Abstract
Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disease characterized by the aggregation of ubiquitinated proteins in affected motor neurons. Recent studies have identified several new molecular constituents of ALS-linked cellular aggregates, including FUS, TDP-43, OPTN, UBQLN2 and the translational product of intronic repeats in the gene C9ORF72. Mutations in the genes encoding these proteins are found in a subgroup of ALS patients and segregate with disease in familial cases, indicating a causal relationship with disease pathogenesis. Furthermore, these proteins are often detected in aggregates of non-mutation carriers and those observed in other neurodegenerative disorders, supporting a widespread role in neuronal degeneration. The molecular characteristics and distribution of different types of protein aggregates in ALS can be linked to specific genetic alterations and shows a remarkable overlap hinting at a convergence of underlying cellular processes and pathological effects. Thus far, self-aggregating properties of prion-like domains, altered RNA granule formation and dysfunction of the protein quality control system have been suggested to contribute to protein aggregation in ALS. The precise pathological effects of protein aggregation remain largely unknown, but experimental evidence hints at both gain- and loss-of-function mechanisms. Here, we discuss recent advances in our understanding of the molecular make-up, formation, and mechanism-of-action of protein aggregates in ALS. Further insight into protein aggregation will not only deepen our understanding of ALS pathogenesis but also may provide novel avenues for therapeutic intervention.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) is a fatal neurodegenerative disease caused by the loss of both upper and lower motor neurons. Affected patients develop progressive muscle weakness eventually leading to death due to respiratory failure, typically 3–5 years after symptom onset. ALS affects ~2 out of 100,000 individuals per year [76]. In the majority of patients the disease occurs sporadic and is referred to as sporadic ALS (SALS). In 5 % of cases there is a family history of ALS (FALS) [29]. The presence of protein aggregates in affected motor neurons is a characteristic, but still poorly understood hallmark of SALS and FALS patients. Recently, many new ALS causing gene defects have been identified including mutations in HNRNPA1, PFN1, C9ORF72, UBQLN2, OPTN, VCP, FUS and TARDBP [1, 105]. Most of these mutations are rare and cause ALS in a small subgroup of patients. Remarkably, however, the proteins encoded by these genes are present in protein aggregates of a large proportion of non-mutation carriers indicating a more widespread role for their abnormal localization in ALS pathogenesis (Table 1). Moreover, some of these proteins are present in pathological aggregates of other neurodegenerative disorders such as frontotemporal lobar degeneration (FTLD), spinocerebellar ataxia (SCA), Huntington’s disease, Alzheimer’s disease and inclusion body myositis (IBM), indicating a more general involvement in neurodegeneration.
Despite clear evidence that protein aggregation is central to the pathology of ALS many questions remain about the role, formation and mechanism-of-action of protein aggregates in ALS. What drives deposition of proteins in ALS? Which cellular mechanisms contribute to protein aggregation or are affected by it? Furthermore, what is the role of proteins carrying ALS-associated mutations in aggregate formation? Pathological, cell culture and animal studies are now beginning to provide insights into these important questions. We will give an overview of the characteristics of aggregates observed in motor neurons of ALS patients and the relationship between protein aggregation, neurotoxicity and disease severity in cellular and animal model systems. We will then discuss possible underlying molecular mechanisms in protein aggregation and neuronal degeneration and provide directions for future research. Although an ever-increasing number of proteins is being implicated in ALS pathogenesis, the emphasis here is on the most recently discovered proteins and those present in spinal cord aggregates.
Novel insights into the molecular makeup and formation of cellular aggregates in ALS
The central pathological hallmark of ALS is the presence of cytoplasmic inclusions or aggregates in degenerating motor neurons and surrounding oligodendrocytes. Inclusions are not restricted to the spinal cord but also present in other brain regions such as the frontal and temporal cortices, hippocampus and cerebellum [1]. The predominant aggregates found in ALS patients are ubiquitinated aggregates that are classified as either Lewy body-like hyaline inclusions or skein-like inclusions. At the ultrastructural level, Lewy body-like or skein-like inclusions appear as randomly oriented filaments covered by fine granules [78, 129, 166]. Additional subclasses of aggregates found in ALS are Bunina bodies, which are small eosinophilic ubiquitin-negative inclusions [158] and round hyaline inclusions without a halo. Bunina bodies consist of amorphous electron-dense material surrounded by tubular and vesicular structures [158]. Furthermore, neurofilamentous inclusions are found in the axon hillock in close proximity to ubiquitinated inclusions. Other cellular abnormalities include the presence of mitochondrial vacuolization, fragmentation of the Golgi apparatus and abnormalities at the neuromuscular junction. In 1993, SOD1 was the first protein to be identified to aggregate in FALS cases carrying a mutation in the SOD1 gene [167]. Later, mutations in VAPB were also shown to cause ALS in a group of FALS patients [154]. Due to exponential development of genetic techniques, several new proteins have been identified to be involved in ALS pathophysiology during the past few years, including TDP-43, FUS, OPTN, UBQLN2 and C9ORF72. In the following sections, we will discuss for each of these proteins the characteristics of the aggregated protein, their physiological functions and effects in ALS disease models.
TAR DNA-binding protein 43 (TDP43)
Following the identification of SOD1 aggregates in a small subset of ALS patients, a breakthrough was achieved in 2006 with the identification of TDP-43 as a major component of ubiquitinated inclusions in FTLD and ALS cases [6, 150]. Non-mutated TDP-43 is found in aggregates in spinal cord motor neurons, hippocampal and frontal cortex neurons and glial cells in all SALS patients and the vast majority of SOD-1-negative FALS patients, but not in SOD1 related ALS [133, 181] (Table 1). In frontotemporal dementia (FTD), TDP-43 aggregates are present in the most common subtype of the disease, FTLD with ubiquitinated inclusions, now referred to as FTLD-TDP [150]. ALS and FTD show a remarkable overlap at the genetic, symptomatic and pathological level and they may actually reflect two ends of a disease spectrum [33, 66]. TDP-43 is also found to accumulate in Alzheimer’s disease, Lewy body disease and SCA2, secondary to other molecules [3, 7, 60, 82] and even in normal control subjects over the age of 65 [67].
Normally TDP-43 predominantly localizes to the nucleus. In ALS patients, cytoplasmic aggregation is often accompanied by nuclear clearing of the protein [133, 150]. Furthermore, the protein is cleaved in C-terminal fragments of 18–26 and 35 kDa and the full-length protein and 18–26 kDa fragments are hyperphosphorylated [78, 91, 150]. Genetic studies have identified mutations in the gene encoding TDP-43, TARDBP, in 1–2 % of FALS and SALS cases [100, 192]. In non-ALS inclusions TDP-43 is also cleaved and hyperphosphorylated. However, banding patterns of TDP-43 cleavage products are distinct between ALS and FTLD-B and other FTLD subtypes [78, 91, 189]. Furthermore, the distribution of the TDP-43-positive aggregates is disease-specific with, for example, involvement of spinal cord motor neurons in ALS and a more widespread distribution in the brain in FTLD [12]. Mutations in TARDBP are unique to ALS and are not found in other neurodegenerative disorders [28, 168, 192] with the exception of a small number of FTD cases [18, 23, 24, 34, 35, 38, 111, 145].
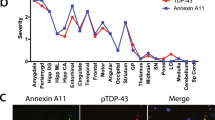
TDP-43 is a DNA and RNA binding protein that binds around 30 % of the mouse transcriptome with a preference for long UG-rich sequences [163, 185] (Fig. 1). Target sequences are mainly intronic, but also include non-coding RNAs and 3′ UTRs. In line with this, TDP-43 plays a role in nuclear RNA metabolism including splicing, transcriptional repression, miRNA synthesis, mRNA nucleo-cytoplasmic shuttling and RNA transport [117]. The majority of TDP-43 mutations identified to date are localized in the C-terminal glycine-rich domain of the protein (Fig. 1). This domain binds other hnRNPs and is important for the splicing activity of TDP-43 [41].
Schematic representation of the domain organization of TDP-43, FUS, ATXN2, OPTN and UBQLN2. Different protein domains are indicated in different colors (see legend). The location of ALS-associated genetic alterations is depicted by sticks (mutations) or arrowheads (repeats). For C9ORF72 intronic and exonic regions are depicted; an intronic hexanucleotide repeat is causative of ALS/FTD (arrowhead). Boxes show a selection of relevant interactors for each of the depicted proteins
Purified TDP-43 is prone to aggregation and this aggregation tendency depends on its C-terminal domain and is enhanced by ALS-linked TDP-43 mutations [94, 95]. In line with endogenous staining patterns, overexpression of TDP-43 causes a predominant nuclear localization [99, 130, 176, 199]. In addition, a small proportion of primary neuronal cells shows cytoplasmic localization, which is enhanced when TDP-43 mutants are overexpressed [13, 54, 73, 99]. Recently, the first induced pluripotent stem cell (iPSC)-derived motor neurons were generated from fibroblasts of patients carrying a M337V TARDBP mutation [22]. Similar to control, the M337V TARDBP line showed predominant nuclear localization of TDP-43 but with additional granular staining in soma and neurites [22].
Overexpression of WT TDP-43 has a toxic effect in yeast and cultured primary neurons derived from rat or mouse embryo’s. This effect is even more prominent when mutant TDP-43 is overexpressed [13, 54, 73, 95, 99]. In most animal studies, overexpression of WT or mutant TDP-43 induces a motor phenotype and reduces life span, but results so far are limited to the toxic effects of TDP-43 overexpression [12] (Supplementary Table). Although some studies report on increased toxicity in mutant as compared to WT transgenic lines [9, 180, 222], other studies do not report such differences [177, 212, 213]. As the toxic effects of TDP-43 are clearly dose-dependent [177, 208, 213], some of these results may be dependent on the level of expression rather than on TDP-43 mutation-specific toxic effects. Furthermore, while it has been reported that TDP-43 WT overexpression has motor neuron-specific toxic effects coinciding with the presence of nuclear and cytoplasmic inclusions and pathological phosphorylation and cleavage of TDP-43 [89, 208, 213], overexpression of TDP-43 can be toxic without aggregate formation [9, 205].
Several factors modulate TDP-43 toxicity. First, the TDP-43 C-terminal fragments that are found in ALS spinal cord aggregates are particularly aggregation-prone and toxic when overexpressed [13, 54, 88, 156, 221]. Second, the cytoplasmic distribution of TDP-43 has been reported to be related to cell death whereas inclusion body formation or nuclear TDP-43 levels were not [13]. In line with this, mutations of the nuclear localization signal (NLS) of TDP-43 confer toxicity while NES mutations abrogate toxicity [13]. Third, phosphorylation of TDP-43 may modulate aggregation and toxicity of the protein [26, 126]. In C. elegans blockage of TDP-43 phosphorylation ameliorates the neurodegenerative effect of ALS-associated TDP-43 mutants [126, 127]. In contrast, another study reported that mutation of phosphorylation sites increased aggregate formation in cells and in Drosophila, whereas hyperphosphorylation reduced aggregation and toxicity [123]. In addition, mutation of TDP-43 phosphorylation sites does not affect C-terminal fragment formation, the formation of cytoplasmic inclusions or survival in cells [51, 153, 221]. Thus, although it is clear that TDP-43 plays a crucial and central role in ALS pathology many questions remain about its pathological mechanisms in ALS.
Fused in sarcoma/translocated in liposarcoma (FUS/TLS)
The discovery of ALS-linked mutations in TDP-43 fueled the identification of mutations in another RNA binding protein, FUS, in FALS patients [116, 196]. A series of genetic studies of large ALS cohorts showed that FUS mutations account for 4 % of FALS and 1 % of SALS cases and that these are in part associated with young-onset disease [117]. Pathological examination of post-mortem tissue of FUS mutation carriers shows predominant degeneration of lower motor neurons with FUS-positive cytoplasmic inclusions and a normal distribution of TDP-43, thereby distinguishing them from other ALS cases [72, 80, 116, 196]. The precise pattern of FUS-immunoreactivity in ALS cases without FUS mutations is still unclear (Table 1). Some studies report that FUS is not present in SALS patients and SOD1 FALS [87, 149, 196], while others show FUS-positive inclusions with signals for TDP-43, p62 and ubiquitin in all SALS and FALS cases, except for SOD1 mutation carriers [48, 103]. In contrast to TDP-43, biochemical analyses show that mutant FUS protein itself is not ubiquitinated, hyperphosphorylated or cleaved. However, the protein is enriched in the insoluble fraction of FTLD-FUS brains [149]. Furthermore, nuclear clearing of FUS is not as evident as observed for TDP-43.
