Abstract
Recombination rate data are presented for three populations of grape based on framework genetic linkage maps developed with simple-sequence repeat markers. These linkage maps were constructed from different Vitis species and represent three genetic backgrounds. The first population is pure Vitis vinifera, derived from a cross of the European cultivars Riesling and Cabernet Sauvignon. The second is an interspecific cross between two commercially used rootstock cultivars of different North American Vitis species parentage, Ramsey (Vitis champinii) and Riparia Gloire (Vitis riparia). The third population, D8909-15 (Vitis rupestris × (Vitis arizonica/Vitis girdiana form)) × F8909-17 (V. rupestris × (V. arizonica/Vitis candicans form)), is an F1 from two half-sibs. Genome-wide and chromosome-wide recombination rates varied across the three populations and among the six Vitis parents. Global recombination rates in the parents of the third F1 population, with a complex Vitis background, were significantly reduced. In the first and third populations, the recombination rate was significantly greater in the male parent. Specific genome locations with frequent heterogeneity in recombination were identified, suggesting that recombination rates are not equal across the Vitis genome. The identification of regions with suppressed or high recombination will aid grape breeders and geneticists who rely on recombination events to introgress disease resistance genes from the genomes of wild Vitis species, develop fine-scale genetic maps, and clone disease resistance genes.

Similar content being viewed by others
References
Adam-Blondon A-F, Roux C, Claux D, Butterlin G, Merdinoglu D, This R (2004) Mapping 245 SSR markers on the Vitis vinifera genome: a tool for grape genetics. Theor Appl Genet 109:1017–1027
Akhunov ED, Goodyear AW, Geng S, Qi L-L, Echalier B, Gill BS et al (2003) The organization and rate of evolution of wheat genomes are correlated with recombination rates along chromosome arms. Genome Res 13:753–763
Anderson LK, Hooker KD, Stack SM (2001) The distribution of early recombination nodules on zygotene bivalents from plants. Genetics 159:1259–1269
Anderson LK, Lai A, Stack SM, Rizzon C, Gaut BS (2006) Uneven distribution of expressed sequence tag loci on maize pachytene chromosomes. Genome Res 16:115–122
Barker CL, Donald T, Pauquet J, Ratnaparkhe MB, Bouquet A, Adam-Blondon A-F Thomas MR, Dry I (2005) Genetic and physical mapping of the grapevine powdery mildew resistance gene, Run1, using a bacterial artificial chromosome library. Theor Appl Genet 111:370–377
Baudry E, Kerdelhué C, Innan H, Stephan W (2001) Species and recombination effects on DNA variability in the tomato genus. Genetics 158:1725–1735
Busso CS, Liu CJ, Hash CT, Witcombe JR, Devos KM, de Wet JMJ, Gale MD (1995) Analysis of recombination rate in female and male gametogenesis in pear millet (Pennisetum glaucum) using RFLP markers. Theor Appl Genet 90:242–246
Chakravarti A, Laher LK, Reefer JE (1991) A maximum likelihood method for estimating genome length using genetic linkage data. Genetics 128:175–182
Dalbó MA, Ye GN, Weeden NG, Steinkellner H, Sefc KM, Reisch BI (2000) A gene controlling sex in grapevines placed on a molecular-based genetic map. Genome 43:333–340
Devaux P, Kilian A, Kleinhofs A (1995) Comparative mapping of the barley genome with male and female recombination-derived double haploid populations. Mol Gen Genet 249:600–608
de Vicente MC, Tanksley SD (1991) Genome-wide reduction in recombination of backcross progeny derived from male versus female gametes in an interspecific cross of tomato. Theor Appl Genet 83:173–178
Di Gaspero G, Peterlunger E, Testolin R, Edwards KJ, Cipriani G (2000) Conservation of microsatellite loci within the genus Vitis. Theor Appl Genet 106:163–172
