Abstract
The current study aims to exploit the zero-cost inducer wheat bran (WB), an agro-industrial byproduct, for production of alkaline protease (ALK-PR23) by the hyper producer psychrotolerant Lysinibacillus sphaericus Strain AA6 EMCCN3080 for the first time ever. Incubation temperature (25 °C), yeast extract concentration, agitation speed (150 rpm), and aeration ratio [1 volume (liquid):5 volume (Erlenmeyer flask)] provoked ALK-PR23 production; OVAT inferences. The pH, yeast extract, and (NH4)2SO4 levels substantively triggered ALK-PR23 production as deduced from Plackett–Burman design. Incubation time (3 days) and WB [2% (w/v)] were the optimal values inducing positive significant influence on ALK-PR23 as conferred from steepest ascent experiments. Yeast extract (0.446% w/v), (NH4)2SO4 (0.339% w/v), and pH (6.872) prompted ALK-PR23 (592.5 U/mL) with an impressive 98-fold enhancement; Box-Behnken design and ridge steepest ascent path implications. The laboratory validation of the model achieved 100% of the predicted value. Laboratory data would present an eco-friendly, cheap, efficient approach towards concurrent WB recycling and massive production of alkaline protease (ALK-PR23) from L. sphaericus Strain AA6 EMCCN3080. Present data would greatly encourage unveiling biochemical characteristics of ALK-PR23 in prospective studies.
Graphical Abstract

Similar content being viewed by others
Statement of Novelty
This is the first article addressing the production of alkaline protease by a local Egyptian psychrotolerant bacterial strain namely Lysinibacillus sphaericus Strain AA6 EMCCN3080 through bio-valorization of the raw-agro-industrial byproduct wheat bran (WB), as a zero-cost substrate using an empirical statistical approach, with an impressive fold of enhancement (98 times) compared to other orthologues of alkaline protease producers reported in the literature.
Introduction
According to the FAO's forecast for world wheat, overall wheat consumption is expected to reach 779 million tons in 2022–2023, an increase of 0.8% from the level in 2021/2022. Consequently, millions of tons of the raw-agro-industrial byproduct wheat bran (WB) are generated annually worldwide. The chemical composition of WB has 13–18% protein [1]. Several studies have addressed the likely utilization of WB in several fields such as enzyme production [2], metabolites production [3], biofuels production [4] heavy metal removal [5], and food-feed additive [6, 7]. Despite the various fields available for up-recycling of WB, its surplus greatly does necessitate the indispensable need to look for additional fields for efficient utilization of WB.
Proteases are a significant class of enzymes with numerous industrial consumptions in different sectors such as proteinous/keratinuous waste treatment, detergent industry, textile industry, leather industry, baking industry, bioremediation, etc. [8]. Statistics of sales in global enzyme markets do indicate that proteases represent the highest percent of enzyme sales annually [9]. Despite a great panel of microorganisms (e.g., Bacillus spp. [10, 11], Serratia spp. [12, 13], Pseudomonas spp. [14, 15], Vibrio spp. [16], Streptomyces spp. [17, 18], Aspergillus spp. [19, 20], Penicillium spp. [21, 22], etc..) have been reported to be protease hyper-producers, searching for novel microbial sources for proteases is a worthy task for co-lowering the expenditure of protease industrialization and improving the yield.
Lysinibacillus is a recently re-classified genus of Bacillus that is psychrotolerant [23]. The biotechnological potential of the Lysinibacillus genus in the insects’ biocontrol [24, 25], production of significant biomolecules/enzymes with industrial prospects [24, 26], and environmental bioremediation have attracted much attention in recent years [27]. Currently, the microbial whole genome sequencing projects launched by NCBI (National center for Biotechnology Information) have been providing updated thorough information about the genome mining of Lysinibacillus spp. (e.g., L. macrolides, L. xylanilyticus, L. boronitolerans, L. varians, L. contaminans, L. sphaericus, L. fusiformis, etc.). Despite the presence of several putative protease sequences from Lysinibacillus spp., indicated by the 'GenBank database, protease production has been reported from L. fusiformis [28] only so far. Consequently, targeting the Lysinibacillus spp., as possible promising candidates for proteases production would expectedly pull the attention of researchers worldwide to unravel the nature of protease from Lysinibacillus spp. Searching for novel hyper-protease producers would necessitate the indispensable need to perform rigorous statistical optimization procedure (Plackett–Burman designs and response surface methodology) to maximize the yield of the protease from its wildtype source. Numerous studies have addressed the potential of statistical modeling towards realizing satisfactory levels of primary and secondary microbial metabolites from wildtype producers [29,30,31,32,33,34].
In this context, the goal of the present study is tailoring a low cost effective medium, using wheat bran as zero-cost substrate, and applying rigorous statistical designs to maximize the protease production by L. sphaericus Strain AA6 EMCCN3080, a local psychrotolerant strain for the first time ever.
Materials and Methods
Wheat Bran: Raw Agro-industrial Byproduct
Wheat bran (WB), employed in this study as a zero-cost alkaline protease inducer, was obtained from a regional farm in Alexandria City, Egypt. The WB was subjected to oven dryness at 60 °C. Then it was conserved in sterile plastic bags at room temperature until being utilized.
Bacterial Strain
Lysinibacillus sphaericus Strain AA6 EMCCN3080, a local psychrotolerant bacterium, was previously isolated from soil sample taken from a local garden, Alexandria City, Egypt during winter season. Shortly, five grams of soil collected from the local garden in the Institute of Graduate studies and Research, Alexandria University, Egypt was added to 30 mL LB (Lauria–Bertani) broth in 250 mL Erylenemyer flask for enrichment purposes. Then, this flask was incubated overnight at 30 °C with agitation speed of 180 rpm (New Brunswick Incubator Shaker, USA). Next day, a serial dilution from the enriched culture was performed using saline (0.85% w/v) as a diluent. One hundred microliter (100 µL) of an appropriate dilution was withdrawn to be plated on LB agar plates followed by incubation overnight at 30 °C. The raised separate colonies were subjected to successive subculturing on LB agar plates for purification purposes using streaking technique. The proteolytic capabilities of the single purified colonies was checked on 1% (w/v) casein LB agar plates followed by incubation overnight at 30 °C. The appearance of clear zones around the colonies was an indicative of proteolytic capability of the tested bacterial isolate (data not shown).
