Abstract
Steroidal resource occupies a vital proportion in the pharmaceutical industry attributing to their important therapeutic effects on fertility, anti-inflammatory and antiviral activities. Currently, microbial transformation from phytosterol has become the dominant strategy of steroidal drug intermediate synthesis that bypasses the traditional chemical route. Mycobacterium sp. serve as the main industrial microbial strains that are capable of introducing selective functional modifications of steroidal intermediate, which has become an indispensable platform for steroid biomanufacturing. By reviewing the progress in past two decades, the present paper concentrates mainly on the microbial rational modification aspects that include metabolic pathway editing, key enzymes engineering, material transport pathway reinforcement, toxic metabolic intermediates removal and byproduct reconciliation. In addition, progress on omics analysis and direct genetic manipulation are summarized and classified that may help reform the industrial hosts with more efficiency. The paper provides an insightful present for steroid biomanufacturing especially on the current trends and prospects of mycobacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Steroidal compounds (steroids) are terpenoid lipids with four cycloalkane rings (A-D) as a unique nucleus, which usually harbors a side chain at C-17 (Fig. 1A). They are widespread in nature as important cell membrane components and hormone precursors, which also gives them precious biological resources value. The common structural and stereochemical skeleton of these compounds can be further modified or substituted with different side chain groups, resulting in multiple biological functions (Wollam and Antebi 2011). About three hundred steroid-based drugs have been approved for clinical application, which makes them the second largest category of pharmaceuticals (Donova 2017). Most recently, the cheap and common steroid dexamethasone was found to use in COVID-19 severely infected patients significantly reduced the mortality by about one-third of patients under respiratory assistance (Ledford 2020). Steroid drugs are mainly divided into the categories of sex hormone and adrenocortical hormone analogs, due to their wide range of therapeutic applications, the steroidal drug market demand has become a strong driver for the fast-growing production scale (Fernandez-Cabezon et al. 2018). Steroidal drugs are derived from a variety of steroid intermediates, which contain a gonane core with different side-chains at C-17 as illustrated in Fig. 1A. Examples include androst-4-ene-3,17-dione (AD), androsta-1,4-diene-3,17-dione (ADD), 9α-hydroxy-4-androstene-3,17-dione (9-OH-AD), 22-hydroxy-23,24-bisnorchol-4-ene-3-one (4-HBC), 22-hydroxy-23,24-bisnorchol-1,4-diene-3-one (1,4-HBC), progesterone (PG), and testosterone (TS), among many others.
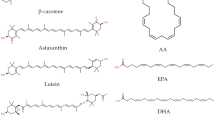
Structures of key steroidal intermediates, pharmaceuticals and phytosterols. A. Steroidal core structure gonane and basic steroidal precursors. B. Diosgenin is an important raw material for chemical synthesis of steroids and some active pharmaceutical ingredients. C. Four main components and structures of phytosterols. AD androst-4-ene-3,17-dione, ADD androsta-1,4-diene-3,17-dione, 9-OH-AD 9α-hydroxy-4-androstene-3,17-dione, 4-HBC 22-hydroxy-23,24-bisnorchol-4-ene-3-one, 1,4-HBC 22-hydroxy-23,24-bisnorchol-1,4-diene-3-one, PG progesterone and TS testosterone
The preparation of steroid intermediates mainly includes chemical methods and microbial transformation. The main raw materials for chemical conversion are diosgenin, a key raw material extracted from Dioscorea species, and the chemically produced 16-dehydropregnenolone acetate (Fig. 1B), as the central intermediates for the synthesis of a large number of steroidal drugs (Baruah et al. 2015; Herraiz 2017) (Fig. 1C). Although chemical synthesis was once absolutely dominant in the field, the limitations of raw material sources and the cumbersome process steps were still difficult to solve (Gupta and Mahajan 2018). In recent decades, the traditional methods have gradually been replaced by biotechnology due to its mild reaction conditions, low cost and environmental friendliness (Giorgi et al. 2019). With the advance of genetic engineering, directed modification of steroids by microbial transformation has become a new trend in recent years.
The main phytosterols in such mixtures, including campesterol, sitosterol, brassicasterol and stigmasterol, are very similar in structure compared with diosgenin and steroidal drugs (Fig. 1D), and have gradually substituted diosgenin as the raw material for the synthesis of steroidal intermediates (Liu et al. 2018a). The phytosterols are abundant, cheap, and easy to obtain from the leftovers of vegetable oil extractions, which makes them a stable and high-quality resource for the production of steroidal drugs (Fernandes and Cabral 2007). Microbial conversion of phytosterols is an effective way to synthesize important steroidal intermediates. Many microorganisms can catabolize steroidal intermediates as carbon sources. Mycobacterium sp. exhibited a superior capability of cholesterol uptake and side-chain degradation, which leads to the accumulation of steroid-based drug intermediates during microbial transformation (Zhao et al. 2021). Compared with other microorganisms, abundant genes and enzymes related to sterol catabolism have been identified in Mycobacterium sp., indicating its great potential for bio-based steroid production on an industrial scale (Behra et al. 2019; Xu et al. 2016).
In this paper, research progress on steroid bioconversion by Mycobacteria neoaurum (M. neoaurum) was reviewed since 2000, with special attention paid to the rational development of mycobacteria as a biorefinery factory. Rational aspects of the modification of steroid transformation pathways, including the exploitation of monooxygenase resources, editing of biosynthetic routes, metabolic engineering and reinforcement of material uptake and transport, are all discussed. New trends in omics analysis and genetic manipulation are also covered, with an emphasis on the rational development of M. neoaurum.
Biotransformation route from sterols to steroidal intermediates
Phytosterols are better sustainable resources for the conventional route of cholesterol-side chain degradation due to their low price and comparatively higher conversion rate (Sripalakit et al. 2006). The mapping and analysis of the complicated sterol catabolic pathway has always been one of the research hotspots in the field, involving a variety of enzymes and reactions, some of which show discrepancies among different species, which precluded the elucidation of the entire metabolic pathway (Wilbrink 2011).
With the development of bioinformatics, the genetic information of Actinomycetes was annotated, and the mystery on the functional genes involved in the general sterol catabolism route has been gradually unveiled (Donova 2017; Giorgi et al. 2019). As shown in Fig. 2, cholesterol is firstly conversed to 4-cholesten-3-one by cholesterol oxidase (chox, ChOx) or 3β-hydroxysteroid oxidase (hsd, 3β-HSD) that was responsible for the initiation of bacterial cholesterol catabolism (Fig. 2M1). The side-chain degradation process was confirmed to be consistent with the fatty acid oxidation pathway (Kendall et al. 2007). Steroid C26-monooxygenase (cyp125, cyp142) is proposed to mainly catalyze side-chain to form a terminal carboxyl group (Fig. 2M2). After the terminal acylation of C-27, the cholesterol side-chain is activated by the terminal CoA thioesterification, and then the carboxyacyl-CoA of C-27 enters the β-oxidation reaction, consisting of acyl-CoA dehydrogenation, alkenyl-CoA hydration, β-hydroxyacyl-CoA dehydrogenation and β-ketoacyl-CoA sulfurization sequentially (Niu et al. 2019) (Fig. 2M3–M4). Module 5 shows the key intermediates synthesis in which aldolases and propionyl-CoA reductase are applied to generate 4-HBC, which can be further synthesized into PG. In the others pathway, the side-chain is completely truncated to produce AD, ADD and 9-OH-AD, and 3-ketosteroid Δ1-dehydrogenases (KstD) and 3-Ketosteroid-9α-hydroxylase (KSH) that play a crucial role in initiating the opening of the steroid nucleus (Li et al. 2019; Rohman and Dijkstra 2021).
Key intermediates and relevant bioconversion steps in the degradation of cholesterol by mycobacteria (Donova and Egorova, 2012). Cholesterol is a classic substrate for sterol metabolism in mycobacteria, C3 is oxidized and isomerized by cholesterol oxidase initially. Three rounds of degradation of the side chain are launched by C27 monooxygenase, similar to the β-oxidation reaction of lipid, and release propionyl-CoA and acetyl-CoA in the process. In addition to the normal reaction, a special pathway involving aldolase and reductase are also drawn for synthesizing 4-HBC. The products obtained from the metabolism will further modify the sterane nucleus under the action of hydroxylase and dehydrogenase. Splitting of the ring are not displayed in the sketch, and finally cholesterol can be degraded to water and carbon dioxide
The complete microbial degradation of a sterol requires a large array of steroid-degrading enzymes with different specificities. Through the combination of metabolic modification and genetic engineering technology, researchers hope to obtain excellent strains that can efficiently transform phytosterols into high-value steroidal intermediates. The corresponding products and intermediates of steroids obtained via cholesterol or phytosterol side chain degradation are listed in Table 1, the recently reported yield indicated the current research level and industrialization potential in this field. Since most of the classical enzymes involved in sterol catabolism have been summarized in previous studies (Donova and Egorova 2012; Zhao et al. 2021), we focused on the most recent advances on the key enzymes involved in the initiation of sterol catabolism in this paper.