Di Gaspero G, Cipriani G, Adam-Blondon A-F, Testolin R (2007) Linkage maps of grapevine displaying the chromosomal locations of 420 microsatellite markers and 82 markers for R-gene candidates. Theor Appl Genet 114:1249–1263
Doligez A, Adam Blondon A-F, Cipriani G, Di Gaspero G, Laucou V, Merdinoglu D et al (2006) An integrated SSR map of grapevine based on five mapping populations. Theor Appl Genet 113:369–382
Dooner HK, Martinez-Ferez IM (1997) Germinal excisions of the maize transposon activator do not stimulate meiotic recombination or homology-dependent repair at the bz locus. Genetics 147:1923–1932
Doucleff M, Jin Y, Gao F, Riaz S, Krivanek AF, Walker MA (2004) A genetic linkage map of grape, utilizing Vitis rupestris and Vitis arizonica. Theor Appl Genet 109:1178–1187
Drouaud J, Camilleri C, Bourguignon P-Y, Canaguier A et al (2006) Variation in crossing-over rates across chromosome 4 of Arabidopsis thaliana reveals the presence of meiotic recombination “hot spots”. Genome Res 16:106–114
Dvorák J, Luo M-C, Yang Z-L (1998) Restriction fragment length polymorphism and divergence in the genomic regions of high and low recombination in self-fertilizing and cross-fertilizing Aegilops species. Genetics 148:423–434
Fischer BM, Salakhutdinov I, Akkurt M, Eibach R, Edwards KJ, Töpfer R, Zyprian EM (2004) Quantitative trait locus analysis of fungal disease resistance factors on a molecular map of grapevine. Theor Appl Genet 108:501–515
Ganal MW, Tanksley SD (1996) Recombination around the Tm2a and Mi resistance genes in different crosses of Lycopersicon peruvianum. Theor Appl Genet 92:101–108
Gaut BS, Wright SI, Rizzon C, Dvorak J, Anderson LK (2007) Recombination: an underappreciated factor in the evolution of plant genomes. Nat Rev Genet 8:77–84
Grando MS, Bellin D, Edwards KJ, Pozzi C, Stefanini M, Velasco R (2003) Molecular linkage maps of Vitis vinifera L. and Vitis riparia Mchx. Theor Appl Genet 106:1213–1224
Groover AT, Williams CG, Devey ME, Lee JM, Neale DB (1995) Sex-related differences in meiotic recombination frequency in Pinus taeda. J Hered 86:157–158
Hemmat M, Weeden NF, Manganaris AG, Lawson DM (1994) Molecular marker linkage map for apple. J Hered 85:4–11
Jaillon O, Aury JM, Noel B, Policriti A, Clepet C, Casagrande A, Choisne N, Aubourg S, Vitulo N, Jubin C, Vezzi A, Legeai F, Hugueney P, Dasilva C, Horner D, Mica E, Jublot D, Poulain J, Bruyere C, Billault A, Segurens B, Gouyvenoux M, Ugarte E, Cattonaro F, Anthouard V, Vico V, Del Fabbro C, Alaux M, Di Gaspero G, Dumas V, Felice N, Paillard S, Juman I, Moroldo M, Scalabrin S, Canaguier A, Le Clainche I, Malacrida G, Durand E, Pesole G, Laucou V, Chatelet P, Merdinoglu D, Delledonne M, Pezzotti M, Lecharny A, Scarpelli C, Artiguenave F, Pe ME, Valle G, Morgante M, Caboche M, Adam-Blondon AF, Weissenbach J, Quetier F, Wincker P (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–467
Jensen-Seaman MI, Furey TS, Payseur BA, Lu Y, Roskin KM, Chen CF, Thomas MA, Haussler D, Jacob H (2004) Comparative recombination rates in the rat, mouse, and human genomes. Genome Res 14:528–538
Kraft T, Säll T, Magnusson-Rading I, Nilsson N-O, Halldén C (1998) Positive correlation between recombination rates and levels of genetic variation in natural populations of sea beet (Beta vulgaris subsp. maritima). Genetics 150:1239–1244
Lamoureux D, Bernole A, Le Clainche I, Tual S, Thareau V, Paillard S, Legeai F, Dossat C, Wincker P, Oswald M, Merdinoglu D, Vignault C, Delrot S, Caboche M, Chalhoub B, Adam-Blondon A-F (2006) Anchoring of a large set of markers onto a BAC library for the development of a draft physical map of the grapevine genome. Theor Appl Genet 113:344–356
