Abstract
Key message
A novel powdery mildew resistance gene Pm2Mb from Aegilops biuncialis was transferred into common wheat and mapped to chromosome 2MbL bin FL 0.49–0.66 by molecular cytogenetic analysis of 2Mb recombinants.
Abstract
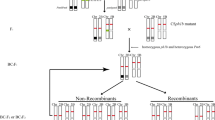
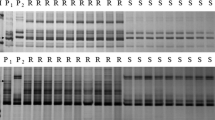
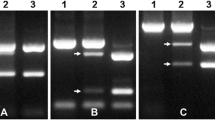
Aegilops biuncialis, a wild relative of common wheat, is highly resistant to powdery mildew. Previous studies identified that chromosome 2Mb in Chinese Spring (CS)-Ae. biuncialis 2Mb disomic addition line TA7733 conferred high resistance to powdery mildew, and the resistance gene was temporarily designated as Pm2Mb. In this study, a total of 65 CS-Ae. biuncialis 2Mb recombinants were developed by ph1b-induced homoeologous recombination and they were grouped into 12 different types based on the presence of different markers of 2Mb-specificity. Segment sizes and breakpoints of each 2Mb recombinant type were further characterized using in situ hybridization and molecular marker analyses. Powdery mildew responses of each type were assessed by inoculation of each 2Mb recombinant-derived F2 progenies using the isolate E05. Combined analyses of in situ hybridization, molecular markers and powdery mildew resistance data of the 2Mb recombinants, the gene Pm2Mb was cytologically located to an interval of FL 0.49–0.66 in the long arm of 2Mb, where 19 2Mb-specific markers were located. Among the 65 2Mb recombinants, T-11 (T2DS.2DL-2MbL) and T-12 (Ti2DS.2DL-2MbL-2DL) contained a small 2MbL segment harboring Pm2Mb. Besides, a physical map of chromosome 2Mb was constructed with 70 2Mb-specific markers in 10 chromosomal bins and the map showed that submetacentric chromosome 2Mb of Ae. biuncialis was rearranged by a terminal intrachromosomal translocation. The newly developed 2Mb recombinants with powdery mildew resistance, the 2Mb-specific molecular markers and the physical map of chromosome 2Mb will benefit wheat disease breeding as well as fine mapping and cloning of Pm2Mb.





Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. Common wheat CS, Ae. comosa accession TA2102, CS-Ae. biuncialis 2Mb disomic addition line TA7733 and CS ph1b mutant TA3809 are maintained by Wheat Genetic Resource Center at Kansas State University, USA, and Henan Agricultural University, China. Sixty-five CS-Ae. biuncialis 2Mb recombinants maintained by Henan Agricultural University, China. All plant materials in this study are available for research purposes inside and outside the country from the corresponding author by request.
References
Chen PD, Qi LL, Zhou B, Zhang SZ, Liu DJ (1995) Development and molecular cytogenetic analysis of wheat-Haynaldia villosa 6VS/6AL translocation lines specifying resistance to powdery mildew. Theor Appl Genet 91:1125–1128
Chen F, Jia H, Zhang X, Qiao L, Li X, Zheng J, Guo H, Powers C, Yan L, Chang Z (2019) Positional cloning of PmCH1357 reveals the origin and allelic variation of the Pm2 gene for powdery mildew resistance in wheat. Crop J 7:771–783
Damania AB, Pecetti L (1990) Variability in a collection of Aegilops species and evaluation for yellow rust resistance at two locations in northern Syria. J Genet Breed 44:97–102
Darko E, Khalil R, Dobi Z, Kovacs V, Szalai G, Janda T, Molnar I (2020) Addition of Aegilops biuncialis chromosomes 2M or 3M improves the salt tolerance of wheat in different way. Sci Rep 10:22327
Dong ZJ, Tian XB, Ma C, Xia Q, Wang BL, Chen QF, Sehgal SK, Friebe B, Li HH, Liu WX (2020) Physical mapping of Pm57, a powdery mildew resistance gene derived from Aegilops searsii. Int J Mol Sci 21:322