Significance of monooxygenase resources for steroid modification
Steroid C27 monooxygenase: initiation of side-chain degradation
Microbial degradation of steroids encompasses three processes, including the oxidation and isomerization of the sterane core by 3β-hydroxysteroid dehydrogenases or cholesterol oxidase, degradation and oxidation of the sterol side chain, and finally decomposition of the steroid nucleus (Fernandez-Cabezon et al. 2017). The degradation of the side chain is crucial for sterol metabolism. Although many enzymes and cofactors participate in the process, this step is mainly driven by steroid C27 monooxygenases belonging to different cytochrome P450 families (Rosloniec et al. 2009).
CYP125 is demonstrated to be crucial both in hydroxylating cholest-4-en-3-one at C27 and oxidizing it into cholest-4-en-3-one-27-carboxylic acid. Therefore, reinforce of CYP125 activity can enhance the transformation ability of steroids. The CYP 125 family has a " letterbox" active center, which can accommodate sterol molecules with a polycyclic structure, and a funnel-shaped entrance near the active-site heme. The alkyl chain of the substrate extends downward along the narrow junction funnel, and the terminal methyl carbon of the side chain is brought in proximity of the heme iron for oxidation (McLean et al. 2010). Accordingly, the CYP 125 level was upregulated with the increase of cholesterol (Brengel et al. 2016). Deletion of the gene greatly affected the growth of mycobacteria utilizing cholesterol as carbon source (Capyk et al. 2009). Overexpression of CYP 125–3 in M. neoaurum M3 increased the specific activity by 22%, and the NAD+/NADH ratio was increased by 31%, the yield of AD was improved approximately 18% in the biotransformation of phytosterol (Su et al. 2018). In addition, three isoenzymes (SMO1, SMO2, SMO3) of steroid C27 monooxygenase were identified in M. neoaurum JC-12. The strongest catalytic activity was exhibited by steroid C27 monooxygenase 2, and when it was co-expressed with ChOx and KstD, the production of ADD reached 20 g/L (Shao et al. 2019a). A whole-cell catalyst for the production of AD overexpressing cholesterol oxidase and steroid C27 monooxygenase exhibited 88.6% capacity improvement, and the highest reported AD yield of 25.8 g/L was achieved in a two-step bioprocess (Chang et al. 2020). Hence, in the process of transformation of steroid by mycobacteria, C27 hydroxylation of the side chain by CYP 125 is both the initial step of side chain degradation and the rate-limiting step of cell growth, which plays an important role in metabolism (Ouellet et al. 2010).
Nucleus hydroxylation for the activation of steroid molecules
Sequencing analysis of Mycobacterium tuberculosis (M. tuberculosis) revealed that there were twenty P450s in its genome. These cytochrome P450s have been designated with systematic “CYP” numbers, such as CYP 125, CYP 142, CYP 106A, CYP 109, CYP 154, CYP 260 and other families (Szaleniec et al. 2018). Many CYPs have also been used in the reaction of steroid modification, and the obtained new steroid resources have more medicinal value. The important sites of the steroidal nucleus, such as C6, C7, C11, C12, C14, C15, C16, C17 and C25, are also a focus of continuous research (Fig. 3). For example, 25-hydroxycholesterol possesses a significant antiviral and counteract inflammation activity (Diczfalusy, 2013; Tricarico et al. 2019), and could be produced from cholesterol by CYP109E1 from Bacillus megaterium (Putkaradze et al. 2019). CYP11A1 can biosynthesize pregnenolone by diametrically catalyzing the side-chain hydroxylation and cleavage of cholesterol or phytosterols, which avoids multiple rounds of step-by-step degradation (Zhang et al. 2019a). CYP154C5, CYP109E1 were found to convert testosterone into 16α-hydroxy-testosterone, which is an important precursor of steroidal drugs in the pharmaceutical industry (Bracco et al. 2013; Jóźwik et al. 2016). Similarly, 17α-hydroxyprogesterone is an important pharmaceutical intermediate in the production of megestrol, hydrocortisone acetate, betamethasone, dexamethasone and other drugs (Gonzalez et al. 2018). In recent years, 17α-hydroxylases (CYP17A) were used to convert 17α-hydroxyprogesterone. At present, CYP17A2 had been successfully expressed in Pichia pastoris and used for the synthesis of 7α-hydroxyprogesterone, but the production was only 120.9 mg/L (Liu et al. 2022a).
Cytochrome P450s with major target sites on steroid substrates. The steroidal precursors are modified by hydroxylation for different functions. C17 and C11 of hydroxylation were particularly concerned owing to its significance for steroids activation. Atoms in red represent the first critical hydroxyl group, and atoms in blue represent the second critical hydroxyl group. C6-hydroxylase: CYP4A21, CYP260B1; C7-hydroxylase: CYP7A2, CYP2A1; C11-hydroxylase: CYP106A2, CYP11B; C12-hydroxylase: CYP8B1; C14-hydroxylase: P450lun; C15-hydroxylase: CYP106A1, CYP106A2; C16-hydroxylase: CYP154C2, CYP109A1; C17-hydroxylase: CYP17A1, CYP17A2; C25-hydroxylase: CYP27A1, CYP109E1, CYP46A1; C26/C27-hydroxylase: CYP125A1, CYP142A1
The hydroxylation of C7, C11, C14 has great significance in the pharmaceutical industry (Chen et al. 2019; Milecka-Tronina et al. 2014; Schiffer et al. 2015). To extend the steroidal bioconversion pathway, exploration and screening of hydroxylase with higher activity and selectivity towards more active steroids and their functional expression in Mycobacterium is regarded as novel direction for the rational development of mycobacteria cells in the future (Donova 2021). Therefore, it is of great significance that we carry out more research on steroid oxidation by CYP enzymes, which will help to expand the application of cholesterol derivatives.
Enhancement of substrate availability
Reaction engineering for improving the accessibility of sterol substrates
The metabolic pathway is only one of the limiting factors for sterol transformation, low substrate solubility and transport barriers are also important constraints (Xiao et al. 2020). The problems of sterol emulsion have not been completely resolved, since the particles disperse poorly and require a complicated biocompatibility balance (Gao 2016). To improve the bioavailability of sterols by the application of emulsion technologies so that the mass transfer interface area is amplified (McClements 2012), researchers mainly add materials such as surfactants (Tween 80, Triton X-100), cyclodextrins (CDs) and their modified variants (2-hydroxylpropyl-β-cyclodextrin, HP-β-CD, methyl-β-cyclodextrin) (Shao et al. 2015). There are many shortcomings in these strategies, since surfactants interfere with cell growth and make wastewater treatment more challenging, while the cost of cyclodextrins remains prohibitive for industrial application (Mancilla et al. 2018).
In the process of mycobacterial growth and metabolism, the addition of vancomycin, glycine, ethambutol, polyethyleneimine and cyclodextrin was found to change the permeability of the cell membrane or cell wall to enhance the trans-membrane delivery of sterols (Wang et al. 2006; Malaviya and Gomes 2008). For example, glycine was found to interfere with the formation of the peptidoglycan network structure. The resulting incomplete cell wall is conducive to breaking the osmotic barrier to promote the transport of sterols and other lipid substances, and thereby increase the accumulation of steroidal intermediates. Notably, most of these studies enhance the permeability of hydrophobic substances by altering the phenotype of cell wall, while the genetic regulation mechanism of sterol transport was seldomly addressed. This aspect had been effectively supplemented by omics analysis and molecular biology technology.
Enhancement of transport channels in the cell membrane
The structure of envelope in mycobacteria is another key factor for improving the accessibility of sterol substrates, not only maintains the life activities, but also represents an osmotic barrier for material transfer. The mycobacterial cell wall is mainly composed of peptidoglycan, arabinogalactan and the very-long-chain mycolic acids (Fig. 4). In these unusual core complexes, mycolic acid endows the cell wall with hydrophobic properties, while the crosslinking of peptidoglycan gives the cell wall its density, these structures play different functions in the process of transport (Kieser and Rubin 2014). With the development of omics technology, many genes regulating substrate transport and the osmotic barrier function of the membrane have been discovered. An analysis of the genomes of M. tuberculosis and M. smegmatis also found similar gene clusters with mce4 homologs. Their genome respectively accommodated 4 and 6 gene clusters corresponding to mce operons, and each of mce loci contained 2 yrbE and 6 mce genes, which encoded two integral membrane proteins. The membrane proteins have numerous similarities with the ATP-binding cassette transporters, which are closely related to cell membrane remodeling and cholesterol assimilation (Kumar et al. 2005).
