Abstract
Understanding the abundance, diversity, and distribution of TEs in genomes is crucial to understand genome structure, function, and evolution. Advances in whole-genome sequencing techniques, as well as in bioinformatics tools, have increased our ability to detect and analyze the transposable element content in genomes. In addition to reference genomes, we now have access to population datasets in which multiple individuals within a species are sequenced. In this chapter, we highlight the recent advances in the study of TE population dynamics focusing on fruit flies and humans, which represent two extremes in terms of TE abundance, diversity, and activity. We review the most recent methodological approaches applied to the study of TE dynamics as well as the new knowledge on host factors involved in the regulation of TE activity. In addition to transposition rates, we also focus on TE deletion rates and on the selective forces that affect the dynamics of TEs in genomes.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
- Long-read sequencing
- Transposition rates
- Self-regulation
- Effective population size
- Adaptation
- Horizontal transfer
1 Transposable Elements Are Abundant and Active Genome Denizens
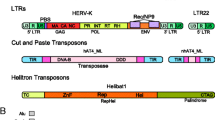
Transposable elements (TEs) are short DNA sequences, typically from a few hundred bp to ~10 kb long, which have the ability to move around in the genome by generating new copies of themselves. In addition to active autonomous elements, genomes also contained nonautonomous elements that can be mobilized by the enzymatic machinery of active TEs from the same family. Additionally, genomes contain TEs that cannot be mobilized anymore due to accumulation of mutations in their sequences [1]. TEs are an ancient, extremely diverse, and exceptionally active component of genomes. TEs have been found in virtually all organisms studied so far including bacteria, archaea, fungi, protists, plants, and animals [2,3,4,5]. The main TE groups, class I and class II, are present in all kingdoms, revealing their persistence over evolutionary time [2]. These two classes of TEs differ in their transposition intermediates: while class I TEs transpose through RNA intermediates, class II TEs transpose directly as DNA. TEs within each class are further classified into (1) different orders, based on their insertion mechanism, structure, and encoded proteins; (2) different superfamilies, based on their replication strategy and on presence and size of target site duplications; and (3) different families, based on sequence conservation [2, 3]. Piegu et al. [1] criticized the current classification system, which accounts for sequence homology, structural features, and target site duplications, because it does not always take into account the evolutionary origins of the TEs [1,2,3]. As a consequence, phylogenetically unrelated classes or subclasses of TEs are grouped [1]. Piegu et al. [1] also suggested that a more inclusive classification that includes prokaryotic and eukaryotic TE classes should be considered. Recently, Arkhipova [6] proposed a TE classification system based on the replicative, integrative, and structural components of TEs, which integrates different aspects of all the existing classification systems [6].
TEs constitute a substantial albeit variable (from ~1% to almost 90%) proportion of genomes [7, 8] (Fig. 1). The identification methods, as well as the sequencing and assembly methods, have an important effect in the TE content estimation [4, 9,10,11]. In some cases, the TE-generated fraction of genomes is likely to be underestimated because methods for detecting TEs in genomic sequences are necessarily biased toward younger and more easily recognizable TEs. Indeed, new tools developed in recent years are able to identify TEs that remained hidden until now [4, 11]. As an example, when the human genome was first sequenced, ~40–45% of the genome was identifiable TEs, 5% was genes and other functional sequences (functional RNAs or regulatory regions), and the remaining ~50% of the genome had no identifiable origin [12]. de Koning et al. [13] using a highly sensitive new strategy named P-cloud found that at least 66–69% of the human genome is identifiable as repetitive sequences, most of them derived from TEs [13]. In Drosophila melanogaster, third-generation sequencing techniques (3GS) have allowed the detection of 37% more TE insertions in chromosome 2L compared to previously available short-read sequencing estimates (see below) [14]. In other Drosophila species such as D. buzzatii, the TE content has also been updated from 6% to 11%, thanks to the recent availability of whole-genome sequences [15].
TE content in the genome of different organisms expressed as percentage of the genome: Homo sapiens (~45% [12], >66% [13]) Mus musculus [143], Saccharomyces cerevisiae [144], Arabidopsis thaliana [145], Pyrococcus furiosus [146], Clostridium difficile [147], Danio rerio [133], Kryptolebias marmoratus [148], Bombyx mori [149], Hypothenemus hampei [150], Drosophila melanogaster (11%, [68], ~20% [69]), Pseudozyma antarctica, and Laccaria bicolor [151]. Zea mays [152] and Fritillaria imperialis [8]. All estimates were obtained with homology-based methods except [13] that uses P-cloud and [69] that uses de novo approaches
As mentioned above, TEs are extremely active genomic denizens that are able to generate mutations of a great diversity of types [16,17,18,19,20,21]. TE-induced mutations range from subtle regulatory mutations to gross genomic rearrangements and often have phenotypic effects of a complexity that is not achievable by point mutations (Fig. 2). Among others, TEs can affect the expression of nearby genes by adding new splice sites, adenylation signals, promoters, or transcription factor binding sites [22,23,24]. TEs can also be targets of epigenetic histone modifications that spread into adjacent genes affecting their expression [25, 26]. In addition to transcriptional changes, TEs have been shown to affect translation regulation when they are transcribed within a mRNA [27,28,29], to contribute to protein-coding regions both at the transcript and at the protein level [30,31,32,33,34,35], and TE-encoded proteins have been domesticated and are part of host genes [17, 36,37,38,39,40]. TE excision can lead to DNA deletions [41], and TE insertion can result in adding DNA through 3′ and, less frequently, through 5′ transduction [42, 43]. Finally, ectopic recombination between TEs causes deletions, duplications, and sequence rearrangements. Two recent studies in the human genome identified 516 chromosome rearrangements potentially generated by LINE-LINE nonallelic homologous recombination and 78 HERV-mediated rearrangements [44, 45]. Both studies used the annotations of LINEs and HERVs in the reference genome and look for evidence of rearrangements induced by these TEs using clinical databases of copy number variants containing information from thousands of patients. In addition to being associated with diseases [24, 46,47,48,49], the number of TE-induced mutations associated with positive effects on fitness-related traits also continues to increase both in humans and in Drosophila [50,51,52,53,54,55,56,57,58,59,60,61,62,63].
Effects of TEs on the host genome. (a) TEs can affect the expression and/or structure of genes. Exons are represented as blue boxes and TEs as green boxes. (1) A TE inserted in the upstream region of a gene can add insulator sequences, transcription factor-binding site (TFBS), or can disrupt an existing promoter gene; (2) A TE inserted in an intron can truncate the mRNA or induce alternative splicing; (3) A TE inserted in the downstream region of a gene can add microRNA binding sites or alter the polyadenylation site; (4) A TE inserted in the exon of a gene can lead to exonization of the TE or to transcript truncation; (5) the whole domain of a TE protein could insert in the coding region of a gene generating a chimeric gene with host and TE domains [5, 21]. In addition to these changes that depend on where the TE is inserted and on the sequences that the TE is adding, TEs can also alter the posttranslational modifications of histones. (b) TEs could also induce translation repression by generating secondary structure in the 3′ UTR of genes that leads to changes in the localization of the mRNA. This secondary structure could bind to one of the protein components of paraspeckle (P54 nrb) and translocate to paraspeckle, a group of subnuclear bodies, avoiding moving out of the nucleus. However, the same secondary structure could bind to the dsRNA-binding protein Staufen 1 (STAU1) and in this case translocate to cytoplasm. Once in the cytoplasm, the secondary structure could bind to STAU1 again allowing translation, but under some situations mRNA could bind to the ds-RNA- dependent protein kinase (PKR) repressing translation [23]. (c) Ectopic recombination between TE copies (green boxes with yellow arrows) in the same orientation can lead to deletions when recombination takes place between copies located on the same chromatid (1) or deletions and duplications when recombination takes place between copies in different chromosomes (2) (recombination between two nonhomologous chromosomes should lead to a translocation). Ectopic recombination between TE copies in opposite orientation leads to inversion of the DNA between the two TEs (3)
Overall, recent advances in sequencing technologies and in TE detection methods showed that, as expected, the TE content is higher than previously estimated. These new data also provided further evidence for the impact of TEs in genome function and genome structure. Thus, it is still indisputable that a thorough understanding of TE population dynamics is essential for the understanding of the eukaryotic genome structure, function, and evolution.
