Abstract
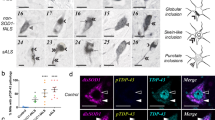
While the pathogenesis of amyotrophic lateral sclerosis (ALS) remains to be clearly delineated, there is mounting evidence that altered RNA metabolism is a commonality amongst several of the known genetic variants of the disease. In this study, we evaluated the expression of 10 ALS-associated proteins in spinal motor neurons (MNs) in ALS patients with mutations in C9orf72 (C9orf72GGGGCC-ALS; n = 5), SOD1 (mtSOD1-ALS; n = 9), FUS/TLS (mtFUS/TLS-ALS; n = 2), or TARDBP (mtTDP-43-ALS; n = 2) and contrasted these to cases of sporadic ALS (sALS; n = 4) and familial ALS without known mutations (fALS; n = 2). We performed colorimetric immunohistochemistry (IHC) using antibodies against TDP-43, FUS/TLS, SOD1, C9orf72, ubiquitin, sequestosome 1 (p62), optineurin, phosphorylated high molecular weight neurofilament, peripherin, and Rho-guanine nucleotide exchange factor (RGNEF). We observed that RGNEF-immunoreactive neuronal cytoplasmic inclusions (NCIs) can co-localize with TDP-43, FUS/TLS and p62 within spinal MNs. We confirmed their capacity to interact by co-immunoprecipitations. We also found that mtSOD1-ALS cases possess a unique IHC signature, including the presence of C9orf72-immunoreactive diffuse NCIs, which allows them to be distinguished from other variants of ALS at the level of light microscopy. These findings support the hypothesis that alterations in RNA metabolism are a core pathogenic pathway in ALS. We also conclude that routine IHC-based analysis of spinal MNs may aid in the identification of families not previously suspected to harbor SOD1 mutations.







Similar content being viewed by others
References
Al-Sarraj S, King A, Troakes C et al (2011) p62 positive, TDP-43 negative, neuronal cytoplasmic and intranuclear inclusions in the cerebellum and hippocampus define the pathology of C9orf72-linked FTLD and MND/ALS. Acta Neuropathol 122:691–702
Arai T, Hasegawa M, Akiyama H et al (2006) TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun 351:602–611
Bergeron C, Beric-Maskarel K, Muntasser S et al (1994) Neurofilament light and polyadenylated mRNA levels are decreased in amyotrophic lateral sclerosis motor neurons. J Neuropathol Exp Neurol 53:221–230
Boeve BF, Boylan KB, Graff-Radford NR et al (2012) Characterization of frontotemporal dementia and/or amyotrophic lateral sclerosis associated with the GGGGCC repeat expansion in C9ORF72. Brain 135:765–783
Bosco DA, Morfini G, Karabacak NM et al (2010) Wild-type and mutant SOD1 share an aberrant conformation and a common pathogenic pathway in ALS. Nat Neurosci 13:1396–1403
Canete-Soler R, Wu J, Zhai J, Shamim M, Schlaepfer WW (2001) p190RhoGEF Binds to a destabilizing element in the 3′ untranslated region of light neurofilament subunit mRNA and alters the stability of the transcript. J Biol Chem 276:32046–32050
Carpenter S (1968) Proximal axonal enlargement in motor neuron disease. Neurology 18:841–851
Chen YZ, Bennett CL, Huynh HM et al (2004) DNA/RNA helicase gene mutations in a form of juvenile amyotrophic lateral sclerosis (ALS4). Am J Hum Genet 74:1128–1135
Chio A, Borghero G, Restagno G et al (2012) Clinical characteristics of patients with familial amyotrophic lateral sclerosis carrying the pathogenic GGGGCC hexanucleotide repeat expansion of C9ORF72. Brain 135:784–793
Cooper-Knock J, Hewitt C, Highley JR et al (2012) Clinico-pathological features in amyotrophic lateral sclerosis with expansions in C9ORF72. Brain 135:751–764
DeJesus-Hernandez M, Mackenzie IR, Boeve BF et al (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72:245–256