The bacterial strain was subjected to biochemical identification using VITEK 2 compact system (Biomérieux, Boston, MA, USA) and molecular identification through 16S rRNA nucleotide sequence analysis. Concisely, the G-spinTM Genomic DNA Extraction Kit for Bacteria (iNtRON Co., Korea) was used to extract the genomic DNA from the bacterial isolate in accordance with the manufacturer's instructions. Using the universal 16S rRNA primer pair of 27 F (5′- AGAGTTTGATCCTGGCTCAG-3′) and 1492 R (5′- GGTTACCTTGTTACGACTT-3′), the complete length 16S rRNA gene was amplified by polymerase chain reaction (PCR) [35]. A previously described method was used for the PCR [29]. The PCR reaction mixture (50 μL) was made up by the addition of 3 μL (30 ng genomic DNA), 25 μL of MyTaqTM Red Mix (Meridian Bioscience Inc., Ohio, USA), 0.5 μM of each forward and reverse primer and 19 μL of nuclease free water. The following PCR conditions were established for the Thermocycler (Biometra, Analytik Jena Co., Germany): initial denaturation at 95 °C for 5 min, 30 cycles, each cycle consisting of denaturation at 94 °C for 30 s, annealing at 55 °C for 45 s, extension at 72 °C for 1.5 min, and a final extension at 72 °C for 10 min. GeneJET PCR purification kit (Thermo Fisher Scientific Co., Massachusetts, USA) was used to purify the PCR product in accordance with the manufacturer's instructions. The primer set (27 F and 1492 R) and 1100 R (5′- AGGGTTGCGCTCGTTG-3′) were then applied for two-directional sequencing of the purified PCR product by Macrogen, Seoul, Korea. Using BLASTN (Basic Local Alignment Search Tool for Nucleotide) of NCBI, the resulting 16S rRNA nucleotide sequence was searched against rRNA/ITS databases. Using the CLC Sequence Viewer 8.0, multiple sequence alignment was conducted among the query sequence and the top seventeen subject sequences that strongly matched with the query sequence. The phylogenetic relationship of the bacterial isolate in question to other closely related bacterial species was evaluated using a neighbor-joining phylogenetic tree constructed by CLC Sequence Viewer 8.0. The 16S rRNA nucleotide sequence of the bacterial strain in question was deposited in GenBank under the accession number MT108450.1.
The growth temperature profile of the bacterial strain was tested at different incubation temperatures: 15, 25, 30, and 37 °C. Shortly, a single colony of an overnight culture of the bacterial strain, cultivated on LB agar, was used to inoculate 50 mL LB (Lauria–Bertani Broth) in 250 mL Erlenmeyer flask. The inoculated broth (seed culture) was incubated at 25 °C for an overnight with agitation speed of 150 rpm (New Brunswick Incubator Shaker, USA). After that, 1% (v/v) of the overnight seed culture was used to inoculate four LB broths, 50 mL broth in 250 mL Erlenmeyer flask each, followed by incubation at different incubation temperatures: 15, 25, 30, and 37 °C with agitation speed of 180 rpm for 18 h. The optical density of all cultures was measured at 600 nm (Evolution ™ 201/220 UV–visible spectrophotometer, Thermo Scientific™, US) after 18 h.
Cultivating Conditions and Protease Production Media
Short-term storage, activation, and seed culture preparation of the hyper- alkaline protease producer L. sphaericus Strain AA6 EMCCN3080 were conducted using Luria–Bertani broth (LB) at 25 °C, pH 7.0, and 180 rpm overnight. The alkaline protease activity was designated from now and later ALK-PR23.
Herein, four core ALK-PR23 production media were used: (a) minimal medium based-distilled water (in g/100 mL): 0.2 KH2PO4, 0.6 K2HPO4, 0.14 (NH4)2SO4, 0.01 MgSO4·7H2O, 2.0 agar, and 1.0 casein), (b) modified minimal medium (in g/100 mL): 0.1 yeast extract, 0.2 KH2PO4, 0.6 K2HPO4, 0.14 (NH4)2SO4, 0.01 MgSO4·7H2O, and 2.0 WB), (c) modified minimal medium (in g/mL): 0.1 peptone, 0.2 KH2PO4, 0.6 K2HPO4, 0.14 (NH4)2SO4, 0.01 MgSO4·7H2O, and 2.0 WB), and modified minimal medium (in g/100 mL): 0.2 KH2PO4, 0.6 K2HPO4, 0.14 (NH4)2SO4, 0.01 MgSO4·7H2O, and 2.0 WB). The production of ALK-PR23 was conducted at 25 °C, pH 7.0, 3 days, and 150 rpm unless otherwise stated.
Alkaline Protease (ALK-PR23) Assay
The ALK-PR23 activity was estimated according to a previously reported procedure [36] using 0.5% (w/v) casein (HiMedia, India) as a substrate. Briefly, 0.5 mL of 0.5% (w/v) casein in 100 mL (0.05 M glycine–NaOH buffer, pH 10.0) was mixed with 0.5 mL of the crude enzyme and incubated at 37 °C for 30 min. The reaction mixture was terminated by adding 0.5 mL 0f (20%) trichloroacetic acid (TCA) (Sigma-Aldrich Co., USA). After that, the solution was centrifuged for 10 min at 10,000 rpm in a microcentrifuge (Centrifuge 5420, Eppendorf, Hamburg, Germany) and the supernatant was transferred to new Eppendorf tubes and the absorbance was measured at 280 nm (UV–visible spectrophotometer, Shimadzu, Japan). The control reactions were performed as mentioned above, except the crude enzyme was added after the addition of the TCA. A standard curve of tyrosine was established. All assays were conducted in triplicates. One arbitrary unit (1U) of enzyme activity was defined as the amount of enzyme that liberates 1 μmol tyrosine from the casein substrate per minute under the assay conditions specified.
Optimization of Alkaline ALK-PR23 Production
The production of ALK-PR23 by the pschyrotolerant, alkaline protease hyper-producer L. sphaericus Strain AA6 EMCCN3080 in submerged state fermentation with WB as the inducer was performed using an empirical, sequential, statistical approach: one variable at a t time (OVAT), Plackett–Burman Design (PBD), and Box-Behnken Design (BBD).
OVAT Methodology
Herein, the examined physiochemical determinants were core production medium (with and without yeast extract and/or peptone), agitation speed (100, 150, and 180 rpm), aeration ratio (1:2.5, 1:5, and 1:10 (volume of liquid: volume of Erlenmeyer flask)), and incubation temperature (20, 25, 30, and 37 °C). Three core ALK-PR23 production media were tested: (a) modified minimal medium with yeast extract and WB, (b) modified minimal medium with peptone and WB, and (c) modified minimal medium with WB containing neither yeast extract nor peptone. The OVAT experiments were carried out in 250 mL Erlenmeyer flasks. An overnight seed culture of the alkaline protease hyper-producer L. sphaericus Strain AA6 EMCCN3080 was used to inoculate all the aforementioned production media with 1% (v/v) inoculum size. All cultures were incubated in a shaking incubator for 72 h at a 150 rpm agitation speed. All experimental trials were carried out in triplicates. The key determinants of ALK-PR23 production, deduced from OVAT experiments, were selected as the base for further tailoring the ALK-PR23 core—production medium in the next optimization experiments.
Plackett–Burman Design (PBD)
Here, the full factorial Plackett–Burman design [37] was used to assess the potential linear effects of seven independent variables on the production of ALK-PR23, including WB concentration (X1), yeast extract concentration (X2), ammonium sulphate concentration (X3), magnesium sulphate concentration (X4), initial pH of the production medium (X5), incubation time (X6), and inoculum size (X7). The Minitab 17.3 software was used to create a full factorial design matrix consisting of twelve experimental trials, as shown in Table 1. Each variable was specified as having two coded values: − 1 (low level) and + 1 (high level). The polynomic equation of a first order (Eq. 1) was validated in order to represent all likely linear influences imposed by the examined independent variables on ALK-PR23 production.
An overnight seed culture of the alkaline protease hyper-producer L. sphaericus Strain AA6 EMCCN3080 was used to inoculate all the experimental trials settled in Table 1. The experimental trials were carried out in 250 mL Erlenmeyer flasks, with a working capacity of 50 mL production medium, at an agitation speed of 150 rpm and at the best experimental conditions deduced from OVAT experiments. The final step of the optimization strategy, BBD, further investigated the independent variables revealing substantial concerns on ALK-PR23 production as deduced from PBD. While the independent variables that had no discernible effects on the production of ALK-PR23 were left at their initial settings in the upcoming optimization trials.