2 Drosophila and Humans: Two Extremes in TE Diversity and Population Dynamics
Much of the detailed information on TE evolution still comes from two species with the best-studied genomes: fruit flies (D. melanogaster) and humans. Fortunately, these two genomes represent two extremes in terms of TE diversity and population dynamics and thus give a reasonably diverse picture of the TE evolution and dynamics. For the rest of this chapter, we focus primarily on these two genomes and will highlight the similarities and differences observed between them.
As mentioned above, the human reference genome has millions of TE copies, with 66–69% of the genome mostly derived from TE sequences [13]. Two human retrotransposable element (class I) families, LINE1 (L1, long interspersed nuclear element 1) and Alu, account for 60% of all interspersed repeat sequences. The vast majority of the TEs in the human genome are fixed, and most families are inactive. However, some elements of the main families of human endogenous retrovirus (HERV-K) and LINE1 elements show autonomous transposition. Meanwhile, elements of Alu and the hybrid SVA elements formed by SINEs (short interspersed nuclear elements), VNTRs (variable number tandem repeat), and Alus show nonautonomous activity [64,65,66].
In contrast, the fruit fly D. melanogaster reference genome contains only thousands of individual TE copies (5416 TE copies in FlyBase R6.04) accounting for only ~5.5% of the euchromatin [67]. If the missing percentage of TEs detected in chromosome 2L is similar in other chromosomes, the euchromatin TE content might be higher (~ 8.7%) [14]. If heterochromatin is also included, TEs account for 11–20% of the D. melanogaster genome [68, 69]. D. melanogaster TEs belong to approximately 100 diverse families of both class I and class II elements [69, 70]. Each family consists of 1–304 copies with no dominant family corresponding to the majority of TEs. The only exception is INE-1 family that contains ~2000 copies and has been inactive for the past ~3–4.6 million years [71,72,73]. The majority of TE families are considered to be active in Drosophila: individual TE copies are generally polymorphic in the population and show a high sequence similarity [69, 70, 74, 75]. Indeed, there is experimental evidence showing that Gypsy and ZAM elements are active [76, 77]. Besides, there is indirect evidence for the activity of 24 D. melanogaster superfamilies based on a whole-genome sequencing experiment of mutation accumulation lines [75] (Table 1).
Why do these two genomes differ so profoundly in content, diversity, and activity of TEs? The answer must lie in different aspects of TE population dynamics within genomes and forces that lead to varying rates of TE family birth and extinction. In the rest of this review, we focus on the state of knowledge of different aspects of TE population dynamics and discuss aspects of TE family evolution. Specifically, we focus on rates of TE transposition, fixation, or loss in human and D. melanogaster populations due to stochastic forces and natural selection for or against TE insertions and forces that affect coexistence of multiple TE families and the standing diversity of TE types (Fig. 3).
Factors that influence the population and evolutionary dynamics of TEs. Our understanding of TE population and evolutionary dynamics is still incomplete. The different factors that affect TE population and evolutionary dynamics are interrelated, new factors have been identified in recent years, and future research is still likely to reveal existence of additional factors
3 Methodology Used to Study TE Population Dynamics
TE dynamics continues to be studied using three main approaches: mathematical modeling, computer simulations, and the analysis of empirical data. Often a combination of these approaches is used to better understand TE abundance, diversity, and distribution (Table 2). Le Rouzic et al. [78] applied the statistical framework originally developed to infer speciation and extinction dynamics in species phylogenies to reconstruct the evolutionary history of TEs [78]. The model allows to estimate and to interpret the pattern of transposition activity that results in different TE copy number distributions [78]. The authors also performed computer simulations to provide reference dynamics that aid in the interpretation of the results obtained (Table 2).
Traditionally, mathematical models considered the relationship between the host and a homogenous group of active TEs. However, the TE content of any genome is a mixed of autonomous and nonautonomous insertions. Xue and Goldenfeld [79] proposed a mathematical model that considers the relationship between nonautonomous and autonomous TEs as a predator-prey dynamic. Unlike previous models that also use the analogy to ecological models, Xue and Goldenfeld model takes into account the molecular level interactions between transposable elements and the small copy number of the active transposons. The model predicts oscillations in the number of TEs in a time scale much longer than the cell replication time, suggesting that the genome stores the predator-prey state during successive generations [79].
TE dynamics have also been analyzed in variable environments [80, 81] (Table 2). Gogolesky et al. [81] proposed a stochastic computational model to analyze the dynamics of active TEs in genomes of sexual diploid organisms under environmental stress. They based their model in the Fisher geometrical model of fitness landscapes. Overall, the authors conclude that the presence of inactive copies of TEs is necessary for the transposition-selection equilibrium of autonomous copies and that the mutator capacity of TEs might be important when host populations face rapid environmental changes [81].
Other recently developed methods analyzed the influence of the mating system in TE dynamics, different modes of selection, or applied branching models for studying the propagation of particular TE classes [82,83,84] (Table 2).
In addition to mathematical modeling and simulations, multiple computational tools have been developed to analyze TEs in sequenced genomes in the last 5 years. While some of these tools aimed at assessing the global abundance and diversity of TEs in the genome, such as dnaPipeTE, or to annotate TEs in assembled genomes, such as REPET, most of them are focused on discovering and/or genotyping individual copies of TEs in the genome using next-generation sequencing (NGS) data [11, 64, 85,86,87,88,89,90]. The diversity of methods available makes it difficult to choose the most appropriate one for the analyses of a given genome. To try to overcome this limitation, Nelson et al. [91] developed an integrated pipeline named McClintock that incorporates six complementary TE detection methods. McClintock generates standardized output for the different TE detection methods, thus facilitating the comparison of the results obtained with the different pipelines, as well as facilitating their installation and use [91]. This and other studies that compared the performance of several tools arrived to the same conclusion: several computational tools should be combined to increase the accuracy of TE analysis [64, 86, 91].
The availability of third-generation sequencing techniques (3GS) should help improve the detection and genotyping of TE insertions. Although 3GS was developed before 2010 [92], it has only been in the last few years when this technique has started to be used [14, 93]. Chakraborty et al. [14] reported the assembly of a D. melanogaster genome from a Zimbabwe strain using long-read single molecule real-time sequencing with 147X coverage. Among several novel structural variants described, they identified 37% additional TE insertions in the 2L chromosome compared with a previous study that used 70X coverage of short reads [14, 94]. 3GS technologies have also been applied to the sequencing of human genomes, although a detailed analysis of TE content based on long-read data has not been performed yet [95,96,97].
Recently, Disdero and Filée [98] introduced the first tool that uses long-read sequences to identify TE insertions in the D. melanogaster genome: LoRTE [98]. The authors argue that available software based on short reads fail to correctly identify TEs that are present in highly repetitive regions of the genome, while long-read technologies should allow us to identify all TEs in a given genome. LoRTE, developed in Python, verifies presence and/or absence of previously annotated TEs and can also detect new insertions not previously annotated in the reference genome. LoRTE is able to work with low-coverage sequences (<10X) providing an efficient accurate TE annotation in a cost-effective manner [98].