Dong Y, Xu D, Xu XW, Ren Y, Gao FM, Song J, Jia AL, Hao YF, He ZH, Xia XC (2022) Fine mapping of QPm.caas-3BS, a stable QTL for adult-plant resistance to powdery mildew in wheat (Triticum aestivum L.). Theor Appl Genet 135:1083–1099
Du P, Zhuang LF, Wang YZ, Yuan L, Wang Q, Wang DR, Dawadondup TLJ, Shen J, Xu HB, Zhao H, Chu CG, Qi ZJ (2017) Development of oligonucleotides and multiplex probes for quick and accurate identification of wheat and Thinopyrum bessarabicum chromosomes. Genome 60:93–103
Endo TR, Gill BS (1996) The deletion stocks of common wheat. J Hered 87:295–307
Farkas A, Molnar I, Dulai S, Rapi S, Oldal V, Cseh A, Kruppa K, Molnar-Lang M (2014) Increased micronutrient content (Zn, Mn) in the 3Mb(4B) wheat-Aegilops biuncialis substitution and 3Mb.4BS translocation identified by GISH and FISH. Genome 57:61–67
Gyawali Y, Zhang W, Chao S, Xu S, Cai X (2019) Delimitation of wheat ph1b deletion and development of ph1b-specific DNA markers. Theor Appl Genet 132:195–204
He HG, Du HN, Liu RK, Liu TL, Yang LJ, Gong SJ, Tang ZX, Du HM, Liu C, Han R, Sun WH, Wang L, Zhu SY (2021) Characterization of a new gene for resistance to wheat powdery mildew on chromosome 1RL of wild Secale sylvestre. Theor Appl Genet 134:887–896
Huang J, Zhao ZH, Song FJ, Wang XM, Xu HX, Huang Y, An DG, Li HJ (2012) Molecular detection of a gene effective against powdery mildew in the wheat cultivar Liangxing 66. Mol Breed 30:1737–1745
Huang XY, Zhu MQ, Zhuang LF, Zhang SY, Wang JJ, Chen XJ, Wang DR, Chen JY, Bao YG, Guo J, Zhang JL, Feng YG, Chu CG, Du P, Qi ZJ, Wang HG, Chen PD (2018) Structural chromosome rearrangements and polymorphisms identified in Chinese wheat cultivars by high-resolution multiplex oligonucleotide FISH. Theor Appl Genet 131:1967–1986
Ishikawa G, Nakamura T, Ashida T, Saito M, Nasuda S, Endo TR, Wu JZ, Matsumoto T (2009) Localization of anchor loci representing five hundred annotated rice genes to wheat chromosomes using PLUG markers. Theor Appl Genet 118:499–514
Juroszek P, von Tiedemann A (2013) Climate change and potential future risks through wheat diseases: a review. Eur J Plant Pathol 136:21–33
Li GQ, Fang TL, Zhang HT, Xie CJ, Li HJ, Yang T, Nevo E, Fahima T, Sun QX, Liu ZY (2009) Molecular identification of a new powdery mildew resistance gene Pm41 on chromosome 3BL derived from wild emmer (Triticum turgidum var. dicoccoides). Theor Appl Genet 119:531–539
Li SJ, Lin ZS, Liu C, Wang K, Du LP, Ye XG (2017) Development and comparative genomic mapping of Dasypyrum villosum 6V#4S-specific PCR markers using transcriptome data. Theor Appl Genet 130:2057–2068
Li HH, Dong ZJ, Ma C, Tian XB, Xiang ZG, Xia Q, Ma PT, Liu WX (2019) Discovery of powdery mildew resistance gene candidates from Aegilops biuncialis chromosome 2Mb based on transcriptome sequencing. PLoS ONE 14:e0220089
Li HH, Dong ZJ, Ma C, Xia Q, Tian XB, Sehgal S, Koo DH, Friebe B, Ma PT, Liu WX (2020) A spontaneous wheat-Aegilops longissima translocation carrying Pm66 confers resistance to powdery mildew. Theor Appl Genet 133:1149–1159
Liu Z, Sun Q, Ni Z, Yang T (1999) Development of SCAR markers linked to the Pm21 gene conferring resistance to powdery mildew in common wheat. Plant Breed 118:215–219
Liu WX, Koo DH, Xia Q, Li CX, Bai FQ, Song YL, Friebe B, Gill BS (2017) Homoeologous recombination-based transfer and molecular cytogenetic mapping of powdery mildew-resistant gene Pm57 from Aegilops searsii into wheat. Theor Appl Genet 130:841–848