FUS has also been detected in aggregates found in other neurodegenerative disorders including FTLD, Huntington’s disease and SCA indicating that disruption of WT FUS function is associated with neurodegenerative disease in general [50, 149]. However, in contrast to ALS, FTLD patients with FUS inclusions only rarely harbor genetic alterations in FUS [195]. Interestingly, FUS-positive inclusions in FTLD cases are immunoreactive for TAF15 and EWS, other members of the FET family of RNA binding proteins, and for transportin-1 [147, 151]. Protein aggregates in ALS cases with FUS mutations do not stain for these proteins.
FUS is a nuclear RNA binding protein and preferentially binds pre-mRNA at intronic sites, but also long non-coding RNAs, exons and 3′UTRs [84] (Fig. 1). Most of the ALS-linked FUS mutations reported to date are localized in the NLS of the protein resulting in impaired nuclear transport of FUS [52]. FUS mediates a wide range of cellular processes including DNA repair, transcription, splicing and miRNA processing [117]. The protein shuttles between the nucleus and the cytoplasm to function in the transport of mRNA [224].
Purified FUS is aggregation prone, but in contrast to TDP-43 this property may not be influenced by ALS-linked mutations [179]. In yeast, cytosolic aggregation of FUS depends on its N-terminal, RRM and first RGG domains (Fig. 1). Toxicity is dependent on the N-terminal and first RGG domains [97, 179]. It should be noted, however, that the NLS of FUS is not fully recognized in yeast suggesting that this domain may still be involved in protein aggregation. Following overexpression in eukaryotic cells, WT FUS localizes to the nucleus, whereas FUS proteins carrying ALS mutations in the NLS form cytoplasmic aggregates, thereby mimicking human disease [116, 196]. This cytoplasmic relocalization is observed in some but not all FUS animal models reported to date [25, 32, 42, 86, 119, 142, 198, 203, 211] (Supplementary Table).
Deletion of the NES in FUS strongly reduces toxicity of mutant FUS in Drosophila [119], suggesting that the cytoplasmic localization of mutant FUS confers toxicity. However, another study reported that deletion of the NLS completely blocks FUS toxicity, as did the addition of a NES [211]. Blocking the RNA binding capacity of FUS also abolishes FUS toxicity in yeast [178] and ameliorates mutant FUS toxicity in Drosophila [42] (Supplementary Table). Although further work is needed to reveal to which extent mislocalization of mutant FUS contributes to disease, recent studies show that mutant FUS triggers stress granule formation and loss of nuclear GEMs, as will be discussed in more detail in the next section.
Optineurin (OPTN)
A study on Japanese ALS patients from consanguineous marriages reported mutations in OPTN to be associated with disease [135]. Although genetic variation in OPTN is rare in ALS patients in other populations, pathologic studies confirm a role for OPTN in ALS. In SALS cases, OPTN is present in cytoplasmic skein-like inclusions and colocalizes with ubiquitin, TDP-43, and possibly FUS [46, 85, 103, 135, 159] (Table 1). Conflicting results are obtained with respect to OPTN immunoreactivity in SOD1 and FUS mutation carriers. Two studies detected colocalization of OPTN with SOD1-positive inclusions in SOD1 mutation carriers [103, 135], while two other studies did not [46, 85]. Similarly, two studies reported OPTN immunoreactivity in FUS mutation carriers [92, 103], while a second group could not detect OPTN-positive inclusions [85].
OPTN is present in inclusions in several other neurodegenerative diseases such as ALS with dementia, Huntington’s disease, Alzheimer’s disease, Parkinson’s disease, Creutzfeldt-Jakob disease, multiple system atrophy and Pick disease [159].
OPTN functions as an inhibitor of NFκB-signaling [223], acts as an autophagy receptor [206] and participates in the regulation of vesicular trafficking and maintenance of the Golgi apparatus [169] (Fig. 1). Purified OPTN is not aggregation prone although strong overexpression in yeast results in aggregation of the protein and in toxicity [113]. This toxicity requires the Rab8 binding region but not the ubiquitin-binding domain of OPTN. Mutations identified in ALS patients so far include truncation mutations thought to act through loss-of-function mechanisms and missense mutations [17, 45, 90, 135, 141, 191]. OPTN E478G, carrying a mutation in the UBAN domain, looses its ability to bind K63-polyubiquitin or linear-polyubiquitin chains [206] and fails to inhibit NFκB [135]. Whereas exogenous WT OPTN localizes to LC3-positive vesicles upon autophagy induction [174, 206], OPTN E478G does not [206]. As the ubiquitin binding capacity of OPTN is necessary to serve as an LC3 adaptor, this likely reflects a loss of binding to autophagosomes [206]. However, whether the inability of OPTN E478G to bind ubiquitin is related to ALS pathophysiology is unknown. It is interesting to note that homozygous knock-in mice expressing a OPTN D477N mutant, which also lacks ubiquitin binding capacity, do not display an ALS-like phenotype [70]. The OPTN truncation mutants reported in ALS patients have been proposed to cause decreased OPTN protein levels. In this light it is interesting that knockdown of OPTN results in motor neuron phenotypes in zebrafish [110]. Whether incorporation of OPTN in ALS aggregates in non-mutation carriers also results in loss of function of the protein or merely reflects its role as an autophagic receptor warrants further investigation.
Ubiquilin-2 (UBQLN2)
Dysfunction of the ubiquitin–proteasome system (UPS) has been linked to ALS based on a variety of functional studies. The contribution of this process to motor neuron degeneration is further underlined by the recent identification of mutations in UBQLN2 in X-linked ALS/FTD [47]. In human spinal cord autopsy material of UBQLN2 mutation carriers, skein-like inclusions are positive for UBQLN2, ubiquitin, p62, TDP-43, FUS and OPTN but not SOD1 [47, 207] (Table 1). In cases with ALS-dementia with or without UBQLN2 mutations, UBQLN2-positive inclusions are found in the hippocampus which are absent in ALS cases without dementia indicating that UBQLN2 aggregation and neurodegeneration are linked [47]. Skein-like inclusions in spinal cord tissue from SALS and FALS patients with unknown mutations or mutations in SOD1, TDP-43 or FUS also stain positive for UBQLN2 [47, 207]. It is currently unknown whether UBQLN2 is present in aggregates in other neurodegenerative diseases and a first study did not detect mutations in UBQLN2 in FTD [79]. Whether the presence of UBQLN2 in ALS aggregates reflects a cellular attempt for protein degradation or is related to dysfunction of protein degradation pathways needs to be further investigated.
The exact function of UBQLN2 is unknown, but it has been implicated in protein degradation via both UPS and autophagy and in G-protein coupled receptor endocytosis [121] (Fig. 1). Overexpression of mutant UBQLN2 has been shown to result in impaired UPS function [47]. Most, but not all ALS-associated mutations identified in UBQLN2 to date involve proline residues in its PXX region, which is thought to be important for protein–protein-interactions. Further research is needed to determine how mutant UBQLN2 impairs protein degradation systems and which proteins are affected.
Ataxin-2 (ATXN2)
A yeast screen for modifiers of TDP-43 toxicity recently led to the discovery that extended polyQ repeats in ATXN2 are associated with ALS [53]. While ATXN2 normally harbors 21 or 22 polyQ repeats, and a polyQ length of 34 and higher is known to cause SCA2 [120], polyQ lengths between 27 and 33 are associated with ALS [53]. Spinal cord tissue of SALS patients shows an increased cytoplasmic accumulation of ATXN2, as compared to controls, but there is no difference between patients with normal or extended polyQ repeats [53] (Table 1). Furthermore, ATXN2 and TDP-43 colocalize in cytoplasmic inclusions in FTLD [53], and FUS and ATXN2 have been reported to colocalize in ALS [56].
ATXN2 functions in mRNA polyadenylation, stress granule formation, polyribosome assembly and miRNA synthesis [136, 157, 171] (Fig. 1). The pathological effect of extended ATXN2 polyQ repeats is likely due to a gain-of-function mechanism as ATXN2 knockout mice do not show overt neurological deficits [104]. In contrast to SCA2, the polyQ repeats associated with ALS are not pure, i.e., containing only CAG codons. Rather, they are composed of CAG codons interrupted by CAA codons [216]. As both codons encode for the same amino acid, the pathogenic effect of ATXN2 polyQ repeat extensions may reside at the mRNA level.
Overexpression of ATXN2 with intermediate length polyQ repeats does not affect ATXN2 localization [53]. However, there is evidence that subcellular distribution of overexpressed WT TDP-43 or mutant FUS is altered upon overexpression of ATXN2 with intermediate length polyQ repeats [53, 56, 152]. Furthermore, ATXN2 with intermediate length polyQ repeats enhances stress-induced activation of caspase-3 as well as cleavage and phosphorylation of TDP-43 [77]. So, there is evidence that ATXN2 intermediate polyQ repeat modulates ALS pathophysiology via its RNA-dependent interaction with FUS and TDP-43, but further research is needed to dissect the underlying molecular mechanisms.
C9ORF72
An intronic hexanucleotide repeat expansion in C9ORF72 was recently identified as the most prevalent cause of ALS, FTD and ALS-FTD [44, 164]. While in the wild type situation the gene harbors fewer than 25 repeats, the repeat region can be extended to several hundred or thousand repeats [15]. Although extended repeat lengths can also be detected in control cases, repeat expansions are strongly associated with ALS and FTD. Extended repeat lengths have also been reported in some cases of Alzheimer’s disease and Huntington disease-like syndrome [15, 134, 209]. C9ORF72 is a protein with unknown function, but shows homology to differentially expressed in normal and neoplasia (DENN), which is a GDP/GTP exchange factor (GEF) that activates Rab GTPases [122, 217] (Fig. 1). The expression pattern of C9ORF72 is unaltered in expanded repeat carriers, although the specificity of the available C9ORF72 antibodies is subject to debate (Table 1). TDP-43-negative, p62- and UBQLN-positive cytoplasmic and nuclear inclusions in the hippocampus, frontotemporal neocortex and cerebellum distinguish expanded repeat from non-expanded repeat carriers [2, 27]. Fascinatingly, these characteristic inclusions contain poly dipeptide repeat proteins generated by non-ATG-initiated translation from the C9ORF72 intronic repeat region [10, 144]. Whether and how these dipeptide repeat proteins mediate pathogenic effects are unknown and they probably represent one of several pathogenic mechanisms in C9ORF72-associated ALS. As a second mechanism, C9ORF72 RNA molecules containing extended repeats may accumulate and sequester RNA binding proteins preventing these proteins from exerting their crucial functions. Similar aggregation of mutant RNAs is observed in other repeat expansion disorders. In support of this, C9ORF72-containing RNA foci have been observed in 25 % of spinal and frontal cortical neurons of expanded repeat carriers compared to 1 % in controls [44]. This observation has, however, not been confirmed in a second, independent study [175]. It has been shown that the GGGGCC repeats present in C9ORF72 bind several RNA binding proteins and that one of these, hnRNPA3, localizes to the p62-positive/TDP-43-negative cytoplasmic inclusions observed in repeat carriers [143]. As a third mechanism, the repeat expansion may result in haploinsufficiency due to impaired transcription or splicing. This is supported by the finding that C9ORF72 protein levels are reduced in patients with increased repeat lengths [44, 68, 144, 193]. Finally, in a number of C9ORF72 repeat carriers with FTLD, tau pathology has been observed suggesting that the C9ORF72 repeat may influence tau protein [20, 106]. Further research is needed to address these different mechanisms in relation to ALS disease pathogenesis.