4 Rates of Transposition
4.1 Empirical Estimates of the Rates of Transposition in Drosophila and Humans
Transposition rates in D. melanogaster have been traditionally estimated empirically by in situ hybridization and by using PCR approaches. The activation of TEs following intra- and interspecific hybridization has been studied in different Drosophila species [99,100,101]. For example, Vela et al. [100] estimated transpositions rates in D. buzzatii-D. koepferae interspecific hybrid flies by in situ hybridization [100]. They found that hybrids showed at least one order of magnitude higher transposition rates than parental lines for at least three TE families [100]. Robillard et al. [102] estimated transposition rates by qPCR in an experimental evolution study in which a TE insertion was introduced in a strain lacking insertions from that particular family [102]. In the first generations after the introduction of the TE insertion, the transposition rate was 0.33–0.45 per copy per generation, while in the following generations, transposition rates were reduced at least one order of magnitude per copy per generation. These values represent the first steps in the invasion of a TE in a genome that is faster than the rate of transposition when measured in natural populations [102].
In the first edition of this chapter [103], we anticipated that NGS would allow studying transposition rates in a deeper and more accurate way. Indeed, recent studies have taken advantage of NGS data to estimate transposition rates in D. melanogaster. Rahman et al. [89] estimated using NGS data the transposition rate in the reference strain by comparing two available genomes that were sequenced with ~15 years difference. The average transposition rate for TEs belonging to different families was 7 × 10−5, which is on the same order of magnitude as the previously reported rates (~10−4–10−5). Furthermore, they confirmed the prediction of increased transposition rate in inbred lines: they estimated a higher average number of TE insertions in lab strains inbred for more generations compared with strains inbred for a smaller number of generations [89]. Adrion et al. [75] estimated spontaneous insertion and deletion rates in D. melanogaster mutation accumulation lines [75]. The authors identified 24 active superfamilies and estimated genome-wide insertion rates to be higher than deletion rates: 2.11 × 10−9 vs. 1.37 × 10−10 per site per generation, respectively. Superfamily-specific rates of insertion varied from 0 to 5.13 × 10−3 insertions per copy per generation and were within the range of previously estimated rates [75] (Table 1).
In humans, previous studies estimated the transposition rate as in 1 in 95 to 1 in 250 births for L1, 1 in 20 births for Alu insertions, and 1 in 916 births for SVA retrotransposons [104,105,106,107]. Although there are several recent studies that estimate transposition rate in humans using NGS data, they all focused on somatic transposition in the brain or in tumor samples [47, 48, 90].
4.2 Transposition Control Mechanisms
Understanding the mechanisms controlling the transposition of TEs is central to our understanding of TE dynamics. Many different mechanisms of TE regulation have been described [43, 108, 109]. In this section, we will highlight recent advances in both TE self-regulation and regulation by host factors.
4.2.1 TE Self-Regulation
Self-regulation of transposition was first described in prokaryotes and soon after in TEs involved in hybrid dysgenesis in Drosophila [110]. Recent studies have cast some doubt on one of the self-regulation mechanisms described: transposase overproduction inhibition. The transposase overproduction inhibition mechanism regulates the transposition of IS630-Tc1-mariner piggyBac and hobo-AC-Tam (hAT) superfamilies [111, 112]. However, several studies reported contradictory results suggesting that transposase inhibition by overproduction does not always happen [113]. Bire et al. [113] suggested that some works failed to detect transposase inhibition because cellular cofactors are necessary to execute this regulation system, and as such it can only be detected in in vivo experiments [113]. However, Woodard et al. [114] showed that aggregation of transposase proteins produces filamentous structures (rodlets) in the nucleus in a host independent manner [114]. The authors further showed that a decline in transposition occurs after transposase concentrations are high enough for filamentous structures to be visible [114]. Thus, it is still not clear why some in vitro experiments failed to detect transposase overproduction inhibition [114].
4.2.2 Regulation by Host Factors
Small RNAs, such as small-interfering RNAs (siRNAs) and piwi-interacting RNAs (piRNAs), are well-known to play an essential role in silencing TEs and preventing transposition. Several recent reviews highlight the monumental progress in this field [115,116,117,118,119]. In addition to posttranscriptional regulation of TEs, small RNAs are involved in transcriptional regulation as well. In mouse, piRNAs are required for de novo methylation and silencing of TEs [120]. In Drosophila, Piwi proteins repress transcription and correlate with an increase in repressive chromatin marks at loci targeted by piRNAs [121].
While the role of siRNAs and piRNAs has been established for several years, a role of micro RNAs (miRs) in suppressing the mobility of retrotransposons was only recently described [122]. The authors showed that mir-128 binds to L1 RNA and represses its integration in humans [122].
New studies have also provided evidence for the role in TE repression of proteins previously known for their roles in other cellular processes such as interferon-stimulated proteins, the tumor suppressor p53, and the longevity regulating protein SIRT6. Several interferon-stimulated genes, such as the Moloney leukemia virus 10 (MOV10), the zinc-finger antiviral protein (ZAP), and the 3′ repair exonuclease 1 (TREX1), which are associated with virus response, have been recently involved in the inhibition of L1 activity [66, 123]. Recently, it has also been shown that the p53 transcription factor, which is involved in stress response networks and acts to restrict oncogenesis, also restricts retrotransposon activity in zebra fish, flies, and humans [124]. The authors showed that p53 interacts with components of the piwi-interacting RNA to suppress retrotransposition [124]. Finally, the longevity regulating protein SIRT6 is also involved in retrotransposon repression by coordinating their packaging into transcriptionally repressive heterochromatin. SIRT6 binds to the 5′ UTR region of retrotransposons and mono-ADP ribosylates the Krüppel-associated protein 1 (KAP1) facilitating the interaction of KAP1 with the heterochromatin protein 1α (HP1α) leading to chromatin compaction [125].
5 Rate of Fixation and Frequency Distribution
5.1 Natural Selection Against TE Insertions
Natural selection and stochastic processes influence both the rate of fixation and the frequency distribution of TEs in populations. The efficiency of selection depends on the effective population size, which largely differs between Drosophila and humans: >108 and ~104, respectively [126, 127]. Thus, while in Drosophila the high efficiency of selection should led to the removal of slightly deleterious TE insertions, in humans, these insertions may accumulate in the genome. Indeed most of the TE sequences in the human genome are remnants of ancient insertions [12].
A review by Barrón et al. [128] explored the latest insights on the nature of selection acting against the deleterious effects of TEs in D. melanogaster populations [128]. More recently, Kofler et al. [129] analyzed intraspecific TE dynamics between D. melanogaster and D. simulans populations to shed light on the long-term evolution of TEs [129]. They confirmed that most of the TEs are present at low frequencies in D. melanogaster and showed that the same pattern is present in D. simulans. Based on computer simulations showing that 50% of the TE families have temporally heterogeneous transposition rates, and on the differences in TE composition between populations of the same species, the authors suggested that TE activity has recently increased in the two species. They proposed that the demographic history of both species, with a recent colonization of different environments, could be the cause of the high TE activity detected [129].
In humans, a recent study took advantage of the 1000 Genome Project data that reports 16,192 polymorphic TEs to perform the most complete TE dynamics analysis to date [130]. Most of the polymorphic TEs were found to be present at very low frequencies: >93% of TEs showed <5% allele frequency in 26 human populations. These results confirm that overall polymorphic TE insertions are deleterious in humans as was previously suggested with smaller family-specific datasets [131].
