Abstract
Key message
Using the ph1b mutant, the recombination frequency between the homoeologous region of 2B and 2G was significantly increased. By this, we narrowed Pm6 to a 0.9 Mb physical region.
Abstract
The powdery mildew (Pm) resistance gene Pm6 from Triticum timopheevii (2n = 48, AAGG) was mapped to the long arm of chromosome 2G and introduced into common wheat in the form of 2B–2G introgressions. The introgression line IGV1-465 has the shortest 2G segment, which is estimated 37 Mb in size when referring to 2BL genome reference of Chinese Spring (CS). The further fine mapping of Pm6 was impeded by the inhibition of allogeneic chromosome recombination between 2B and 2G in the Pm6 region. In the present study, to overcome 2B/2G recombination suppression, a ph1b-based strategy was employed to produce introgressions with reduced 2G fragments for the fine mapping of Pm6. IGV1-465 was crossed and backcrossed to the CSph1b mutant to produce plants with increased 2B/2G chromosome pairing frequency at the Pm6 region. A total of 182 allogeneic recombinants were obtained through two-round screening, i.e., first round of screening of 820 BC1F2:3 progenies using the flanking markers CIT02g-14/CIT02g-19 and second round of screening of 642 BC1F2:4 progenies using the flanking markers CIT02g-13/CIT02g-18, respectively. Through marker analysis using 30 chromosome 2G-specific markers located in the Pm6 region, the identified recombinants were divided into 14 haplotypes. Pm resistance evaluation of these haplotypes enabled us to narrow Pm6 to a 0.9 Mb physical region of 2BL, flanked by markers CIT02g-20 and CIT02g-18. Six wheat varieties containing Pm6 were identified from a natural population, and they showed increased Pm resistance. This implied Pm6 is still effective, especially when used in combination with other Pm resistance genes.




Similar content being viewed by others
References
Bennett F (1984) Resistance to powdery mildew in wheat: a review of its use in agriculture and breeding programmes. Plant Pathol 33:279–300. https://doi.org/10.1111/j.1365-3059.1984.tb01324.x
Cao A, Xing L, Wang X, Yang X, Wang X, Sun Y et al (2011) Serine/threonine kinase gene Stpk-V, a key member of powdery mildew resistance gene Pm21, confers powdery mildew resistance in wheat. Proc Natl Acad Sci 108:7727–7732. https://doi.org/10.1073/pnas.1016981108
Chen T, Xiao J, Xu J, Wan W, Qin B, Cao A et al (2016) Two members of TaRLK family confer powdery mildew resistance in common wheat. BMC Plant Biol 16:27. https://doi.org/10.1186/s12870-016-0713-8
Costamilan L, Trigo E (2005) Variability of the wheat powdery mildew pathogen Blumeria graminis f. sp. tritici in the 2003 crop season. Fitopatol Bras 30:420–422. https://doi.org/10.1590/s0100-41582005000400015
Giorgi B (1978) A homoeologous pairing mutant isolated in Triticum durum cv. Cappelli. Mutat Breed Newsl 11:4–5
Herrera-Foessel S, Singh R, Lillemo M, Huerta-Espino J, Bhavani S, Singh S et al (2014) Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theor Appl Genet 127:781–789. https://doi.org/10.1007/s00122-013-2256-9
Hurni S, Brunne S, Buchmann G, Herren G, Jordan T, Krukowski P et al (2013) Rye Pm8 and wheat Pm3 are orthologous genes and show evolutionary conservation of resistance function against powdery mildew. Plant J 76:957–969. https://doi.org/10.1111/tpj.12345
Ishikawa G, Yonemaru J, Saito M, Nakamura T (2007) PCR-based landmark unique gene (PLUG) markers effectively assign homoeologous wheat genes to A, B and D genomes. BMC Genom 8:135. https://doi.org/10.1186/1471-2164-8-135
