com), which was co-funded by the EU seventh Framework Programme.
Funding
This work was funded by the USDA’s National Institute of Food and Agriculture (NIFA)–Specialty Crop Research Initiative projects “RosBREED: Enabling marker-assisted breeding in Rosaceae” (2009-51181-05808) and “RosBREED: Combining disease resistance with horticultural quality in new rosaceous cultivars” (2014-51181-22378), and USDA NIFA Hatch projects 0211277 and 1014919.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by D. Chagné
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Online Resource 1
Full-sib families used in this study, with complete SNP data, phenotypic data, and verified pedigrees. Families 1–4, 10–11, and 13 were part of RosBREED’s Crop Reference Set as described in Peace et al. (2014); compared to numbers in that previous study, the number of offspring in all of those families except 1 and 4 were approximately doubled by including WSU’s Breeding Pedigree Set here. Families 6–9 and 15–16 were from the WSU’s Breeding Pedigree Set. The small families of 5, 12, and 14 arose due to unintended paternity for particular offspring originally assigned to one of the other families and were maintained as the genomes of their parents were already well represented by the other families. A 17th family, ‘Aurora Golden Gala’ × ‘Granny Smith’ (intended paternity was ‘Cripps Pink’) with 12 offspring, was excluded because of insufficient representation of one of the homologs of ‘Granny Smith’ (while the other homolog was well represented through ‘Cripps Pink’ offspring) (PDF 15 kb)
Online Resource 2
Usual and unusual changes in fruit acidity observed over three storage and ripening periods, resulting in data curation adjustments. A, B. Most offspring showed a steady decrease in acidity over storage, such as for two shown here. C–F. Some offspring showed an irregular pattern within or among years, up to 3.5 mg/L across a 10-week storage period rather than a decrease (C–E), sometimes preceded by an extraordinary decrease (C, D). Those data points within a 1.5 mg/L change (F) were not converted to missing as they were considered to have a strong possibly of being due to valid experimental variation among samples or between years (PDF 114 kb)
Online Resource 3
Offspring with unexpected large increases in TA after storage in one or more of the three evaluation years. Yellow-marked numbers correspond to a TA increase of more than 1.5 mg/L. Orange-marked numbers are the corresponding data points causing the discrepant increases (PDF 62 kb)
Online Resource 4
Offspring with unexpected low TA after the 20wk storage treatment in one of the three evaluation years. The orange-marked data point was converted to missing (PDF 18 kb)
Online Resource 5
Offspring with large variability in TA at harvest (H storage treatment). Blue-marked data points were made missing because they deviated more than 2 mg/L from the other two data points (PDF 62 kb)
Online Resource 6
Original and curated phenotypic and genotypic data for 200 offspring of 16 F1 full-sib families used in QTL analyses. SNP genotypes in blue cells were imputed (XLSX 59 kb)
Online Resource 7
FlexQTL parameter settings and justifications for values chosen (PDF 105 kb)
Online Resource 8
Correlation plots for TA at harvest across all full-sib family individuals for each pair of years and the associated Pearson’s correlation coefficients before data curation (upper panels) and after data curation (lower panels). Red dots indicate pairs of TA data for which one value was made missing during data curation (PDF 335 kb)
Online Resource 9
TA distributions across families for each of three storage treatments and each of three years (A = 2010, B = 2011, C = 2012). Storage treatments were H (0 weeks of cold storage), 10wk (10 weeks of cold storage and one week at room temperature), and 20wk (20 weeks of cold storage and one week at room temperature). Numbers on the side of each plot represent corresponding sample sizes (PDF 1025 kb)
Online Resource 10
QTL detection for at-harvest acidity. Chromosome-wise 2lnBF values for the NQTL model versus NQTL-1 model (NQTL = 1 or 2) for three years (2010–2012) and two replicative runs per year (R1 and R2). Note that the negative values indicate the favoring of the sparser QTL model. Values ≥5 are in bold. Each combination had two replicates indicated by R1 and R2 (differing in starting seeds for the MCMC simulation) (PDF 48 kb)