5.2 TE-Induced Adaptations
Several recent reviews have compiled results that showcase the adaptive role of TEs [19, 24, 50, 59, 128]. We would like to highlight the recent discovery of a TE in a fish-like marine chordate that encodes RAG-like proteins with endonuclease-transposase activity [39]. This discovery provides evidence that supports the TE origin hypothesis for the adaptive immune system in jawed vertebrates [39]. Two other recent publications provide experimental evidence for a role of TEs as providers of functional transcription factor binding sites (TFBS) involved in immune response and in cell pluripotency [50, 132]. A recent study linked ERV elements in humans with the interferon response pathway [50]. The authors showed that ERVs carrying enhancers have been co-opted to activate different genes involve in inflammatory response activated by interferon. This example shows how the exaptation of one family of TEs could shape a transcriptional network to activate different genes with one trigger system [50]. Sundaram et al. [132] reported mouse-specific TEs that contain multiple transcription factor binding sites for pluripotency transcription factors. The majority of the TEs were experimentally shown to exhibit enhancer activity in mouse embryonic stem cells including an in silico reconstructed ancestral TE. This latter result suggests that ancestral TEs already had transcriptional regulatory sites [132].
In Drosophila, the adaptive role of several TEs has also been identified. Most of the TEs characterized so far are involved in stress response: viral infection and xenobiotics (Doc1420, [60, 61]), oxidative stress (FBti0018880, [53]), xenobiotic stress (Accord, [62, 63], and FBti0019627, [52]), cold stress (FBti0019985, [55]), and heavy metal stress (FBti0019170, [56]), while FBti0019386 insertion was associated with faster developmental time [54]. Some of these adaptive insertions have been shown to affect gene expression through different molecular mechanisms, such as affecting the polyadenylation site choice [52], and adding TFBS [53], while others have been associated with gene duplication [60, 62].
6 Rate of Loss
A recent study estimated genome-wide and superfamily-specific TE deletion rates in D. melanogaster inbred lines [75]. The authors found that most of the deletions involved retrotransposon elements suggesting that the deletions were due to ectopic recombination instead of excision. Deletion rates were smaller than insertion rates estimated in the same inbred lines [75].
In vertebrates, lineage-specific differences in TE deletion rates have been reported [133]. A possible explanation for this observation is that the success of some families results in a competition for the genome resources leading to the elimination of other TE families [133].
In addition to TE deletion rates, DNA loss rates should also be considered. In the human linage, estimates of DNA loss are smaller than estimates of DNA gain, 650 Mb vs. 815 Mb [134], while in D. melanogaster, the rate of DNA loss is higher than the rate of DNA gain [135,136,137].
7 Horizontal Transfer of TE Insertions
In addition to parent to offspring transmission, TEs can also be horizontally transferred [138,139,140,141]. By combining simulation and analytical approaches, Groth and Blumenstiel [142] suggested that exposure rate to new TE families through horizontal transfer can be an important determinant of TE genomic content when the effects of drift in a population are weak [142]. Thus, larger populations are expected to carry a higher TE content if population exposure rate is proportional to population size [142]. So far, most of the evidence for TE horizontal transfer comes from closely related and geographically close species [140]. There are several examples of horizontal transfer of TEs in Drosophila species, while so far horizontal transfer of TEs has not been described in humans [138].
8 Conclusion
Recent years have seen an increase in the number of reference genome sequences available as well as of population genome datasets. The availability of all these genome sequences and the development of new bioinformatics tools have allowed us to update our previous estimates of genomic TE content that have increased both in humans and in D. melanogaster. These data has also allowed us to gather more evidence for the functional impact, both detrimental and beneficial, of TE insertions. Thus, it is still indisputable that understanding TE population dynamics is essential to understand genome structure, genome function, and genome evolution.
New methods developed to analyze the dynamics of TEs in populations have shed light on the interplay between autonomous and nonautonomous TE copies, TE invasion dynamics, and how the mating system influences the dynamics of TEs in genomes. We have also considerably advanced our knowledge on the host factors that regulate TE activity as well as in the genome features that influence TE dynamics (Fig. 3). Finally, differences in effective population sizes that affect the efficiency of selection against new TE insertions and differences in the rates of TE loss between humans and D. melanogaster can still be considered two important factors that contribute to the different abundance, diversity, and activity of TEs in this two species [103].
9 Questions
-
How differences in the rate of DNA loss can affect the evolutionary dynamics of TEs?
-
Why host regulation of transposition is relevant for TE dynamics?
-
Which is the most important factor explaining the differences in TE content, diversity, and activity between humans and Drosophila?
-
Have the next-generation sequencing (NGS) technologies allowed us to identify all the TEs in a given genome?
-
How does the interaction between active and inactive copies of TEs affect TE dynamics?
References
Piegu B, Bire S, Arensburger P, Bigot Y (2015) A survey of transposable element classification systems--a call for a fundamental update to meet the challenge of their diversity and complexity. Mol Phylogenet Evol 86:90–109
Wicker T, Sabot F, Hua-Van A, Bennetzen JL, Capy P, Chalhoub B, Flavell A, Leroy P, Morgante M, Panaud O, Paux E, SanMiguel P, Schulman AH (2007) A unified classification system for eukaryotic transposable elements. Nat Rev Genet 8(12):973–982
Kapitonov VV, Jurka J (2008) A universal classification of eukaryotic transposable elements implemented in Repbase. Nat Rev Genet 9(5):411–412. author reply 414
Bao W, Kojima KK, Kohany O (2015) Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob DNA 6:11
Hua-Van A, Le Rouzic A, Boutin TS, Filee J, Capy P (2011) The struggle for life of the genome’s selfish architects. Biol Direct 6:19
Arkhipova IR (2017) Using bioinformatic and phylogenetic approaches to classify transposable elements and understand their complex evolutionary histories. Mob DNA 8:19
Touchon M, Rocha EP (2007) Causes of insertion sequences abundance in prokaryotic genomes. Mol Biol Evol 24(4):969–981