Molecular mechanisms underlying protein aggregation in ALS
When studying ALS pathophysiology it is essential, but very difficult, to distinguish cause and consequence in the cellular cascades driving protein aggregation. The recent discovery of new, disease-associated mutations that trigger protein aggregation or stability represent unique opportunities to further dissect the effect and mechanism-of-action of protein aggregation in ALS. Several important questions need to be addressed including (1) how are proteins sequestered into ALS aggregates and (2) how do these aggregates affect neuronal function? Several models addressing these questions have emerged and will be discussed below.
Low complexity domains in ALS proteins with aggregation-prone properties
FUS, TDP-43 and other RNA binding proteins (RBPs) contain domains with similarity to yeast prion domains (Figs. 1, 2). These domains are enriched for asparagine, glutamine, tyrosine and glycine residues and can adopt two conformational states: an unfolded and an aggregated state. Prion proteins in an aggregated state can sequester prion proteins in an unfolded state to adopt the aggregation-prone confirmation and as such aggregation can spread. It has been hypothesized that in ALS aggregation may propagate from one cell to the other in a comparable fashion [107, 162]. Aggregation of FUS and TDP-43 has been shown to rely on regions resembling prion domains [94, 102, 179] and mutations in TDP-43 associated with ALS occur mainly in its prion-like region (Fig. 1). Furthermore, there is direct evidence that ALS proteins share features with yeast prion proteins, e.g., that aggregated FUS or TDP-43 can sequester native protein [62, 64, 102, 215]. Finally, mutations in the prion-like domain of the RBP hnRNPA1 segregate with disease in FALS [105]. Yeast prions form amyloid deposits characterized by a secondary beta-sheet structure and stain positive for amyloid dyes. Although short synthetic TDP-43 and FUS peptides form amyloid-like fibrils in vitro [31, 73, 102] full-length TDP-43 does not [95]. Rather, purified full-length FUS and TDP-43 form pore-like oligomers and fibrils resembling the ultrastructure of the skein-like inclusions [95, 179]. However, although initial reports emphasized that ALS inclusions lack features of amyloid [148], two recent studies found that some TDP-43 positive inclusions stain positive for amyloid dyes [21, 166]. Furthermore, to date evidence for cell-to-cell spread of ALS aggregated proteins is lacking. Finally, toxicity induced by TDP-43 and FUS depends not only on prion-like domains but also on RNA binding properties [42, 53, 94, 179, 199]. In all, accumulating evidence hints at an important role for prion-like mechanisms in ALS pathogenesis but further studies are needed to uncover their precise mechanism-of-action and pathological effects.
Cellular mechanisms linked to protein aggregation in ALS. ALS-associated mutations result in cytoplasmic mislocalization or increased aggregation tendency thereby increasing the risk for aberrant aggregation. Proteins with domains of low complexity (prion-like domains) such as FUS, TDP-43 and HNRNPA1 are though to be intrinsically aggregation prone. Many of these proteins participate in RNA granule formation. ALS-associated mutants alter RNA granule formation, thereby interfering with the local translation of RNA. Moreover, sequestration of ALS-associated proteins and their interactors into cytoplasmic aggregates may result in a loss of function. Protein degradation by the UPS and autophagy is essential for the clearance of ubiquitinated proteins. Dysfunction of these systems, as has been suggested for mutant UBQLN2, OPTN and VCP, can lead to proteins deposits
Stress granule formation and ALS protein aggregation
Although FUS and TDP-43 have a predominant nuclear localization, both proteins rapidly shuttle between the nucleus and the cytoplasm and are actively transported into axons and colocalize with other RBPs [16, 54, 63, 186, 202, 224]. RBPs regulate local translation of mRNAs by forming granular RNA–protein complexes in which translation is repressed [5]. Several types of RNA granules can be distinguished: stress granules (SG), processing bodies (PBs) and so called neuronal or transport granules. These granules are highly dynamic structures and interaction and exchange of RBPs and transcripts occur between the different types of granules [5]. SGs or PBs are formed following polysome disassembly in response to stress, while neuronal granules serve in the transport of mRNA to their site of local translation [5]. Interestingly, it has been suggested that the aggregation-prone prion-like domains of RBPs assist in the dynamic movement of RBPs in and out of RNA granules [102].
ALS-associated mutations in TDP-43, FUS and ATXN2 are being linked to SGs (Fig. 2). SG formation depends on the RRM domains and C-terminal region of TDP-43, the same domains required for TDP-43 toxicity [19, 37]. Although TDP-43 is dispensable for SG formation [19, 37, 131], depletion of TDP-43 inhibits SG formation [137]. WT TDP-43 co-immunoprecipitates and colocalizes with stress granule markers, but the effect of mutant TDP-43 is less clear [37, 43, 49, 61, 83, 131, 137, 138, 160]. While two studies reported an increase in the number of cells containing SGs upon TDP-43 mutant overexpression [49, 131], another study failed to observe such an effect [137]. A fourth study did not detect cytoplasmic signals for WT or mutant TDP-43 and consequently no colocalization with SG markers, although a TDP-43 NLS mutant clearly localized to SGs [19]. Thus, WT TDP-43 likely plays a role in SG formation with some evidence for altered SG formation associated with mutant TDP-43.
In contrast to TDP-43, mutant but not WT FUS localizes to SGs [19, 25, 52, 108, 152, 197]. Whether mutant FUS induces SG formation or is recruited to SGs following stress is unclear [108, 152, 197]. Similarly, studies reporting the effect of FUS mutations or truncations on SG recruitment show conflicting results [19, 102, 108]. However, proteins involved in SG formation have been identified as modifiers of FUS toxicity in yeast screens [97, 179].
ATXN2 localizes to PBs and SGs following cellular stress and its Lsm domain and PAM2 motif are required for this localization [152, 157] (Fig. 1). It is unknown whether ATXN2 directly regulates RNA granule formation or function. However, overexpression of ATXN2 31Q increases the cytoplasmic localization of TDP-43 following heat shock [53] and increases the cytoplasmic localization of mutant FUS, but without affecting SG formation [56].
In conclusion, FUS, TDP-43 and ATXN2 have all been implicated in SG formation and there is evidence that alterations in SG formation are associated with ALS-associated mutant proteins. Interestingly, HNRNPA1 localizes to SGs as well and ALS-associated mutations in HNRNPA1 augment its incorporation into SGs [105]. However, while the formation of RNA granules is normally a reversible process, in ALS disrupted RNA granule formation is hypothesized to result in insoluble aggregates. The most convincing evidence for this comes from pathological examination of ALS spinal cords. Although one study did not detect colocalization of TDP-43 positive inclusions with SG markers in SALS motor neurons [37], three other studies, using more sensitive methods, convincingly did [19, 131, 200]. Furthermore, the SG markers PABP-1 and eIF4G colocalize with FUS aggregates in spinal cord of FUS mutation carriers [52]. Whether these aggregates have primarily formed as SGs or reflect a more general sequestration of interacting proteins in FUS and TDP-43 aggregates (see below), remains to be shown. Furthermore, how reversible SGs develop into insoluble aggregates is unknown. Increased cytoplasmic localization of FUS and TDP-43, increased aggregation tendency of mutant RBPs and phosphorylation of RBPs have been suggested to regulate their localization to SGs [102, 108, 138] and may be altered in ALS. An altered RNA binding preference of mutant RBPs could also underlie altered RNA granule formation [84]. How RNA granule formation and disassembly are dysregulated in ALS and which transcripts are affected by these processes will undoubtedly be a focus of further investigations.
Protein sequestering
Cytoplasmic aggregation of ALS proteins at ectopic sites in the cell may prevent these proteins from executing their normal function (Fig. 2). If this mechanism would be solely responsible for ALS pathogenesis, gene knockdown or knockout is expected to result in strong motor neuron phenotypes. Unfortunately, FUS and TDP-43 are essential for normal development and knockout of FUS or TDP-43 results in premature death in mice [81, 112, 115, 172]. There is some evidence that knockdown of FUS or TDP-43 results in motor dysfunction in Drosophila and zebrafish models [58, 98, 99, 128, 170, 203] (Supplementary Table). However, the relevance of these findings in the context ALS is unclear as knockdown often triggers abnormalities outside the nervous system as well and motor neuron degeneration is not consistently observed. Additional models are needed to study the effect of reduced levels of proteins like FUS and TDP-43 but it seems unlikely that reduced levels of these proteins alone cause ALS.
On the other hand, FUS and TDP-43 aggregation could exert a toxic effect via sequestration of multiple binding partners or even interactomes essential for neuronal function. FUS and TDP-43 interact and colocalize with many different proteins including SMN, gemin proteins and small nuclear ribonucleoprotein particles (snRNPs) [188, 214] (Fig. 1). The SMN complex can be detected in the cytoplasm, but also in nuclear foci called GEMs. Knockdown of FUS or TDP-43 results in the loss of GEMs [173, 188]. Furthermore, a reduced number of GEMs is observed in ALS spinal cord motor neurons [188], fibroblasts derived from FUS or TDP-43 mutation carriers [214] and ALS mouse models [65, 101, 173]. Therefore, sequestration of SMN in FUS or TDP-43 cytoplasmic aggregates could affect SMN levels and function.
The idea that aggregation of proteins, such as TDP-43, leads to the sequestering of other essential proteins is supported by a recent study identifying the lariat debranching enzyme Dbr1 as a modifier of TDP-43 toxicity [8]. Dbr1 normally mediates the degradation of intronic lariats. Knockdown of Dbr1 triggers the transport of excessive lariat RNAs into the cytoplasm where they bind TDP-43. Since this leads to a reduction in cellular toxicity it was proposed that knockdown of Dbr1 reduces TDP-43 toxicity by capturing TDP-43 and thereby diminishing sequestration of RNAs and RBPs in TDP-43 aggregates. Although currently functional evidence for RNA and protein sequestering by ALS-associated protein aggregates is scarce, these recent findings warrant a more extensive study of the ability of ALS-associated protein aggregates to sequester RNAs and RBPs and of the effect this has on motor neuron homeostasis.
Dysfunction of protein degradation pathways
Molecular chaperones, the UPS, and the autophagy-lysosome system function to monitor protein quality and protect cells from dysfunctional, malfolded or denatured proteins. The presence of ubiquitin, p62 and molecular chaperones in ALS aggregates implicates a role for all these three systems in ALS pathophysiology (Fig. 2).
Chaperone molecules assist in protein folding under physiological circumstances and prevent protein aggregation in response to stress. They also assist in protein degradation by the proteasome or autophagy. Chaperones, such as heat shock proteins, are upregulated in ALS spinal cord [4] and present in motor neuron aggregates [14, 204]. Interestingly, upregulation of molecular chaperones increases solubility and reduces toxicity of FUS [139] and TDP-43, especially of TDP-43 C-terminal fragments [40, 71]. In line with this, knockdown of molecular chaperones increases accumulation of pTDP-43 C-terminal fragments [220] and enhances toxicity of TDP-43 overexpression [219]. Thus, chaperone molecules are likely to play a significant role in aggregation of ALS proteins and in their toxic effects.