The structure of cell envelope in mycobacteria refer to the model of Jackson's cell capsule (Jackson et al., 2020). The cell envelope is made up of periplasm, outer membrane and capsule. The component of periplasm is peptidoglycan and arabinose and their biosynthesis is regulated by pbpA/B, araR, embR genes respectively, which further affects the efficiency of cholesterol uptake. kasB, ephD, aceE and MmpL3 are relevant to the extension of mycolic acid and trehalose. Lipoarabinomannan and lipomannan are important backbones in the capsule, which are elaborated and modified covalently by succinate, methythioxylose, mannan and glucan. To provide detailed information, the corresponding schematic modules and structures were not drawn to scale
The mce4 genes have been confirmed to form part of the operon of M. tuberculosis for the uptake and utilization of cholesterol. Similarly, the mce4 operon is also involved in the cholesterol transport of M. smegmatis (Klepp et al. 2012). Moreover, it was found that any of the mce4 mutants could not grow on cholesterol. Hence, these genes are necessary for cholesterol uptake, while the homologous genes of other mce operons did not have the ability to replace them (Garcia-Fernandez et al. 2017). The Mce4 transporters were recently found to form a multiprotein complex that is essential for the mycobacterial cells to take up cholesterol. At the same time, it was found that Mce4A might bind to Mce systems, including Mce1 (Rank et al. 2021). When mceG, yrbE4A and yrbE4B from M. tuberculosis, which respectively encode energy-related factors and two integral membrane proteins, were expressed in Mycobacterium sp. MS136-GAB, they increased the product yield by 20% compared with the wild-type strain, indicated that the system promoted the mass transfer of phytosterols (He et al. 2018).
Regulation of cell envelope
In recent years, the mycobacterial cell wall structure has also been widely investigated because of its unique function in sterol uptake. The mmpl3 gene encodes a trehalose transmembrane transporter, and is also an important gene affecting mycobacteria cell envelope assembly (Tahlan et al. 2012). When the expression of mmpL3 was reduced in M. neoaurum ATCC 25795, the cell permeability was increased by 23.4%, and the uptake of sterols was enhanced by 15.6% (Xiong et al. 2017). This is only one of the strategies that can be used to change the cell permeability by modifying the mycobacterial membrane and membrane proteins to affect the uptake and transport of substrates. The pbpAB genes were modified to change the structure of the peptidoglycan layer, and the yield of 4-HBC could be increased by 28% compared with the wild type (Sun et al. 2019a).
What’s more, the mycolic acid layer can also be changed. The deletion of the nonessential gene kasB, which encodes β-ketoacyl acyl carrier synthetase, resulted in the disorder of the proportion of mycolic acids in M. neoaurum ATCC 25795. Furthermore, the cell permeability was increased by about twofold, and the yield was significantly increased by 2.38-fold, greatly reducing the conversion time (Xiong et al. 2020b). embC converts lipomannan to lipoarabinomannan in M. tuberculosis. The knockout of the embC resulted in a dysfunction of the cell envelope and increase of cell permeability, increasing sterol uptake by 52.4% over the parental strain (Xiong et al. 2020a). Recent reports indicate that LmeA and SucT mediate the maturation of lipoarabinomannan and can change the hydrophobicity and stiffness of the capsule (Palcekova et al. 2019; Rahlwes et al. 2020). Modulation of the mycobacterial cell envelope had not only become a new research hotspot, but also strengthened the sterol production performance of the strains from another aspect.
Coordination strategy of toxic metabolic intermediates
Rational modulation of propionyl-CoA
Mycobacterium sp. can use cholesterol as a carbon source for growth, or convert sterols into steroids, what mainly involves the degradation of the sterol side chain and the decomposition of the core rings, whereby propionyl-CoA and acetyl-CoA are produced in the metabolic pathway (Uhia et al. 2012). The propionyl-CoA can maintain metabolic balance and provide energy in Mycobacterium sp., such as acylation of amino acids, metabolism of lipids (Crowe et al. 2017; Xu et al. 2020). However, studies have shown that when M. smegmatis and M. tuberculosis grow on propionate, the cells will accumulate excessive amounts of propionyl-CoA, which acts as a toxic intermediate that triggers metabolic stress and inhibits growth (Lee et al. 2013). Therefore, the detoxification of the propionyl-CoA pool is essential. There are three pathways for the degradation of toxic propionyl-CoA, including the methyl citrate cycle (MCC), the vitamin B12 dependent methylmalonyl pathway (MMP) and the synthesis of methyl-branched chain lipids in the cell envelope (Liu et al. 2018c). MCC is characterized by the initial condensation of propionyl-CoA and oxaloacetate to form methyl citrate, which is converted into pyruvate and succinate to achieve the goal of detoxification after several reactions (Otzen et al. 2014).
Based on these findings, in order to improve the ability of microbial transformation of AD, it is necessary to relieve the toxicity of propionyl-CoA. In M. neoaurum, there is a complete MCC pathway encoded by a prp gene cluster including prpB, prpC, and prpD encoding key enzymes, as well as a transcriptional regulator encoded by prpR. It was demonstrated that the expression of prpR and related genes (prpDBC) could be enhanced by the phosphate regulator RegX3 in MCC (Pei et al. 2021). Overexpression of prpBCDR significantly increased the transcription level of the prp gene cluster and conversion of AD. Compared with the wild type, prpR could effectively reduce the level of propionyl-CoA and improve the cell viability. Furthermore, the nitrogen transcription regulator GlnR inhibited the transcription of the prp operon in nitrogen- limited medium. The deletion of glnR brought about the enhancement of the prp gene cluster transcription and expression, and promoted propionate detoxification (Zhang et al. 2020).
MMP is another detoxification strategy for propionyl-CoA. Firstly, propionyl-CoA is carboxylated into methyl malonyl-CoA, and then isomerized to non-toxic succinyl-CoA. The key enzyme in this metabolic process is biotin dependent propionyl-CoA carboxylase (PCC) (Savvi et al. 2008). For example, increasing the expression of pccB enhanced the biotransformation ability of phytosterols (Zhou et al. 2019b). Furthermore, the pccB and prpR genes were overexpressed to construct a citrate-based ATP futile cycle (AFC) and pyruvate-based AFC. These cycles could reduce the accumulation of ATP and propionyl-CoA in cells, enhancing cell viability and increasing the AD conversion rate (Zhou et al. 2020). In conclusion, propionyl-CoA is one of the key factors restricting the growth and production efficiency of host cells in the process of microbial transformation of phytosterols into steroids. The research and application of detoxification pathways will help improve the conversion rate of sterol substrates.
Metabolic detoxication of reactive oxygen species
Intracellular reactive oxygen species (ROS), such as superoxide radical, hydrogen peroxide (H2O2), hydroxyl radical and lipid peroxides are mainly produced by NADPH oxidase, xanthine oxidase, or during electron transfer in the respiration process (Shao et al. 2017; Yeware et al. 2017). ROS has traditionally been regarded as a regulator of many cellular functions, such as transcription, direct oxidative modification, protein turnover, protein interaction and enzyme modification (Finkel 2011). However, excessive accumulation of ROS leads to oxidative stress and damages a variety of biological molecules, including DNA, proteins, and lipids (Tyagi et al. 2015).
Most mycobacteria have evolved antioxidant systems to resist the damage of ROS. There are two frequently-used detoxification mechanisms, based either on oxygen scavenging enzymes, such as catalase (CAT), catalase peroxidase and superoxide dismutase, or on low molecular weight mercaptans, such as glutathione (GSH), fungal mercaptan (MSH) and ergothioneine (EGT) (Richard-Greenblatt et al. 2015; Sao Emani et al. 2018). When Mycobacterium sp. utilizes sterols, a series of redox reactions will occur inside the cell, which requires a large number of NAD+ and FAD molecules, producing ROS beyond the normal threshold (Shao et al. 2017). Excessive ROS brings about oxidative stress on the metabolism and subsequently damages various biological molecules, and ultimately injures cells critically (Imlay 2003), thus both the cell growth and steroidal bio-synthetic capability of mycobacteria was severely inhibited. Therefore, it is an important strategy to reduce ROS level for enhancing steroid transformation.
Enhancing type II NADH dehydrogenase in M. neoaurum effectively increased the NAD+/NADH ratio and ATP level, and also effectively reduced the oxidative stress, resulting in greater cell viability. After enhancing NADH dehydrogenase, ROS was decreased by 42.3%, resulting in a 54.2% increase of viability, thus substantially raising the productivity of AD (Zhou et al. 2019a). NADH oxidase and catalase genes from Bacillus subtilis were overexpressed to eliminate the toxic H2O2 accumulated during sterol bioconversion, which improved the sterol transformation efficiency by 80% compared to the initial strain (Shao et al. 2019b). Mycobacterial endogenous antioxidants, ROS scavenging enzymes and low molecular weight mercaptans were used to construct detoxification pathways. The combination of CAT, MSH and EGT could boost cell viability by 54.2% and increase 4-HBC production by 47.5% (Sun et al. 2019b). Thus, normal ROS levels were restored by skillful metabolic adjustment, which may be employed as an effective strategy for enhancing sterol bioconversion.