Ambrozova K, Mandakova T, Bures P, Neumann P, Leitch IJ, Koblizkova A, Macas J, Lysak MA (2011) Diverse retrotransposon families and an AT-rich satellite DNA revealed in giant genomes of Fritillaria lilies. Ann Bot 107(2):255–268
Chaisson MJ, Wilson RK, Eichler EE (2015) Genetic variation and the de novo assembly of human genomes. Nat Rev Genet 16(11):627–640
Treangen TJ, Salzberg SL (2011) Repetitive DNA and next-generation sequencing: computational challenges and solutions. Nat Rev Genet 13(1):36–46
Flutre T, Duprat E, Feuillet C, Quesneville H (2011) Considering transposable element diversification in de novo annotation approaches. PLoS One 6(1):e16526
Human Genome Sequencing Consortium (2001) Initial sequencing and analysis of the human genome. Nature 409(6822):860–921
de Koning AP, Gu W, Castoe TA, Batzer MA, Pollock DD (2011) Repetitive elements may comprise over two-thirds of the human genome. PLoS Genet 7(12):e1002384
Chakraborty M, VanKuren NW, Zhao R, Zhang X, Kalsow S, Emerson JJ (2018) Hidden genetic variation shapes the structure of functional elements in Drosophila. Nat Genet 50(1):20–25
Rius N, Guillen Y, Delprat A, Kapusta A, Feschotte C, Ruiz A (2016) Exploration of the Drosophila buzzatii transposable element content suggests underestimation of repeats in Drosophila genomes. BMC Genomics 17:344
Kidwell MG, Lisch DR (2000) Transposable elements and host genome evolution. Trends Ecol Evol 15(3):95–99
Feschotte C, Pritham EJ (2007) DNA transposons and the evolution of eukaryotic genomes. Annu Rev Genet 41:331–368
Cowley M, Oakey RJ (2013) Transposable elements re-wire and fine-tune the transcriptome. PLoS Genet 9(1):e1003234
Casacuberta E, Gonzalez J (2013) The impact of transposable elements in environmental adaptation. Mol Ecol 22(6):1503–1517
Belyayev A (2014) Bursts of transposable elements as an evolutionary driving force. J Evol Biol 27(12):2573–2584
Rebollo R, Romanish MT, Mager DL (2012) Transposable elements: an abundant and natural source of regulatory sequences for host genes. Annu Rev Genet 46:21–42
Feschotte C (2008) Transposable elements and the evolution of regulatory networks. Nat Rev Genet 9(5):397–405
Elbarbary RA, Lucas BA, Maquat LE (2016) Retrotransposons as regulators of gene expression. Science 351(6274):aac7247
Chuong EB, Elde NC, Feschotte C (2017) Regulatory activities of transposable elements: from conflicts to benefits. Nat Rev Genet 18(2):71–86
Lippman Z, Gendrel AV, Black M, Vaughn MW, Dedhia N, McCombie WR, Lavine K, Mittal V, May B, Kasschau KD, Carrington JC, Doerge RW, Colot V, Martienssen R (2004) Role of transposable elements in heterochromatin and epigenetic control. Nature 430(6998):471–476
Sentmanat MF, Elgin SC (2012) Ectopic assembly of heterochromatin in Drosophila melanogaster triggered by transposable elements. Proc Natl Acad Sci U S A 109(35):14104–14109
Capshew CR, Dusenbury KL, Hundley HA (2012) Inverted Alu dsRNA structures do not affect localization but can alter translation efficiency of human mRNAs independent of RNA editing. Nucleic Acids Res 40(17):8637–8645
Fitzpatrick T, Huang S (2012) 3′-UTR-located inverted Alu repeats facilitate mRNA translational repression and stress granule accumulation. Nucleus 3(4):359–369
Liu WM, Chu WM, Choudary PV, Schmid CW (1995) Cell stress and translational inhibitors transiently increase the abundance of mammalian SINE transcripts. Nucleic Acids Res 23(10):1758–1765
Makalowski W, Mitchell GA, Labuda D (1994) Alu sequences in the coding regions of mRNA: a source of protein variability. Trends Genet 10(6):188–193
Gotea V, Makalowski W (2006) Do transposable elements really contribute to proteomes? Trends Genet 22(5):260–267
Wu M, Li L, Sun Z (2007) Transposable element fragments in protein-coding regions and their contributions to human functional proteins. Gene 401(1-2):165–171
Charng YC, Liu LD (2013) The extent of Ds1 transposon to enrich transcriptomes and proteomes by exonization. Bot Stud 54(1):14
Mandal AK, Pandey R, Jha V, Mukerji M (2013) Transcriptome-wide expansion of non-coding regulatory switches: evidence from co-occurrence of Alu exonization, antisense and editing. Nucleic Acids Res 41(4):2121–2137
Hoen DR, Bureau TE (2015) Discovery of novel genes derived from transposable elements using integrative genomic analysis. Mol Biol Evol 32(6):1487–1506
Huda A, Bushel PR (2013) Widespread exonization of transposable elements in human coding sequences is associated with epigenetic regulation of transcription. Transcr Open Access 1(1)
Abascal F, Tress ML, Valencia A (2015) Alternative splicing and co-option of transposable elements: the case of TMPO/LAP2alpha and ZNF451 in mammals. Bioinformatics 31(14):2257–2261
Lin L, Jiang P, Park JW, Wang J, Lu ZX, Lam MP, Ping P, Xing Y (2016) The contribution of Alu exons to the human proteome. Genome Biol 17:15
Huang S, Tao X, Yuan S, Zhang Y, Li P, Beilinson HA, Zhang Y, Yu W, Pontarotti P, Escriva H, Le Petillon Y, Liu X, Chen S, Schatz DG, Xu A (2016) Discovery of an active RAG transposon illuminates the origins of V(D)J recombination. Cell 166(1):102–114
Shaheen M, Williamson E, Nickoloff J, Lee SH, Hromas R (2010) Metnase/SETMAR: a domesticated primate transposase that enhances DNA repair, replication, and decatenation. Genetica 138(5):559–566
Nordborg M, Walbot V (1995) Estimating allelic diversity generated by excision of different transposon types. Theor Appl Genet 90(6):771–775
Moran JV, DeBerardinis RJ, Kazazian HH Jr (1999) Exon shuffling by L1 retrotransposition. Science 283(5407):1530–1534
Goodier JL, Kazazian HH Jr (2008) Retrotransposons revisited: the restraint and rehabilitation of parasites. Cell 135(1):23–35
Campbell IM, Gambin T, Dittwald P, Beck CR, Shuvarikov A, Hixson P, Patel A, Gambin A, Shaw CA, Rosenfeld JA, Stankiewicz P (2014) Human endogenous retroviral elements promote genome instability via non-allelic homologous recombination. BMC Biol 12:74
Startek M, Szafranski P, Gambin T, Campbell IM, Hixson P, Shaw CA, Stankiewicz P, Gambin A (2015) Genome-wide analyses of LINE-LINE-mediated nonallelic homologous recombination. Nucleic Acids Res 43(4):2188–2198
Hancks DC, Kazazian HH Jr (2012) Active human retrotransposons: variation and disease. Curr Opin Genet Dev 22(3):191–203
Helman E, Lawrence MS, Stewart C, Sougnez C, Getz G, Meyerson M (2014) Somatic retrotransposition in human cancer revealed by whole-genome and exome sequencing. Genome Res 24(7):1053–1063
Evrony GD, Lee E, Park PJ, Walsh CA (2016) Resolving rates of mutation in the brain using single-neuron genomics. Elife 5:e12966
Payer LM, Steranka JP, Yang WR, Kryatova M, Medabalimi S, Ardeljan D, Liu C, Boeke JD, Avramopoulos D, Burns KH (2017) Structural variants caused by Alu insertions are associated with risks for many human diseases. Proc Natl Acad Sci U S A 114(20):E3984–E3992
Chuong EB, Elde NC, Feschotte C (2016) Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 351(6277):1083–1087