Ubiquitination not only marks proteins for degradation but also mediates intracellular signaling, e.g., NFκB activation [114]. As proteins present in ALS aggregates are ubiquitinated it has been postulated that they are marked for degradation by the UPS, but eventually deposit when dysfunctional protein levels exceed UPS capacity. In line with this, examination of ALS spinal cord tissue shows that ubiquitination occurs before accumulation starts and that inclusion formation inversely correlates to the number of motor neurons, indicating that deposition of ubiquitinated proteins relates to toxicity [69]. Consistently, proteasome inhibition has been found to increase endogenous TDP-43 levels, blocks degradation of overexpressed TDP-43 C-terminal fragments and enhances toxicity [88, 155, 194, 220]. Recently, motor neuron-specific knockout of the proteasome subunit Rpt3 was reported to result in the loss of spinal motor neurons and locomotor dysfunction in mice [183]. Interestingly, these mice contained basophilic, hyaline and skein-like inclusions positive for TDP-43, FUS, UBQLN2 and OPTN, indicating that dysfunction of the proteasome itself is sufficient to induce aggregation of ALS-associated proteins and motor neuron degeneration. The fact that mutations in UBQLN2 cause ALS and are associated with impaired UPS function further underlines the notion that proteasomal degradation does not only modulate but also may play a causative role in ALS pathogenesis.
p62 serves as an adaptor of autophagic degradation by binding both polyubiquitinated proteins and LC3, an important autophagosomal marker. Autophagy mediates the degradation of TDP-43 C-terminal fragments, but not full-length protein [30, 93, 182]. Interestingly, treatment of transgenic TDP-43 mice with autophagic activators reduces locomotor dysfunction, learning and memory deficits, and neuron loss [201]. This is accompanied by a decrease in cytosolic TDP-43 inclusions and in insoluble full-length and truncated TDP-43. This indicates that autophagic clearance of TDP-43 reduces neurotoxicity. Knockout mice lacking Atg7 or Atg5 in the central nervous system, genes essential for autophagy, display movement disorders, widespread neurodegeneration and ubiquitin-positive inclusions in a variety of brain regions [75, 109]. Motor neuron-specific knockout of these genes results in ubiquitinated and p62-positive inclusions, but do not stain for TDP-43, FUS, OPTN or UBQLN2 and do not result in motor neuron death [183]. This would suggest that autophagic disruption does not primarily underlie ALS pathogenesis. However, mutations in genes encoding autophagy regulators have been associated with ALS, i.e., VCP, p62, CHMP2B and UBQLN2 [47, 57, 96, 161]. Patients with mutations in SQSTM1, the gene encoding p62, show large round p62-positive inclusions in motor neurons with additional TDP-43 deposits which are p62-negative [184]. Furthermore, TDP-43 has been identified as a modifier of mutant VCP toxicity [165] and ubiquilins have been reported to modulate TDP-43 toxicity in Drosophila [74]. In conclusion, components of the protein degradation pathways have emerged as important modulators of protein aggregation and toxicity in ALS. Furthermore, mutations in genes encoding these components have been associated with ALS indicating a causal role in the disease. Future studies are needed to investigate how these processes are affected in ALS, as they may represent potential therapeutic targets.
Conclusions
Protein aggregation in affected motor neurons is a central hallmark of ALS, and recent genetic, cellular and histological studies have enlarged our understanding of the molecular composition of these aggregates. This has led to the identification of new ALS causing genes, has linked the composition of aggregates to specific genetic defects, and has provided starting points for further investigation of underlying molecular pathways. Unique pathological features have now been identified that distinguish FUS, SOD1 and C9ORF72 mutation carriers from other ALS cases. It is likely that with future discoveries further classification will be possible. TDP-43, FUS, p62, OPTN and UBQLN2 show a widespread distribution in ALS-linked aggregates. This could reflect a general role in pathogenesis or, on the contrary, question their specific relevance to disease. However, the fact that mutations in the genes encoding these proteins segregate with disease in FALS supports the idea that their dysfunction is linked to motor neuron degeneration and disease pathogenesis. The proteins identified to be present in ALS aggregates play a role in a wide range of cellular processes with a marked overlapping role for TDP-43, FUS and ATXN2 in RNA metabolism and for OPTN, UBQLN2 and VCP in protein quality control and degradation (Fig. 2). How a disturbance of these ubiquitously expressed proteins can result in motor-neuron-specific degeneration remains an unresolved issue in the field of ALS research. Although cellular and animal models confirm a role for aggregation in ALS, results are often contradictory and models fully recapitulating ALS pathogenesis are mostly lacking. This may in part be explained by the fact that many of these models rely on overexpression of the protein. New model systems, such as iPSC-generated patient-derived cell lines or inducible animal models, may help to overcome these problems. So far, TDP-43 and FUS have been investigated most extensively; future studies on the role of OPTN, UBQLN2, ATXN2 and C9ORF72 will further enlarge our understanding of the cellular processes underlying ALS. Processes underlying aggregation in ALS include enhanced intrinsic aggregation propensity of ALS proteins, RNA granule dysregulation and dysfunction of protein degradation pathways. A further understanding of these processes will not only deepen our understanding of ALS pathogenesis, but also may aid the development of novel therapeutic strategies for this disease.
References
Al-Chalabi A, Jones A, Troakes C, King A, Al-Sarraj S, van den Berg LH (2012) The genetics and neuropathology of amyotrophic lateral sclerosis. Acta Neuropathol 124:339–352. doi:10.1007/s00401-012-1022-4
Al-Sarraj S, King A, Troakes C et al (2011) p62 positive, TDP-43 negative, neuronal cytoplasmic and intranuclear inclusions in the cerebellum and hippocampus define the pathology of C9orf72-linked FTLD and MND/ALS. Acta Neuropathol 122:691–702. doi:10.1007/s00401-011-0911-2
Amador-Ortiz C, Lin WL, Ahmed Z et al (2007) TDP-43 immunoreactivity in hippocampal sclerosis and Alzheimer’s disease. Ann Neurol 61:435–445. doi:10.1002/ana.21154
Anagnostou G, Akbar MT, Paul P, Angelinetta C, Steiner TJ, de Belleroche J (2010) Vesicle associated membrane protein B (VAPB) is decreased in ALS spinal cord. Neurobiol Aging 31:969–985. doi:10.1016/j.neurobiolaging.2008.07.005
Anderson P, Kedersha N (2009) RNA granules: post-transcriptional and epigenetic modulators of gene expression. Nat Rev Mol Cell Biol 10:430–436. doi:10.1038/nrm2694
Arai T, Hasegawa M, Akiyama H et al (2006) TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun 351:602–611. doi:10.1016/j.bbrc.2006.10.093
Arai T, Mackenzie IR, Hasegawa M et al (2009) Phosphorylated TDP-43 in Alzheimer’s disease and dementia with Lewy bodies. Acta Neuropathol 117:125–136. doi:10.1007/s00401-008-0480-1
Armakola M, Higgins MJ, Figley MD et al (2012) Inhibition of RNA lariat debranching enzyme suppresses TDP-43 toxicity in ALS disease models. Nat Genet 44:1302–1309. doi:10.1038/ng.2434
Arnold ES, Ling S-C, Huelga SC et al (2013) ALS-linked TDP-43 mutations produce aberrant RNA splicing and adult-onset motor neuron disease without aggregation or loss of nuclear TDP-43. Proc Natl Acad Sci USA 110:E736–E745. doi:10.1073/pnas.1222809110
Ash PEA, Bieniek KF, Gendron TF et al (2013) Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron 77:639–646. doi:10.1016/j.neuron.2013.02.004
Ash PEA, Zhang YJ, Roberts CM et al (2010) Neurotoxic effects of TDP-43 overexpression in C. elegans. Hum Mol Genet 19:3206–3218. doi:10.1093/hmg/ddq230
Baloh RH (2011) TDP-43: the relationship between protein aggregation and neurodegeneration in amyotrophic lateral sclerosis and frontotemporal lobar degeneration. FEBS J 278:3539–3549. doi:10.1111/j.1742-4658.2011.08256.x
Barmada SJ, Skibinski G, Korb E, Rao EJ, Wu JY, Finkbeiner S (2010) Cytoplasmic mislocalization of TDP-43 is toxic to neurons and enhanced by a mutation associated with familial amyotrophic lateral sclerosis. J Neurosci 30:639–649. doi:10.1523/JNEUROSCI.4988-09.2010
Basso M, Samengo G, Nardo G et al (2009) Characterization of detergent-insoluble proteins in ALS indicates a causal link between nitrative stress and aggregation in pathogenesis. PLoS ONE 4:e8130. doi:10.1371/journal.pone.0008130.t002
Beck J, Poulter M, Hensman D et al (2013) Large C9orf72 hexanucleotide repeat expansions are seen in multiple neurodegenerative syndromes and are more frequent than expected in the UK population. Am J Hum Genet 92:345–353. doi:10.1016/j.ajhg.2013.01.011
Belly A, Moreau-Gachelin F, Sadoul R, Goldberg Y (2005) Delocalization of the multifunctional RNA splicing factor TLS/FUS in hippocampal neurones: exclusion from the nucleus and accumulation in dendritic granules and spine heads. Neurosci Lett 379:152–157. doi:10.1016/j.neulet.2004.12.071
Belzil VV, Daoud H, Desjarlais A et al (2011) Analysis of OPTN as a causative gene for amyotrophic lateral sclerosis. Neurobiol Aging 32(555):e513–e554. doi:10.1016/j.neurobiolaging.2010.10.001
Benajiba L, Le Ber I, Camuzat A et al (2009) TARDBP mutations in motoneuron disease with frontotemporal lobar degeneration. Ann Neurol 65:470–473. doi:10.1002/ana.21612
Bentmann E, Neumann M, Tahirovic S, Rodde R, Dormann D, Haass C (2012) Requirements for stress granule recruitment of fused in sarcoma (FUS) and TAR DNA-binding protein of 43 kDa (TDP-43). J Biol Chem 287:23079–23094. doi:10.1074/jbc.M111.328757
Bieniek KF, Murray ME, Rutherford NJ et al (2013) Tau pathology in frontotemporal lobar degeneration with C9ORF72 hexanucleotide repeat expansion. Acta Neuropathol 125:289–302. doi:10.1007/s00401-012-1048-7
Bigio EH, Wu JY, Deng HX et al (2013) Inclusions in frontotemporal lobar degeneration with TDP-43 proteinopathy (FTLD-TDP) and amyotrophic lateral sclerosis (ALS), but not FTLD with FUS proteinopathy (FTLD-FUS), have properties of amyloid. Acta Neuropathol 125:463–465. doi:10.1007/s00401-013-1089-6
