Abstract
Key message
Transcriptome-based gene expression analysis identifies many critical salt-responsive genes in radish and facilitates further dissecting the molecular mechanism underlying salt stress response.
Abstract
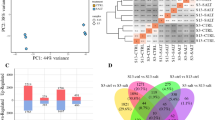
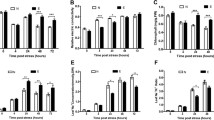
Salt stress severely impacts plant growth and development. Radish, a moderately salt-sensitive vegetable crop, has been studied for decades towards the physiological and biochemical performances under salt stress. However, no systematic study on isolation and identification of genes involved in salt stress response has been performed in radish, and the molecular mechanism governing this process is still indistinct. Here, the RNA-Seq technique was applied to analyze the transcriptomic changes on radish roots treated with salt (200 mM NaCl) for 48 h in comparison with those cultured in normal condition. Totally 8709 differentially expressed genes (DEGs) including 3931 up- and 4778 down-regulated genes were identified. Functional annotation analysis indicated that many genes could be involved in several aspects of salt stress response including stress sensing and signal transduction, osmoregulation, ion homeostasis and ROS scavenging. The association analysis of salt-responsive genes and miRNAs exhibited that 36 miRNA–mRNA pairs had negative correlationship in expression trends. Reverse-transcription quantitative PCR (RT-qPCR) analysis revealed that the expression profiles of DEGs were in line with results from the RNA-Seq analysis. Furthermore, the putative model of DEGs and miRNA-mediated gene regulation was proposed to elucidate how radish sensed and responded to salt stress. This study represents the first comprehensive transcriptome-based gene expression profiling under salt stress in radish. The outcomes of this study could facilitate further dissecting the molecular mechanism underlying salt stress response and provide a valuable platform for further genetic improvement of salt tolerance in radish breeding programs.








Similar content being viewed by others
Abbreviations
- DEG:
-
Differentially expressed gene
- RT-qPCR:
-
Reverse-transcription quantitative PCR
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- GO:
-
Gene ontology
- RPKM:
-
Reads per kilobase per million reads
- FDR:
-
False discovery rate
- TF:
-
Transcription factor
References
Agarwal PK, Shukla PS, Gupta K, Jha B (2013) Bioengineering for salinity tolerance in plants: state of the art. Mol Biotechnol 54:102–123
Álvarez S, Sánchez-Blanco M (2014) Long-term effect of salinity on plant quality, water relations, photosynthetic parameters and ion distribution in Callistemon citrinus. Plant Biol 16:757–764
Ara H, Sinha AK (2015) Role of mitogen-activated protein kinase cascade in combating abiotic stress in plants. In: Pandey GK (ed) Elucidation of abiotic stress signaling in plants. Springer, New York, pp 207–229
Ashraf M, Foolad MR (2007) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59:206–216
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanisms, and function. Cell 116:281–297
Bazihizina N, Barrett-Lennard EG, Colmer TD (2012) Plant growth and physiology under heterogeneous salinity. Plant Soil 354:1–19
Boursiac Y, Chen S, Luu DT, Sorieul M, van den Dries N, Maurel C (2005) Early effects of salinity on water transport in Arabidopsis roots. Molecular and cellular features of aquaporin expression. Plant Physiol 139:790–805
Cabot C, Sibole JV, Barceló J, Poschenrieder C (2014) Lessons from crop plants struggling with salinity. Plant Sci 226:2–13
Chen H, Jiang JG (2010) Osmotic adjustment and plant adaptation to environmental changes related to drought and salinity. Environ Rev 18:309–319
Dai X, Zhao PX (2011) psRNATarget: a plant small RNA target analysis server. Nucleic Acids Res 39:W155–W159
Dai X, Xu Y, Ma Q, Xu W, Wang T, Xue Y, Chong K (2007) Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol 143:1739–1751