Omics-guided studies provide new insights
Multi-omics strategies are valuable tools in the search for key genes and proteins of metabolic pathways. In the wake of the continuous development and convergence of internet and biotechnology, the application of bioinformatics is becoming increasingly widespread (Shtratnikova et al. 2021). The mycobacteria research can be divided into two categories. One is the research on pathogenic mycobacteria such as M. tuberculosis (Malone et al. 2018). Through the means of bioinformatics, the mechanism of infection, therapeutic drugs and metabolic pathways are continuously being explored (Goff et al. 2020). Another branch focuses on non-pathogenic mycobacteria (such as M. smegmatis and M. neoaurum) which have industrial potential and can be used to produce steroids (Uhia et al. 2012) (Table 2). Multi-omics data analysis of samples from different growth conditions or life cycles is not only help understand the biology of the strains, but is also conducive to the mining of key genes and new metabolic pathways, aiming to achieve the goal of industrialization (Li et al. 2016).
Genomics and transcriptomics assist the rational improvement of mycobacteria
Genome and transcriptome sequencing can reveal the gene expression changes that may be responsible for steroid catabolism of M. neoaurum. On the basis of significant differences in omics data, combined with single nucleotide polymorphisms and dynamic sRNA data, we can understand gene associations and interactions, as well as the regulatory mechanism of small molecules in various pathways, such as lipid transport and metabolism, amino acid metabolism, signal transduction, cell envelope function and ATP biosynthesis (Liu et al. 2016a, b). For example, a high yield and substrate resistant mutation of M. neoaurum MN4 was obtained by physical and chemical mutagenesis and screening. Experimental data from the combined omics analysis between the wild type and mutation revealed that both ChOx and enzymes involved in β-oxidation were mutated (Xu et al. 2017). By comparing and analyzing the genome of Mycobacterium sp. VKM, researchers found several key enzymes of steroid catabolism, such as KSH and KstD (Bragin et al. 2013). These results not only confirmed the hypotheses raised by previous studies, but also guided the design of experiments to increase production (Rohman and Dijkstra 2021; Zhang et al. 2019b).
Furthermore, transcriptome analysis was applied more frequently in sterol biodegradation research. The high-throughput transcriptome sequencing of Mycobacterium sp. VKM Ac-1817D grown on phytosterols showed that core genes encoding KSH and KstD exhibited tenfold of upregulation (Bragin et al. 2019). The changes in the transcriptome and proteome of M. neoaurum ATCC 25795 were used to panoramically analyze the sterol transformation process, substrate uptake, carbon metabolism, cell wall and membrane synthesis, regulatory changes of efflux protein and ATP at different levels, which provided targets for the removal of by-products and increase of sterol uptake (Liu et al. 2018b, c).
Profiling of biotransformation intermediates using metabolomics
Many studies investigated the metabolome of mycobacteria, but they mainly focused on the physiological networks, pathogenic mechanisms and novel drug targets of M. tuberculosis (Rego et al. 2021). By comparison, the research on the metabolome of M. neoaurum is sparse. Tween 80, a detergent that promotes the dissolution of sterols and reduces the aggregation of bacterial cells, is conventionally added to the medium for steroid transformation. Metabolomics revealed that Tween 80 induced the upregulation of multiple pathways in central carbon metabolism and affected the synthesis of triacylglycerols, proteinogenic amino acids and nucleotide precursors (Pietersen et al. 2020). The in-depth influence of detergents on the synthesis of steroids at the metabolic level is worth exploring. In addition to these results, propionyl-CoA, ROS, and 4-ene-3-ketosteroids were identified as toxic molecules that inhibit the synthesis of 9α-OH-AD, and an in-situ adsorption scheme improved the production efficiency by 23.15% (Wang et al. 2021). Development of omics not only rapidly promoted the technological development of the biosynthesis of steroids, but also provided genetic data for constructing efficient pathways and designing biological cell factories.
Advances in genetic manipulation approaches of mycobacteria
General genetic manipulation
Non-pathogenic M. smegmatis and M. neoaurum are widely applied in the development of steroid biotransformation, but wild-type bacteria usually have a long growth period, large accumulation of by-products and low product yield, so it is necessary to conduct extensive genetic engineering to obtain high-yield strains (Loraine and Smith 2017). Genetic manipulations can be divided into three broad strategies: (I) Gene replacement based on homologous recombination and resistance marker screening; (II) Gene silencing based on homologous sequences and controllable promoters; (III) CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) editing technology, which was recently also introduced in Mycobacterium sp. (Choudhary et al. 2016).
It is difficult and inefficient to eliminate resistance markers to obtain label-free mutants. To solve this problem, two strategies have been developed, based on suicide plasmids carrying screening gene cassettes and mutually dependent or incompatible plasmids (Pashley et al. 2003). These vectors can be easily eliminated after silencing the target gene. However, there are still many problems such as multiple transformation steps and low efficiency of screening. In addition to the influence of resistance markers, the introduction of exogenous genes into mycobacterial hosts will cause a greater burden on growth and metabolism, which is not conducive to the implementation of genetic manipulation (Brown and Parish, 2006). Genetic manipulation for activation or silencing at appropriate times can effectively relieve the growth burden of the host.
Precise genome editing
The emergence of CRISPR greatly simplified gene editing on the chromosome. CRISPR-Cas is a natural defense system found in many bacteria and archaea, which can selectively cleave exogenous invading DNA (Han and She 2017; Makarova and Koonin 2015). It has become the “No.1 scissors” of genetic engineering, and its inventors were awarded the Nobel Prize in physiology and medicine in 2020. Before plasmids can be introduced into the acceptor strain, it needs alkali or ultraviolet treatment to improve the success rates of transformation and recombination (Song et al. 2015). The time-consuming traditional tools limited the research on mycobacteria. CRISPR-Cas can shorten the time of genetic operation, reduce the difficulty of marker selection, and remove the restriction of homologous arm length (Doudna and Charpentier 2014). Consequently, the application of the CRISPR system in mycobacteria is becoming increasingly popular.
Many successful cases have been demonstrated in various mycobacteria. For example, the introduction of codon optimized dCas9 and sgRNA from Streptococcus pyogenes into M. tuberculosis resulted in complete inhibition of single or multiple target genes (Tang and Fu 2018). Using the CRISPR-Cas9 system, the expression of sepF was successfully inhibited in M. smegmatis. The inhibition rate is as high as 98%, and no off-target effects were found (Xiao et al. 2019). Moreover, CRISPR-Cas9 combined with non-homologous end joining (NHEJ) could achieve efficient frameshift mutation of genes in M. marinum and M. tuberculosis, which accelerated the research on these lethal pathogens (Meijers et al. 2020). However, CRISPR-Cas9 has many limitations, such as toxicity, single strand cleavage and low efficiency in GC rich recognition regions. CRISPR-Cas12a (Cpf1) edits bacterial genomes more effectively, providing a new tool for special hosts that cannot accept Cas9. It has different recognition target sites and the ability to cut double stranded DNA, as well as lower host toxicity (Tang and Fu 2018; Yan et al. 2017).
When different Cas effector proteins were tested in M. smegmatis, the expression and editing effect of FnCpf1-cg was significantly better than that of Cas9. The efficiency of FnCpf1 combined with the NHEJ system was as high as 70%, which greatly reduced the genetic operation time (Sun et al. 2018). CRISPR-Cas12a assisted recombination technology was also applied to steroidal biosynthesis. M. smegmatis strains with inactivated kstD1, kstD2 and kstD3 were successfully constructed, and accumulated 2.86 g/L 9α-OH-AD (Meng et al. 2020). Unfortunately, there are few reports on the application of CRISPR-Cas systems in M. neoaurum. In 2021, it was reported that M. neoaurum could be edited using Cas12 in combination with NHEJ or HR-mediated DSB repair. CRISPR-Cas12a was able to delete target fragments of 1–24 kb with a maximum efficiency of 70%. Combined with a suicide plasmid system, the probability of single crossover increased to 100% (Liu et al. 2022b). CRISPR-Cas has many advantages in genetic operation, and there are still many expectations on how to help reform industrial hosts for the production of steroids.
Conclusions and future prospects
The research on the transformation of sterols into steroids by Mycobacterium sp. has been ongoing for decades. Although the direction and content of these studies are very extensive, and many breakthroughs have been made, there are still many problems that have not been solved. For example, (I) the low solubility of the substrates is one of the most important factors limiting the microbial uptake and transformation of sterols; (II) More than ten enzymes are involved in the degradation of the sterol side chain, which lack unabridged analysis of their catalytic mechanisms and isozyme relationships; (III) There are still no efficient, systematic and simple genetic operation tools; (V) There is no ideal host that can overcome the low material transport efficiency and slow growth. Accordingly, many new research strategies are needed in the future.