Gonzalez J, Lenkov K, Lipatov M, Macpherson JM, Petrov DA (2008) High rate of recent transposable element-induced adaptation in Drosophila melanogaster. PLoS Biol 6(10):e251
Mateo L, Ullastres A, Gonzalez J (2014) A transposable element insertion confers xenobiotic resistance in Drosophila. PLoS Genet 10(8):e1004560
Guio L, Barron MG, Gonzalez J (2014) The transposable element Bari-Jheh mediates oxidative stress response in Drosophila. Mol Ecol 23(8):2020–2030
Ullastres A, Petit N, Gonzalez J (2015) Exploring the phenotypic space and the evolutionary history of a natural mutation in Drosophila melanogaster. Mol Biol Evol 32(7):1800–1814
Merenciano M, Ullastres A, de Cara MA, Barron MG, Gonzalez J (2016) Multiple independent retroelement insertions in the promoter of a stress response gene have variable molecular and functional effects in drosophila. PLoS Genet 12(8):e1006249
Le Manh H, Guio L, Merenciano M, Rovira Q, Barron MG, Gonzalez J (2017) Natural and laboratory mutations in kuzbanian are associated with zinc stress phenotypes in Drosophila melanogaster. Sci Rep 7:42663
McCue AD, Nuthikattu S, Reeder SH, Slotkin RK (2012) Gene expression and stress response mediated by the epigenetic regulation of a transposable element small RNA. PLoS Genet 8(2):e1002474
Schrader L, Kim JW, Ence D, Zimin A, Klein A, Wyschetzki K, Weichselgartner T, Kemena C, Stokl J, Schultner E, Wurm Y, Smith CD, Yandell M, Heinze J, Gadau J, Oettler J (2014) Transposable element islands facilitate adaptation to novel environments in an invasive species. Nat Commun 5:5495
Shapiro JA (2017) Exploring the read-write genome: mobile DNA and mammalian adaptation. Crit Rev Biochem Mol Biol 52(1):1–17
Magwire MM, Bayer F, Webster CL, Cao C, Jiggins FM (2011) Successive increases in the resistance of Drosophila to viral infection through a transposon insertion followed by a duplication. PLoS Genet 7(10):e1002337
Aminetzach YT, Macpherson JM, Petrov DA (2005) Pesticide resistance via transposition-mediated adaptive gene truncation in Drosophila. Science 309(5735):764–767
Schmidt JM, Good RT, Appleton B, Sherrard J, Raymant GC, Bogwitz MR, Martin J, Daborn PJ, Goddard ME, Batterham P, Robin C (2010) Copy number variation and transposable elements feature in recent, ongoing adaptation at the Cyp6g1 locus. PLoS Genet 6(6):e1000998
Daborn PJ, Yen JL, Bogwitz MR, Le Goff G, Feil E, Jeffers S, Tijet N, Perry T, Heckel D, Batterham P, Feyereisen R, Wilson TG, ffrench-Constant RH (2002) A single p450 allele associated with insecticide resistance in Drosophila. Science 297(5590):2253–2256
Rishishwar L, Marino-Ramirez L, Jordan IK (2017) Benchmarking computational tools for polymorphic transposable element detection. Brief Bioinform 18:908
Rishishwar L, Wang L, Clayton EA, Marino-Ramirez L, McDonald JF, Jordan IK (2017) Population and clinical genetics of human transposable elements in the (post) genomic era. Mob Genet Elements 7(1):1–20
Goodier JL (2016) Restricting retrotransposons: a review. Mob DNA 7:16
Bergman CM, Quesneville H, Anxolabehere D, Ashburner M (2006) Recurrent insertion and duplication generate networks of transposable element sequences in the Drosophila melanogaster genome. Genome Biol 7(11):R112
Sessegolo C, Burlet N, Haudry A (2016) Strong phylogenetic inertia on genome size and transposable element content among 26 species of flies. Biol Lett 12(8):20160407
Quesneville H, Bergman CM, Andrieu O, Autard D, Nouaud D, Ashburner M, Anxolabehere D (2005) Combined evidence annotation of transposable elements in genome sequences. PLoS Comput Biol 1(2):166–175
Kaminker JS, Bergman CM, Kronmiller B, Carlson J, Svirskas R, Patel S, Frise E, Wheeler DA, Lewis SE, Rubin GM, Ashburner M, Celniker SE (2002) The transposable elements of the Drosophila melanogaster euchromatin: a genomics perspective. Genome Biol 3(12):RESEARCH0084
Kapitonov VV, Jurka J (2003) Molecular paleontology of transposable elements in the Drosophila melanogaster genome. Proc Natl Acad Sci U S A 100(11):6569–6574
Singh ND, Petrov DA (2004) Rapid sequence turnover at an intergenic locus in Drosophila. Mol Biol Evol 21(4):670–680
Yang HP, Barbash DA (2008) Abundant and species-specific DINE-1 transposable elements in 12 Drosophila genomes. Genome Biol 9(2):R39
Petrov DA, Fiston-Lavier AS, Lipatov M, Lenkov K, Gonzalez J (2011) Population genomics of transposable elements in Drosophila melanogaster. Mol Biol Evol 28(5):1633–1644
Adrion JR, Song MJ, Schrider DR, Hahn MW, Schaack S (2017) Genome-wide estimates of transposable element insertion and deletion rates in Drosophila melanogaster. Genome Biol Evol 9(5):1329–1340
Kim A, Terzian C, Santamaria P, Pelisson A, Purd’homme N, Bucheton A (1994) Retroviruses in invertebrates: the gypsy retrotransposon is apparently an infectious retrovirus of Drosophila melanogaster. Proc Natl Acad Sci U S A 91(4):1285–1289
Leblanc P, Desset S, Giorgi F, Taddei AR, Fausto AM, Mazzini M, Dastugue B, Vaury C (2000) Life cycle of an endogenous retrovirus, ZAM, in Drosophila melanogaster. J Virol 74(22):10658–10669
Le Rouzic A, Payen T, Hua-Van A (2013) Reconstructing the evolutionary history of transposable elements. Genome Biol Evol 5(1):77–86
Xue C, Goldenfeld N (2016) Stochastic predator-prey dynamics of transposons in the human genome. Phys Rev Lett 117(20):208101
Startek M, Le Rouzic A, Capy P, Grzebelus D, Gambin A (2013) Genomic parasites or symbionts? Modeling the effects of environmental pressure on transposition activity in asexual populations. Theor Popul Biol 90:145–151
Gogolesky K, Startek A, Gambin A, Le Rouzic A (2016) Modelling the proliferation of transposable elements in populations under environmental stress. arXiv. arXiv:1611.04812
Boutin TS, Le Rouzic A, Capy P (2012) How does selfing affect the dynamics of selfish transposable elements? Mob DNA 3:5
Kijima TE, Innan H (2013) Population genetics and molecular evolution of DNA sequences in transposable elements. I. A simulation framework. Genetics 195(3):957–967
Moulin S, Seux N, Chretien S, Guyeux C, Lerat E (2017) Simulation-based estimation of branching models for LTR retrotransposons. Bioinformatics 33(3):320–326
Goubert C, Modolo L, Vieira C, ValienteMoro C, Mavingui P, Boulesteix M (2015) De novo assembly and annotation of the Asian tiger mosquito (Aedes albopictus) repeatome with dnaPipeTE from raw genomic reads and comparative analysis with the yellow fever mosquito (Aedes aegypti). Genome Biol Evol 7(4):1192–1205
Ewing AD (2015) Transposable element detection from whole genome sequence data. Mob DNA 6:24
Kofler R, Gomez-Sanchez D, Schlotterer C (2016) PoPoolationTE2: comparative population genomics of transposable elements using pool-seq. Mol Biol Evol 33(10):2759–2764
Fiston-Lavier AS, Barron MG, Petrov DA, Gonzalez J (2015) T-lex2: genotyping, frequency estimation and re-annotation of transposable elements using single or pooled next-generation sequencing data. Nucleic Acids Res 43(4):e22