Bilican B, Serio A, Barmada SJ et al (2012) Mutant induced pluripotent stem cell lines recapitulate aspects of TDP-43 proteinopathies and reveal cell-specific vulnerability. Proc Natl Acad Sci USA 109:5803–5808. doi:10.1073/pnas.1202922109
Borghero G, Floris G, Cannas A et al (2011) A patient carrying a homozygous p.A382T TARDBP missense mutation shows a syndrome including ALS, extrapyramidal symptoms, and FTD. Neurobiol Aging 32(2327):e2321–e2325. doi:10.1016/j.neurobiolaging.2011.06.009
Borroni B, Bonvicini C, Alberici A et al (2009) Mutation within TARDBP leads to frontotemporal dementia without motor neuron disease. Hum Mutat 30:E974–E983. doi:10.1002/humu.21100
Bosco DA, Lemay N, Ko HK et al (2010) Mutant FUS proteins that cause amyotrophic lateral sclerosis incorporate into stress granules. Hum Mol Genet 19:4160–4175. doi:10.1093/hmg/ddq335
Braak H, Ludolph A, Thal DR, Del Tredici K (2010) Amyotrophic lateral sclerosis: dash-like accumulation of phosphorylated TDP-43 in somatodendritic and axonal compartments of somatomotor neurons of the lower brainstem and spinal cord. Acta Neuropathol 120:67–74. doi:10.1007/s00401-010-0683-0
Brettschneider J, Deerlin VM, Robinson JL et al (2012) Pattern of ubiquilin pathology in ALS and FTLD indicates presence of C9ORF72 hexanucleotide expansion. Acta Neuropathol 123:825–839. doi:10.1007/s00401-012-0970-z
Brouwers N, Bettens K, Gijselinck I et al (2010) Contribution of TARDBP to Alzheimer’s disease genetic etiology. J Alzheimers Dis 21:423–430. doi:10.3233/JAD-2010-100198
Byrne S, Walsh C, Lynch C et al (2011) Rate of familial amyotrophic lateral sclerosis: a systematic review and meta-analysis. J Neurol Neurosurg Psychiatry 82:623–627. doi:10.1136/jnnp.2010.224501
Caccamo A, Majumder S, Deng JJ, Bai Y, Thornton FB, Oddo S (2009) Rapamycin rescues TDP-43 mislocalization and the associated low molecular mass neurofilament instability. J Biol Chem 284:27416–27424. doi:10.1074/jbc.M109.031278
Chen AKH, Lin RYY, Hsieh EZJ et al (2010) Induction of amyloid fibrils by the C-terminal fragments of TDP-43 in amyotrophic lateral sclerosis. J Am Chem Soc 132:1186–1187. doi:10.1021/ja9066207
Chen Y, Yang M, Deng J et al (2011) Expression of human FUS protein in Drosophila leads to progressive neurodegeneration. Protein Cell 2:477–486. doi:10.1007/s13238-011-1065-7
Chen-Plotkin AS, Lee VM, Trojanowski JQ (2010) TAR DNA-binding protein 43 in neurodegenerative disease. Nat Rev Neurol 6:211–220. doi:10.1038/nrneurol.2010.18
Chiang HH, Andersen PM, Tysnes OB, Gredal O, Christensen PB, Graff C (2012) Novel TARDBP mutations in Nordic ALS patients. J Hum Genet 57:316–319. doi:10.1038/jhg.2012.24
Chio A, Calvo A, Moglia C et al (2010) Amyotrophic lateral sclerosis-frontotemporal lobar dementia in 3 families with p.Ala382Thr TARDBP mutations. Arch Neurol 67:1002–1009. doi:10.1001/archneurol.2010.173
Collins M, Riascos D, Kovalik T et al (2012) The RNA-binding motif 45 (RBM45) protein accumulates in inclusion bodies in amyotrophic lateral sclerosis (ALS) and frontotemporal lobar degeneration with TDP-43 inclusions (FTLD-TDP) patients. Acta Neuropathol 124:717–732. doi:10.1007/s00401-012-1045-x
Colombrita C, Zennaro E, Fallini C et al (2009) TDP-43 is recruited to stress granules in conditions of oxidative insult. J Neurochem 111:1051–1061. doi:10.1111/j.1471-4159.2009.06383.x
Corrado L, Ratti A, Gellera C et al (2009) High frequency of TARDBP gene mutations in Italian patients with amyotrophic lateral sclerosis. Hum Mutat 30:688–694. doi:10.1002/humu.20950
Couthouis J, Hart MP, Shorter J et al (2011) A yeast functional screen predicts new candidate ALS disease genes. Proc Natl Acad Sci USA 108:20881–20890. doi:10.1073/pnas.1109434108
Crippa V, Sau D, Rusmini P et al (2010) The small heat shock protein B8 (HspB8) promotes autophagic removal of misfolded proteins involved in amyotrophic lateral sclerosis (ALS). Hum Mol Genet 19:3440–3456. doi:10.1093/hmg/ddq257
D’Ambrogio A, Buratti E, Stuani C et al (2009) Functional mapping of the interaction between TDP-43 and hnRNP A2 in vivo. Nucleic Acids Res 37:4116–4126. doi:10.1093/nar/gkp342
Daigle JG, Lanson NA, Smith RB et al (2013) RNA-binding ability of FUS regulates neurodegeneration, cytoplasmic mislocalization and incorporation into stress granules associated with FUS carrying ALS-linked mutations. Hum Mol Genet 22:1193–1205. doi:10.1093/hmg/dds526
Dammer EB, Fallini C, Gozal YM et al (2012) Coaggregation of RNA-binding proteins in a model of TDP-43 proteinopathy with selective RGG motif methylation and a role for RRM1 ubiquitination. PLoS ONE 7:e38658. doi:10.1371/journal.pone.0038658.t005
DeJesus-Hernandez M, Mackenzie IR, Boeve BF et al (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72:245–256. doi:10.1016/j.neuron.2011.09.011
Del Bo R, Tiloca C, Pensato V et al (2011) Novel optineurin mutations in patients with familial and sporadic amyotrophic lateral sclerosis. J Neurol Neurosurg Psychiatry 82:1239–1243. doi:10.1136/jnnp.2011.242313
Deng H-X, Bigio EH, Zhai H et al (2011) Differential involvement of optineurin in amyotrophic lateral sclerosis with or without SOD1 mutations. Arch Neurol 68:1057–1061. doi:10.1001/archneurol.2011.178
Deng HX, Chen W, Hong S-T et al (2011) Mutations in UBQLN2 cause dominant X-linked juvenile and adult-onset ALS and ALS/dementia. Nature 477:211–215. doi:10.1038/nature10353
Deng HX, Zhai H, Bigio EH et al (2010) FUS-immunoreactive inclusions are a common feature in sporadic and non-SOD1 familial amyotrophic lateral sclerosis. Ann Neurol 67:739–748. doi:10.1002/ana.22051
Dewey CM, Cenik B, Sephton CF et al (2011) TDP-43 is directed to stress granules by sorbitol, a novel physiological osmotic and oxidative stressor. Mol Cell Biol 31:1098–1108. doi:10.1128/MCB.01279-10
Doi H, Koyano S, Suzuki Y, Nukina N, Kuroiwa Y (2010) The RNA-binding protein FUS/TLS is a common aggregate-interacting protein in polyglutamine diseases. Neurosci Res 66:131–133. doi:10.1016/j.neures.2009.10.004
Dormann D, Capell A, Carlson AM et al (2009) Proteolytic processing of TAR DNA binding protein-43 by caspases produces C-terminal fragments with disease defining properties independent of progranulin. J Neurochem 110:1082–1094. doi:10.1111/j.1471-4159.2009.06211.x
Dormann D, Rodde R, Edbauer D et al (2010) ALS-associated fused in sarcoma (FUS) mutations disrupt Transportin-mediated nuclear import. EMBO J 29:2841–2857. doi:10.1038/emboj.2010.143
Elden AC, Kim H-J, Hart MP et al (2010) Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature 466:1069–1075. doi:10.1038/nature09320
Fallini C, Bassell GJ, Rossoll W (2012) The ALS disease protein TDP-43 is actively transported in motor neuron axons and regulates axon outgrowth. Hum Mol Genet 21:3703–3718. doi:10.1093/hmg/dds205
Farg MA, Soo KY, Walker AK et al (2012) Mutant FUS induces endoplasmic reticulum stress in amyotrophic lateral sclerosis and interacts with protein disulfide-isomerase. Neurobiol Aging 33:2855–2868. doi:10.1016/j.neurobiolaging.2012.02.009
Farg MA, Soo KY, Warraich ST, Sundaramoorthy V, Blair IP, Atkin JD (2013) Ataxin-2 interacts with FUS and intermediate-length polyglutamine expansions enhance FUS-related pathology in amyotrophic lateral sclerosis. Hum Mol Genet 22:717–728. doi:10.1093/hmg/dds479
Fecto F, Siddique T (2011) Making connections: pathology and genetics link amyotrophic lateral sclerosis with frontotemporal lobe dementia. J Mol Neurosci 45:663–675. doi:10.1007/s12031-011-9637-9
Feiguin F, Godena VK, Romano G, D’Ambrogio A, Klima R, Baralle FE (2009) Depletion of TDP-43 affects Drosophila motoneurons terminal synapsis and locomotive behavior. FEBS Lett 583:1586–1592. doi:10.1016/j.febslet.2009.04.019
Fiesel FC, Voigt A, Weber SS et al (2010) Knockdown of transactive response DNA-binding protein (TDP-43) downregulates histone deacetylase 6. EMBO J 29:209–221. doi:10.1038/emboj.2009.324
Freeman SH, Spires-Jones T, Hyman BT, Growdon JH, Frosch MP (2008) TAR-DNA binding protein 43 in Pick disease. J Neuropathol Exp Neurol 67:62–67. doi:10.1097/nen.0b013e3181609361
Freibaum BD, Chitta RK, High AA, Taylor JP (2010) Global analysis of TDP-43 interacting proteins reveals strong association with RNA splicing and translation machinery. J Proteome Res 9:1104–1120. doi:10.1021/pr901076y
Fuentealba RA, Udan M, Bell S et al (2010) Interaction with polyglutamine aggregates reveals a Q/N-rich domain in TDP-43. J Biol Chem 285:26304–26314. doi:10.1074/jbc.M110.125039
Fujii R (2005) TLS facilitates transport of mRNA encoding an actin-stabilizing protein to dendritic spines. J Cell Sci 118:5755–5765. doi:10.1242/jcs.02692
Furukawa Y, Kaneko K, Watanabe S, Yamanaka K, Nukina N (2011) A seeding reaction recapitulates intracellular formation of Sarkosyl-insoluble transactivation response element (TAR) DNA-binding protein-43 inclusions. J Biol Chem 286:18664–18672. doi:10.1074/jbc.M111.231209
Gertz B, Wong M, Martin LJ (2012) Nuclear localization of human SOD1 and mutant SOD1-specific disruption of survival motor neuron protein complex in transgenic amyotrophic lateral sclerosis mice. J Neuropathol Exp Neurol 71:162–177. doi:10.1097/NEN.0b013e318244b635
Geser F, Prvulovic D, O’Dwyer L et al (2011) On the development of markers for pathological TDP-43 in amyotrophic lateral sclerosis with and without dementia. Prog Neurobiol 95:649–662. doi:10.1016/j.pneurobio.2011.08.011
Geser F, Robinson JL, Malunda JA et al (2010) Pathological 43-kDa transactivation response DNA-binding protein in older adults with and without severe mental illness. Arch Neurol 67:1238–1250. doi:10.1001/archneurol.2010.254
Gijselinck I, Van Langenhove T, van der Zee J et al (2012) A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol 11:54–65. doi:10.1016/S1474-4422(11)70261-7
Giordana MT, Piccinini M, Grifoni S et al (2010) TDP-43 redistribution is an early event in sporadic amyotrophic lateral sclerosis. Brain Pathol 20:351–360. doi:10.1111/j.1750-3639.2009.00284.x
Gleason CE, Ordureau A, Gourlay R, Arthur JSC, Cohen P (2011) Polyubiquitin binding to optineurin is required for optimal activation of TANK-binding kinase 1 and production of interferon β. J Biol Chem 286:35663–35674. doi:10.1074/jbc.M111.267567
Gregory JM, Barros TP, Meehan S, Dobson CM, Luheshi LM (2012) The aggregation and neurotoxicity of TDP-43 and its ALS-associated 25 kDa fragment are differentially affected by molecular chaperones in Drosophila. PLoS ONE 7:e31899. doi:10.1371/journal.pone.0031899.g005