Synthetic biology and bioinformatics are currently very popular disciplines. Combined with emerging technologies, they will not only encourage technological innovation in the research of steroids, but also promote the efficient development of industrialization processes for steroid biosynthesis (Fernandez-Cabezon et al. 2018). By taking the high-speed trains of bioinformatics, researchers can deeply analyze the interaction relationship of genes, proteins, regulatory factors, and metabolic networks at the individuality or global level using massive data, and point out the direction for accurate mining and rational improvement of strains. Using the technology of synthetic biology, sterol conversion systems will be modularized and componentized, skillfully transplanted and assembled into a more efficient host. This will enable researchers to reconstruct the metabolic pathway, build an efficient cell factory, and break through the traditional boundaries. Artificial cell synthesis instead of natural extraction will achieve great economic benefits and contribute to industrial upgrading tremendously.
Existing mycobacteria cell factories cannot meet the requirements of the steroid industry revolution thoroughly. Nowadays, even though the biocatalytic potential of mycobacteria towards steroids is gradually disclosed, the development of an ideal mycobacterial chassis strain is still a pressing problem. An industrial chassis strain must have numerous robust features, such as low nutritional requirements and short growth cycle, efficient substrate uptake and product efflux, high copy and correct translation of genetic information, knock out or silencing of by-product genes, flexible and modular gene circuit design, as well as strong environmental adaptability and tolerance to substrates and products.
The metabolism of steroids in mycobacteria is highly complex. The optimization of metabolic pathways will greatly increase the flux of steroid synthesis and reduce the difficulty of regulation. Related strategies include the mining and heterologous expression of aldolases or high-performance hydroxylases. It is not only beneficial to improve the biosynthesis efficiency of steroids, but also to achieve the synthesis of related derivatives. What's more, enzymes that are difficult to adapt can be directed or artificially created to modify steroids. Recent research on the change of catalytic sites in the directed evolution of enzymes can serve as inspiration (Grobe et al. 2021).
Cholesterol is a model substrate for the study of sterol metabolism, but it is too expensive for large-scale industrial production of steroids. At present, phytosterols are the main raw materials for steroid production, but they also have many disadvantages, such as complex composition, limited sources and many by-products. There were many reports on the synthesis of lanosterol, ergosterol, campesterol and cholesterol from glucose by yeast (Xu and Li 2020; Zhang et al. 2017). Therefore, combining the advantages of yeasts in the synthesis of polycyclic substances with the mycobacterial characteristics in the degradation and utilization of sterols holds promise to completely solve the problem of substrate availability in the future.
References
Baruah D, Das RN, Konwar D (2015) Facile green synthesis of 16-dehydropregnenolone acetate (16-DPA) from diosgenin. Synthetic Commun 46:79–84. https://doi.org/10.1080/00397911.2015.1121280
Behra PRK, Pettersson BMF, Ramesh M et al (2019) Insight into the biology of Mycobacterium mucogenicum and Mycobacterium neoaurum clade members. Sci Rep 9:19259. https://doi.org/10.1038/s41598-019-55464-5
Bracco P, Janssen DB, Schallmey A (2013) Selective steroid oxyfunctionalisation by CYP154C5, a bacterial cytochrome P450. Microb Cell Fact 12:95. https://doi.org/10.1186/1475-2859-12-95
Bragin EY, Shtratnikova VY, Dovbnya DV et al (2013) Comparative analysis of genes encoding key steroid core oxidation enzymes in fast-growing Mycobacterium spp. strains. J Steroid Biochem Mol Biol 138:41–53. https://doi.org/10.1016/j.jsbmb.2013.02.016
Bragin EY, Shtratnikova VY, Schelkunov MI et al (2019) Genome-wide response on phytosterol in 9-hydroxyandrostenedione-producing strain of Mycobacterium sp. VKM Ac-1817D. BMC Biotechnol 19:39. https://doi.org/10.1186/s12896-019-0533-7
Brengel C, Thomann A, Schifrin A et al (2016) Discovery and biophysical evaluation of first low nanomolar hits targeting CYP125 of M. tuberculosis. ChemMedChem 11:2385–2391. https://doi.org/10.1002/cmdc.201600361
Brown AC, Parish T (2006) Instability of the acetamide-inducible expression vector pJAM2 in Mycobacterium tuberculosis. Plasmid 55:81–86. https://doi.org/10.1016/j.plasmid.2005.06.005
Capyk JK, Kalscheuer R, Stewart GR et al (2009) Mycobacterial cytochrome p450 125 (CYP125) catalyzes the terminal hydroxylation of C27 steroids. J Biol Chem 284:35534–35542. https://doi.org/10.1074/jbc.M109.072132
Chang H, Zhang H, Zhu L et al (2020) A combined strategy of metabolic pathway regulation and two-step bioprocess for improved 4-androstene-3,17-dione production with an engineered Mycobacterium neoaurum. Biochem Eng J. https://doi.org/10.1016/j.bej.2020.107789
Chen J, Tang JL, Xi YY et al (2019) Production of 14 alpha-hydroxysteroids by a recombinant Saccharomyces cerevisiae biocatalyst expressing of a fungal steroid 14 alpha-hydroxylation system. Appl Microbiol Biot 103:8363–8374. https://doi.org/10.1007/s00253-019-10076-x
Choudhary E, Lunge A, Agarwal N (2016) Strategies of genome editing in mycobacteria: achievements and challenges. Tuberculosis (edinb) 98:132–138. https://doi.org/10.1016/j.tube.2016.03.005
Crowe AM, Casabon I, Brown KL et al (2017) Catabolism of the last two steroid rings in Mycobacterium tuberculosis and other bacteria. Mbio. https://doi.org/10.1128/mBio.00321-17
Diczfalusy U (2013) On the formation and possible biological role of 25-hydroxycholesterol. Biochimie 95:455–460. https://doi.org/10.1016/j.biochi.2012.06.016
Donova MV (2017) Steroid bioconversions. Methods Mol Biol 1645:1–13. https://doi.org/10.1007/978-1-4939-7183-1_1
Donova M (2021) Microbial steroid production technologies: current trends and prospects. Microorganisms. https://doi.org/10.3390/microorganisms10010053
Donova MV, Egorova OV (2012) Microbial steroid transformations: current state and prospects. Appl Microbiol Biotechnol 94:1423–1447. https://doi.org/10.1007/s00253-012-4078-0
Doudna JA, Charpentier E (2014) Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346:1258096. https://doi.org/10.1126/science.1258096
Fernandes P, Cabral JMS (2007) Phytosterols: applications and recovery methods. Bioresource Technol 98:2335–2350. https://doi.org/10.1016/j.biortech.2006.10.006
Fernandez-Cabezon L, Garcia-Fernandez E, Galan B et al (2017) Molecular characterization of a new gene cluster for steroid degradation in Mycobacterium smegmatis. Environ Microbiol 19:2546–2563. https://doi.org/10.1111/1462-2920.13704
Fernandez-Cabezon L, Galan B, Garcia JL (2018) New insights on steroid biotechnology. Front Microbiol 9:958. https://doi.org/10.3389/fmicb.2018.00958
Finkel T (2011) Signal transduction by reactive oxygen species. J Cell Biol 194:7–15. https://doi.org/10.1083/jcb.201102095
Gao XQ (2016) Enhanced steroid metabolites production by resting cell phytosterol bioconversion. Chem Biochem Eng Q 29:567–573. https://doi.org/10.15255/cabeq.2014.2098
Garcia-Fernandez J, Papavinasasundaram K, Galan B et al (2017) Molecular and functional analysis of the mce4 operon in Mycobacterium smegmatis. Environ Microbiol 19:3689–3699. https://doi.org/10.1111/1462-2920.13869
Giorgi V, Menendez P, Garcia-Carnelli C (2019) Microbial transformation of cholesterol: reactions and practical aspects-an update. World J Microbiol Biotechnol 35:131. https://doi.org/10.1007/s11274-019-2708-8
Goff A, Cantillon D, Muraro Wildner L et al (2020) Multi-omics technologies applied to tuberculosis drug discovery. Appl Sci. https://doi.org/10.3390/app10134629
Gonzalez E, Johnson KM, Pallan PS et al (2018) Inherent steroid 17alpha,20-lyase activity in defunct cytochrome P450 17A enzymes. J Biol Chem 293:541–556. https://doi.org/10.1074/jbc.RA117.000504