Järve K, Peusha H, Tsymbalova J, Tamm S, Devos K, Enno T (2000) Chromosomal location of a Triticum timopheevii-derived powdery mildew resistance gene transferred to common wheat. Genome 43:377–381. https://doi.org/10.1139/g99-141
Jayatilake D, Tucker E, Bariana H, Kuchel H, Edwards J, McKay A et al (2013) Genetic mapping and marker development for resistance of wheat against the root lesion nematode Pratylenchus neglectus. BMC Plant Biol 13:230. https://doi.org/10.1186/1471-2229-13-230
Ji J, Wang H, Cao A, Wang S, Zhuang L, Chen P et al (2006) STS markers for powdery mildew resistance gene Pm6 in wheat. Euphytica 2008 163(2):159–165. https://doi.org/10.1007/s10681-007-9578-0
Jia J, Devos K, Chao S, Miller T, Reader S, Gale M (1996) RFLP-based maps of the homoeologous group-6 chromosomes of wheat and their application in the tagging of Pm12, a powdery mildew resistance gene transferred from Aegilops speltoides to wheat. Theor Appl Genet 92:559–565. https://doi.org/10.1007/bf00224558
Jorgensen J, Jensen C (1973) Gene Pm6 for resistance to powdery mildew in wheat. Euphytica. https://doi.org/10.1007/BF00022656
Kilian B, Ozkan H, Deusch O, Effgen S, Brandolini A, Kohl J et al (2007) Independent wheat B and G genome origins in outcrossing Aegilops progenitor haplotypes. Mol Biol Evol 24:217–227. https://doi.org/10.1093/molbev/msl151
Lagudah E, Krattinger S, Herrera-Foessel S, Singh R, Huerta-Espino J, Spielmeyer W et al (2009) Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor Appl Genet 119:889–898. https://doi.org/10.1007/s00122-009-1097-z
Li H, Wang X, Song F, Wu C, Wu X, Zhang N et al (2011) Response to powdery mildew and detection of resistance genes in wheat cultivars from China. Acta Agron Sin 37:943–954. https://doi.org/10.1016/s1875-2780(11)60026-6
Li J, Zhou R, Endo T, Stein N (2018) High-throughput development of SSR marker candidates and their chromosomal assignment in rye (Secale cereale L.). Plant Breed 137:561–572. https://doi.org/10.1111/pbr.12619
Li G, Cowger C, Wang X, Carver B, Xu X (2019) Characterization of Pm65, a new powdery mildew resistance gene on chromosome 2AL of a facultative wheat cultivar. Theor Appl Genet 132(9):2625–2632. https://doi.org/10.1007/s00122-019-03377-2
Lillemo M, Asalf B, Singh R, Huerta-Espino J, Chen X, He Z et al (2008) The adult plant rust resistance loci Lr34/Yr18 and Lr46/Yr29 are important determinants of partial resistance to powdery mildew in bread wheat line Saar. Theor Appl Genet 116:1155–1166. https://doi.org/10.1007/s00122-008-0743-1
Lukaszewski A, Rybka K, Korzun V, Malyshev S, Lapinski B, Whitkus R (2004) Genetic and physical mapping of homoeologous recombination points involving wheat chromosome 2B and rye chromosome 2R. Genome 47:36–45. https://doi.org/10.1139/g03-089
Maxwell J, Lyerly J, Cowger C, Marshall D, Brown-Guedira G, Murphy J (2009) MlAG12: a Triticum timopheevii-derived powdery mildew resistance gene in common wheat on chromosome 7AL. Theor Appl Genet 119:1489–1495. https://doi.org/10.1007/s00122-009-1150-y
Mayer K, Rogers J, Dolezel J, Pozniak C, Eversole K, Feuillet C et al (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:6194. https://doi.org/10.1126/science.1251788
McIntosh R, Dubcovsky J, Rogers W, Morris C, Appels R, Xia X (2013) Catalogue of gene symbols for wheat. In: KOMUGI-integrated wheat science database at http://www.shigen.nig.ac.jp/wheat/komugi/genes/download.jsp