Deng HX, Bigio EH, Zhai H et al (2011) Differential involvement of optineurin in amyotrophic lateral sclerosis with or without SOD1 mutations. Arch Neurol 68:1057–1061
Deng HX, Chen W, Hong ST et al (2011) Mutations in UBQLN2 cause dominant X-linked juvenile and adult-onset ALS and ALS/dementia. Nature 477:211–215
Deng HX, Zhai H, Bigio EH et al (2010) FUS-immunoreactive inclusions are a common feature in sporadic and non-SOD1 familial amyotrophic lateral sclerosis. Ann Neurol 67:739–748
Droppelmann CA, Keller BA, Campos-Melo D, Volkening K, Strong MJ (2012) Rho guanine nucleotide exchange factor is a NFL mRNA destabilizing factor that forms cytoplasmic inclusions in amyotrophic lateral sclerosis. Neurobiol Aging. doi:10/1016/jneurobiolaging.2012.06.021
Elden AC, Kim HJ, Hart MP et al (2010) Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature 466:1069–1075
Fiesel FC, Kahle PJ (2011) TDP-43 and FUS/TLS: cellular functions and implications for neurodegeneration. FEBS J 278:3550–3568
Forsberg K, Andersen PM, Marklund SL, Brannstrom T (2011) Glial nuclear aggregates of superoxide dismutase-1 are regularly present in patients with amyotrophic lateral sclerosis. Acta Neuropathol 121:623–634
Forsberg K, Jonsson PA, Andersen PM et al (2010) Novel antibodies reveal inclusions containing non-native SOD1 in sporadic ALS patients. PLoS ONE 5:e11552
Ge WW, Wen W, Strong W, Leystra-Lantz C, Strong MJ (2005) Mutant copper-zinc superoxide dismutase binds to and destabilizes human low molecular weight neurofilament mRNA. J Biol Chem 280:118–124
Geser F, Prvulovic D, O’Dwyer L et al (2011) On the development of markers for pathological TDP-43 in amyotrophic lateral sclerosis with and without dementia. Prog Neurobiol 95:649–662
Greenway MJ, Andersen PM, Russ C et al (2006) ANG mutations segregate with familial and ‘sporadic’ amyotrophic lateral sclerosis. Nat Genet 38:411–413
Hortobagyi T, Troakes C, Nishimura AL et al (2011) Optineurin inclusions occur in a minority of TDP-43 positive ALS and FTLD-TDP cases and are rarely observed in other neurodegenerative disorders. Acta Neuropathol 121:519–527
Ince PG, Highley JR, Kirby J et al (2011) Molecular pathology and genetic advances in amyotrophic lateral sclerosis: an emerging molecular pathway and the significance of glial pathology. Acta Neuropathol 122:657–671
Ito H, Fujita K, Nakamura M et al (2011) Optineurin is co-localized with FUS in basophilic inclusions of ALS with FUS mutation and in basophilic inclusion body disease. Acta Neuropathol 121:555–557
Kabashi E, Agar JN, Strong MJ, Durham HD (2012) Impaired proteasome function in sporadic amyotrophic lateral sclerosis. Amyotroph Lateral Scler 13:367–371
Kato S, Saito M, Hirano A, Ohama E (1999) Recent advances in research on neuropathological aspects of familial amyotrophic lateral sclerosis with superoxide dismutase 1 gene mutations: neuronal Lewy body-like hyaline inclusions and astrocytic hyaline inclusions. Histol Histopathol 14:973–989
Kwiatkowski TJ Jr, Bosco DA, Leclerc AL et al (2009) Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323:1205–1208
Kwong LK, Neumann M, Sampathu DM, Lee VM, Trojanowski JQ (2007) TDP-43 proteinopathy: the neuropathology underlying major forms of sporadic and familial frontotemporal lobar degeneration and motor neuron disease. Acta Neuropathol 114:63–70
Lagier-Tourenne C, Polymenidou M, Cleveland DW (2010) TDP-43 and FUS/TLS: emerging roles in RNA processing and neurodegeneration. Hum Mol Genet 19:R46–R64
Lee T, Li YR, Ingre C et al (2011) Ataxin-2 intermediate-length polyglutamine expansions in European ALS patients. Hum Mol Genet 20:1697–1700