Grobe S, Badenhorst CPS, Bayer T et al (2021) Engineering regioselectivity of a p450 monooxygenase enables the synthesis of ursodeoxycholic acid via 7 beta-hydroxylation of lithocholic acid. Angew Chem Int Edit 60:753–757. https://doi.org/10.1002/anie.202012675
Gupta P, Mahajan A (2018) Sustainable approaches for steroid synthesis. Environ Chem Lett 17:879–895. https://doi.org/10.1007/s10311-018-00845-x
Han W, She Q (2017) CRISPR History: discovery, characterization, and prosperity. Prog Mol Biol Transl Sci 152:1–21. https://doi.org/10.1016/bs.pmbts.2017.10.001
He K, Sun H, Song H (2018) Engineering phytosterol transport system in Mycobacterium sp. strain MS136 enhances production of 9alpha-hydroxy-4-androstene-3,17-dione. Biotechnol Lett 40:673–678. https://doi.org/10.1007/s10529-018-2520-9
Herraiz I (2017) Chemical pathways of corticosteroids, industrial synthesis from sapogenins. Methods Mol Biol 1645:15–27. https://doi.org/10.1007/978-1-4939-7183-1_2
Hu Y, Wang D, Wang X et al (2020) A recycled batch biotransformation strategy for 22-hydroxy-23,24-bisnorchol-4-ene-3-one production from high concentration of phytosterols by mycobacterial resting cells. Biotechnol Lett 42:2589–2594. https://doi.org/10.1007/s10529-020-02991-1
Imlay JA (2003) Pathways of oxidative damage. Annu Rev Microbiol 57:395–418. https://doi.org/10.1146/annurev.micro.57.030502.090938
Jackson M, Stevens CM, Zhang L et al (2020) Transporters involved in the biogenesis and functionalization of the mycobacterial cell envelope. Chem Rev 121:5124–5157. https://doi.org/10.1021/acs.chemrev.0c00869
Jóźwik IK, Kiss FM, Gricman Ł et al (2016) Structural basis of steroid binding and oxidation by the cytochrome P450 CYP 109E1 from Bacillus megaterium. FEBS J 283:4128–4148. https://doi.org/10.1111/febs.13911
Kendall SL, Withers M, Soffair CN et al (2007) A highly conserved transcriptional repressor controls a large regulon involved in lipid degradation in Mycobacterium smegmatis and Mycobacterium tuberculosis. Mol Microbiol 65:684–699. https://doi.org/10.1111/j.1365-2958.2007.05827.x
Kieser KJ, Rubin EJ (2014) How sisters grow apart: mycobacterial growth and division. Nat Rev Microbiol 12:550–562. https://doi.org/10.1038/nrmicro3299
Klepp LI, Forrellad MA, Osella AV et al (2012) Impact of the deletion of the six mce operons in Mycobacterium smegmatis. Microbes Infect 14:590–599. https://doi.org/10.1016/j.micinf.2012.01.007
Kumar A, Chandolia A, Chaudhry U et al (2005) Comparison of mammalian cell entry operons of mycobacteria: in silico analysis and expression profiling. FEMS Immunol Med Microbiol 43:185–195. https://doi.org/10.1016/j.femsim.2004.08.013
Ledford H (2020) Steroid is first drug shown to prevent deaths from Covid-19. Nature 582:469–469. https://doi.org/10.1038/d41586-020-01824-5
Lee W, VanderVen BC, Fahey RJ et al (2013) Intracellular Mycobacterium tuberculosis exploits host-derived fatty acids to limit metabolic stress. J Biol Chem 288:6788–6800. https://doi.org/10.1074/jbc.M112.445056
Li Q, Ge F, Tan Y et al (2016) Genome-wide transcriptome profiling of Mycobacterium smegmatis MC(2) 155 cultivated in minimal media supplemented with cholesterol, androstenedione or glycerol. Int J Mol Sci. https://doi.org/10.3390/ijms17050689
Li H, Wang XD, Zhou LF et al (2019) Enhancing expression of 3-ketosteroid-9 alpha-hydroxylase oxygenase, an enzyme with broad substrate range and high hydroxylation ability in Mycobacterium sp. LY-1. Appl Biochem Biotech 187:1238–1254. https://doi.org/10.1007/s12010-018-2876-2
Liu M, Zhu ZT, Tao XY et al (2016a) RNA-Seq analysis uncovers non-coding small RNA system of Mycobacterium neoaurum in the metabolism of sterols to accumulate steroid intermediates. Microb Cell Fact 15:64. https://doi.org/10.1186/s12934-016-0462-2
Liu M, Zhu ZT, Tao XY et al (2016b) Single nucleotide polymorphism analysis for the production of valuable steroid intermediates in Mycobacterium neoaurum. Biotechnol Lett 38:1881–1892. https://doi.org/10.1007/s10529-016-2187-z
Liu HH, Xu LQ, Yao K et al (2018a) Engineered 3-ketosteroid 9alpha-hydroxylases in Mycobacterium neoaurum: an efficient platform for production of steroid drugs. Appl Environ Microbiol. https://doi.org/10.1128/AEM.02777-17
Liu M, Xiong LB, Tao X et al (2018b) Integrated transcriptome and proteome studies reveal the underlying mechanisms for sterol catabolism and steroid production in Mycobacterium neoaurum. J Agric Food Chem 66:9147–9157. https://doi.org/10.1021/acs.jafc.8b02714
Liu M, Xiong LB, Tao X et al (2018c) Metabolic adaptation of Mycobacterium neoaurum ATCC 25795 in the catabolism of sterols for producing important steroid intermediates. J Agric Food Chem 66:12141–12150. https://doi.org/10.1021/acs.jafc.8b04777
Liu C, Shao M, Osire T et al (2020) Identification of bottlenecks in 4-androstene-3,17-dione/1,4-androstadiene-3,17-dione synthesis by Mycobacterium neoaurum JC-12 through comparative proteomics. J Biosci Bioeng. https://doi.org/10.1016/j.jbiosc.2020.10.006
Liu C, Chen K, Wang Y et al (2022a) Identification of a novel cytochrome P450 17A2 enzyme catalyzing the C17α hydroxylation of progesterone and its application in engineered Pichia pastoris. Biochem Eng J. https://doi.org/10.1016/j.bej.2021.108264
Liu K, Gao Y, Li ZH et al (2022b) CRISPR-Cas12a assisted precise genome editing of Mycolicibacterium neoaurum. N Biotechnol 66:61–69. https://doi.org/10.1016/j.nbt.2021.10.003
Loraine JK, Smith MCM (2017) Genetic techniques for manipulation of the phytosterol biotransformation strain Mycobacterium neoaurum NRRL B-3805. Methods Mol Biol 1645:93–108. https://doi.org/10.1007/978-1-4939-7183-1_7
Makarova KS, Koonin EV (2015) Annotation and classification of CRISPR-Cas systems. Methods Mol Biol 1311:47–75. https://doi.org/10.1007/978-1-4939-2687-9_4
Malaviya A, Gomes J (2008) Enhanced biotransformation of sitosterol to androstenedione by Mycobacterium sp. using cell wall permeabilizing antibiotics. J Ind Microbiol Biotechnol 35:1235–1239. https://doi.org/10.1007/s10295-008-0419-5
Malone KM, Rue-Albrecht K, Magee DA et al (2018) Comparative 'omics analyses differentiate Mycobacterium tuberculosis and Mycobacterium bovis and reveal distinct macrophage responses to infection with the human and bovine tubercle bacilli. Microb Genom. https://doi.org/10.1099/mgen.0.000163
Mancilla RA, Little C, Amoroso A (2018) Efficient bioconversion of high concentration phytosterol microdispersion to 4-androstene-3,17-dione (AD) by Mycobacterium sp. B3805. Appl Biochem Biotechnol 185:494–506. https://doi.org/10.1007/s12010-017-2665-3
McClements DJ (2012) Crystals and crystallization in oil-in-water emulsions: implications for emulsion-based delivery systems. Adv Colloid Interface Sci 174:1–30. https://doi.org/10.1016/j.cis.2012.03.002
McLean KJ, Belcher J, Driscoll MD et al (2010) The Mycobacterium tuberculosis cytochromes P450: physiology, biochemistry & molecular intervention. Future Med Chem 2:1339–1353. https://doi.org/10.4155/fmc.10.216
Meijers AS, Troost R, Ummels R et al (2020) Efficient genome editing in pathogenic mycobacteria using Streptococcus thermophilus CRISPR1-Cas9. Tuberculosis (edinb) 124:101983. https://doi.org/10.1016/j.tube.2020.101983
Meng L, Xuemei L, Jinhui F et al (2020) Effects of the 3-ketosteroid-Δ1 -dehydrogenase on phytosterol transformation with the accumulation of 9α-hydroxyandrost-4-ene-3,17-dione by Mycobacterium smegmatis. Chin J Appl Environ Biol 26:739–746. https://doi.org/10.19675/j.cnki.1006-687x.2020.05037
Milecka-Tronina N, Kolek T, Swizdor A et al (2014) Hydroxylation of DHEA and its analogues by Absidia coerulea AM93. Can an inducible microbial hydroxylase catalyze 7 alpha- and 7 beta-hydroxylation of 5-ene and 5 alpha-dihydro C-19-steroids? Bioorgan Med Chem 22:883–891. https://doi.org/10.1016/j.bmc.2013.11.050
Niu Y, Ge F, Yang Y et al (2019) Biochemical characterization of acyl-coenzyme A synthetases involved in mycobacterial steroid side-chain catabolism and molecular design: synthesis of an anti-mycobacterial agent. 3 Biotech 9:169. https://doi.org/10.1007/s13205-019-1703-y
Otzen C, Bardl B, Jacobsen ID et al (2014) Candida albicans utilizes a modified beta-oxidation pathway for the degradation of toxic propionyl-CoA. J Biol Chem 289:8151–8169. https://doi.org/10.1074/jbc.M113.517672