Steepest Ascent (or Descent) Path
The initial predictions of some PBDs operating parameters are slightly distant from the real optimum levels; therefore, a strategy is required to quickly conduct experiments in the general area of the optimum. To go swiftly in the direction with the highest rise in the response, the steepest ascent (or descent) strategy is employed. To find the route of greatest improvement, the non-significant variables derived from the PBD design were further optimized. The sharpest ascent (or fall) path in the PBD began from the center (zero level) of the selected variables up until the response showed no further enhancement. Accordingly, the slight non-significant effect of the two independent variables, wheat bran concentration and incubation time would necessitate the need for conducting additional experiments with more different levels to uncover exactly the optimum levels of these variables enforcing optimum ALK-PR23 activity.
Box-Behnken Design (BBD)
To determine the optimal level of each key determinant in combination with the maximal level of ALK-PR23, the key determinants that imposed substantial influences on the level of Alk-PR23 as deduced from PBD were studied using the BBD [38]. Through a design matrix of fifteen experimental runs (generated by Minitab 17.3 software), each independent variable was tested in three coded values indicated as − 1 (low level), 0 (center level), and + 1 (high level) (Table 2). Yeast extract concentration, ammonium sulphate concentration, and initial pH of the production medium were the three independent variables that were studied.
The second-order polynomial Eq. (2) represented all likely configurations of interactions among the investigated independent variables having substantial effects on ALK-PR23 production (2).
For statistical calculations, each independent variable X was coded as Xi according to the Eq. (3). Where Xi is dimensional coded value of the independent variable, xi is the real value of this variable at this coded value, xo is the real value of this variable at the center point (zero level) and ∆xi is the step change value.
The experimental trials were carried out in 250 mL Erlenmeyer flasks, with a working volume of 50 mL production medium, at an agitation speed of 150 rpm and at the best experimental conditions deduced from OVAT and steepest ascent path tests.
Statistical and Ridge Analyses
GraphPad software was used to analyze the data of OVAT experiments. The Minitab software version 17.3 was used to create PBD-BBD matrices and carry out multiple linear and non-linear regressions. To display the three-dimensional surface plots of the response, Statistica software 13.1 was used. Ridge analysis was carried out in this study using the RSM package version 2.10.3 (R development Core team 2021), which is accessible at the Comprehensive R Archive Network (http://CRAN.R-project.org/package=rsm).
Results and Discussion
Lysinibacillus sphaericus Strain AA6 EMCCN3080: Growth Temperature Profile
Lysinibacillus sphaericus Strain AA6 EMCCN3080 exhibited almost quite similar growth pattern upon incubation at different temperatures: 15, 25, 30, and 37 °C after 18 h of incubation (Table S1). The observed optical density at 600 nm of the tested growing cultures at 15, 25, 30, and 37 °C were 0.81 ± 0.03, 0.82 ± 0.037, 0.91 ± 0.05, and 0.89 ± 0.045, respectively. Present data would greatly underpin the psychrotolerant nature of L. sphaericus Strain AA6 EMCCN3080.
Lysinibacillus sphaericus Strain AA6 EMCCN3080: Biochemical and Molecular Profile
The biochemical features of the bacterial strain were displayed in Table S2 using VITEK 2 compact system (Biomérieux). The BLASTN sequence similarity search analysis of the 16S rRNA nucleotide sequence of the alkaline protease hyper -producer bacterial strain in question, against rRNA/ITS database, confirmed its affiliation to L. sphaericus with 98.73% similarity to L. sphaericus DSM28 under the accession number Gb:NR_042073.1. It was further designated on a strain level as L. sphaericus Strain AA6 according to the pre-requisites of 16S rRNA nucleotide sequence submission in GenBank databases. Currently, the bacterial strain L. sphaericus Strain AA6 has been deposited under the accession number EMCCN3080 in the public, Egyptian Microbial Culture Collection Network (EMCCN) at the National Research Center in Cairo, Egypt. Furthermore, the genetic relatedness of L. sphaericus Strain AA6 EMCCN3080 with other closely related species was deciphered by the phylogenetic tree depicted in Fig. 1. The Neighbor-Joining tree, constructed by CLC-Sequence Viewer 8.0, verified the close-relatedness of L. sphaericus Strain AA6 EMCCN3080 to L. sphaericus.
Neighbor-joining tree (constructed using CLC Sequence Viewer 8.0) presenting the phylogenetic association between 16S rRNA sequence of the candidate alkaline protease producing bacterial isolate (AA6) and other 16SrRNA nucleotides sequences fitting to closely related bacterial species. Bootstrap values were presented by numbers on branch nodes (1000 re-samplings). The bacterial isolate AA6 is the quest bacterial strain L. sphaericus Strain AA6 EMCCN3080 and is indicated by red arrow. (Color figure online)
Statistically Optimized Production of ALK-PR23 by L. sphaericus Strain AA6 EMCCN3080
Proteases are one of the main crucial enzymes particularly from an industrial standpoint. Accordingly, the permanent quest for novel members of alkaline proteases is being continuously addressed to fulfil the demands of enzymes markets.
Raw agro-industrial wastes are regarded as a valuable natural source for the manufacturing of enzymes [39]. The primary goal of the current study was to enhance the production of alkaline protease through the valorization of the raw agro-industrial byproduct WB and simultaneously reducing environmental problems caused by unchecked dumping and accumulation of these wastes and their byproducts. Much attention is currently directed towards sustainability which can be achieved through recycling of raw agro-industrial byproducts as an alternative zero-cost substrates for the production value-added bioproducts like bacterial enzymes. In support of the above-mentioned fact, John et al. reported that agricultural leftovers are a good replacement for expensive raw materials used in the production of proteases [40].
This study pointed the WB valorization towards the production of an alkaline protease known as ALK-PR23 by the aid of the psychrotolerant L. sphaericus Strain AA6 EMCCN3080. Moreover, due to the protein components, mineral nutrients, and co-factors that are essential for the development of microorganisms as well as the synthesis of metabolites [41] and to tailor a low-cost effective production medium, WB was chosen as a superlative substrate for the production of proteases in submerged fermentation.
Obviously, the cost of the production medium accounts for much of a bioprocess capital expense. Hence, creating a production medium with minimal expenditure would be the ideal option. Here, the zero-cost inducer WB was effectively exploited to reduce the total expense of the ALK-PR23 production medium.
The reviewed literature has revealed differences in the acceptable and utilized fermentable substrates that result in significant amounts of proteases when various bacterial strains are used. Groundnut oil cake and cabbage leaf were proved to be the best agro-substrate provoking alkaline protease production by the aid of Bacillus subtilis [42]. While gruel and wheat bran did induce the maximum production protease by L. fusiformis C250R [26]. In addition, pretreated cassava waste-stream was utilized to produce high yield of protease from Stenotrophomonas acidaminiphila [43] and Pseudomonas fluorescens W3 [44].
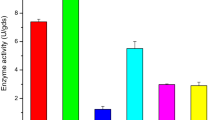
Regarding the type of the ALK-PR23 core production medium, the modified minimal medium containing yeast extract did promote the ultimate significant level (P-value < 0.05) of ALK-PR23 (216.55 ± 4.3 U/mL) in comparison to the other protease levels attained in minimal medium containing no yeast extract (Fig. 2a). One of the sources of amino acids and growth factors for bacteria is yeast extract. It also contains vitamins and co-factors that are necessary for bacterial growth and the synthesis of metabolites.