Lenormand T, Dutheil J (2005) Recombination difference between sexes: a role for haploid selection. PLoS Biol 3:396–402
Lodhi MA, Reisch BI (1995) Nuclear DNA content of Vitis species, cultivars, and other genera of the Vitaceae. Theor Appl Genet 90:11–16
Lodhi MA, Ye GN, Weeden NF, Reisch BI (1995) A molecular marker-based linkage map of Vitis. Genome 38:786–794
Lowe KM, Walker MA (2006) Genetic linkage map of the interspecific grape rootstock cross Ramsey (Vitis champinii) × Riparia Gloire (Vitis riparia). Theor Appl Genet 112:1582–1592
Lu H, Romero-Severson J, Bernardo R (2002) Chromosomal regions associated with segregation distortion in maize. Theor Appl Genet 105:622–628
Maliepaard C, Jansen J, Van Ooijen JW (1997) Linkage analysis in a full-sib family of an outbreeding plant species: overview and consequences for applications. Genet Res 70:237–250
Moran GF, Bell JC, Hilliker AJ (1983) Greater meiotic recombination in male vs. female gametes in Pinus radiata. Heredity 74:62
Nachman MW (2002) Variation in recombination rate across the genome: evidence and implications. Curr Opin Genet Dev 12:657–663
Negi SS, Olmo HP (1966) Sex conversion in male Vitis vinifera L. by a kinin. Science 152:1624–1625
Olmo HP (1995) Grapes. In: Smartt J, Simmonds NW (eds) Evolution of crop plants, 2nd edn. Longman Group, UK
Opperman R, Emmanuel E, Levy AA (2004) The effect of sequence divergence on recombination between direct repeats in Arabidopsis. Genetics 168:2207–2215
Pauquet J, Bouquet A, This P, Adam-Blondon A-F (2001) Establishment of a local map of AFLP markers around the powdery mildew resistance gene Run1 in grapevine and assessment of their usefulness for marker assisted selection. Theor Appl Genet 103:1201–1210
Peng J, Korol AB, Fahima T, Roder M, Ronin YI, Li YC, Nevo E (2000) Molecular genetic maps in wild emmer wheat, Triticum dicoccoides: genome-wide coverage, massive negative interference, and putative quasi-linkage. Genome Res 10:1509–1531
Qi LL, Echalier B, Chao S, Lazo GR, Butler GE et al (2004) A chromosome bin map of 16,000 Expressed sequence tag loci and distribution of genes among the three genomes of polyploid wheat. Genetics 168:701–712
Riaz S, Dangl GS, Edwards KJ, Meredith CP (2004) A microsatellite marker based framework linkage map of Vitis vinifera L. Theor Appl Genet 108:864–872
Riaz S, Krivanek AF, Xu K, Walker MA (2006) Refined mapping of the Pierce’s disease resistance locus, PdR1, and Sex on an extended genetic linkage map of Vitis rupestris × V. arizonica. Theor Appl Genet 113:1317–1329
Robertson DS (1984) Different frequency in the recovery of crossover products from male and female gametes of plants hypoploid for B-A translocations in maize. Genetics 107:117–130
Serre D, Nadon R, Hudson TJ (2005) Large-scale recombination rate patterns are conserved among human populations. Genome Res 15:1547–1552
Sherman JD, Stack SM (1995) Two dimensional spreads of synaptonemal complexes from Solanaceous plants: high resolution recombination nodule map for tomato (Lycopersicon esculentum). Genetics 141:683–708
Stirling B, Newcombe G, Vrebalov J, Bosdet I, Bradshaw HD Jr. (2001) Suppressed recombination around the MXC3 locus, a major gene for resistance to poplar leaf rust. Theor Appl Genet 103:1129–1137
Taylor DR, Ingvarsson PK (2003) Common features of segregation distortion in plants and animals. Genetica 117:27–35
Thomas MR, Matsumoto S, Cain P, Scott NS (1993) Repetitive DNA of grapevine—classes present and sequences suitable for cultivar identification. Theor Appl Genet 86:173–180
Troggio M, Malacarne G, Coppola G, Segala C, Cartwright DA, Pindo M et al (2007) A dense SNP-based genetic linkage map of grapevine (Vitis vinifera L.) anchoring Pinot Noir BAC contigs. Genetics 176:2637–2650