Ouellet H, Guan S, Johnston JB et al (2010) Mycobacterium tuberculosis CYP125A1, a steroid C27 monooxygenase that detoxifies intracellularly generated cholest-4-en-3-one. Mol Microbiol 77:730–742. https://doi.org/10.1111/j.1365-2958.2010.07243.x
Palcekova Z, Angala SK, Belardinelli JM et al (2019) Disruption of the SucT acyltransferase in Mycobacterium smegmatis abrogates succinylation of cell envelope polysaccharides. J Biol Chem 294:10325–10335. https://doi.org/10.1074/jbc.RA119.008585
Pashley CA, Parish T, McAdam RA et al (2003) Gene replacement in mycobacteria by using incompatible plasmids. Appl Environ Microbiol 69:517–523. https://doi.org/10.1128/aem.69.1.517-523.2003
Pei JF, Qi N, Li YX et al (2021) RegX3-mediated regulation of methylcitrate cycle in Mycobacterium smegmatis. Front Microbiol 12:619387. https://doi.org/10.3389/fmicb.2021.619387
Peng H, Wang Y, Jiang K et al (2021) A dual role reductase from phytosterols catabolism enables the efficient production of valuable steroid precursors. Angew Chem Int Ed Engl 60:5414–5420. https://doi.org/10.1002/anie.202015462
Pietersen RD, du Preez I, Loots DT et al (2020) Tween 80 induces a carbon flux rerouting in Mycobacterium tuberculosis. J Microbiol Methods 170:105795. https://doi.org/10.1016/j.mimet.2019.105795
Putkaradze N, Litzenburger M, Hutter MC et al (2019) CYP109E1 from Bacillus megaterium acts as a 24-and 25-hydroxylase for cholesterol. ChemBioChem 20:655–658. https://doi.org/10.1002/cbic.201800595
Rahlwes KC, Osman SH, Morita YS (2020) Role of LmeA, a mycobacterial periplasmic protein, in maintaining the mannosyltransferase MptA and its product lipomannan under stress. mSphere. https://doi.org/10.1128/mSphere.01039-20
Rank L, Herring LE, Braunstein M (2021) Evidence for the mycobacterial mce4 transporter being a multiprotein complex. J Bacteriol 203:e00685-e1620. https://doi.org/10.1128/JB.00685-20
Rego AM, Alves da Silva D, Ferreira NV et al (2021) Metabolic profiles of multidrug resistant and extensively drug resistant Mycobacterium tuberculosis unveiled by metabolomics. Tuberculosis (edinb) 126:102043. https://doi.org/10.1016/j.tube.2020.102043
Richard-Greenblatt M, Bach H, Adamson J et al (2015) Regulation of ergothioneine biosynthesis and its effect on Mycobacterium tuberculosis growth and infectivity. J Biol Chem 290:23064–23076. https://doi.org/10.1074/jbc.M115.648642
Rohman A, Dijkstra BW (2021) Application of microbial 3-ketosteroid Delta(1)-dehydrogenases in biotechnology. Biotechnol Adv 49:107751. https://doi.org/10.1016/j.biotechadv.2021.107751
Rosloniec KZ, Wilbrink MH, Capyk JK et al (2009) Cytochrome P450 125 (CYP125) catalyses C26-hydroxylation to initiate sterol side-chain degradation in Rhodococcus jostii RHA1. Mol Microbiol 74:1031–1043. https://doi.org/10.1111/j.1365-2958.2009.06915.x
Sao Emani C, Williams MJ, Van Helden PD et al (2018) Gamma-glutamylcysteine protects ergothioneine-deficient Mycobacterium tuberculosis mutants against oxidative and nitrosative stress. Biochem Biophys Res Commun 495:174–178. https://doi.org/10.1016/j.bbrc.2017.10.163
Savvi S, Warner DF, Kana BD et al (2008) Functional characterization of a vitamin B-12-dependent methylmalonyl pathway in Mycobacterium tuberculosis: Implications for propionate metabolism during growth on fatty acids. J Bacteriol 190:3886–3895. https://doi.org/10.1128/Jb.01767-07
Schiffer L, Anderko S, Hannemann F et al (2015) The CYP11B subfamily. J Steroid Biochem 151:38–51. https://doi.org/10.1016/j.jsbmb.2014.10.011
Shao M, Zhang X, Rao Z et al (2015) Enhanced production of androst-1,4-diene-3,17-dione by Mycobacterium neoaurum JC-12 using three-stage fermentation strategy. PLoS ONE 10:e0137658. https://doi.org/10.1371/journal.pone.0137658
Shao M, Sha Z, Zhang X et al (2017) Efficient androst-1,4-diene-3,17-dione production by co-expressing 3-ketosteroid-Delta(1)-dehydrogenase and catalase in Bacillus subtilis. J Appl Microbiol 122:119–128. https://doi.org/10.1111/jam.13336
Shao M, Zhang X, Rao Z et al (2019a) Identification of steroid C27 monooxygenase isoenzymes involved in sterol catabolism and stepwise pathway engineering of Mycobacterium neoaurum for improved androst-1,4-diene-3,17-dione production. J Ind Microbiol Biotechnol 46:635–647. https://doi.org/10.1007/s10295-018-02135-5
Shao M, Zhao Y, Liu Y et al (2019b) Intracellular environment improvement of Mycobacterium neoaurum for enhancing androst-1,4-diene-3,17-dione production by manipulating NADH and reactive oxygen species levels. Molecules. https://doi.org/10.3390/molecules24213841
Shtratnikova VY, Sсhelkunov MI, Fokina VV et al (2021) Different genome-wide transcriptome responses of Nocardioides simplex VKM Ac-2033D to phytosterol and cortisone 21-acetate. BMC Biotechnol 21:7. https://doi.org/10.1186/s12896-021-00668-9
Song CW, Lee J, Lee SY (2015) Genome engineering and gene expression control for bacterial strain development. Biotechnol J 10:56–68. https://doi.org/10.1002/biot.201400057
Sripalakit P, Wichai U, Saraphanchotiwitthaya A (2006) Biotransformation of various natural sterols to androstenones by Mycobacterium sp. and some steroid-converting microbial strains. J Mol Catal B Enzym 41:49–54. https://doi.org/10.1016/j.molcatb.2006.04.007
Su L, Shen Y, Xia M et al (2018) Overexpression of cytochrome p450 125 in Mycobacterium: a rational strategy in the promotion of phytosterol biotransformation. J Ind Microbiol Biotechnol 45:857–867. https://doi.org/10.1007/s10295-018-2063-z
Sun B, Yang J, Yang S et al (2018) A CRISPR-Cpf1-assisted non-homologous end joining genome editing system of Mycobacterium smegmatis. Biotechnol J 13:e1700588. https://doi.org/10.1002/biot.201700588
Sun WJ, Liu YJ, Liu HH et al (2019a) Enhanced conversion of sterols to steroid synthons by augmenting the peptidoglycan synthesis gene pbpB in Mycobacterium neoaurum. J Basic Microbiol 59:924–935. https://doi.org/10.1002/jobm.201900159
Sun WJ, Wang L, Liu HH et al (2019b) Characterization and engineering control of the effects of reactive oxygen species on the conversion of sterols to steroid synthons in Mycobacterium neoaurum. Metab Eng 56:97–110. https://doi.org/10.1016/j.ymben.2019.09.004
Szaleniec M, Wojtkiewicz AM, Bernhardt R et al (2018) Bacterial steroid hydroxylases: enzyme classes, their functions and comparison of their catalytic mechanisms. Appl Microbiol Biotechnol 102:8153–8171. https://doi.org/10.1007/s00253-018-9239-3
Tahlan K, Wilson R, Kastrinsky DB et al (2012) SQ109 targets MmpL3, a membrane transporter of trehalose monomycolate involved in mycolic acid donation to the cell wall core of Mycobacterium tuberculosis. Antimicrob Agents Chemother 56:1797–1809. https://doi.org/10.1128/AAC.05708-11
Tang Y, Fu Y (2018) Class 2 CRISPR/Cas: an expanding biotechnology toolbox for and beyond genome editing. Cell & Biosci. https://doi.org/10.1186/s13578-018-0255-x
Tricarico PM, Caracciolo I, Gratton R et al (2019) 25-hydroxycholesterol reduces inflammation, viral load and cell death in ZIKV-infected U-87 MG glial cell line. Inflammopharmacology 27:621–625. https://doi.org/10.1007/s10787-018-0517-6
Tyagi P, Dharmaraja AT, Bhaskar A et al (2015) Mycobacterium tuberculosis has diminished capacity to counteract redox stress induced by elevated levels of endogenous superoxide. Free Radic Biol Med 84:344–354. https://doi.org/10.1016/j.freeradbiomed.2015.03.008
Uhia I, Galan B, Kendall SL et al (2012) Cholesterol metabolism in Mycobacterium smegmatis. Environ Microbiol Rep 4:168–182. https://doi.org/10.1111/j.1758-2229.2011.00314.x
Wang Z, Zhao F, Chen D et al (2006) Biotransformation of phytosterol to produce androsta-diene-dione by resting cells of Mycobacterium in cloud point system. Process Biochem 41:557–561. https://doi.org/10.1016/j.procbio.2005.09.014
Wang H, Song S, Peng F et al (2020) Whole-genome and enzymatic analyses of an androstenedione-producing Mycobacterium strain with residual phytosterol-degrading pathways. Microb Cell Fact 19:187. https://doi.org/10.1186/s12934-020-01442-w
Wang D, Zhang J, Cao D-D et al (2021) Identification and in situ removal of an inhibitory intermediate to develop an efficient phytosterol bioconversion process using a cyclodextrin-resting cell system. RSC Adv 11:24787–24793. https://doi.org/10.1039/d1ra02774c
Wilbrink, M.H., 2011. Microbial sterol side chain degradation in Actinobacteria. Dissertation, University of Groningen.