OVAT trials displaying the effect of using four physico-chemical factors on ALK-PR23 production by L. sphaericus Strain AA6 EMCCN3080. a Effect of using different core ALK-PR23 production media. b Effect of using different incubation temperatures (°C). c Effect of using different aeration ratios. d Effect of using different shaking speeds. Data are the average of three readings ± standard error (SE)
Extracellular enzymes including proteases produced by microbes are known to be significantly influenced by physical parameters such as temperature, aeration, agitation and dissolved oxygen. The temperature has a significant impact on how microorganisms evolve, carry out their biosynthesis and produce a variety of primary and secondary compounds including enzymes. Interestingly, results showed that incubation temperature (25 °C) was proven to be the ideal temperature triggering the highest significant level (P-value < 0.05) of ALK-PR23 (225 ± 4.0 U/mL) (Fig. 2b). Remarkably, there was a gradual significant decline (P-value < 0.05) in ALK-PR23 activity when the bacterial strain was incubated above 25 °C (Fig. 2b). In bioprocesses, using low incubation temperatures is clearly preferable to using high incubation temperatures. Low incubation temperatures help prevent protein mis-folding, conserve energy, and lessen the chance of coming into contact with contaminated microorganisms. Reportedly, incubation temperature 31 °C was found to be the optimum temperature for production of protease from S. acidaminiphila [43], while 37 °C was the ideal temperature for protease production from B. subtilis GACAS8 [42] and B. nakamurai PL4 [45].
A critical factor in aerobic submerged fermentation is oxygen transport, which is influenced by aeration and agitation. To achieve the highest yield, the best airflow and agitation combinations must be established [46]. Present data evidenced that conducting the bioprocess with 1:5 aeration ratio did significantly triggered the highest significant level of ALK-PR23 (at P-value < 0.0018) compared to that obtained upon conducting the bioprocess with an aeration ratio of 1:10 (Fig. 2c). Conversely, the level of ALK-PR23 at an aeration ratio of 1:2.5 was significantly lower (P-value < 0.0001) than that obtained at an aeration ratio of 1:5. Our finding revealed that conducting the bioprocess at 150 rpm did incite a higher significant increase (at P-value < 0.0001) in the level of ALK-PR23 compared to that obtained from the bioprocess conducted at 100 rpm. Conversely, no significant effect (at P-value = 0.855) could be evidenced on the level of ALK-PR23 upon conducting the bioprocess at 180 rpm compared to that obtained at 150 rpm (Fig. 2d). Consequently, agitation speed (150 rpm) (Fig. 2d) and aeration ratio (1:5) (Fig. 2c) were the best optimal conditions stimulating the highest significant production of ALK-PR23 (113.62 ± 1.8 U/mL and 177 ± 4.0 U/mL, respectively) compared to the other levels attained using the other settings.
As inferred from OVAT attempts, modified minimal core production medium with yeast extract concentration, incubation temperature (25 °C), agitation speed (150 rpm) and aeration ratio (1:5) were selected as the utmost proper parameters promoting ALK-PR23 production in the subsequent optimization experiments.
Placket-Burman design is a prevailing method that can be used extensively in the conception of biotechnological products to screen significant aspects from a large number of variables [47]. In this study, we employed a statistical approach PBD to screen the influential variables and chose their level to determine the maximum alkaline protease production using L. sphaericus strain AA6 EMCCN3080 in submerged fermentation by wheat bran. Twelve runs were carried out to investigate the impact of the seven factors at two different levels on the generation of proteases (Table 3). Substantial experimental levels of ALK-PR23 ranging from 9.07 to 120.21 U/mL were achieved upon executing twelve empirical PBD experiments as indicated in Table 3. When factors have significance and a negative impact, they are efficient for enzyme production; nonetheless, it is worth mentioning that the required quantity in PB experiments may be less than the indicated low level (− 1). Additionally, when an effect is positive, it is crucial to focus on the high level suggested larger value (+ 1). Pareto chart was outlined by Minitab software 17.3 as an informative tool to show the significance consequences of the seven tested independent variables on the process output (ALK-PR23 production), according to their P-values regardless the magnitude and the sign of the co-efficient for each independent variable (Fig. 3). Additionally, the main effect chart was outlined by Minitab software 17.3 to evaluate the weight (considering the magnitude and sign of the co-efficient), of each tested independent variable in relation to the process outcome (Fig. 4).
The multiple linear regression of PBD data (Table 4) demonstrated that the yeast extract concentration (X2), ammonium sulphate concentration (X3) and initial pH of the ALK-PR23 production medium (X5), imposed substantial gains (P ≤ 0.05) on ALK-PR23 production. The rule of thumb that R2 should be more than 0.7 and within the range of 0.7–0.99 is acceptable for application [48]. The computed co-efficient R2, which was 0.96, served as an estimate of the model validity. According to this R2 value, the regression model could account for 96% of the response variations. The best correlation between experimental and anticipated values was also conferred by the multiple correlation co-efficient R value of 0.98 as it was closer to 1.0. Moreover, the high relevance of the regression model was verified from the high model F-value of 14.01 and low model P-value of 0.01. To figure out the impact of the independent variables on ALK-PR23 production, the regression coefficients were estimated in terms of coded values and were considered to settle the 1st order full polynomial Eq. (4). A graph deciphering the predicted yield and the experimental yield was shown in Fig. S1.
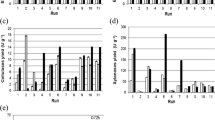
The steepest ascent/descent method is a process for going step-by-step along the path of steepest ascent/descent, or along the path of the greatest increase in the response. Starting in the middle of the PB design, the steepest ascending path travelled along the path that increased WB concentration and incubation time. Data revealed that WB at a concentration of 2% (w/v) was the most proper concentration prompting the highest significant level (P-value < 0.05) of ALK-PR23 (182.97 ± 2.8 U/mL) (Fig. 5a). Incubation time is one of the ultimate and substantial factors influencing any bioprocess including the production of enzymes. Furthermore, the interval of time required for any bioprocessing depends on a variety of factors, including the type of producer strain (bacteria or fungi), the mode of fermentation (batch, fed-batch, continuous), the type of fermentation (submerged liquid state fermentation or solid state fermentation), the composition of the substrate, etc. [29].
Figure 5b showed that maximum ALK-PR23 production (201.62 ± 0.5 U/mL) was achieved 3 days after the culture onset, indicating a correlation between enzyme production and bacterial growth. The findings demonstrate that L. sphaericus Strain AA6 EMCCN3080 growth was followed by maximal protease production, although with a slight delay that was likely caused by the time it took for the bacterial strain to acclimatize to the agro-industrial byproduct or waste [49].
As deduced from the steepest ascent experiments, WB concentration and incubation time settled at 2% (w/v) and 3 days, respectively were found to be the optimum values that prompted the utmost increase in ALK-PR23 production.