Van Oojen JW, Voorips RE (2001) JoinMap 3.0, software for the calculation of genetic linkage maps. Plant Research International, Wageningen, The Netherlands
Velasco R, Zharkikh A, Troggio M, Cartwright DA, Cestaro A, Pruss D, Pindo M, FitzGerald LM, Vezzulli S, Reid J, Malacarne G, Iliev D, Coppola G, Wardell B, Micheletti D, Macalma TM, Facci M, Mitchell JT, Perazzolli M, Eldredge G, Gatto P, Oyzerski R, Moretto M, Gutin N, Stefanini M, Chen Y, Segala C, Davenport C, Demattč L, Mraz A, Battilana J, Stormo K, Costa F, Tao Q, Si-Ammour A, Harkins T, Lackey A, Perbost C, Taillon B, Stella A, Solovyev V, Fawcett JA, Sterck L, Vandepoele K, Grando MS, Toppo S, Moser C, Lanchbury J, Bogden R, Skolnick M, Sgaramella V, Bhatnagar SK, Fontana P, Gutin A, Van de Peer Y, Salamini F, Viola R (2007) A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS ONE 2(12):e1326 doi:10.1371/journal.pone.0001326
Vizir Y, Korol B (1990) Sex difference in recombination frequency in Arabidopsis. Heredity 65:379–383
Waghmare VN, Rong J, Rogers CJ, Pierce GJ, Wendel JF, Paterson AH (2005) Genetic mapping of a cross between Gossypium hirsutum (cotton) and the Hawaiian endemic, Gossypium tomentosum. Theor Appl Genet 111:665–676
Wei F, Gobleman-Werner K, Morrol SM, Kurth J, Mao L, Wing R et al (1999) The Mla (powdery mildew) resistance cluster is associated with three NBS-LRR gene families and suppressed recombination within a 240-kb DNA interval on chromosome 5S (1HS) of barley. Genetics 153:1929–1948
Welter LJ, Göktürk-Baydar N, Akkurt M, Maul E, Eibach R, Töpfer R, Zyprian EM (2007) Genetic mapping and localization of quantitative trait loci affecting fungal disease resistance and leaf morphology in grapevine (Vitis vinifera L). Mol breeding 20:359–374
Wu J, Mizuno H, Hayahi-Tsugane M, Ito Y, Chiden Y, Fujisawa M et al (2003) Physical maps and recombination frequency of six rice chromosomes. Plant J 36:720–730
Xu Y, Zhu L, Xiao J, Huang N, McCouch SR (1997) Chromosomal regions associated with segregation distortion of molecular markers in F2, backcross, double haploid and recombinant inbred populations of rice (Oryza sativa L.). Mol Gen Genet 253:535–545
Xu K, Riaz S, Roncoroni NC, Jin Y, Hu R, Zhour R, Walker MA (2008) Genetic and QTL analysis of resistance to Xiphinena index in a grapevine cross. Theor Appl Genet 116:305–311
Zhang L, Gaut BS (2003) Does recombination shape the distribution and evolution of tandemly arrayed genes (TAGs) in the Arabidopsis thaliana genome? Genome Res 13:2533–2540
Acknowledgements
The authors gratefully acknowledge project funding from the California Grape Rootstock Improvement Commission, the California Department of Food and Agriculture Pierce’s Disease Board, and the Louis P. Martini Endowed Chair funds. We are also thankful for scholarship support to K.M. Lowe from the Tchelistcheff Fund, the American Society of Enology and Viticulture, the American Wine Society, and UC Davis Viticulture and Enology scholarships.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Velasco
Electronic Supplementary Material
Below is the link to the electronic supplementary material:
Supplemental Fig. 1
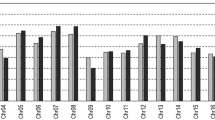
Pair-wise comparisons of the number of common marker intervals with significantly different recombination frequencies (RFs) between each Vitis map (Fig. 1a–d). Linkage groups (1–19) are presented on the X-axis, and significant marker intervals are on the Y-axis. (JPG 1.68 MB)
Rights and permissions
About this article
Cite this article
Lowe, K.M., Riaz, S. & Walker, M.A. Variation in recombination rates across Vitis species. Tree Genetics & Genomes 5, 71–80 (2009). https://doi.org/10.1007/s11295-008-0187-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-008-0187-4