Wollam J, Antebi A (2011) Sterol regulation of metabolism, homeostasis, and development. Annu Rev Biochem 80:885–916. https://doi.org/10.1146/annurev-biochem-081308-165917
Xiao J, Jia H, Pan L et al (2019) Application of the CRISPRi system to repress sepF expression in Mycobacterium smegmatis. Infect Genet Evol 72:183–190. https://doi.org/10.1016/j.meegid.2018.06.033
Xiao X, He J-K, Guan Y-X et al (2020) Effect of cholinium amino acids ionic liquids as cosolvents on the bioconversion of phytosterols by Mycobacterium sp. resting cells. ACS Sustain Chem Eng 8:17124–17132. https://doi.org/10.1021/acssuschemeng.0c05296
Xiong LB, Liu HH, Xu LQ et al (2017) Improving the production of 22-hydroxy-23,24-bisnorchol-4-ene-3-one from sterols in Mycobacterium neoaurum by increasing cell permeability and modifying multiple genes. Microb Cell Fact 16:89. https://doi.org/10.1186/s12934-017-0705-x
Xiong LB, Liu HH, Song XW et al (2020a) Improving the biotransformation of phytosterols to 9alpha-hydroxy-4-androstene-3,17-dione by deleting embC associated with the assembly of cell envelope in Mycobacterium neoaurum. J Biotechnol 323:341–346. https://doi.org/10.1016/j.jbiotec.2020.09.019
Xiong LB, Liu HH, Zhao M et al (2020b) Enhancing the bioconversion of phytosterols to steroidal intermediates by the deficiency of kasB in the cell wall synthesis of Mycobacterium neoaurum. Microb Cell Fact 19:80. https://doi.org/10.1186/s12934-020-01335-y
Xu S, Li Y (2020) Yeast as a promising heterologous host for steroid bioproduction. J Ind Microbiol Biotechnol 47:829–843. https://doi.org/10.1007/s10295-020-02291-7
Xu LQ, Liu YJ, Yao K et al (2016) Unraveling and engineering the production of 23,24-bisnorcholenic steroids in sterol metabolism. Sci Rep 6:21928. https://doi.org/10.1038/srep21928
Xu LX, Yang HL, Kuang MA et al (2017) Comparative genomic analysis of Mycobacterium neoaurum MN2 and MN4 substrate and product tolerance. 3Biotech 7:181. https://doi.org/10.1007/s13205-017-0818-2
Xu JY, Zhao L, Xu Y et al (2020) Dynamic characterization of protein and posttranslational modification levels in mycobacterial cholesterol catabolism. mSystems. https://doi.org/10.1128/mSystems.00424-19
Yan MY, Yan HQ, Ren GX et al (2017) CRISPR-Cas12a-assisted recombineering in bacteria. Appl Environ Microbiol. https://doi.org/10.1128/AEM.00947-17
Yao K, Wang FQ, Zhang HC et al (2013) Identification and engineering of cholesterol oxidases involved in the initial step of sterols catabolism in Mycobacterium neoaurum. Metab Eng 15:75–87. https://doi.org/10.1016/j.ymben.2012.10.005
Yao K, Xu LQ, Wang FQ et al (2014) Characterization and engineering of 3-ketosteroid- big up tri, open1-dehydrogenase and 3-ketosteroid-9alpha-hydroxylase in Mycobacterium neoaurum ATCC 25795 to produce 9alpha-hydroxy-4-androstene-3,17-dione through the catabolism of sterols. Metab Eng 24:181–191. https://doi.org/10.1016/j.ymben.2014.05.005
Yeware AM, Shurpali KD, Athalye MC et al (2017) Superoxide generation and its involvement in the growth of Mycobacterium smegmatis. Front Microbiol 8:105. https://doi.org/10.3389/fmicb.2017.00105
Zhang Y, Wang Y, Yao MD et al (2017) Improved campesterol production in engineered Yarrowia lipolytica strains. Biotechnol Lett 39:1033–1039. https://doi.org/10.1007/s10529-017-2331-4
Zhang R, Liu X, Wang Y et al (2018) Identification, function, and application of 3-ketosteroid Delta1-dehydrogenase isozymes in Mycobacterium neoaurum DSM 1381 for the production of steroidic synthons. Microb Cell Fact 17:77. https://doi.org/10.1186/s12934-018-0916-9
Zhang R, Zhang Y, Wang Y et al (2019a) Pregnenolone overproduction in Yarrowia lipolytica by integrative components pairing of the cytochrome p450scc system. Acs Synth Biol 8:2666–2678. https://doi.org/10.1021/acssynbio.9b00018
Zhang X, Zhu M, Han R et al (2019b) A novel 3-phytosterone-9alpha-hydroxylase oxygenation component and its application in bioconversion of 4-androstene-3,17-dione to 9alpha-hydroxy-4-androstene-3,17-dione coupling with a nadh regeneration formate dehydrogenase. Molecules. https://doi.org/10.3390/molecules24142534
Zhang Y, Zhou X, Wang X et al (2020) Improving phytosterol biotransformation at low nitrogen levels by enhancing the methylcitrate cycle with transcriptional regulators PrpR and GlnR of Mycobacterium neoaurum. Microb Cell Fact 19:13. https://doi.org/10.1186/s12934-020-1285-8
Zhao A, Zhang X, Li Y et al (2021) Mycolicibacterium cell factory for the production of steroid-based drug intermediates. Biotechnol Adv 53:107860. https://doi.org/10.1016/j.biotechadv.2021.107860
Zhou X, Zhang Y, Shen Y et al (2019a) Efficient production of androstenedione by repeated batch fermentation in waste cooking oil media through regulating NAD(+)/NADH ratio and strengthening cell vitality of Mycobacterium neoaurum. Bioresour Technol 279:209–217. https://doi.org/10.1016/j.biortech.2019.01.144
Zhou X, Zhang Y, Shen Y et al (2019b) Economical production of androstenedione and 9alpha-hydroxyandrostenedione using untreated cane molasses by recombinant mycobacteria. Bioresour Technol 290:121750. https://doi.org/10.1016/j.biortech.2019.121750
Zhou X, Zhang Y, Shen Y et al (2020) Efficient repeated batch production of androstenedione using untreated cane molasses by Mycobacterium neoaurum driven by ATP futile cycle. Bioresour Technol 309:123307. https://doi.org/10.1016/j.biortech.2020.123307
Funding
This research was supported by the National Key Research and Development Project of China (2019YFA0905300).
Author information
Authors and Affiliations
Contributions
All the authors were involved in the concept and design of this review. XXW: Conceptualization and original draft eriting. XK: Review and editing. ZQL: Review, editing and supervision. YGZ: Review, editing and supervision.
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, XX., Ke, X., Liu, ZQ. et al. Rational development of mycobacteria cell factory for advancing the steroid biomanufacturing. World J Microbiol Biotechnol 38, 191 (2022). https://doi.org/10.1007/s11274-022-03369-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03369-3