In the following optimization experiments, the independent variables (magnesium sulphate concentration and inoculum size) were left at their initial values because they clearly had no significant effects on the production of ALK-PR23. While the independent variables, yeast extract concentration (X2), ammonium sulphate concentration (X3) and initial pH of the production medium (X5) were submitted to BBD to pinpoint their ideal levels, evaluate their interrelation, and realize the highest measure of ALK-PR23. Moreover, Box-Behnken experimental strategy was used to generate a quadratic model consisting of fifteen runs with five center points. Table 5 describes the design matrix of three investigated parameters at different levels (− 1, 0, + 1) with experimental and theoretical expected response outcomes. The significant experimental levels of ALK-PR23, which were attained after executing fifteen empirical BBD trials ranged from 69.22 to 621.25 U/mL. The three tested independent variables yeast extract concentration (X2), ammonium sulphate concentration (X3) and initial pH of the production medium (X5) did enforce a significant influence (P ≤ 0.05) on the response in in the form of quadratic interaction (X2.X2), (X3.X3), (X5.X5) and cross interaction (X2.X3) (Table 6). The regression model coefficient R2 value of 0.93 indicated that it could account for 93% of the response unevenness. Also, as it gets closer to 1.0, the multiple correlation co-efficient R value of 0.965 does show the best correlation between the experimental and predictable values. On the other contrary, the model’s strong significance was deduced from its low P-value of 0.019 and high F-value of 7.55. The second order full polynomial Eq. (5) was developed using coded values to represent all likely possibilities of interactions within the independent variables and imposing significant impact on the production of ALK-PR23. A graph deciphering the predicted yield and the experimental yield was shown in Fig. S2.
Production of enzymes was influenced by pH of the production medium, which supports stabilizing enzyme structure and is crucial for the movement of substances across cell membranes, which supports cell development and product synthesis [50]. Regarding the positive significant impact of the initial pH of the production medium on ALK-PR23 by L. sphaericus Strain AA6 EMCCN3080, the present findings are in a good agreement with the previous findings on protease by Bacillus sp. KU-K2 (pH = 7) [11], B. nakamurai (pH = 6) PL4 [45] and S. acidaminiphila (pH = 7) [43]. Conversely, the optimum pH for protease production from B. licheniformis ALW1 and Citrococcus sp. was observed to be at 8 and 10, respectively [51, 52]
The nitrogen source also had an impact on the enzyme synthesis [53]. Therefore, by investigating the impact of ammonium sulphate as an inorganic nitrogen source in the current study, it was discovered that it had a positive significant impact on the maximum activity of ALK-PR23. Lakshmi et al. [54] reported a similar outcome for the alkaline protease generation by B. licheniformis MTCC NO. 7053.
In an attempt to reach the optimized conditions for ALK-PR23 production by L. sphaericus Strain AA6 EMCCN3080, the features of the response were studied by portraying the three-dimensional surface plots to better understand the link between the three tested variables, as well as to establish the optimal of the examined parameters in order to achieve the highest protease production. (Fig. 6a–c). The three-dimensional surface plots provide a proof that the stationary point has a saddle-like shape. Herein, the maximum stationary point could not be realized from the analysis of the three-dimensional surface plots. This would therefore make it mandatory to explore the maximum stationary point using the steepest ascent path via ridge analysis and the canonical path. The type of the predictable stationary point a saddle, minimum, or maximum was further confirmed and delineated by the ridge analysis. The eigenvectors and eigenvalues could provide proof of the response shape. The eigenvector could be used to infer the surface curvatures, however the real indication of the surface shape in these directions lies in the signs and magnitude of the eigenvalues. Earlier explanations of the eigenvalue concept and its mathematical relevance can be found in Myers (1976) [55].
Three-dimensional surface plots displaying the effect of three independent variables (X2: yeast extract concentration, X3: (NH4)2SO4 concentration, and X5: initial pH of the production medium) and their interactions on ALK-PR23 production by L. sphaericus Strain AA6 EMCCN3080. a Effect of using different levels of X2 and X3 at a constant level of X5. b Effect of using different levels of X2 and X5 at a constant level of X3. c Effect of using different levels of X3 and X5 at a constant level of X2. Independent variables in terms of coded values
The response downward and upward curvatures may be determined by the negative and positive eigenvalues, respectively (1st rule of Myers). The amplitude of an eigenvalue in its maximum absolute value implies the response curvature in the relevant direction, according to 2nd rule of Myers.
The RSM package analysis of the RSM data demonstrates that the ALK-PR23 production's RSM model has the following eigenvalues: [ λ2 = − 93.13221, λ3 = − 191.59843, and λ5 = − 266.97478]. According to Myers’ rules, the response would have a downward curvature in the direction of X5 (initial pH of the production medium), with the largest absolute eigenvalue. Additionally, the RSM package solver differentiated the second polynomial Eq. (5) and found that the projected stationary point was situated at X2 = 0.2473901, X3 = − 0.3197334, and X5 = − 0.1327876. The anticipated stationary point revealed two inferences. The stationary point was localized inside the design constraints, which was the first deduction. The anticipated stationary point was a maximum stationary point, according to the second deduction. By taking calculated steps in the right way starting from the design's center ridge analysis provided additional confirmation for the predictor combination set of the three evaluated independent variables (Table 7). The greatest anticipated level of the response (592.430 U/mL), as shown in Table 7, was produced by the combination set of predictors (X2 = 0.231, X3 = − 0.301, and X5 = − 0.128), localized at a distance of 0.4 from the design's center. The stationary point predicted by the ridge analysis was almost identical to the result of the RSM package's solver function. A drop in the response level, the acquisition of predicator combination sets situated outside the design constraints, and the prediction of three-dimensional surface plots with downward curvatures were all outcomes of going further in the positive direction from the center. The final inference took into account how closely the three-dimensional surface plots direction matched the three-dimensional plots shown in Fig. 6a–c. According to the model's laboratory validation, the actual experimental maximum concentration of ALK-PR23 was 593.6 U/mL, obtaining 100% validity. By the end of the optimization strategy, the optimal conditions for realizing the ultimate level of ALK-PR23 (i.e., 592.5 U/mL) were modified minimal medium containing WB 2% (w/v), yeast extract (0.446%w/v), (NH4)2SO4 (0.339%w/v), pH 6.872, 25 °C, and 3 days.
When compared to its level before the optimization technique, L. sphaericus Strain AA6 EMCCN3080 production of ALK-PR23 had increased 98-fold by its completion.
The high yield of the bioproduct and the low capital cost of this bioprocess are used to assess the economic viability of any industrialized bioprocess. The high capability of the chosen producer strain, at most, serves as the key determinant of a bioproduct's adequate yield in a specific bioprocess. Yet, to increase the yield of any bioprocess, it should be subjected to rigorous statistical optimization techniques. Here, a three-step sequential statistical strategy using OVAT, PBD, and BBD was implemented to maximize the yield of ALK-PR23 by L. sphaericus Strain AA6 EMCCN3080. Reportedly, experimental designs have been employed frequently to maximize the fermentation parameters responsible for enzyme synthesis [29,30,31], production of microbial pigments[32], antibiotics biodegradation [34], biosynthesis of chito-oligosaccharides [56], and biosynthesis pectic-oligosaccharides [57]. The 98-fold increase in ALK-PR23 yield over the actual yield achieved before optimization did much to support the feasibility of the employed optimization procedure to get a substantially improved and adequate yield of ALK-PR23.