Groen EJN, van Es MA, van Vught PWJ et al (2010) FUS mutations in familial amyotrophic lateral sclerosis in the Netherlands. Arch Neurol 67:224–230. doi:10.1001/archneurol.2009.329
Guo W, Chen Y, Zhou X et al (2011) An ALS-associated mutation affecting TDP-43 enhances protein aggregation, fibril formation and neurotoxicity. Nat Struct Mol Biol 18:822–830. doi:10.1038/nsmb.2053
Hanson KA, Kim SH, Wassarman DA, Tibbetts RS (2010) Ubiquilin modifies TDP-43 toxicity in a Drosophila model of amyotrophic lateral sclerosis (ALS). J Biol Chem 285:11068–11072. doi:10.1074/jbc.C109.078527
Hara T, Nakamura K, Matsui M et al (2006) Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nat Cell Biol 441:885–889. doi:10.1038/nature04724
Hardiman O, van den Berg LH, Kiernan MC (2011) Clinical diagnosis and management of amyotrophic lateral sclerosis. Nat Rev Neurol 7:639–649. doi:10.1038/nrneurol.2011.153
Hart MP, Gitler AD (2012) ALS-associated ataxin 2 polyQ expansions enhance stress-induced caspase 3 activation and increase TDP-43 pathological modifications. J Neurosci 32:9133–9142. doi:10.1523/JNEUROSCI.0996-12.2012
Hasegawa M, Arai T, Nonaka T et al (2008) Phosphorylated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Ann Neurol 64:60–70. doi:10.1002/ana.21425
Hernández I, Espinosa A, Real LM et al (2012) Molecular evaluation of human ubiquilin 2 gene PXX domain in familial frontotemporal dementia patients. J Neurol 259:2488–2490. doi:10.1007/s00415-012-6568-5
Hewitt C, Kirby J, Highley JR et al (2010) Novel FUS/TLS mutations and pathology in familial and sporadic amyotrophic lateral sclerosis. Arch Neurol 67:455–461. doi:10.1001/archneurol.2010.52
Hicks GG, Singh N, Nashabi A et al (2000) FUS deficiency in mice results in defective B-lymphocyte development and activation, high levels of chromosomal instability and perinatal death. Nat Genet 24:175–179. doi:10.1038/72842
Higashi S, Iseki E, Yamamoto R et al (2007) Concurrence of TDP-43, tau and alpha-synuclein pathology in brains of Alzheimer’s disease and dementia with Lewy bodies. Brain Res 1184:284–294. doi:10.1016/j.brainres.2007.09.048
Higashi S, Kabuta T, Nagai Y, Tsuchiya Y, Akiyama H, Wada K (2013) TDP-43 associates with stalled ribosomes and contributes to cell survival during cellular stress. J Neurochem. doi:10.1111/jnc.12194
Hoell JI, Larsson E, Runge S et al (2011) RNA targets of wild-type and mutant FET family proteins. Nat Struct Mol Biol 18:1428–1431. doi:10.1038/nsmb.2163
Hortobagyi T, Troakes C, Nishimura AL et al (2011) Optineurin inclusions occur in a minority of TDP-43 positive ALS and FTLD-TDP cases and are rarely observed in other neurodegenerative disorders. Acta Neuropathol 121:519–527. doi:10.1007/s00401-011-0813-3
Huang C, Zhou H, Tong J et al (2011) FUS transgenic rats develop the phenotypes of amyotrophic lateral sclerosis and frontotemporal lobar degeneration. PLoS Genet 7:e1002011. doi:10.1371/journal.pgen.1002011.g006
Huang EJ, Zhang J, Geser F et al (2010) Extensive FUS-immunoreactive pathology in juvenile amyotrophic lateral sclerosis with basophilic inclusions. Brain Pathol 20:1069–1076. doi:10.1111/j.1750-3639.2010.00413.x
Igaz LM, Kwong LK, Chen-Plotkin A et al (2009) Expression of TDP-43 C-terminal fragments in vitro recapitulates pathological features of TDP-43 proteinopathies. J Biol Chem 284:8516–8524. doi:10.1074/jbc.M809462200
Igaz LM, Kwong LK, Lee EB et al (2011) Dysregulation of the ALS-associated gene TDP-43 leads to neuronal death and degeneration in mice. J Clin Invest 121:726–738. doi:10.1172/JCI44867DS1
Iida A, Hosono N, Sano M et al (2012) Novel deletion mutations of OPTN in amyotrophic lateral sclerosis in Japanese. Neurobiol Aging 33(1843):e1819–e1824. doi:10.1016/j.neurobiolaging.2011.12.037
Inukai Y, Nonaka T, Arai T et al (2008) Abnormal phosphorylation of Ser409/410 of TDP-43 in FTLD-U and ALS. FEBS Lett 582:2899–2904. doi:10.1016/j.febslet.2008.07.027
Ito H, Fujita K, Nakamura M et al (2011) Optineurin is co-localized with FUS in basophilic inclusions of ALS with FUS mutation and in basophilic inclusion body disease. Acta Neuropathol 121:555–557. doi:10.1007/s00401-011-0809-z
Jinwal UK, Abisambra JF, Zhang J et al (2012) Cdc37/Hsp90 protein complex disruption triggers an autophagic clearance cascade for TDP-43 protein. J Biol Chem 287:24814–24820. doi:10.1074/jbc.M112.367268
Johnson BS, McCaffery JM, Lindquist S, Gitler AD (2008) A yeast TDP-43 proteinopathy model: exploring the molecular determinants of TDP-43 aggregation and cellular toxicity. Proc Natl Acad Sci USA 105:6439–6444. doi:10.1073/pnas.0802082105
Johnson BS, Snead D, Lee JJ, McCaffery JM, Shorter J, Gitler AD (2009) TDP-43 is intrinsically aggregation-prone, and amyotrophic lateral sclerosis-linked mutations accelerate aggregation and increase toxicity. J Biol Chem 284:20329–20339. doi:10.1074/jbc.M109.010264
Johnson JO, Mandrioli J, Benatar M et al (2010) Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 68:857–864. doi:10.1016/j.neuron.2010.11.036
Ju S, Tardiff DF, Han H et al (2011) A yeast model of FUS/TLS-dependent cytotoxicity. PLoS Biol 9:e1001052. doi:10.1371/journal.pbio.1001052
Kabashi E, Bercier V, Lissouba A et al (2011) FUS and TARDBP but not SOD1 interact in genetic models of amyotrophic lateral sclerosis. PLoS Genet 7:e1002214. doi:10.1371/journal.pgen.1002214.t001
Kabashi E, Lin L, Tradewell ML et al (2010) Gain and loss of function of ALS-related mutations of TARDBP (TDP-43) cause motor deficits in vivo. Hum Mol Genet 19:671–683. doi:10.1093/hmg/ddp534
Kabashi E, Valdmanis PN, Dion P et al (2008) TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat Genet 40:572–574. doi:10.1038/ng.132
Kariya S, Re DB, Jacquier A, Nelson K, Przedborski S, Monani UR (2012) Mutant superoxide dismutase 1 (SOD1), a cause of amyotrophic lateral sclerosis, disrupts the recruitment of SMN, the spinal muscular atrophy protein to nuclear Cajal bodies. Hum Mol Genet 21:3421–3434. doi:10.1093/hmg/dds174
Kato M, Han TW, Xie S et al (2012) Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149:753–767. doi:10.1016/j.cell.2012.04.017
Keller BA, Volkening K, Droppelmann CA, Ang L-C, Rademakers R, Strong MJ (2012) Co-aggregation of RNA binding proteins in ALS spinal motor neurons: evidence of a common pathogenic mechanism. Acta Neuropathol 124:733–747. doi:10.1007/s00401-012-1035-z
Kiehl T-R, Nechiporuk A, Figueroa KP, Keating MT, Huynh DP, Pulst S-M (2006) Generation and characterization of Sca2 (ataxin-2) knockout mice. Biochem Biophys Res Commun 339:17–24. doi:10.1016/j.bbrc.2005.10.186
Kim HJ, Kim NC, Wang Y-D et al (2013) Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature 495(7442):467–473. doi:10.1038/nature11922
King A, Al-Sarraj S, Troakes C et al (2013) Mixed tau, TDP-43 and p62 pathology in FTLD associated with a C9ORF72 repeat expansion and p.Ala239Thr MAPT (tau) variant. Acta Neuropathol 125:303–310. doi:10.1007/s00401-012-1050-0
King OD, Gitler AD, Shorter J (2012) The tip of the iceberg: RNA-binding proteins with prion-like domains in neurodegenerative disease. Brain Res 1462:61–80. doi:10.1016/j.brainres.2012.01.016
Kino Y, Washizu C, Aquilanti E et al (2011) Intracellular localization and splicing regulation of FUS/TLS are variably affected by amyotrophic lateral sclerosis-linked mutations. Nucleic Acids Res 39:2781–2798. doi:10.1093/nar/gkq1162
Komatsu M, Waguri S, Chiba T et al (2006) Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441:880–884. doi:10.1038/nature04723
Korac J, Schaeffer V, Kovacevic I et al (2012) Ubiquitin-independent function of optineurin in autophagic clearance of protein aggregates. J Cell Sci. doi:10.1242/jcs.114926
Kovacs GG, Murrell JR, Horvath S et al (2009) TARDBP variation associated with frontotemporal dementia, supranuclear gaze palsy, and chorea. Mov Disord 24:1843–1847. doi:10.1002/mds.22697
Kraemer BC, Schuck T, Wheeler JM et al (2010) Loss of murine TDP-43 disrupts motor function and plays an essential role in embryogenesis. Acta Neuropathol 119:409–419. doi:10.1007/s00401-010-0659-0
Kryndushkin D, Ihrke G, Piermartiri TC, Shewmaker F (2012) A yeast model of optineurin proteinopathy reveals a unique aggregation pattern associated with cellular toxicity. Mol Microbiol 86:1531–1547. doi:10.1111/mmi.12075
Kulathu Y, Komander D (2012) Atypical ubiquitylation—the unexplored world of polyubiquitin beyond Lys48 and Lys63 linkages. Nat Rev Mol Cell Biol 13:508–523. doi:10.1038/nrm3394
Kuroda M, Sok J, Webb L et al (2000) Male sterility and enhanced radiation sensitivity in TLS(−/−) mice. EMBO J 19:453–462. doi:10.1093/emboj/19.3.453
Kwiatkowski TJ Jr, Bosco DA, Leclerc AL et al (2009) Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323:1205–1208. doi:10.1126/science.1166066
Lagier-Tourenne C, Polymenidou M, Cleveland DW (2010) TDP-43 and FUS/TLS: emerging roles in RNA processing and neurodegeneration. Hum Mol Genet 19:R46–R64. doi:10.1093/hmg/ddq137
Laird AS, Van Hoecke A, De Muynck L et al (2010) Progranulin is neurotrophic in vivo and protects against a mutant TDP-43 induced axonopathy. PLoS ONE 5:e13368. doi:10.1371/journal.pone.0013368.g005
Lanson NA, Maltare A, King H et al (2011) A Drosophila model of FUS-related neurodegeneration reveals genetic interaction between FUS and TDP-43. Hum Mol Genet 20:2510–2523. doi:10.1093/hmg/ddr150
Lastres-Becker I, Rub U, Auburger G (2008) Spinocerebellar ataxia 2 (SCA2). Cerebellum 7:115–124. doi:10.1007/s12311-008-0019-y
Lee DY, Brown EJ (2012) Ubiquilins in the crosstalk among proteolytic pathways. Biol Chem 393:441–447. doi:10.1515/hsz-2012-0120
Levine TP, Daniels RD, Gatta AT, Wong LH, Hayes MJ (2013) The product of C9orf72, a gene strongly implicated in neurodegeneration, is structurally related to DENN Rab-GEFs. Bioinformatics 29:499–503. doi:10.1093/bioinformatics/bts725
Li H-Y, Yeh P-A, Chiu H-C, Tang C-Y, Tu BP-H (2011) Hyperphosphorylation as a defense mechanism to reduce TDP-43 aggregation. PLoS ONE 6:e23075. doi:10.1371/journal.pone.0023075.g009
Li Y, Ray P, Rao EJ et al (2010) A Drosophila model for TDP-43 proteinopathy. Proc Natl Acad Sci USA 107:3169–3174. doi:10.1073/pnas.0913602107
Li Z, Lu Y, Xu XL, Gao FB (2012) The FTD/ALS-associated RNA-binding protein TDP-43 regulates the robustness of neuronal specification through microRNA-9a in Drosophila. Hum Mol Genet 22:218–225. doi:10.1093/hmg/dds420