Online Resource 11
Genetic position (cM) and physical position of the SNP markers associated with the acidity QTLs on LG 8 and 16, the gene ALMT1 (MDP0000252114) reported to represent Ma (Bai et al. 2012; Ma et al. 2015a, b), and the SSR marker CH01c06 (Liebhard et al. 2002). Genetic positions are according to Howard et al. (2017). Physical coordinates are according to the GDDH13 genome sequence v1.1 (Daccord et al. 2017) (PDF 26 kb)
Online Resource 12
QTL detection for after-storage acidity. Chromosome-wise 2lnBF values for the NQTL model versus NQTL-1 model (NQTL = 1 or 2) for three years (2010–2012) and two replicative runs per year (R1 and R2). Note that the negative values indicate the favoring of the sparser QTL model. Values ≥5 are in bold. Each combination had two replicates indicated by R1 and R2 (differing in starting seeds for the MCMC simulation) (PDF 56 kb)
Online Resource 13
Estimated Ma and Ma3 genotypes of important breeding parents for two replicative simulation runs for each of three storage treatments and each of three years of phenotypic data. QTL genotypes are characterized by their number of Q alleles: 0 (qq), 1 (Qq), and 2 (QQ). inc = inconclusive. Genetic positions refer to the center of 2 cM bins (PDF 422 kb)
Online Resource 14
Haploblock genotypes of ‘Jonathan’ and F2-26829-2-2 for the 16.6–60.7 cM region of LG 8. The shared extended haplotype across multiple successive haploblocks is marked by a box and stretches approximately 36 cM. Haploblock data came from van de Weg et al. (2018), with genetic positions were according to Di Pierro et al. (2016). These aggregate-SNP haploblocks were used here rather than single SNPs for ease of visual description. The outermost flanking SNPs common to this study were ss475879074 of HB-08-22 and ss475882898 of HB-08-56.7. These two markers span 35 cM here and span 33 cM in Howard et al. (2017). Of the 12 SNP markers that represented the LG 8 QTL in this study (Online Resource 11), three are shared with van de Weg et al. (2018), jointly stretching approximately 10.5 cM which distance is consistent with the 10.6 cM in this study (Online Resource 11) (PDF 230 kb)
Online Resource 15
Determination of the effect of the W1 haplotypes at Ma3, derived from ‘McIntosh’ and the father of ‘Golden Delicious’ (alleles D-Ma3 and F-ma3 in Table 2, respectively). Mean TA content (left) and sample sizes (right) are shown for sets of offspring carrying one or the other haplotype and differing for their background Q-allele dose at both loci of Ma and Ma3, for each of three storage periods and three years. The highest average of each data pair is in blue font (PDF 38 kb)
Online Resource 16
Phenotypic variances explained by Ma and Ma3 under an additive genetic model for each of the simulations where the estimated parental QTL genotypes matched the consensus genotypes (PDF 17 kb)
Online Resource 17
Genetic positions and parental sources of LG 8 acidity QTLs reported in the literature (PDF 16 kb)
Online Resource 18
Estimated Ma and Ma3 genotypes of important breeding parents from QTL discovery analyses without (A) and with (B) a co-factor for Ma-Q2 as approximated by SNP genotypes of marker ss475882553. Results are presented for two replicative simulation runs (R1 and R2) with FlexQTL™ on data from each of three treatment-year combinations (H and 10wk storage treatments). QTL genotypes are characterized by their number of Q alleles: 0 (qq), 1 (Qq), and 2 (QQ). inc = inconclusive. Genetic positions refer to the center of 2 cM bins (PDF 85 kb)
Online Resource 19
Overview of germplasm representation of parental haplotypes for the Ma and Ma3 regions. For each haplotype, the first and last two SNPs of the haplotype are presented. The lowest number within rows indicates the representation of the full haplotype (ignoring rare double recombinants within), and is given in bold. Haplotype labels are explained in Table 3 (PDF 34 kb)
Online Resource 20
Numbers of informative meioses across the studied germplasm for Ma and Ma3 under the initial bi-allelic QTL model. Mat = maternal, Pat = paternal (PDF 22 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Verma, S., Evans, K., Guan, Y. et al. Two large-effect QTLs, Ma and Ma3, determine genetic potential for acidity in apple fruit: breeding insights from a multi-family study. Tree Genetics & Genomes 15, 18 (2019). https://doi.org/10.1007/s11295-019-1324-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-019-1324-y