Deinlein U, Stephan AB, Horie T, Luo W, Xu G, Schroeder JI (2014) Plant salt-tolerance mechanisms. Trends Plant Sci 19:371–379
Diao Y, Xu H, Li G, Yu A, Yu X, Hu W, Zheng X, Li S, Wang Y, Hu Z (2014) Cloning a glutathione peroxidase gene from Nelumbo nucifera and enhanced salt tolerance by overexpressing in rice. Mol Biol Rep 41:4919–4927
Ding HD, Zhang XH, Xu SC, Sun LL, Jiang MY, Zhang A, Jin YG (2009) Induction of protection against paraquat-induced oxidative damage by abscisic acid in maize leaves is mediated through mitogen-activated protein kinase. J Integr Plant Biol 51:961–972
Diray-Arce J, Clement M, Gul B, Khan MA, Nielsen BL (2015) Transcriptome assembly, profiling and differential gene expression analysis of the halophyte Suaeda fruticosa provides insights into salt tolerance. BMC Genom 16:353
Droillard MJ, Boudsocq M, Barbier-Brygoo H, Laurière C (2002) Different protein kinase families are activated by osmotic stresses in Arabidopsis thaliana cell suspensions: involvement of the MAP kinases AtMPK3 and AtMPK6. FEBS Lett 527:43–50
Droillard MJ, Boudsocq M, Barbier-Brygoo H, Laurière C (2004) Involvement of MPK4 in osmotic stress response pathways in cell suspensions and plantlets of Arabidopsis thaliana: activation by hypoosmolarity and negative role in hyperosmolarity tolerance. FEBS Lett 574:42–48
Golldack D, Lüking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30:1383–1391
Grattan S, Hanson B, Salinity S (2006) Crop salt tolerance. Agricultural salinity and drainage Univ of California Irrigation Program, Davis. http://hos.ufl.edu/sites/default/files/faculty/gdliu/HansonGrattan2006_0.pdf. Accessed 23 Nov 2013
Hoang XLT, Thu NBA, Thao NP, Tran LSP (2014) Transcription factors in abiotic stress responses: Their potentials in crop improvement. In: Ahmad P, Wani MR, Azooz MM, Tran LSP (eds) Improvement of crops in the era of climatic changes. Springer, New York, pp 337–366
Kakumanu A, Ambavaram MM, Klumas C, Krishnan A, Batlang U, Myers E, Grene R, Pereira A (2012) Effects of drought on gene expression in maize reproductive and leaf meristem tissue revealed by RNA-Seq. Plant Physiol 160:846–867
Kim MC, Chung WS, Yun DJ, Cho MJ (2009) Calcium and calmodulin-mediated regulation of gene expression in plants. Mol Plant 2:13–21
Kitashiba H, Li F, Hirakawa H, Kawanabe T, Zou Z, Hasegawa Y, Tonosaki K, Shirasawa S, Fukushima A, Yokoi S (2014) Draft sequences of the radish (Raphanus sativus L.) genome. DNA Res 21:481–490
Koyro HW (2006) Effect of salinity on growth, photosynthesis, water relations and solute composition of the potential cash crop halophyte Plantago coronopus (L.). Environ Exp Bot 56:136–146
Liu H, Kohane IS (2009) Tissue and process specific microRNA-mRNA co-expression in mammalian development and malignancy. PLoS One 4:e5436
Liu Y, Zhang S (2004) Phosphorylation of 1-aminocyclopropane-1-carboxylic acid synthase by MPK6, a stress-responsive mitogen-activated protein kinase, induces ethylene biosynthesis in Arabidopsis. Plant Cell 16:3386–3399
Liu YB, Liu ML, Li XR, Cao B, Ma XF (2014) Identification of differentially expressed genes in leaf of Reaumuria soongorica under PEG-Induced drought stress by digital gene expression profiling. PLoS One 9:e94277
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Ludwig AA, Romeis T, Jones JD (2004) CDPK-mediated signalling pathways: specificity and cross-talk. J Exp Bot 55:181–188
Luo X, Wu J, Li Y, Nan Z, Guo X, Wang Y, Zhang A, Wang Z, Xia G, Tian Y (2013) Synergistic effects of GhSOD1 and GhCAT1 overexpression in cotton chloroplasts on enhancing tolerance to methyl viologen and salt stresses. PLoS One 8:e54002
Marcelis L, Van Hooijdonk J (1999) Effect of salinity on growth, water use and nutrient use in radish (Raphanus sativus L.). Plant Soil 215:57–64
Mišić D, Šiler B, Živković JN, Simonović A, Maksimović V, Budimir S, Janošević D, Đuričković M, Nikolić M (2012) Contribution of inorganic cations and organic compounds to osmotic adjustment in root cultures of two Centaurium species differing in tolerance to salt stress. Plant Cell Tissue Org 108:389–400