Liachko NF, Guthrie CR, Kraemer BC (2010) Phosphorylation promotes neurotoxicity in a Caenorhabditis elegans model of TDP-43 proteinopathy. J Neurosci 30:16208–16219. doi:10.1523/JNEUROSCI.2911-10.2010
Liachko NF, McMillan PJ, Guthrie CR, Bird TD, Leverenz JB, Kraemer BC (2013) CDC7 inhibition blocks pathological TDP-43 phosphorylation and neurodegeneration. Ann Neurol. doi:10.1002/ana.23870
Lin M-J, Cheng C-W, Shen CKJ (2011) Neuronal function and dysfunction of Drosophila dTDP. PLoS ONE 6:e20371. doi:10.1371/journal.pone.0020371
Lin W-L, Dickson DW (2008) Ultrastructural localization of TDP-43 in filamentous neuronal inclusions in various neurodegenerative diseases. Acta Neuropathol 116:205–213. doi:10.1007/s00401-008-0408-9
Ling S-C, Albuquerque CP, Han JS et al (2010) ALS-associated mutations in TDP-43 increase its stability and promote TDP-43 complexes with FUS/TLS. Proc Natl Acad Sci USA 107:13318–13323. doi:10.1073/pnas.1008227107
Liu-Yesucevitz L, Bilgutay A, Zhang Y-J et al (2010) Tar DNA binding protein-43 (TDP-43) associates with stress granules: analysis of cultured cells and pathological brain tissue. PLoS ONE 5:e13250. doi:10.1371/journal.pone.0013250.t001
Lu Y, Ferris J, Gao F-B (2009) Frontotemporal dementia and amyotrophic lateral sclerosis-associated disease protein TDP-43 promotes dendritic branching. Mol Brain 2:30. doi:10.1186/1756-6606-2-30
Mackenzie IRA, Bigio EH, Ince PG et al (2007) Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann Neurol 61:427–434. doi:10.1002/ana.21147
Majounie E, Abramzon Y, Renton AE et al (2012) Repeat expansion in C9ORF72 in Alzheimer’s disease. N Engl J Med 366:283–284. doi:10.1056/NEJMc1113592
Maruyama H, Morino H, Ito H et al (2010) Mutations of optineurin in amyotrophic lateral sclerosis. Nature 465:223–226. doi:10.1038/nature08971
McCann C, Holohan EE, Das S et al (2011) The ataxin-2 protein is required for microRNA function and synapse-specific long-term olfactory habituation. Proc Natl Acad Sci USA 108:E655–E662. doi:10.1073/pnas.1107198108
McDonald KK, Aulas A, Destroismaisons L et al (2011) TAR DNA-binding protein 43 (TDP-43) regulates stress granule dynamics via differential regulation of G3BP and TIA-1. Hum Mol Genet 20:1400–1410. doi:10.1093/hmg/ddr021
Meyerowitz J, Parker SJ, Vella LJ et al (2011) C-Jun N-terminal kinase controls TDP-43 accumulation in stress granules induced by oxidative stress. Mol Neurodegener 6:57. doi:10.1186/1750-1326-6-57
Miguel L, Avequin T, Delarue M et al (2012) Accumulation of insoluble forms of FUS protein correlates with toxicity in Drosophila. Neurobiol Aging 33(1008):e1001–e1015. doi:10.1016/j.neurobiolaging.2011.10.008
Miguel L, Frébourg T, Campion D, Lecourtois M (2011) Both cytoplasmic and nuclear accumulations of the protein are neurotoxic in Drosophila models of TDP-43 proteinopathies. Neurobiol Dis 41:398–406. doi:10.1016/j.nbd.2010.10.007
Millecamps S, Boillee S, Chabrol E et al (2011) Screening of OPTN in French familial amyotrophic lateral sclerosis. Neurobiol Aging 32(557):e511–e553. doi:10.1016/j.neurobiolaging.2010.11.005
Mitchell JC, McGoldrick P, Vance C et al (2012) Overexpression of human wild-type FUS causes progressive motor neuron degeneration in an age- and dose-dependent fashion. Acta Neuropathol 125:273–288. doi:10.1007/s00401-012-1043-z
Mori K, Lammich S, Mackenzie IRA et al (2013) hnRNP A3 binds to GGGGCC repeats and is a constituent of p62-positive/TDP43-negative inclusions in the hippocampus of patients with C9orf72 mutations. Acta Neuropathol 125:413–423. doi:10.1007/s00401-013-1088-7
Mori K, Weng SM, Arzberger T et al (2013) The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 339:1335–1338. doi:10.1126/science.1232927
Mosca L, Lunetta C, Tarlarini C et al (2012) Wide phenotypic spectrum of the TARDBP gene: homozygosity of A382T mutation in a patient presenting with amyotrophic lateral sclerosis, Parkinson’s disease, and frontotemporal lobar degeneration, and in neurologically healthy subject. Neurobiol Aging 33(1846):e1841–e1844. doi:10.1016/j.neurobiolaging.2012.01.108
Murakami T, Yang SP, Xie L et al (2011) ALS mutations in FUS cause neuronal dysfunction and death in Caenorhabditis elegans by a dominant gain-of-function mechanism. Hum Mol Genet 21:1–9. doi:10.1093/hmg/ddr417
Neumann M, Bentmann E, Dormann D et al (2011) FET proteins TAF15 and EWS are selective markers that distinguish FTLD with FUS pathology from amyotrophic lateral sclerosis with FUS mutations. Brain J Neurol 134:2595–2609. doi:10.1093/brain/awr201
Neumann M, Kwong LK, Sampathu DM, Trojanowski JQ, Lee VM (2007) TDP-43 proteinopathy in frontotemporal lobar degeneration and amyotrophic lateral sclerosis: protein misfolding diseases without amyloidosis. Arch Neurol 64:1388–1394. doi:10.1001/archneur.64.10.1388
Neumann M, Rademakers R, Roeber S, Baker M, Kretzschmar HA, Mackenzie IRA (2009) A new subtype of frontotemporal lobar degeneration with FUS pathology. Brain J Neurol 132:2922–2931. doi:10.1093/brain/awp214
Neumann M, Sampathu DM, Kwong LK et al (2006) Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314:130–133. doi:10.1126/science.1134108
Neumann M, Valori CF, Ansorge O et al (2012) Transportin 1 accumulates specifically with FET proteins but no other transportin cargos in FTLD-FUS and is absent in FUS inclusions in ALS with FUS mutations. Acta Neuropathol 124:705–716. doi:10.1007/s00401-012-1020-6
Nihei Y, Ito D, Suzuki N (2012) Roles of ataxin-2 in pathological cascades mediated by TAR DNA-binding protein 43 (TDP-43) and fused in sarcoma (FUS). J Biol Chem 287:41310–41323. doi:10.1074/jbc.M112.398099
Nishimoto Y, Ito D, Yagi T, Nihei Y, Tsunoda Y, Suzuki N (2010) Characterization of alternative isoforms and inclusion body of the TAR DNA-binding protein-43. J Biol Chem 285:608–619. doi:10.1074/jbc.M109.022012
Nishimura AL, Mitne-Neto M, Silva HCA et al (2004) A mutation in the vesicle-trafficking protein VAPB causes late-onset spinal muscular atrophy and amyotrophic lateral sclerosis. Am J Hum Genet 75:822–831. doi:10.1086/425287
Nonaka T, Arai T, Buratti E, Baralle FE, Akiyama H, Hasegawa M (2009) Phosphorylated and ubiquitinated TDP-43 pathological inclusions in ALS and FTLD-U are recapitulated in SH-SY5Y cells. FEBS Lett 583:394–400. doi:10.1016/j.febslet.2008.12.031
Nonaka T, Kametani F, Arai T, Akiyama H, Hasegawa M (2009) Truncation and pathogenic mutations facilitate the formation of intracellular aggregates of TDP-43. Hum Mol Genet 18:3353–3364. doi:10.1093/hmg/ddp275
Nonhoff U, Ralser M, Welzel F et al (2007) Ataxin-2 interacts with the DEAD/H-box RNA helicase DDX6 and interferes with P-bodies and stress granules. Mol Biol Cell 18:1385–1396. doi:10.1091/mbc.E06-12-1120
Okamoto K, Mizuno Y, Fujita Y (2008) Bunina bodies in amyotrophic lateral sclerosis. Neuropathology 28:109–115. doi:10.1111/j.1440-1789.2007.00873.x
Osawa T, Mizuno Y, Fujita Y, Takatama M, Nakazato Y, Okamoto K (2011) Optineurin in neurodegenerative diseases. Neuropathology 31:569–574. doi:10.1111/j.1440-1789.2011.01199.x
Parker SJ, Meyerowitz J, James JL et al (2012) Endogenous TDP-43 localized to stress granules can subsequently form protein aggregates. Neurochem Int 60:415–424. doi:10.1016/j.neuint.2012.01.019
Parkinson N, Ince PG, Smith MO et al (2006) ALS phenotypes with mutations in CHMP2B (charged multivesicular body protein 2B). Neurology 67:1074–1077. doi:10.1212/01.wnl.0000231510.89311.8b
Polymenidou M, Cleveland DW (2011) The seeds of neurodegeneration: prion-like spreading in ALS. Cell 147:498–508. doi:10.1016/j.cell.2011.10.011
Polymenidou M, Lagier-Tourenne C, Hutt KR et al (2011) Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat Neurosci 14:459–468. doi:10.1038/nn.2779
Renton AE, Majounie E, Waite A et al (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72:257–268. doi:10.1016/j.neuron.2011.09.010
Ritson GP, Custer SK, Freibaum BD et al (2010) TDP-43 mediates degeneration in a novel Drosophila model of disease caused by mutations in VCP/p97. J Neurosci 30:7729–7739. doi:10.1523/JNEUROSCI.5894-09.2010
Robinson JL, Geser F, Stieber A et al (2013) TDP-43 skeins show properties of amyloid in a subset of ALS cases. Acta Neuropathol 125:121–131. doi:10.1007/s00401-012-1055-8
Rosen DR, Siddique T, Patterson D et al (1993) Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362:59–62. doi:10.1038/362059a0
Rutherford NJ, Zhang Y-J, Baker M et al (2008) Novel mutations in TARDBP (TDP-43) in patients with familial amyotrophic lateral sclerosis. PLoS Genet 4:e1000193. doi:10.1371/journal.pgen.1000193
Sahlender DA, Roberts RC, Arden SD et al (2005) Optineurin links myosin VI to the Golgi complex and is involved in Golgi organization and exocytosis. J Cell Biol 169:285–295. doi:10.1083/jcb.200501162
Sasayama H, Shimamura M, Tokuda T et al (2012) Knockdown of the Drosophila fused in sarcoma (FUS) homologue causes deficient locomotive behavior and shortening of motoneuron terminal branches. PLoS ONE 7:e39483. doi:10.1371/journal.pone.0039483.t001
Satterfield TF (2006) Ataxin-2 and its Drosophila homolog, ATX2, physically assemble with polyribosomes. Hum Mol Genet 15:2523–2532. doi:10.1093/hmg/ddl173
Sephton CF, Good SK, Atkin S et al (2010) TDP-43 is a developmentally regulated protein essential for early embryonic development. J Biol Chem 285:6826–6834. doi:10.1074/jbc.M109.061846
Shan X, Chiang P-M, Price DL, Wong PC (2010) Altered distributions of Gemini of coiled bodies and mitochondria in motor neurons of TDP-43 transgenic mice. Proc Natl Acad Sci USA 107:16325–16330. doi:10.1073/pnas.1003459107
Shen X, Ying H, Qiu Y et al (2011) Processing of optineurin in neuronal cells. J Biol Chem 286:3618–3629. doi:10.1074/jbc.M110.175810
Simon-Sanchez J, Dopper EGP, Cohn-Hokke PE et al (2012) The clinical and pathological phenotype of C9ORF72 hexanucleotide repeat expansions. Brain J Neurol 135:723–735. doi:10.1093/brain/awr353
Sreedharan J, Blair IP, Tripathi VB et al (2008) TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319:1668–1672. doi:10.1126/science.1154584
Stallings NR, Puttaparthi K, Luther CM, Burns DK, Elliott JL (2010) Progressive motor weakness in transgenic mice expressing human TDP-43. Neurobiol Dis 40:404–414. doi:10.1016/j.nbd.2010.06.017
Sun Y, Yang P, Zhang Y et al (2011) A genome-wide RNAi screen identifies genes regulating the formation of P bodies in C. elegans and their functions in NMD and RNAi. Protein Cell 2:918–939. doi:10.1007/s13238-011-1119-x