Mullan D, Mirzaghaderi G, Walker E, Colmer TD, Francki M (2009) Development of wheat–Lophopyrum elongatum recombinant lines for enhanced sodium ‘exclusion’ during salinity stress. Theor Appl Genet 119:1313–1323. https://doi.org/10.1007/s00122-009-1136-9
Neu C, Stein N, Keller B (2002) Genetic mapping of the Lr20-Pm1 resistance locus reveals suppressed recombination on chromosome arm 7AL in hexaploid wheat. Genome 45:737–744. https://doi.org/10.1139/g02-040
Perugini L, Murphy J, Marshall D, Brown-Guedira G (2008) Pm37, a new broadly effective powdery mildew resistance gene from Triticum timopheevii. Theor Appl Genet 116:417–425. https://doi.org/10.1007/s00122-007-0679-x
Peusha H, Enno T, Priilinn O (2000) Chromosomal location of powdery mildew resistance genes and cytogenetic analysis of meiosis in common wheat cultivar Meri. Hereditas 132:29–34. https://doi.org/10.1111/j.1601-5223.2000.00029.x
Purnhauser L, Bóna L, Láng L (2011) Occurrence of 1BL. 1RS wheat-rye chromosome translocation and of Sr36/Pm6 resistance gene cluster in wheat cultivars registered in Hungary. Euphytica 179:287–295. https://doi.org/10.1007/s10681-010-0312-y
Qi L, Friebe B, Zhang P, Gill B (2007) Homoeologous recombination, chromosome engineering and crop improvement. Chromosome Res 15:3–19. https://doi.org/10.1007/s10577-006-1108-8
Qi L, Pumphrey M, Friebe B, Chen P, Gill B (2008) Molecular cytogenetic characterization of alien introgressions with gene Fhb3 for resistance to Fusarium head blight disease of wheat. Theor Appl Genet 117:1155–1166. https://doi.org/10.1007/s00122-008-0853-9
Qin B, Cao A, Wang H, You F, Liu Y et al (2011) Collinearity-based marker mining for the fine mapping of Pm6, a powdery mildew resistance gene in wheat. Theor Appl Genet 123:207–218. https://doi.org/10.1007/s00122-011-1577-9
Rey M, Calderón M, Prieto P (2015) The use of the ph1b mutant to induce recombination between the chromosomes of wheat and barley. Front Plant Sci 6:160. https://doi.org/10.3389/fpls.2015.00160
Riley R, Chapman V (1958) Genetic control of the cytologically diploid behaviour of hexaploid wheat. Nature 182:713. https://doi.org/10.1038/182713a0
Sanchez-Martin J, Steuernagel B, Ghosh S, Herren G, Hurni S, Adamski N et al (2016) Rapid gene isolation in barley and wheat by mutant chromosome sequencing. Genome Biol 17:221. https://doi.org/10.1186/s13059-016-1082-1
Sears E (1977) Genetics society of Canada award of excellence lecture an induced mutant with homoeologous pairing in common wheat. Can J Genet Cytol 19:585–593. https://doi.org/10.1139/g77-063
Sun H, Hu J, Song W, Qiu D, Cui L, Wu P et al (2018) Pm61: a recessive gene for resistance to powdery mildew in wheat landrace Xuxusanyuehuang identified by comparative genomics analysis. Theor Appl Genet 131:2085–2097. https://doi.org/10.1007/s00122-018-3135-1
Švec M, Miklovičová M (1998) Structure of populations of wheat powdery mildew (Erysiphe graminis DC f. sp. tritici Marchal) in Central Europe in 1993–1996: I. Dynamics of virulence. Eur J Plant Pathol 104:537–544. https://doi.org/10.1023/a:1008642816326
Tan C, Li G, Cowger C, Carver B, Xu X (2018a) Characterization of Pm59, a novel powdery mildew resistance gene in Afghanistan wheat landrace PI 181356. Theor Appl Genet 131:1145–1152. https://doi.org/10.1007/s00122-018-3067-9
Tan C, Li G, Cowger C, Carver B, Xu X (2018b) Characterization of Pm63, a powdery mildew resistance gene in Iranian landrace PI 628024. Theor Appl Genet 132:1137. https://doi.org/10.1007/s00122-018-3265-5