Li X, Lu L, Bush DJ et al (2009) Mutant copper-zinc superoxide dismutase associated with amyotrophic lateral sclerosis binds to adenine/uridine-rich stability elements in the vascular endothelial growth factor 3′-untranslated region. J Neurochem 108:1032–1044
Liu HN, Sanelli T, Horne P et al (2009) Lack of evidence of monomer/misfolded superoxide dismutase-1 in sporadic amyotrophic lateral sclerosis. Ann Neurol 66:75–80
Lu L, Wang S, Zheng L et al (2009) Amyotrophic lateral sclerosis-linked mutant SOD1 sequesters Hu antigen R (HuR) and TIA-1-related protein (TIAR): implications for impaired post-transcriptional regulation of vascular endothelial growth factor. J Biol Chem 284:33989–33998
Lu L, Zheng L, Viera L et al (2007) Mutant Cu/Zn-superoxide dismutase associated with amyotrophic lateral sclerosis destabilizes vascular endothelial growth factor mRNA and downregulates its expression. J Neurosci 27:7929–7938
Mackenzie IR, Bigio EH, Ince PG et al (2007) Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann Neurol 61:427–434
Majounie E, Renton AE, Mok K et al (2012) Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurol 11:323–330
Maruyama H, Morino H, Ito H et al (2010) Mutations of optineurin in amyotrophic lateral sclerosis. Nature 465:223–226
Menzies FM, Grierson AJ, Cookson MR et al (2002) Selective loss of neurofilament expression in Cu/Zn superoxide dismutase (SOD1) linked amyotrophic lateral sclerosis. J Neurochem 82:1118–1128
Migheli A, Pezzulo T, Attanasio A, Schiffer D (1993) Peripherin immunoreactive structures in amyotrophic lateral sclerosis. Lab Invest 68:185–191
Moisse K, Strong MJ (2006) Innate immunity in amyotrophic lateral sclerosis. Biochim Biophys Acta 1762:1083–1093
Moisse K, Volkening K, Leystra-Lantz C et al (2009) Divergent patterns of cytosolic TDP-43 and neuronal progranulin expression following axotomy: implications for TDP-43 in the physiological response to neuronal injury. Brain Res 1249:202–211
Nakano T, Nakaso K, Nakashima K, Ohama E (2004) Expression of ubiquitin-binding protein p62 in ubiquitin-immunoreactive intraneuronal inclusions in amyotrophic lateral sclerosis with dementia: analysis of five autopsy cases with broad clinicopathological spectrum. Acta Neuropathol 107:359–364
Neumann M, Sampathu DM, Kwong LK et al (2006) Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314:130–133
Okamoto Y, Ihara M, Urushitani M et al (2011) An autopsy case of SOD1-related ALS with TDP-43 positive inclusions. Neurology 77:1993–1995
Osawa T, Mizuno Y, Fujita Y et al (2011) Optineurin in neurodegenerative diseases. Neuropathology 31:569–574
Rademakers R, Neumann M, Mackenzie IR (2012) Advances in understanding the molecular basis of frontotemporal dementia. Nat Rev Neurol 8(8):423–434
Renton AE, Majounie E, Waite A et al (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72:257–268
Robertson J, Sanelli T, Xiao S et al (2007) Lack of TDP-43 abnormalities in mutant SOD1 transgenic mice shows disparity with ALS. Neurosci Lett 420:128–132
Rosen DR, Siddique T, Patterson D et al (1993) Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362:59–62
Schiffer D, Autilio-Gambetti L, Chio A et al (1991) Ubiquitin in motor neuron disease: study at the light and electron microscope. J Neuropathol Exp Neurol 50:463–473
Shibata N, Asayama K, Hirano A, Kobayashi M (1996) Immunohistochemical study on superoxide dismutases in spinal cords from autopsied patients with amyotrophic lateral sclerosis. Dev Neurosci 18:492–498
Shin J (1998) P62 and the sequestosome, a novel mechanism for protein metabolism. Arch Pharm Res 21:629–633