The strategy of optimization (tree-like optimization method) is a multi-step approach working in a hierarchy like scaffold. Each step in the tree- like optimization method is a strict prerequisite for the next step. In other words, the shortcomings included in a certain step would be amended in the next step of the optimization strategy. Herein, the approach of OVAT, the root of the tree-like optimization method, is powerful in testing the key determinants controlling the ALK-PR23 production (process outcome) such as physical factors (agitation speed, aeration rate, and incubation temperature) and chemical factors (e.g., core medium composition (type of organic nitrogen source). Thus, it helped reduce the number of key determinants that would be studied in the next step of the optimization strategy (PBD). Herein, OVAT helped address the foundations (i.e., broadlines) of the bioprocess of ALK-PR23 production. However, OVAT approach is not a significant approach to test the independent variable within a given range of concentrations. It would necessitate multiple trials that are absolutely waste of time, waste of effort, and too expensive without realizing the ultimate level of the process outcome (ALK-PR23 production). Additionally, the yield of the process outcome in the OVAT step is a preliminary yield as long the interactions among variables are completely masked (i.e., disregarded). The next step in the tree-like optimization method (PBD) is a robust approach in investigating the co-linearity relationship among an unlimited number of independent variables regarding the concentration of each variable within an empirical defined range of concentration (design’s constrains) and the process outcome. The goal of PBD (i.e., the stem of the tree-like optimization method), is to unravel the nature of linear relationship, either in a linear descending manner or a linear ascending manner, between each independent variable and the process outcome in one experiment. Herein, the PBD model approved that the independent variables yeast extract concentration, ammonium sulphate concentration, and initial pH of the production medium exhibited positive significant influence (a linear ascending approach) on ALK-PR23 production. Likewise, OVAT approach, the PBD does not consider the interactions among the tested independent variables. Consequently, the yield derived from PBD experiments is not the ultimate one. The next step in the tree-like optimization method is BBD (i.e., tree branches). The BBD aims to localize the optimal level of each key determinant imposing significant influence on the process outcome. Herein, BBD could not localize the optimal concentration of the three key determinants: yeast extract concentration, ammonium sulphate concentration, and initial pH of the production medium in conjugation with the ultimate level of ALK-PR23. As shown in Fig. 6, the stationary point (ultimate ALK-PR23 level) could not be realized. Conversely, the predicted stationary point by BBD was nearby the stationary point. Herein, the predicted nearby stationary point predicted by BBD was saddle point (i.e., a flat plateau region where several predictor combination sets were localized in conjugation with almost quite similar ultimate level of ALK-PR23). To realize the maximum stationary point with the possible lowest optimal levels of predictor combination sets, ridge analysis was carried out. Herein, ridge analysis, last step in the tree-like optimization method, the leaf of the tree and the path end of the optimization method) did succeed to localize the optimal levels of the three key determinants: yeast extract concentration, ammonium sulphate concentration, and initial pH of the production medium along with the ultimate level of ALK-PR23 (i.e., 592.5 U/mL). Despite the advantages and shortcomings encountered in each step in the tree-like optimization method, the comparison among these steps, regarding the obtained yield in each step, are not justified yet as long the optimization process is a multistep and each step has a mandatory role.
The maximum yield of ALK-PR23, produced by L. sphaericus Strain AA6 EMCCN3080, reached 592.5 U/mL. Extremely lower levels of proteases were formerly reported by other bacterial strains including B. nakamurai PL4 (0.3 U/mL) [45], Citrococcus sp. (23.21 U/mL) [51] B. subtilis PTCC 1254 (117.43 U/mL) [58] Bacillus sp. NRD9 (138.17 U/mL) [59], L. fusiformis AU01 (35.6 U/mL) [60], B. aryabhattai Ab15-ES (247.84 U/mL) [61] and B. licheniformis ALW1 (22.9 U/mL) [52].
Conclusions
With the help of L. sphaericus Strain AA6 EMCCN3080, a pschyrotolerant locally isolated bacterial strain, the present study has revealed how to efficiently utilize the raw agro-industrial byproduct wheat bran as a zero-cost substrate and achieve superior protease levels compared to those obtained from other reported homologue producers for the first time. The scaling up of protease production by L. sphaericus Strain AA6 EMCCN3080 in prospective investigations is supported by the present data.
Data Availability
All data are available and included in the article.
References
Hell, J., Kneifel, W., Rosenau, T., Boehmdorfer, S.: Analytical techniques for the elucidation of wheat bran constituents and their structural features with emphasis on dietary fiber—a review. Trends Food Sci. Technol. 35, 102–113 (2014)
Javed, M.M., Zahoor, S., Shafaat, S., Mehmooda, I., Gul, A., Rasheed, H., Bukhari, S.A.I., Aftab, M.N., Haq, I.: Wheat bran as a brown gold: nutritious value and its biotechnological applications. Afr. J. Microbiol. Res. 6, 724–733 (2012)
Farzana, K., Shah, S.N., Butt, F.B., Awan, S.B.: Biosynthesis of bacitracin in solid-state fermentation by Bacillus licheniformis using defatted oil seed cakes as substrate. Pak. J. Pharm. Sci. 18, 55–57 (2005)
Palmarola-Adrados, B., Chotěborská, P., Galbe, M., Zacchi, G.: Ethanol production from non-starch carbohydrates of wheat bran. Bioresour. Technol. 96, 843–850 (2005)
Singh, K.K., Hasan, S.H., Talat, M., Singh, V.K., Gangwar, S.K.: Removal of Cr (VI) from aqueous solutions using wheat bran. Chem. Eng. J. 151, 113–121 (2009)
De Kock, S., Taylor, J., Taylor, J.R.N.: Effect of heat treatment and particle size of different brans on loaf volume of brown bread. LWT-Food Sci. Technol. 32, 349–356 (1999)
Rosenfelder, P., Eklund, M., Mosenthin, R.: Nutritive value of wheat and wheat by-products in pig nutrition: a review. Anim. Feed Sci. Technol. 185, 107–125 (2013)
Razzaq, A., Shamsi, S., Ali, A., Ali, Q., Sajjad, M., Malik, A., Ashraf, M.: Microbial proteases applications. Front. Bioeng. Biotechnol 7, 110 (2019)
Singhal, P., Nigam, V.K., Vidyarthi, A.S.: Studies on production, characterization and applications of microbial alkaline proteases. Int. J. Adv. Biotechnol. Res. 3, 653–669 (2012)
da Silva Bernardo, B., Kopplin, B.W., Daroit, D.J.: Bioconversion of fish scales and feather wastes by Bacillus sp. CL18 to obtain protease and bioactive hydrolysates. Waste Biomass Valoriz. (2022). https://doi.org/10.1007/s12649-022-01907-6