Rahman R, Chirn GW, Kanodia A, Sytnikova YA, Brembs B, Bergman CM, Lau NC (2015) Unique transposon landscapes are pervasive across Drosophila melanogaster genomes. Nucleic Acids Res 43(22):10655–10672
Treiber CD, Waddell S (2017) Resolving the prevalence of somatic transposition in Drosophila. Elife 6:e28297
Nelson MG, Linheiro RS, Bergman CM (2017) McClintock: an integrated pipeline for detecting transposable element insertions in whole genome shotgun sequencing data. G3 (Bethesda) 7:2763
McCarthy A (2010) Third generation DNA sequencing: Pacific Biosciences’ single molecule real time technology. Chem Biol 17(7):675–676
McCoy RC, Taylor RW, Blauwkamp TA, Kelley JL, Kertesz M, Pushkarev D, Petrov DA, Fiston-Lavier AS (2014) Illumina TruSeq synthetic long-reads empower de novo assembly and resolve complex, highly-repetitive transposable elements. PLoS One 9(9):e106689
Cridland JM, Macdonald SJ, Long AD, Thornton KR (2013) Abundance and distribution of transposable elements in two Drosophila QTL mapping resources. Mol Biol Evol 30(10):2311–2327
Sudmant PH, Rausch T, Gardner EJ, Handsaker RE, Abyzov A, Huddleston J, Zhang Y, Ye K, Jun G, Fritz MH, Konkel MK, Malhotra A, Stutz AM, Shi X, Casale FP, Chen J, Hormozdiari F, Dayama G, Chen K, Malig M, Chaisson MJP, Walter K, Meiers S, Kashin S, Garrison E, Auton A, Lam HYK, Mu XJ, Alkan C, Antaki D, Bae T, Cerveira E, Chines P, Chong Z, Clarke L, Dal E, Ding L, Emery S, Fan X, Gujral M, Kahveci F, Kidd JM, Kong Y, Lameijer EW, McCarthy S, Flicek P, Gibbs RA, Marth G, Mason CE, Menelaou A, Muzny DM, Nelson BJ, Noor A, Parrish NF, Pendleton M, Quitadamo A, Raeder B, Schadt EE, Romanovitch M, Schlattl A, Sebra R, Shabalin AA, Untergasser A, Walker JA, Wang M, Yu F, Zhang C, Zhang J, Zheng-Bradley X, Zhou W, Zichner T, Sebat J, Batzer MA, McCarroll SA, Genomes Project C, Mills RE, Gerstein MB, Bashir A, Stegle O, Devine SE, Lee C, Eichler EE, Korbel JO (2015) An integrated map of structural variation in 2,504 human genomes. Nature 526(7571):75–81
Huddleston J, Chaisson MJP, Steinberg KM, Warren W, Hoekzema K, Gordon D, Graves-Lindsay TA, Munson KM, Kronenberg ZN, Vives L, Peluso P, Boitano M, Chin CS, Korlach J, Wilson RK, Eichler EE (2017) Discovery and genotyping of structural variation from long-read haploid genome sequence data. Genome Res 27(5):677–685
Pendleton M, Sebra R, Pang AW, Ummat A, Franzen O, Rausch T, Stutz AM, Stedman W, Anantharaman T, Hastie A, Dai H, Fritz MH, Cao H, Cohain A, Deikus G, Durrett RE, Blanchard SC, Altman R, Chin CS, Guo Y, Paxinos EE, Korbel JO, Darnell RB, McCombie WR, Kwok PY, Mason CE, Schadt EE, Bashir A (2015) Assembly and diploid architecture of an individual human genome via single-molecule technologies. Nat Methods 12(8):780–786
Disdero E, Filee J (2017) LoRTE: detecting transposon-induced genomic variants using low coverage PacBio long read sequences. Mob DNA 8:5
Kelleher ES, Edelman NB, Barbash DA (2012) Drosophila interspecific hybrids phenocopy piRNA-pathway mutants. PLoS Biol 10(11):e1001428
Vela D, Fontdevila A, Vieira C, Garcia Guerreiro MP (2014) A genome-wide survey of genetic instability by transposition in Drosophila hybrids. PLoS One 9(2):e88992
Romero-Soriano V, Modolo L, Lopez-Maestre H, Mugat B, Pessia E, Chambeyron S, Vieira C, Garcia Guerreiro MP (2017) Transposable element misregulation is linked to the divergence between parental piRNA pathways in drosophila hybrids. Genome Biol Evol 9(6):1450–1470
Robillard E, Le Rouzic A, Zhang Z, Capy P, Hua-Van A (2016) Experimental evolution reveals hyperparasitic interactions among transposable elements. Proc Natl Acad Sci U S A 113(51):14763–14768
Gonzalez J, Petrov DA (2012) Evolution of genome content: population dynamics of transposable elements in flies and humans. Methods Mol Biol 855:361–383
Cordaux R, Hedges DJ, Herke SW, Batzer MA (2006) Estimating the retrotransposition rate of human Alu elements. Gene 373:134–137
Ewing AD, Kazazian HH Jr (2010) High-throughput sequencing reveals extensive variation in human-specific L1 content in individual human genomes. Genome Res 20(9):1262–1270
Huang CR, Schneider AM, Lu Y, Niranjan T, Shen P, Robinson MA, Steranka JP, Valle D, Civin CI, Wang T, Wheelan SJ, Ji H, Boeke JD, Burns KH (2010) Mobile interspersed repeats are major structural variants in the human genome. Cell 141(7):1171–1182
Xing J, Zhang Y, Han K, Salem AH, Sen SK, Huff CD, Zhou Q, Kirkness EF, Levy S, Batzer MA, Jorde LB (2009) Mobile elements create structural variation: analysis of a complete human genome. Genome Res 19(9):1516–1526
Ernst C, Odom DT, Kutter C (2017) The emergence of piRNAs against transposon invasion to preserve mammalian genome integrity. Nat Commun 8(1):1411
McCullers TJ, Steiniger M (2017) Transposable elements in Drosophila. Mob Genet Elements 7(3):1–18
Charlesworth B, Charlesworth D (1983) The population dynamics of transposable elements. Genet Res 42(1):1–27
Lohe AR, Hartl DL (1996) Autoregulation of mariner transposase activity by overproduction and dominant-negative complementation. Mol Biol Evol 13(4):549–555
Grabundzija I, Irgang M, Mates L, Belay E, Matrai J, Gogol-Doring A, Kawakami K, Chen W, Ruiz P, Chuah MK, VandenDriessche T, Izsvak Z, Ivics Z (2010) Comparative analysis of transposable element vector systems in human cells. Mol Ther 18(6):1200–1209
Bire S, Casteret S, Arnaoty A, Piegu B, Lecomte T, Bigot Y (2013) Transposase concentration controls transposition activity: myth or reality? Gene 530(2):165–171
Woodard LE, Downes LM, Lee YC, Kaja A, Terefe ES, Wilson MH (2017) Temporal self-regulation of transposition through host-independent transposase rodlet formation. Nucleic Acids Res 45(1):353–366
Wheeler BS (2013) Small RNAs, big impact: small RNA pathways in transposon control and their effect on the host stress response. Chromosome Res 21(6-7):587–600
Clark JP, Lau NC (2014) Piwi proteins and piRNAs step onto the systems biology stage. Adv Exp Med Biol 825:159–197
Toth KF, Pezic D, Stuwe E, Webster A (2016) The piRNA pathway guards the germline genome against transposable elements. Adv Exp Med Biol 886:51–77
Yang F, Xi R (2017) Silencing transposable elements in the Drosophila germline. Cell Mol Life Sci 74(3):435–448
Luo S, Lu J (2017) Silencing of transposable elements by piRNAs in drosophila: an evolutionary perspective. Genomics Proteomics Bioinformatics 15(3):164–176
Aravin AA, Sachidanandam R, Bourc’his D, Schaefer C, Pezic D, Toth KF, Bestor T, Hannon GJ (2008) A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell 31(6):785–799
Le Thomas A, Rogers AK, Webster A, Marinov GK, Liao SE, Perkins EM, Hur JK, Aravin AA, Toth KF (2013) Piwi induces piRNA-guided transcriptional silencing and establishment of a repressive chromatin state. Genes Dev 27(4):390–399
Hamdorf M, Idica A, Zisoulis DG, Gamelin L, Martin C, Sanders KJ, Pedersen IM (2015) miR-128 represses L1 retrotransposition by binding directly to L1 RNA. Nat Struct Mol Biol 22(10):824–831