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628
Moustafa K, AbuQamar S, Jarrar M, Al-Rajab AJ, Trémouillaux-Guiller J (2014) MAPK cascades and major abiotic stresses. Plant Cell Rep 33:1217–1225
Munns R, Gilliham M (2015) Salinity tolerance of crops—what is the cost? New Phytol. doi:10.1111/nph.13519
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Nakagami H, Pitzschke A, Hirt H (2005) Emerging MAP kinase pathways in plant stress signalling. Trends Plant Sci 10:339–346
Naliwajski M, Skłodowska M (2014) The oxidative stress and antioxidant systems in cucumber cells during acclimation to salinity. Biol Plant 58:47–54
Nounjan N, Nghia PT, Theerakulpisut P (2012) Exogenous proline and trehalose promote recovery of rice seedlings from salt-stress and differentially modulate antioxidant enzymes and expression of related genes. J Plant Physiol 169:596–604
O’Rourke JA, Yang SS, Miller SS, Bucciarelli B, Liu J, Rydeen A, Bozsoki Z, Uhde-Stone C, Tu ZJ, Allan D (2013) An RNA-Seq transcriptome analysis of orthophosphate-deficient white lupin reveals novel insights into phosphorus acclimation in plants. Plant Physiol 161:705–724
Oh SJ, Song SI, Kim YS, Jang HJ, Kim SY, Kim M, Kim YK, Nahm BH, Kim JK (2005) Arabidopsis CBF3/DREB1A and ABF3 in transgenic rice increased tolerance to abiotic stress without stunting growth. Plant Physiol 138:341–351
Planchet E, Verdu I, Delahaie J, Cukier C, Girard C, More`re-Le Paven MC, Limami AM (2014) Abscisic acid-induced nitric oxide and proline accumulation in independent pathways under water-deficit stress during seedling establishment in Medicago truncatula. J Exp Bot 65:2161–2170
Qi XH, Xu XW, Lin XJ, Zhang WJ, Chen XH (2012) Identification of differentially expressed genes in cucumber (Cucumis sativus L.) root under waterlogging stress by digital gene expression profile. Genomics 99:160–168
Qiu QS, Barkla BJ, Vera-Estrella R, Zhu JK, Schumaker KS (2003) Na+/H+ exchange activity in the plasma membrane of Arabidopsis. Plant Physiol 132:1041–1052
Saibi W, Feki K, Mahmoud RB, Brini F (2015) Durum wheat dehydrin (DHN-5) confers salinity tolerance to transgenic Arabidopsis plants through the regulation of proline metabolism and ROS scavenging system. Planta 242:1187–1194
Šamajová O, Plíhal O, Al-Yousif M, Hirt H, Šamaj J (2013) Improvement of stress tolerance in plants by genetic manipulation of mitogen-activated protein kinases. Biotechnol Adv 31:118–128
Sarwat M, Ahmad P, Nabi G, Hu X (2013) Ca2+ signals: the versatile decoders of environmental cues. Crit Rev Biotechnol 33:97–109
Shen X, Wang Z, Song X, Xu J, Jiang C, Zhao Y, Ma C, Zhang H (2014) Transcriptomic profiling revealed an important role of cell wall remodeling and ethylene signaling pathway during salt acclimation in Arabidopsis. Plant Mol Biol 86:303–317
Sottosanto JB, Saranga Y, Blumwald E (2007) Impact of AtNHX1, a vacuolar Na+/H+ antiporter, upon gene expression during short- and long-term salt stress in Arabidopsis thaliana. BMC Plant Biol 7:18
Srivastava A, Rai A, Patade V, Suprasanna P (2013) Calcium signaling and its significance in alleviating salt stress in plants. In: Ahmad P, Azooz MM, Prasad MNV (eds) Salt stress in plants. Springer, New York, pp 197–218
Sun X, Xu L, Wang Y, Yu R, Zhu X, Luo X, Gong Y, Wang R, Limera C, Zhang K, Liu L (2015) Identification of novel and salt-responsive miRNAs to explore miRNA-mediated regulatory network of salt stress response in radish (Raphanus sativus L.). BMC Genom 16:197
Takahashi F, Yoshida R, Ichimura K, Mizoguchi T, Seo S, Yonezawa M, Maruyama K, Yamaguchi-Shinozaki K, Shinozaki K (2007) The mitogen-activated protein kinase cascade MKK3-MPK6 is an important part of the jasmonate signal transduction pathway in Arabidopsis. Plant Cell 19:805–818
Türkan I, Demiral T (2009) Recent developments in understanding salinity tolerance. Environ Exp Bot 67:2–9