Luo PG, Zhang HY, Shu K, Wu XH, Zhang HQ, Ren ZL (2009) The physiological genetic effffects of 1BL/1RS translocated chromosome in “stay green” wheat cultivar CN17. Can J Plant Sci 89:1–10
Makkouk K, Ghulam W, Comeau A (1994) Resistance to barley yellow dwarf luteovirus in Aegilops species. Can J Plant Sci 74:631–634
McIntosh R, Dubcovsky J, Rogers W, Morris C, Xia XC (2017) Catalogue of gene symbols for wheat. 2017. Supplement Annu Wheat Newsl 53:107–128
Nasuda S, Friebe B, Busch W, Kynast RG, Gill BS (1998) Structural rearrangement in chromosome 2M of Aegilops comosa has prevented the utilization of the Compair and related wheat-Ae. comosa translocations in wheat improvement. Theor Appl Genet 96:780–785
Niu Z, Klindworth DL, Yu G, Friesen TL, Chao S, Jin Y, Cai X, Ohm JB, Rasmussen JB, Xu SS (2014) Development and characterization of wheat lines carrying stem rust resistance gene Sr43 derived from Thinopyrum ponticum. Theor Appl Genet 127:969–980
Reynolds M, Foulkes J, Furbank R, Griffiths S, King J, Murchie E, Parry M, Slafer M (2012) Achieving yield gains in wheat. Plant Cell Environ 35:1799–1823
Riley R, Chapman V (1958) Genetic control of the cytologically diploid behaviour of hexaploid wheat. Nature 182:713–715
Said M, Holusova K, Farkas A, Ivanizs L, Gaal E, Capal P, Abrouk M, Martis-Thiele MM, Kalapos B, Bartos J, Friebe B, Dolezel J, Molnar I (2021) Development of DNA markers from physically mapped loci in Aegilops comosa and Aegilops umbellulata using single-gene FISH and chromosome sequences. Front Plant Sci 12:689031
Schneider A, Linc G, Molnar I, Molnar-Lang M (2005) Molecular cytogenetic characterization of Aegilops biuncialis and its use for the identification of 5 derived wheat-Aegilops biuncialis disomic addition lines. Genome 48:1070–1082
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675
Sears E (1977) Genetics society of Canada award of excellence lecture an induced mutant with homoeologous pairing in common wheat. Can J Genet Cytol 19:585–593
Sun H, Hu J, Song W, Qiu D, Cui L, Wu P, Zhang H, Liu H, Yang L, Qu Y, Li Y, Li T, Cheng W, Zhou Y, Liu Z, Li J, Li H (2018) Pm61: a recessive gene for resistance to powdery mildew in wheat landrace Xuxusanyuehuang identified by comparative genomics analysis. Theor Appl Genet 131:2085–2097
Sun YY, Lyu MJ, Han HM, Zhou SH, Lu YQ, Liu WH, Yang XM, Li XQ, Zhang JP, Liu X, Li LH (2021) Identification and fine mapping of alien fragments associated with enhanced grain weight from Agropyron cristatum chromosome 7P in common wheat backgrounds. Theor Appl Genet 134:3759–3772
Wan WT, Xiao J, Li ML, Tang X, Wen MX, Cheruiyot AK, Li YB, Wang HY, Wang X (2020) Fine mapping of wheat powdery mildew resistance gene Pm6 using 2B/2G homoeologous recombinants induced by the ph1b mutant. Theor Appl Genet 133:1265–1275
Wang XW, Lai JR, Chen LH, Liu GT (1998) Molecular identification for Chinese Spring ph1b mutant. Sci Agric Sinca 31:31–34
Wang KY, Lin ZS, Wang L, Wang K, Shi QH, Du LP, Ye XG (2018) Development of a set of PCR markers specific to Aegilops longissima chromosome arms and application in breeding a translocation line. Theor Appl Genet 131:13–25
Xie JZ, Guo GH, Wang Y, Hu TZ, Wang LL, Li JT, Qiu D, Li YH, Wu QH, Lu P, Chen YX, Dong LL, Li MM, Zhang HZ, Zhang PP, Zhu KY, Li BB, Deal KR, Huo NX, Zhang Y, Luo MC, Liu SZ, Gu YQ, Li HJ, Liu ZY (2020) A rare single nucleotide variant in Pm5e confers powdery mildew resistance in common wheat. New Phytol 228:1011–1026