Ariumi Y (2016) Guardian of the human genome: host defense mechanisms against LINE-1 retrotransposition. Front Chem 4:28
Wylie A, Jones AE, D’Brot A, Lu WJ, Kurtz P, Moran JV, Rakheja D, Chen KS, Hammer RE, Comerford SA, Amatruda JF, Abrams JM (2016) p53 genes function to restrain mobile elements. Genes Dev 30(1):64–77
Van Meter M, Kashyap M, Rezazadeh S, Geneva AJ, Morello TD, Seluanov A, Gorbunova V (2014) SIRT6 represses LINE1 retrotransposons by ribosylating KAP1 but this repression fails with stress and age. Nat Commun 5:5011
Karasov T, Messer PW, Petrov DA (2010) Evidence that adaptation in Drosophila is not limited by mutation at single sites. PLoS Genet 6(6):e1000924
Park L (2011) Effective population size of current human population. Genet Res (Camb) 93(2):105–114
Barron MG, Fiston-Lavier AS, Petrov DA, Gonzalez J (2014) Population genomics of transposable elements in Drosophila. Annu Rev Genet 48:561–581
Kofler R, Nolte V, Schlotterer C (2015) Tempo and mode of transposable element activity in drosophila. PLoS Genet 11(7):e1005406
Rishishwar L, Tellez Villa CE, Jordan IK (2015) Transposable element polymorphisms recapitulate human evolution. Mob DNA 6:21
Boissinot S, Davis J, Entezam A, Petrov D, Furano AV (2006) Fitness cost of LINE-1 (L1) activity in humans. Proc Natl Acad Sci U S A 103(25):9590–9594
Sundaram V, Choudhary MN, Pehrsson E, Xing X, Fiore C, Pandey M, Maricque B, Udawatta M, Ngo D, Chen Y, Paguntalan A, Ray T, Hughes A, Cohen BA, Wang T (2017) Functional cis-regulatory modules encoded by mouse-specific endogenous retrovirus. Nat Commun 8:14550
Chalopin D, Naville M, Plard F, Galiana D, Volff JN (2015) Comparative analysis of transposable elements highlights mobilome diversity and evolution in vertebrates. Genome Biol Evol 7(2):567–580
Kapusta A, Suh A, Feschotte C (2017) Dynamics of genome size evolution in birds and mammals. Proc Natl Acad Sci U S A 114(8):E1460–E1469
Leushkin EV, Bazykin GA, Kondrashov AS (2013) Strong mutational bias toward deletions in the Drosophila melanogaster genome is compensated by selection. Genome Biol Evol 5(3):514–524
Petrov DA, Lozovskaya ER, Hartl DL (1996) High intrinsic rate of DNA loss in Drosophila. Nature 384(6607):346–349
Petrov DA, Hartl DL (1998) High rate of DNA loss in the Drosophila melanogaster and Drosophila virilis species groups. Mol Biol Evol 15(3):293–302
Loreto EL, Carareto CM, Capy P (2008) Revisiting horizontal transfer of transposable elements in Drosophila. Heredity (Edinb) 100(6):545–554
Schaack S, Gilbert C, Feschotte C (2010) Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution. Trends Ecol Evol 25(9):537–546
Peccoud J, Loiseau V, Cordaux R, Gilbert C (2017) Massive horizontal transfer of transposable elements in insects. Proc Natl Acad Sci U S A 114(18):4721–4726
Peccoud J, Cordaux R, Gilbert C (2018) Analyzing horizontal transfer of transposable elements on a large scale: challenges and prospects. Bioessays 40(2)
Groth SB, Blumenstiel JP (2017) Horizontal transfer can drive a greater transposable element load in large populations. J Hered 108(1):36–44
Mouse Genome Sequencing Consortium (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420(6915):520–562
Kim JM, Vanguri S, Boeke JD, Gabriel A, Voytas DF (1998) Transposable elements and genome organization: a comprehensive survey of retrotransposons revealed by the complete Saccharomyces cerevisiae genome sequence. Genome Res 8(5):464–478
Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408(6814):796–815
Filee J, Siguier P, Chandler M (2007) Insertion sequence diversity in archaea. Microbiol Mol Biol Rev 71(1):121–157
Sebaihia M, Peck MW, Minton NP, Thomson NR, Holden MT, Mitchell WJ, Carter AT, Bentley SD, Mason DR, Crossman L, Paul CJ, Ivens A, Wells-Bennik MH, Davis IJ, Cerdeno-Tarraga AM, Churcher C, Quail MA, Chillingworth T, Feltwell T, Fraser A, Goodhead I, Hance Z, Jagels K, Larke N, Maddison M, Moule S, Mungall K, Norbertczak H, Rabbinowitsch E, Sanders M, Simmonds M, White B, Whithead S, Parkhill J (2007) Genome sequence of a proteolytic (Group I) Clostridium botulinum strain Hall A and comparative analysis of the clostridial genomes. Genome Res 17(7):1082–1092
Rhee JS, Choi BS, Kim J, Kim BM, Lee YM, Kim IC, Kanamori A, Choi IY, Schartl M, Lee JS (2017) Diversity, distribution, and significance of transposable elements in the genome of the only selfing hermaphroditic vertebrate Kryptolebias marmoratus. Sci Rep 7:40121
Osanai-Futahashi M, Suetsugu Y, Mita K, Fujiwara H (2008) Genome-wide screening and characterization of transposable elements and their distribution analysis in the silkworm, Bombyx mori. Insect Biochem Mol Biol 38(12):1046–1057
Hernandez-Hernandez EM, Fernandez-Medina RD, Navarro-Escalante L, Nunez J, Benavides-Machado P, Carareto CMA (2017) Genome-wide analysis of transposable elements in the coffee berry borer Hypothenemus hampei (Coleoptera: Curculionidae): description of novel families. Mol Genet Genomics 292(3):565–583
Castanera R, Borgognone A, Pisabarro AG, Ramirez L (2017) Biology, dynamics, and applications of transposable elements in basidiomycete fungi. Appl Microbiol Biotechnol 101(4):1337–1350
Tenaillon MI, Hufford MB, Gaut BS, Ross-Ibarra J (2011) Genome size and transposable element content as determined by high-throughput sequencing in maize and Zea luxurians. Genome Biol Evol 3:219–229
Boissinot S, Sookdeo A (2016) The evolution of LINE-1 in vertebrates. Genome Biol Evol 8(12):3485–3507
Quadrana L, Bortolini Silveira A, Mayhew GF, LeBlanc C, Martienssen RA, Jeddeloh JA, Colot V (2016) The Arabidopsis thaliana mobilome and its impact at the species level. Elife 5:e15716
Stuart T, Eichten SR, Cahn J, Karpievitch YV, Borevitz JO, Lister R (2016) Population scale mapping of transposable element diversity reveals links to gene regulation and epigenomic variation. Elife 5:e20777
Yan L, Gu YH, Tao X, Lai XJ, Zhang YZ, Tan XM, Wang H (2014) Scanning of transposable elements and analyzing expression of transposase genes of sweet potato Ipomoea batatas. PLoS One 9(3):e90895
Acknowledgment
We thank the reviewers for providing constructive comments on a previous version of this manuscript. This work has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (H2020-ERC-2014-CoG-647900) and from the Spanish Ministry of Economy and Competitiveness/FEDER (BFU2014-57779-P).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2019 The Author(s)
About this protocol
Cite this protocol
Guio, L., González, J. (2019). New Insights on the Evolution of Genome Content: Population Dynamics of Transposable Elements in Flies and Humans. In: Anisimova, M. (eds) Evolutionary Genomics. Methods in Molecular Biology, vol 1910. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9074-0_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9074-0_16
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9073-3
Online ISBN: 978-1-4939-9074-0
eBook Packages: Springer Protocols