Tao W, Liu D, Liu J, Feng Y, Chen P (2000) Genetic mapping of the powdery mildew resistance gene Pm6 in wheat by RFLP analysis. Theor Appl Genet 100:564–568. https://doi.org/10.1007/s001220050074
The International Wheat Genome Sequencing Consortium (IWGSC) (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science. https://doi.org/10.1126/science.aar7191
Thiel T, Michalek W, Varshney R, Graner A (2003) Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106:411–422. https://doi.org/10.1007/s00122-002-1031-0
Vechet L (2006) Reaction of winter wheat cultivars and breeding lines to Blumeria graminis f. sp. tritici. Plant Protect Sci 42:15–20. https://doi.org/10.17221/2691-PPS
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang B et al (2014) Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796. https://doi.org/10.1111/pbi.12183
Wang H, Dai K, Xiao J, Yuan C, Zhao R, Dolezal J et al (2017) Development of intron targeting (IT) markers specific for chromosome arm 4VS of Haynaldia villosa by chromosome sorting and next-generation sequencing. BMC Genom 18:167. https://doi.org/10.1186/s12864-017-3567-z
Xiao J, Dai K, Fu L, Vrana J, Kubalakova M, Wan W et al (2017) Sequencing flow-sorted short arm of Haynaldia villosa chromosome 4V provides insights into its molecular structure and virtual gene order. BMC Genom 18:791. https://doi.org/10.1186/s12864-017-4211-7
Xie W, Ben-David R, Zeng B, Dinoor A, Xie C, Sun Q et al (2012) Suppressed recombination rate in 6VS/6AL translocation region carrying the Pm21 locus introgressed from Haynaldia villosa into hexaploid wheat. Mol Breed 29:399–412. https://doi.org/10.1007/s11032-011-9557-y
Xin Z, Zhang Z, Chen X, Lin Z, Ma Y, Xu H et al (2001) Development and characterization of common wheat-Thinopyrum intermedium translocation lines with resistance to barley yellow dwarf virus. Wheat Glob Environ Euphytica 2001 119(1–2):163–167. https://doi.org/10.1023/A:1017508915932
Xing L, Hu P, Liu J, Witek K, Zhou S, Xu J et al (2018) Pm21 from Haynaldia villosa encodes a CC-NBS-LRR protein conferring powdery mildew resistance in wheat. Mol Plant. https://doi.org/10.1016/j.molp.2018.02.013
Xue S, Zhang Z, Lin F, Kong Z, Cao Y, Li C et al (2008) A high-density intervarietal map of the wheat genome enriched with markers derived from expressed sequence tags. Theor Appl Genet 117:181–189. https://doi.org/10.1007/s00122-008-0764-9
Yahiaoui N, Srichumpa P, Dudler R, Keller B (2004) Genome analysis at different ploidy levels allows cloning of the powdery mildew resistance gene Pm3b from hexaploid wheat. Plant J 37:528–538. https://doi.org/10.1046/j.1365-313X.2003.01977.x
Yu J, Dake T, Singh S, Benscher D, Li W, Gill B et al (2004) Development and mapping of EST-derived simple sequence repeat markers for hexaploid wheat. Genome 47:805–818. https://doi.org/10.1139/g04-057
Zhang X, Wei X, Xiao J, Yuan C, Wu Y, Cao A et al (2017) Whole genome development of intron targeting (IT) markers specific for Dasypyrum villosum chromosomes based on next-generation sequencing technology. Mol Breed 37:115. https://doi.org/10.1007/s11032-017-0710-0
Zhang R, Fan Y, Kong L, Wang Z, Wu J, Xing L et al (2018) Pm62, an adult-plant powdery mildew resistance gene introgressed from Dasypyrum villosum chromosome arm 2VL into wheat. Theor Appl Genet 131:2613. https://doi.org/10.1007/s00122-018-3176-5