Wahid A, Close T (2007) Expression of dehydrins under heat stress and their relationship with water relations of sugarcane leaves. Biol Plant 51:104–109
Wang Y-P, Li K-B (2009) Correlation of expression profiles between microRNAs and mRNA targets using NCI-60 data. BMC Genom 10:218
Wang XJ, Zhu SY, Lu YF, Zhao R, Xin Q, Wang XF, Zhang DP (2010) Two coupled components of the mitogen-activated protein kinase cascade MdMPK1 and MdMKK1 from apple function in ABA signal transduction. Plant Cell Physiol 51:754–766
Wu CA, Yang GD, Meng QW, Zheng CC (2004) The cotton GhNHX1 gene encoding a novel putative tonoplast Na+/H+ antiporter plays an important role in salt stress. Plant Cell Physiol 45:600–607
Xu H, Li K, Yang F, Shi Q, Wang X (2010) Overexpression of CsNMAPK in tobacco enhanced seed germination under salt and osmotic stresses. Mol Biol Rep 37:3157–3163
Xu L, Wang L, Gong Y, Dai W, Wang Y, Zhu X, Wen T, Liu L (2012) Genetic linkage map construction and QTL mapping of cadmium accumulation in radish (Raphanus sativus L.). Theor Appl Genet 125:659–670
Xu XB, Pan YY, Wang CL, Ying QC, Song HM, Wang HZ (2014) Overexpression of DnWRKY11 enhanced salt and drought stress tolerance of transgenic tobacco. Biologia 69:994–1000
Xu L, Wang Y, Liu W, Wang J, Zhu X, Zhang K, Yu R, Wang R, Xie Y, Zhang W, Gong Y, Liu L (2015) De novo sequencing of root transcriptome reveals complex cadmium-responsive regulatory networks in radish (Raphanus sativus L.). Plant Sci 236:313–323
Yang Y, He M, Zhu Z, Li S, Xu Y, Zhang C, Singer SD, Wang Y (2012) Identification of the dehydrin gene family from grapevine species and analysis of their responsiveness to various forms of abiotic and biotic stress. BMC Plant Biol 12:140
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L, Wang J (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:W293–W297
Yu Y, Huang W, Chen H, Wu G, Yuan H, Song X, Kang Q, Zhao D, Jiang W, Liu Y (2014) Identification of differentially expressed genes in flax (Linum usitatissimum L.) under saline-alkaline stress by digital gene expression. Gene 549:113–122
Zhang JL, Shi H (2013) Physiological and molecular mechanisms of plant salt tolerance. Photosynth Res 115:1–22
Zhang Y, Chen C, Jin XF, Xiong AS, Peng RH, Hong YH, Yao QH, Chen JM (2009) Expression of a rice DREB1 gene, OsDREB1D, enhances cold and high-salt tolerance in transgenic Arabidopsis. BMB Rep 42:486–492
Zhang S, Song G, Gao J, Li Y, Guo D, Fan Q, Sui X, Chu X, Huang C, Liu J (2014) Transcriptome characterization and differential expression analysis of cold-responsive genes in young spikes of common wheat. J Biotechnol 189:48–57
Zhou S, Chen X, Zhang X, Li Y (2008) Improved salt tolerance in tobacco plants by co-transformation of a betaine synthesis gene BADH and a vacuolar Na+/H+ antiporter gene SeNHX1. Biotechnol Lett 30:369–376
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Acknowledgments
This work was in part supported by grants from the National Natural Science Foundation of China (31501759, 31171956, 31372064), Jiangsu Key Laboratory for Horticultural Crop Genetic Improvement (2014017), Key Technology R&D Program of Jiangsu Province, China (BE2013429) and Jiangsu Agricultural Science and Technology Innovation Fund (CN) [JASTIF,CX (12)2006, CX (13)2007].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by W. Harwood.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S9 The DEGs encoding other significant proteins in salt stress response in radish (XLSX 25 kb)
299_2015_1887_MOESM10_ESM.xlsx
Table S10 The detailed information of association analysis between salt-responsive genes and miRNAs in radish (XLSX 22 kb)
Rights and permissions
About this article
Cite this article
Sun, X., Xu, L., Wang, Y. et al. Transcriptome-based gene expression profiling identifies differentially expressed genes critical for salt stress response in radish (Raphanus sativus L.). Plant Cell Rep 35, 329–346 (2016). https://doi.org/10.1007/s00299-015-1887-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-015-1887-5