Sleegers K, Brouwers N, Maurer-Stroh S et al (2008) Progranulin genetic variability contributes to amyotrophic lateral sclerosis. Neurology 71:253–259
Strong MJ (2003) The basic aspects of therapeutics in amyotrophic lateral sclerosis. Pharmacol Ther 98:379–414
Strong MJ (2008) The syndromes of frontotemporal dysfunction in amyotrophic lateral sclerosis. Amyotroph Lateral Scler 9:323–338
Strong MJ (2010) The evidence for altered RNA metabolism in amyotrophic lateral sclerosis (ALS). J Neurol Sci 288:1–12
Strong MJ, Grace GM, Freedman M et al (2009) Consensus criteria for the diagnosis of frontotemporal cognitive and behavioural syndromes in amyotrophic lateral sclerosis. Amyotroph Lateral Scler 10:131–146
Strong MJ, Kesavapany S, Pant HC (2005) The pathobiology of amyotrophic lateral sclerosis: a proteinopathy? J Neuropathol Exp Neurol 64:649–664
Strong MJ, Volkening K, Hammond R et al (2007) TDP43 is a human low molecular weight neurofilament (hNFL) mRNA-binding protein. Mol Cell Neurosci 35:320–327
Sumi H, Kato S, Mochimaru Y et al (2009) Nuclear TAR DNA binding protein 43 expression in spinal cord neurons correlates with the clinical course in amyotrophic lateral sclerosis. J Neuropathol Exp Neurol 68:37–47
Thyagarajan B, Liu Y, Shin S et al (2008) Creation of engineered human embryonic stem cell lines using phiC31 integrase. Stem Cells 26:119–126
Thyagarajan B, Olivares EC, Hollis RP, Ginsburg DS, Calos MP (2001) Site-specific genomic integration in mammalian cells mediated by phage phiC31 integrase. Mol Cell Biol 21:3926–3934
Vadlamudi RK, Joung I, Strominger JL, Shin J (1996) p62, a phosphotyrosine-independent ligand of the SH2 domain of p56lck, belongs to a new class of ubiquitin-binding proteins. J Biol Chem 271:20235–20237
Van Damme P, Veldink JH, van Blitterswijk M et al (2011) Expanded ATXN2 CAG repeat size in ALS identifies genetic overlap between ALS and SCA2. Neurology 76:2066–2072
Vance C, Rogelj B, Hortobagyi T et al (2009) Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323:1208–1211
Volkening K, Leystra-Lantz C, Strong MJ (2010) Human low molecular weight neurofilament (NFL) mRNA interacts with a predicted p190RhoGEF homologue (RGNEF) in humans. Amyotroph Lateral Scler 11:97–103
Wong NK, He BP, Strong MJ (2000) Characterization of neuronal intermediate filament protein expression in cervical spinal motor neurons in sporadic amyotrophic lateral sclerosis (ALS). J Neuropathol Exp Neurol 59:972–982
Zielinski J, Kilk K, Peritz T et al (2006) In vivo identification of ribonucleoprotein-RNA interactions. Proc Natl Acad Sci USA 103:1557–1562
Acknowledgments
This work was supported by the McFeat Family Fund and the Michael Halls Endowment. The authors declare no conflicts of interest. Postmortem spinal cord tissues were provided by the Department of Pathology at the London Health Sciences Center at the University of Western Ontario, London, Canada, and were obtained using standard autopsy consents. We thank Drs Janice Robertson and John Ravits for making available additional mtSOD1 slides. We are thankful to the donors and their families who contributed to this work through the donation of tissues and their consent to diagnostic confirmation and research. We also wish to thank Dr. Stephen Pasternak for his assistance with Bitplane’s Imaris image analysis software.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Keller, B.A., Volkening, K., Droppelmann, C.A. et al. Co-aggregation of RNA binding proteins in ALS spinal motor neurons: evidence of a common pathogenic mechanism. Acta Neuropathol 124, 733–747 (2012). https://doi.org/10.1007/s00401-012-1035-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-012-1035-z