Lomthong, T., Suntornnimit, P., Sakdapetsiri, C., Trakarnpaiboon, S., Sawaengkaew, J., Kitpreechavanich, V.: Alkaline protease production by thermotolerant Bacillus sp. KU-K2, from non-rubber skim latex through the non-sterile system and its enzymatic characterization. Biocatal. Agric. Biotechnol. 46, 102542 (2022)
Wang, S.-L., Chang, T.-J., Liang, T.-W.: Conversion and degradation of shellfish wastes by Serratia sp. TKU016 fermentation for the production of enzymes and bioactive materials. Biodegradation 21, 321–333 (2010)
Vélez-Gómez, J.M., Melchor-Moncada, J.J., Veloza, L.A., Sepúlveda-Arias, J.C.: Purification and characterization of a metalloprotease produced by the C8 isolate of Serratia marcescens using silkworm pupae or casein as a protein source. Int. J. Biol. Macromol. 135, 97–105 (2019)
Dutta, J.R., Dutta, P.K., Banerjee, R.: Optimization of culture parameters for extracellular protease production from a newly isolated Pseudomonas sp. using response surface and artificial neural network models. Process Biochem. 39, 2193–2198 (2004)
Bhagwat, P.K., Jhample, S.B., Dandge, P.B.: Statistical medium optimization for the production of collagenolytic protease by Pseudomonas sp. SUK using response surface methodology. Microbiology (N Y) 84, 520–530 (2015)
Venugopal, M., Saramma, A.V.: Characterization of alkaline protease from Vibrio fluvialis strain VM10 isolated from a mangrove sediment sample and its application as a laundry detergent additive. Process Biochem. 41, 1239–1243 (2006)
De Azeredo, L.A.I., De Lima, M.B., Coelho, R.R.R., Freire, D.M.G.: Thermophilic protease production by Streptomyces sp. 594 in submerged and solid-state fermentations using feather meal. J. Appl. Microbiol. 100, 641–647 (2006)
Al-Dhabi, N.A., Esmail, G.A., Ghilan, A.-K.M., Arasu, M.V., Duraipandiyan, V., Ponmurugan, K.: Characterization and fermentation optimization of novel thermo stable alkaline protease from Streptomyces sp. Al-Dhabi-82 from the Saudi Arabian environment for eco-friendly and industrial applications. J. King Saud Univ.-Sci. 32, 1258–1264 (2020)
Osmolovskiy, A.A., Popova, E.A., Kreyer, V.G., Baranova, N.A., Egorov, N.S.: Vermiculite as a new carrier for extracellular protease production by Aspergillus spp. under solid-state fermentation. Biotechnol. Rep. 29, e00576 (2021)
Mamo, J., Kangwa, M., Fernandez-Lahore, H.M., Assefa, F.: Optimization of media composition and growth conditions for production of milk-clotting protease (MCP) from Aspergillus oryzae DRDFS13 under solid-state fermentation. Braz. J. Microbiol. 51, 571–584 (2020)
Guo, Y., Li, X., Jia, W., Huang, F., Liu, Y., Zhang, C.: Characterization of an intracellular aspartic protease (PsAPA) from Penicillium sp. XT7 and its application in collagen extraction. Food Chem. 345, 128834 (2021)
Guo, Y., Zhou, J., Jia, W., Gao, H., Zhang, H., Zhang, C.: Characterization of a novel milk-clotting aspartic protease from Penicillium sp. and structural explanation for its high Milk-Clotting Index. J. Agric. Food Chem. (2023). https://doi.org/10.1021/acs.jafc.2c07303
Rizvi, A., Ahmed, B., Khan, M.S., Umar, S., Lee, J.: Psychrophilic bacterial phosphate-biofertilizers: a novel extremophile for sustainable crop production under cold environment. Microorganisms 9, 2451 (2021)
Matrawy, A.A., Khalil, A.I., Embaby, A.M.: Molecular study on recombinant cold-adapted, detergent-and alkali stable esterase (EstRag) from Lysinibacillus sp.: a member of family VI. World J. Microbiol. Biotechnol. 38, 217 (2022)
Ahsan, N., Shimizu, M.: Lysinibacillus species: their potential as effective bioremediation, biostimulant, and biocontrol agents. Rev. Agric. Sci. 9, 103–116 (2021). https://doi.org/10.7831/ras.9.0_103
Mechri, S., Kriaa, M., Ben Elhoul Berrouina, M., Omrane Benmrad, M., Zaraî Jaouadi, N., Rekik, H., Bouacem, K., Bouanane-Darenfed, A., Chebbi, A., Sayadi, S., Chamkha, M., Bejar, S., Jaouadi, B.: Optimized production and characterization of a detergent-stable protease from Lysinibacillus fusiformis C250R. Int. J. Biol. Macromol. 101, 383–397 (2017). https://doi.org/10.1016/j.ijbiomac.2017.03.051
Jinal, H.N., Gopi, K., Prittesh, P., Kartik, V.P., Amaresan, N.: Phytoextraction of iron from contaminated soils by inoculation of iron-tolerant plant growth-promoting bacteria in Brassica juncea L. Czern. Environ. Sci. Pollut. Res. 26, 32815–32823 (2019). https://doi.org/10.1007/s11356-019-06394-2
Prabha, M., Divakar, K., Priya, J.D., Selvam, G., Subramanian, B., Gautam, P.: Statistical analysis of production of protease and esterase by a newly isolated Lysinibacillus fusiformis AU01: purification and application of protease in sub-culturing cell lines. Ann. Microbiol. 65, 1–14 (2014). https://doi.org/10.1007/s13213-014-0833-z
Matrawy, A.A., Khalil, A.I., Embaby, A.M.: Bioconversion of bread waste by marine psychrotolerant Glutamicibacter soli strain AM6 to a value-added product: cold-adapted, salt-tolerant, and detergent-stable α-amylase (CA-AM21). Biomass Convers. Biorefin. (2022). https://doi.org/10.1007/s13399-022-02325-3
Matrawy, A.A., Khalil, A.I., Marey, H.S., Embaby, A.M.: Use of wheat straw for value-added product xylanase by Penicillium chrysogenum strain A3 DSM105774. J. Fungi 7, 696 (2021)
Matrawy, A.A., Khalil, A.I., Marey, H.S., Embaby, A.M.: Biovalorization of the raw agro-industrial waste rice husk through directed production of xylanase by Thermomyces lanuginosus strain A3–1 DSM 105773: a statistical sequential model. Biomass Convers. Biorefin. 11, 2177–2189 (2021)
Embaby, A.M., Hussein, M.N., Hussein, A.: Monascus orange and red pigments production by Monascus purpureus ATCC16436 through co-solid state fermentation of corn cob and glycerol: an eco-friendly environmental low cost approach. PLoS ONE 13, e0207755 (2018)
Embaby, A.M., Heshmat, Y., Hussein, A., Marey, H.S.: A sequential statistical approach towards an optimized production of a broad spectrum bacteriocin substance from a soil bacterium Bacillus sp. YAS 1 strain. Sci. World J. 2014, 396304 (2014). https://doi.org/10.1155/2014/396304
Anan, A., Ghanem, K.M., Embaby, A.M., Hussein, A., El-Naggar, M.Y.: Statistically optimized ceftriaxone sodium biotransformation through Achromobacter xylosoxidans strain Cef6: an unusual insight for bioremediation. J. Basic Microbiol. 58, 120–130 (2018). https://doi.org/10.1002/jobm.201700497
Eden, P.A., Schmidt, T.M., Blakemore, R.P., Pace, N.R.: Phylogenetic analysis of Aquaspirillum magnetotacticum using polymerase chain reaction-amplified 16S rRNA-specific DNA. Int. J. Syst. Evol. Microbiol. 41, 324–325 (1991)
Hammami, A., Bayoudh, A., Abdelhedi, O., Nasri, M.: Low-cost culture medium for the production of proteases by Bacillus mojavensis SA and their potential use for the preparation of antioxidant protein hydrolysate from meat sausage by-products. Ann. Microbiol. 68, 473–484 (2018). https://doi.org/10.1007/s13213-018-1352-0
Plackett, R.L., Burman, J.P.: The design of optimum multifactorial experiments. Biometrika 33, 305–325 (1946)
Box, G.E.P., Behnken, D.W.: Some new three level designs for the study of quantitative variables. Technometrics 2, 455–475 (1960)