Sun Z, Diaz Z, Fang X et al (2011) Molecular determinants and genetic modifiers of aggregation and toxicity for the ALS disease protein FUS/TLS. PLoS Biol 9:e1000614. doi:10.1371/journal.pbio.1000614.t002
Swarup V, Phaneuf D, Bareil C et al (2011) Pathological hallmarks of amyotrophic lateral sclerosis/frontotemporal lobar degeneration in transgenic mice produced with TDP-43 genomic fragments. Brain J Neurol 134:2610–2626. doi:10.1093/brain/awr159
Tan C-F, Eguchi H, Tagawa A et al (2007) TDP-43 immunoreactivity in neuronal inclusions in familial amyotrophic lateral sclerosis with or without SOD1 gene mutation. Acta Neuropathol 113:535–542. doi:10.1007/s00401-007-0206-9
Tanji K, Zhang H-X, Mori F, Kakita A, Takahashi H, Wakabayashi K (2012) p62/sequestosome 1 binds to TDP-43 in brains with frontotemporal lobar degeneration with TDP-43 inclusions. J Neurosci Res 90:2034–2042. doi:10.1002/jnr.23081
Tashiro Y, Urushitani M, Inoue H et al (2012) Motor neuron-specific disruption of proteasomes, but not autophagy, replicates amyotrophic lateral sclerosis. J Biol Chem 287:42984–42994. doi:10.1074/jbc.M112.417600
Teyssou E, Takeda T, Lebon V et al (2013) Mutations in SQSTM1 encoding p62 in amyotrophic lateral sclerosis: genetics and neuropathology. Acta Neuropathol 125:511–522. doi:10.1007/s00401-013-1090-0
Tollervey JR, Curk T, Rogelj B et al (2011) Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat Neurosci 14:452–458. doi:10.1038/nn.2778
Tradewell ML, Yu Z, Tibshirani M, Boulanger MC, Durham HD, Richard S (2011) Arginine methylation by PRMT1 regulates nuclear-cytoplasmic localization and toxicity of FUS/TLS harbouring ALS-linked mutations. Hum Mol Genet 21:136–149. doi:10.1093/hmg/ddr448
Tsai KJ, Yang CH, Fang YH et al (2010) Elevated expression of TDP-43 in the forebrain of mice is sufficient to cause neurological and pathological phenotypes mimicking FTLD-U. J Exp Med 207:1661–1673. doi:10.1084/jem.20092164
Tsuiji H, Iguchi Y, Furuya A et al (2013) Spliceosome integrity is defective in the motor neuron diseases ALS and SMA. EMBO Mol Med 5:221–234. doi:10.1002/emmm.201202303
Tsuji H, Arai T, Kametani F et al (2012) Molecular analysis and biochemical classification of TDP-43 proteinopathy. Brain J Neurol 135:3380–3391. doi:10.1093/brain/aws230
Vaccaro A, Tauffenberger A, Aggad D, Rouleau G, Drapeau P, Parker JA (2012) Mutant TDP-43 and FUS cause age-dependent paralysis and neurodegeneration in C. elegans. PLoS ONE 7:e31321. doi:10.1371/journal.pone.0031321.g010
van Blitterswijk M, van Vught PW, van Es MA et al (2012) Novel optineurin mutations in sporadic amyotrophic lateral sclerosis patients. Neurobiol Aging 33(1016):e1011–e1017. doi:10.1016/j.neurobiolaging.2011.05.019
Van Deerlin VM, Leverenz JB, Bekris LM et al (2008) TARDBP mutations in amyotrophic lateral sclerosis with TDP-43 neuropathology: a genetic and histopathological analysis. Lancet Neurol 7:409–416. doi:10.1016/S1474-4422(08)70071-1
van der Zee J, Gijselinck I, Dillen L et al (2013) A pan-European Study of the C9orf72 repeat associated with FTLD: geographic prevalence, genomic instability, and intermediate repeats. Hum Mutat 34:363–373. doi:10.1002/humu.22244
van Eersel J, Ke YD, Gladbach A et al (2011) Cytoplasmic accumulation and aggregation of TDP-43 upon proteasome inhibition in cultured neurons. PLoS ONE 6:e22850. doi:10.1371/journal.pone.0022850.g006
Van Langenhove T, van der Zee J, Sleegers K et al (2010) Genetic contribution of FUS to frontotemporal lobar degeneration. Neurology 74:366–371. doi:10.1212/WNL.0b013e3181ccc732
Vance C, Rogelj B, Hortobagyi T et al (2009) Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323:1208–1211. doi:10.1126/science.1165942
Vance C, Scotter EL, Nishimura AL et al (2013) ALS mutant FUS disrupts nuclear localization and sequesters wild-type FUS within cytoplasmic stress granules. Hum Mol Genet. doi:10.1093/hmg/ddt117
Verbeeck C, Deng Q, DeJesus-Hernandez M et al (2012) Expression of fused in sarcoma mutations in mice recapitulates the neuropathology of FUS proteinopathies and provides insight into disease pathogenesis. Mol Neurodegener 7:53. doi:10.1186/1750-1326-7-53
Voigt A, Herholz D, Fiesel FC et al (2010) TDP-43-mediated neuron loss in vivo requires RNA-binding activity. PLoS ONE 5:e12247. doi:10.1371/journal.pone.0012247.t002
Volkening K, Leystra-Lantz C, Yang W, Jaffee H, Strong MJ (2009) Tar DNA binding protein of 43 kDa (TDP-43), 14–3-3 proteins and copper/zinc superoxide dismutase (SOD1) interact to modulate NFL mRNA stability. Implications for altered RNA processing in amyotrophic lateral sclerosis (ALS). Brain Res 1305:168–182. doi:10.1016/j.brainres.2009.09.105
Wang I-F, Guo B-S, Liu Y-C et al (2012) Autophagy activators rescue and alleviate pathogenesis of a mouse model with proteinopathies of the TAR DNA-binding protein 43. Proc Natl Acad Sci USA 109:15024–15029. doi:10.1073/pnas.1206362109
Wang IF, Wu L-S, Chang H-Y, Shen CKJ (2008) TDP-43, the signature protein of FTLD-U, is a neuronal activity-responsive factor. J Neurochem 105:797–806. doi:10.1111/j.1471-4159.2007.05190.x
Wang J-W, Brent JR, Tomlinson A, Shneider NA, McCabe BD (2011) The ALS-associated proteins FUS and TDP-43 function together to affect Drosophila locomotion and life span. J Clin Invest 121:4118–4126. doi:10.1172/JCI57883DS1
Watanabe M, Dykes-Hoberg M, Cizewski Culotta V, Price DL, Wong PC, Rothstein JD (2001) Histological evidence of protein aggregation in mutant SOD1 transgenic mice and in amyotrophic lateral sclerosis neural tissues. Neurobiol Dis 8:933–941. doi:10.1006/nbdi.2001.0443
Wegorzewska I, Bell S, Cairns NJ, Miller TM, Baloh RH (2009) TDP-43 mutant transgenic mice develop features of ALS and frontotemporal lobar degeneration. Proc Natl Acad Sci USA 106:18809–18814. doi:10.1073/pnas.0908767106
Wild P, Farhan H, McEwan DG et al (2011) Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science 333:228–233. doi:10.1126/science.1205405
Williams KL, Warraich ST, Yang S et al (2012) UBQLN2/ubiquilin 2 mutation and pathology in familial amyotrophic lateral sclerosis. Neurobiol Aging 33:2527 e2523-2510. doi:10.1016/j.neurobiolaging.2012.05.008
Wils H, Kleinberger G, Janssens J et al (2010) TDP-43 transgenic mice develop spastic paralysis and neuronal inclusions characteristic of ALS and frontotemporal lobar degeneration. Proc Natl Acad Sci USA 107:3858–3863. doi:10.1073/pnas.0912417107
Wojtas A, Heggeli KA, Finch N et al (2012) C9ORF72 repeat expansions and other FTD gene mutations in a clinical AD patient series from Mayo Clinic. Am J Neurodegener Dis 1:107–118
Wu LS, Cheng WC, Hou SC, Yan YT, Jiang ST, Shen CK (2010) TDP-43, a neuro-pathosignature factor, is essential for early mouse embryogenesis. Genesis 48:56–62. doi:10.1002/dvg.20584
Xia R, Liu Y, Yang L, Gal J, Zhu H, Jia J (2012) Motor neuron apoptosis and neuromuscular junction perturbation are prominent features in a Drosophila model of FUS-mediated ALS. Mol Neurodegener 7:10. doi:10.1186/1750-1326-7-10
Xu Y-F, Zhang Y-J, Lin W-L et al (2011) Expression of mutant TDP-43 induces neuronal dysfunction in transgenic mice. Mol Neurodegener 6:73. doi:10.1186/1750-1326-6-73
Xu YF, Gendron TF, Zhang YJ et al (2010) Wild-type human TDP-43 expression causes TDP-43 phosphorylation, mitochondrial aggregation, motor deficits, and early mortality in transgenic mice. J Neurosci 30:10851–10859. doi:10.1523/JNEUROSCI.1630-10.2010
Yamazaki T, Chen S, Yu Y et al (2012) FUS-SMN protein interactions link the motor neuron diseases ALS and SMA. Cell Rep 2:799–806. doi:10.1016/j.celrep.2012.08.025
Yang C, Tan W, Whittle C et al (2010) The C-terminal TDP-43 fragments have a high aggregation propensity and harm neurons by a dominant-negative mechanism. PLoS ONE 5:e15878. doi:10.1371/journal.pone.0015878.t001
Yu Z, Zhu Y, Chen-Plotkin AS et al (2011) PolyQ repeat expansions in ATXN2 associated with ALS are CAA interrupted repeats. PLoS ONE 6:e17951. doi:10.1371/journal.pone.0017951.t002
Zhang D, Iyer LM, He F, Aravind L (2012) Discovery of novel DENN proteins: implications for the evolution of eukaryotic intracellular membrane structures and human disease. Front Genet 3:283. doi:10.3389/fgene.2012.00283
Zhang T, Hwang H-Y, Hao H, Talbot C, Wang J (2012) Caenorhabditis elegans RNA-processing protein TDP-1 regulates protein homeostasis and life span. J Biol Chem 287:8371–8382. doi:10.1074/jbc.M111.311977
Zhang T, Mullane PC, Periz G, Wang J (2011) TDP-43 neurotoxicity and protein aggregation modulated by heat shock factor and insulin/IGF-1 signaling. Hum Mol Genet 20:1952–1965. doi:10.1093/hmg/ddr076
Zhang Y-J, Gendron TF, Xu Y-F, Ko L-W, Yen S-H, Petrucelli L (2010) Phosphorylation regulates proteasomal-mediated degradation and solubility of TAR DNA binding protein-43 C-terminal fragments. Mol Neurodegener 5:33. doi:10.1186/1750-1326-5-33
Zhang Y-J, Xu Y-F, Cook C et al (2009) Aberrant cleavage of TDP-43 enhances aggregation and cellular toxicity. Proc Natl Acad Sci USA 106:7607–7612. doi:10.1073/pnas.0900688106
Zhou H, Huang C, Chen H et al (2010) Transgenic rat model of neurodegeneration caused by mutation in the TDP gene. PLoS Genet 6:e1000887. doi:10.1371/journal.pgen.1000887.g008
Zhu G, Wu C-J, Zhao Y, Ashwell JD (2007) Optineurin negatively regulates TNFα- induced NF-κB activation by competing with NEMO for ubiquitinated RIP. Curr Biol 17:1438–1443. doi:10.1016/j.cub.2007.07.041
Zinszner H, Sok J, Immanuel D, Yin Y, Ron D (1997) TLS (FUS) binds RNA in vivo and engages in nucleo-cytoplasmic shuttling. J Cell Sci 110(Pt 15):1741–1750
Acknowledgments
Work in the laboratories of the authors is supported by funding from the European Community’s Health Seventh Framework Programme (FP7/2007-2013) under grant agreement no. 259867, the Netherlands Organization for Health Research and Development (VICI), the Van Meer Stichting and Netherlands ALS Foundation (to L.H.v.d.B), the Netherlands Organization for Health Research and Development (VIDI), Human Frontier Science Program Organization (HFSP-CDA), Dutch Brain Foundation, and Thierry Latran Foundation (to R.J.P.), Prinses Beatrix Fonds and the Adessium Foundation (to L.H.v.d.B. and R.J.P.).
Conflict of interest
The authors declare no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Blokhuis, A.M., Groen, E.J.N., Koppers, M. et al. Protein aggregation in amyotrophic lateral sclerosis. Acta Neuropathol 125, 777–794 (2013). https://doi.org/10.1007/s00401-013-1125-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-013-1125-6