Abstract
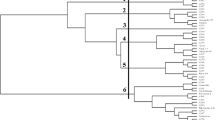
Ten microsatellite markers were used to investigate genetic diversity and genetic structure among 32 accessions of Jatropha curcas. Low levels of average genetic diversity were observed (H E = 0.160). A dendrogram produced by the Unweighted Pair Group Method with Arithmetic Mean (UPGMA) based on Nei’s genetic distances revealed 3 groups among 32 accessions. The genetic differentiation (F ST ) among two groups was significant (P < 0.01). The model-based Bayesian clustering method indicated that a population structure (ΔK) was separated into two groups. The analysis of molecular variance (AMOVA) showed higher variability (63.753%) among groups than within groups (36.247%). These findings could assist in defining the best method of genetic conservation and studies in breeding programs for genetic improvement of J. curcas.
Similar content being viewed by others
References
Achten WMJ, Nielsen LR, Aerts R, Lengkeek AG, Kjaer ED, Trabucco A, Hansen JK, Maes WK, Grudal L, Akennifesi FK, Muys B. 2010. Towards domestication of Jatropha curcas. Biofuels 1: 91–10
Basha SD, George F, Makkar HPS, Becker K, Sujatha M. 2009. A comparative study of biochemical traits and molecular markers for assessment of genetic relationships between Jatropha Curcas L. germplasm from different countries. Plant Sci. 176: 812–823
Benbouza H, Jacquemin JM, Baudoin JP, Mergeai G. 2006. Optimization of a reliable, fast, cheap and sensitive silver staining method to detect SSR markers in polyacrylamide gels. Biotechnol. Agron. Soc. Environ. 10: 77–81
Carvalho CR, Clarindo WR, Praca MM, Araujo FS, Carels N. 2008. Genome size, base composition and karyotype of Jatropha curcas L., an important biofuel plant. Plant Sci. 174: 613–617
Dhakshanamoorthy D, Selvaraj R, Chidambaram ALA. 2011. Induced mutagenesis in Jatropha curcas L. using gamma rays and detection of DNA polymorphism through RAPD marker. Comptes Rendus Biol. 334: 24–30
Doyle JJ, Doyle JL. 1990. Isolation of plant DNA from fresh tissue. Focus 12: 13–15
Evanno G, Regnaut S, Goudet J. 2005. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14: 2611–2620
Excoffier L, Laval G, Schneider S. 2005. Arlequin ver. 3.0: An integrated software package for population genetics data analysis. Evol. Bioinform. Online 1: 47–50
Ghosh A, Chaudhary DR, Reddy MP, Rao SN, Chikara J, Pandya JB. 2007. Prospects for Jatropha methyl ester (biodiesel) in India. Int. J. Environ. Stud. 64: 659–674
Gupta PK, Varshney RK. 2000. The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica 113: 163–185
Heller J. 1996. Physic nut, Jatropha curcas. Promoting the conservation and use of underutilized and neglected crops. Rome, Italy, International Plant Genetic Resources Institute (IPGRI)
Hu JB, Zhou XY, Li JW. 2009. Development of novel chloroplast microsatellite markers for Cucumis from sequence database. Biol. Plant 53: 793–796
Liu K, Muse SV. 2005. PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21: 2128–2129
Nei M. 1973. Analysis of gene diversity in subdivided populations. Proc. Natl. Acad. Sci. USA 70: 3321–3323
Nei M. 1978. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89: 583–590
Peakall R, Smouse PE. 2006. GENALEX 6: genetic analysis in Excel Population genetic software for teaching and research. Mol. Ecol. Notes 6: 288–295
Phumichai C, Doungchan W, Puddhanon P, Jampatong S, Grudloyma P, Kirdsri C, Chunwongse J, Pulam T. 2008. SSR-based and grain yield-based diversity of hybrid maize in Thailand. Field Crops Res. 108: 157–162
Phumichai C, Phumichai T, Kongsiri N, Wongkaew A, Sripichit P, Kaveeta R. 2011. Isolation of 55 microsatellite markers for physic nut (Jatropha curcas L.) and its closely related species. Biol. Plant 55: 387–390
Pritchard JK, Stephens M, Donnelly P. 2000. Inference of population structure using multilocus genotype data. Genetics 155: 945–959
Shen J, Jia X, Ni H, Sun P, Niu S, Chen X. 2010. AFLP analysis of genetic diversity of Jatropha curcas grown in Hainan, China Trees. Trees — Structure and Function 24: 455–462
Sudheer PDVN, Nirali P, Reddy MP, Radhakrishnan T. 2009a. Comparative study of interspecific genetic divergence and phylogenic analysis of genus Jatropha by RAPD and AFLP. Mol. Biol. Rep. 36: 901–907
Sudheer PDVN, Rahman H, Mastan SG, Reddy MP. 2010c. Isolation of novel microsatellites using FIASCO by dual probe enrichment from Jatropha curcas L. and study on genetic equilibrium and diversity of Indian population revealed by isolated microsatellites. Mol. Biol. Rep. 37: 3785–3793
Sudheer PDVN, Singh S, Mastan SG, Patel J, Reddy MP. 2009b. Molecular characterization and identification of markers for toxic and non-toxic varieties of Jatropha curcas L. using RAPD, AFLP and SSR markers. Mol. Biol. Rep. 36: 1357–1364
Sun QB, Li LF, Li Y, Wu GJ, Ge XJ. 2008. SSR and AFLP markers reveal low genetic diversity in the biofuel plant Jatropha curcas in China. Crop Sci. 48: 1865–1871
Takeda Y. 1982. Development study on Jatropha curcas (Sabudum) oil as a substitute for diesel engine oil in Thailand. J. Agric Assoc. China 120: 1–8
Tamura K, Dudley J, Nei M, Kumar S. 2007. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24: 1596–1599
Tanya P, Taeprayoon P, Hadkam Y, Srinives P. 2010. Genetic diversity among Jatropha and Jatropha-related species based on ISSR markers. Plant Mol. Biol. Rep. 29: 252–264
Tatikonda L, Wani SP, Kannan S, Beerelli N, Sreedevi TK, Hoisington DA, Devi P, Varshney RK. 2009. AFLP-based molecular characterization of an elite germplasm collection of Jatropha curcas L., a biofuel plant. Plant Sci. 176: 505–513
Weir BS. 1996. Genetic data analysis II. Sinauer Associates Inc., Sunderland
Wen M, Wang H, Xia Z, Zou M, Lu C, Wang W. 2010. Develop ment of EST-SSR and genomic-SSR markers to assess genetic diversity in Jatropha Curcas L. BMC Res. Notes 3: 1–8
Yeh FC, Yang R, Boyle TJ, Ye Z, Xiyan JM. 2000. PopGene32, Microsoft Windows-based freeware for population genetic analysis, version 1.32. Molecular Biology and Biotechnology Centre, University of Alberta, Edmonton, Alberta, Canada
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Na-ek, Y., Wongkaew, A., Phumichai, T. et al. Genetic diversity of physic nut (Jatropha curcas L.) revealed by SSR markers. J. Crop Sci. Biotechnol. 14, 105–110 (2011). https://doi.org/10.1007/s12892-011-0008-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12892-011-0008-4