Madeira, J.V., Jr., Contesini, F.J., Calzado, F., Rubio, M.V., Zubieta, M.P., Lopes, D.B., de Melo, R.R.: Agro-industrial residues and microbial enzymes: an overview on the eco-friendly bioconversion into high value-added products. In: Biotechnology of microbial enzymes, pp. 475–511. Elsevier, Amsterdam (2017)
John, R.P., Nampoothiri, K.M., Pandey, A.: Simultaneous saccharification and L-(+)-lactic acid fermentation of protease-treated wheat bran using mixed culture of lactobacilli. Biotechnol. Lett. 28, 1823–1826 (2006). https://doi.org/10.1007/s10529-006-9159-7
de Albuquerque, M.F.G., Guimarães, V.M., de Rezende, S.T.: Use of sugar beet flour and wheat bran as carbon source improves the efficiency of Chrysoporthe cubensis enzymes in sugarcane bagasse saccharification. Bioenergy Res. 14, 1147–1160 (2021)
Sathishkumar, R., Ananthan, G., Arun, J.: Production, purification and characterization of alkaline protease by ascidian associated Bacillus subtilis GA CAS8 using agricultural wastes. Biocatal. Agric. Biotechnol. 4, 214–220 (2015). https://doi.org/10.1016/J.BCAB.2014.12.003
Asitok, A., Ekpenyong, M., Takon, I., Antai, S., Ogarekpe, N., Antigha, R., Edet, P., Ben, U., Akpan, A., Antai, A.: Overproduction of a thermo-stable halo-alkaline protease on agro-waste-based optimized medium through alternate combinatorial random mutagenesis of Stenotrophomonas acidaminiphila. Biotechnol. Rep. 35, e00746 (2022)
Wang, Y., Sun, J., Deng, Y., Tu, Y., Niu, H., Cai, W., Han, X.: Whey protein influences the production and activity of extracellular protease from Pseudomonas fluorescens W3. LWT 154, 112865 (2022). https://doi.org/10.1016/J.LWT.2021.112865
Shaikh, I.A., Turakani, B., Malpani, J., Goudar, S.V., Mahnashi, M.H., Hamed Al-Serwi, R., Ghoneim, M.M., El-Sherbiny, M., Abdulaziz Mannasaheb, B., Alsaikhan, F., Sindagimath, V., Khan, A.A., Muddapur, U.M., Azzouz, S., Mohammed, T., Shakeel Iqubal, S.M.: Extracellular protease production, optimization, and partial purification from Bacillus nakamurai PL4 and its applications. J. King Saud Univ. Sci. 35, 102429 (2023). https://doi.org/10.1016/J.JKSUS.2022.102429
Potumarthi, R., Subhakar, C., Jetty, A.: Alkaline protease production by submerged fermentation in stirred tank reactor using Bacillus licheniformis NCIM-2042: effect of aeration and agitation regimes. Biochem. Eng. J. 34, 185–192 (2007). https://doi.org/10.1016/J.BEJ.2006.12.003
Heydari, M., Karimyan, K., Darvishmotevalli, M., Karami, A., Vasseghian, Y., Azizi, N., Ghayebzadeh, M., Moradi, M.: Data for efficiency comparison of raw pumice and manganese-modified pumice for removal phenol from aqueous environments—application of response surface methodology. Data Brief 20, 1942–1954 (2018)
Thite, V.S., Nerurkar, A.S., Baxi, N.N.: Optimization of concurrent production of xylanolytic and pectinolytic enzymes by Bacillus safensis M35 and Bacillus altitudinis J208 using agro-industrial biomass through response surface methodology. Sci. Rep. 10, 1–12 (2020)
Mladenoska, I., Dimitrovski, A.: Microbial production of lipases on media containing vegetable oil waste: process development. Chem. Eng. (2013). https://doi.org/10.3303/CET1334018
Ellaiah, P., Adinarayana, K., Bhavani, Y., Padmaja, P., Srinivasulu, B.: Optimization of process parameters for glucoamylase production under solid state fermentation by a newly isolated Aspergillus species. Process Biochem. 38, 615–620 (2002)
Verma, J., Pandey, S.: Characterization of partially purified alkaline protease secreted by halophilic bacterium Citricoccus sp. isolated from agricultural soil of northern India. Biocatal. Agric. Biotechnol. 17, 605–612 (2019). https://doi.org/10.1016/J.BCAB.2019.01.020
Emran, M.A., Ismail, S.A., Hashem, A.M.: Production of detergent stable thermophilic alkaline protease by Bacillus licheniformis ALW1. Biocatal. Agric. Biotechnol. 26, 101631 (2020). https://doi.org/10.1016/J.BCAB.2020.101631
Ward, N.E.: With dietary modifications, wheat can be used for poultry. Feedstuffs (1995)
Lakshmi, B.K.M., Ratna Sri, P.V., Ambika Devi, K., Hemalatha, K.P.J.: Media optimization of protease production by Bacillus licheniformis and partial characterization of alkaline protease. Int. J. Curr. Microbiol. App. Sci. 3, 650–659 (2014)
Myers, R.H.: Response surface methodology. Edwards Brothers, Ann Arbor (1976)
Embaby, A.M., Melika, R.R., Hussein, A., El-Kamel, A.H., Marey, H.S.: Biosynthesis of chitosan-oligosaccharides (COS) by non-aflatoxigenic Aspergillus sp. strain EGY1 DSM 101520: a robust biotechnological approach. Process Biochem. 64, 16–30 (2018). https://doi.org/10.1016/J.PROCBIO.2017.09.030
Embaby, A.M., Melika, R.R., Hussein, A., El-Kamel, A.H., Marey, H.S.: A novel non-cumbersome approach towards biosynthesis of pectic-oligosaccharides by non-aflatoxigenic Aspergillus sp. section flavi strain EGY1 DSM 101520 through citrus pectin fermentation. PLoS ONE 11, e0167981 (2016)
Jafari, Z., Najafpour, G., Zare, H.: Growth media optimization for production of alkaline protease from industrial wastewater using Bacillus subtilis PTCC 1254. Int. J. Eng. 36, 513–522 (2023). https://doi.org/10.5829/IJE.2023.36.03C.11
Suberu, Y., Akande, I., Samuel, T., Lawal, A., Olaniran, A.: Optimization of protease production in indigenous Bacillus species isolated from soil samples in Lagos, Nigeria using response surface methodology. Biocatal. Agric. Biotechnol. 18, 101011 (2019). https://doi.org/10.1016/J.BCAB.2019.01.049
Suryia Prabha, M., Divakar, K., Deepa Arul Priya, J., Panneer Selvam, G., Balasubramanian, N., Gautam, P.: Statistical analysis of production of protease and esterase by a newly isolated Lysinibacillus fusiformis AU01: purification and application of protease in sub-culturing cell lines. Ann. Microbiol. 65, 33–46 (2015). https://doi.org/10.1007/s13213-014-0833-z
Adetunji, A.I., Olaniran, A.O.: Statistical modelling and optimization of protease production by an autochthonous Bacillus aryabhattai Ab15-ES: a response surface methodology approach. Biocatal. Agric. Biotechnol. 24, 101528 (2020). https://doi.org/10.1016/J.BCAB.2020.101528
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). Not applicable.
Author information
Authors and Affiliations
Contributions
AM wrote the manuscript and performed all laboratory work. AM put the idea of the research, analyzed the lab data, and revised the manuscript. AE put the idea of the research, analyzed the lab data, wrote, and revised the manuscript. HM: performed the statistical ridge analysis. AM and AE approved the final form of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that there is no any conflict of interest.
Consent for Participation
Not applicable.
Consent for Publication
Not applicable.
Ethical Approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Matrawy, A.A., Marey, H.S. & Embaby, A.M. The Agro-industrial Byproduct Wheat Bran as an Inducer for Alkaline Protease (ALK-PR23) Production by Pschyrotolerant Lysinibacillus sphaericus Strain AA6 EMCCN3080. Waste Biomass Valor 15, 1943–1958 (2024). https://doi.org/10.1007/s12649-023-02283-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12649-023-02283-5