Zhang D, Zhu K, Dong L, Liang Y, Li G, Fang T et al (2019) Wheat powdery mildew resistance gene Pm64 derived from wild emmer (Triticum turgidum var. dicoccoides) is tightly linked in repulsion with stripe rust resistance gene Yr5. Crop J. https://doi.org/10.1016/j.cj.2019.03.003
Zhao R, Wang H, Xiao J, Bie T, Cheng S, Jia Q et al (2013) Induction of 4VS chromosome recombinants using the CS ph1b mutant and mapping of the wheat yellow mosaic virus resistance gene from Haynaldia villosa. Theor Appl Genet 126:2921–2930. https://doi.org/10.1007/s00122-013-2181-y
Zou S, Wang H, Li Y, Kong Z, Tang D (2018) The NB-LRR gene Pm60 confers powdery mildew resistance in wheat. New Phytol 218:298–309. https://doi.org/10.1111/nph.14964
Acknowledgements
This research was supported by the Grants from the National Key Research and Development Program (2016YFD0101004, 2016YFD0102001-004), the National Natural Science Foundation of China (Nos. 31571653, 31771782, 31201204), the International Cooperation and Exchange of the National Natural Science Foundation of China (No. 31661143005), the Special Fund of Jiangsu Province for the Transformation of Scientific and Technological Achievements (BA2017138), the Creation of Major New Agricultural Varieties in Jiangsu Province (PZCZ201706), the SAAS Program for Excellent Research Team, the Science and Technology Service Programs of the Chinese Academy of Sciences (KFJ-STS-ZDTP-002), the Jiangsu Agricultural Technology System (JATS) (No. 2019429) and the Key Research and Development Major Project of Ningxia Hui Autonomous Region (No. 2019BBF02022-04).
Author information
Authors and Affiliations
Contributions
WXE and WHY designed the experimental plan. WWT, LML, TX, CAK and WMX performed the experiments. WWT, LML, TX, LYB and WMX managed the materials in the field. WWT, XJ and WXE wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Communicated by Albrecht E. Melchinger.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig.
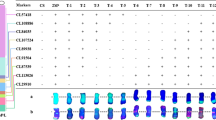
1 The distribution of previously reported markers forPm6by Qin et al. (2011) along the wheat 2B chromosome. The black boxes indicate introgression fragments from 2G of T. timopheevii; the red arrows point to the recombination site in IGV1-466/Prins and IGV1-465/Prins F2 population. Supplementary Fig. 2: The microcolinearity analysis ofPm6region in Tritium crop. Ta, Td, At, Tu and Hv represent common wheat, Triticum dicoccoides, Aegilops tauschii, Triticum urartu and Hordeum vulgare, respectively. The lines link the homologous genes between chromosomes. Supplementary Fig. 3: The flanking marker ofPm6amplified in a natural group contains 387 materials. Recombinant represents a recombinant derived from IGV1-465/CSph1b which obtains the gene of Pm6. Lanes 1 to 6 are six wheat varieties harboring Pm6; lanes 7 to 21 are representatives without Pm6. Supplementary Fig. 4: Identification ofPm6in a natural population and validation of its contribution toPmresistance. A: Comparison of Pm resistance between wheat varieties with and without Pm6, tested with Blumeria graminis f. sp. tritici (Bgt) isolate E26 and Bgt mixture, respectively; B: the geographical distribution of Pm6-containing wheat varieties in China. (PDF 520 kb)
Rights and permissions
About this article
Cite this article
Wan, W., Xiao, J., Li, M. et al. Fine mapping of wheat powdery mildew resistance gene Pm6 using 2B/2G homoeologous recombinants induced by the ph1b mutant. Theor Appl Genet 133, 1265–1275 (2020). https://doi.org/10.1007/s00122-020-03546-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03546-8