Xing LP, Hu P, Liu JQ, Witek K, Zhou S, Xu JF, Zhou WH, Gao L, Huang ZP, Zhang RQ, Wang XE, Chen PD, Wang HY, Jones J, Karafiatova M, Vrana J, Bartos J, Dolezel J, Tian YC, Wu YF, Cao AZ (2018) Pm21 from Haynaldia villosa encodes a CC-NBS-LRR protein conferring powdery mildew resistance in wheat. Mol Plant 11:874–878
Xue SL, Zhang ZZ, Lin F, Kong ZX, Cao Y, Li CJ, Yi HY, Mei MF, Zhu HL, Wu J, Xu HB, Zhao DM, Tian DG, Zhang CQ, Ma ZQ (2008) A high-density intervarietal map of the wheat genome enriched with markers derived from expressed sequence tags. Theor Appl Genet 117:181–189
Zhang W, Cao Y, Zhang M, Zhu X, Ren S, Long Y, Gyawali Y, Chao S, Xu SS, Cai X (2017) Meiotic homoeologous recombination based alien gene introgression in the genomics era of wheat. Crop Sci 57:1189–1198
Zhang W, Zhu X, Zhang M, Chao S, Xu S, Cai X (2018) Meiotic homoeologous recombination-based mapping of wheat chromosome 2B and its homoeologues in Aegilops speltoides and Thinopyrum elongatum. Theor Appl Genet 131:2381–2395
Zhang DY, Zhu KY, Dong LL, Liang Y, Li GQ, Fang TL, Guo GH, Wu QH, Xie JZ, Chen YX, Lu P, Li MM, Zhang HZ, Wang ZZ, Zhang Y, Sun QX, Liu ZY (2019) Wheat powdery mildew resistance gene Pm64 derived from wild emmer (Triticum turgdium var. dicoccoides) is tightly linked in repulsion with stripe rust resistance gene Yr5. Crop J 7:761–770
Zhang RQ, Xiong CX, Mu HQ, Yao RN, Meng XR, Kong LN, Xing LP, Wu JZ, Feng YG, Cao AZ (2021) Pm67, a powdery mildew resistance gene transferred from Dasypyrum villosum chromosome 1V to common wheat (Triticum aestivum L.). Crop J 9:882–888
Zhao H, Zhang W, Wang J, Li FR, Cui F, Ji J, Wang DW, Li JM (2013) Comparative study on drought tolerance of wheat and wheat-Aegilops biuncialis 6Ub addition lines. J Food Agric Environ 11:1046–1052
Zhou JP, Cheng Y, Zhang LL, Yang EN, Liu C, Zheng XL, Deng KJ, Zhu YQ, Zhang Y (2014) Characterization of a new wheat-Aegilops biuncialis 1Mb(1B) substitution line with good quality-associated HMW glutenin subunits. Cereal Res Commun 44:198–205
Zhu KY, Li MM, Wu HB, Zhang DY, Dong LL, Wu QH, Chen YX, Xie JZ, Lu P, Guo GH, Zhang HZ, Zhang PP, Li BB, Li WN, Dong L, Wang QF, Zhu JH, Hu WL, Guo LQ, Wang RG, Yuan CG, Li HJ, Liu ZY, Hua W (2022) Fine mapping of powdery mildew resistance gene MlWE74 derived from wild emmer wheat (Triticum Turgidum ssp. dicoccoides) in an NBS-LRR gene cluster. Theor Appl Genet. https://doi.org/10.1007/s00122-021-04027-2
Acknowledgements
We would like to thank Dr. Pengtao Ma (College of Life and Sciences, Yantai University) for providing Bgt isolate E05.
Funding
This research was financially supported by the National Natural Science Foundation of China (Nos. 31801361 and 31971887), the Scientific and Technological Research Project of Henan Province of China (No. 212102110059) and the Topnotch Talents of Henan Agricultural University (30500939).
Author information
Authors and Affiliations
Contributions
WXL and HHL: conceived the research. WQM, HHL and ZWF: performed the research. CM, YZ, CLW, XBT, QFC, JNM, JQH and SKS: participated in the preparation of the reagents and materials used in this study. HHL, WXL and SKS: wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Evans Lagudah.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Men, W., Fan, Z., Ma, C. et al. Mapping of the novel powdery mildew resistance gene Pm2Mb from Aegilops biuncialis based on ph1b-induced homoeologous recombination. Theor Appl Genet 135, 2993–3003 (2022). https://doi.org/10.1007/s00122-022-04162-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04162-4