Abstract
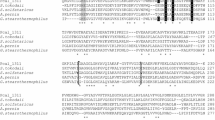
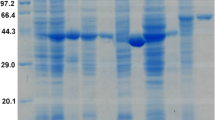
NAD(P)+-dependent alcohol dehydrogenases (ADH) are widely distributed in all phyla. These proteins can be assigned to three nonhomologous groups of isozymes, with group III being highly diverse with regards to catalytic activity and primary structure. Members of group III ADHs share a conserved stretch of amino acid residues important for cofactor binding and metal ion coordination, while sequence identities for complete proteins are highly diverse (<20 to >90 %). A putative group III ADH PaYqhD has been identified in BLAST analysis from the plant pathogenic enterobacterium Pectobacterium atrosepticum. The PaYqhD gene was expressed in the heterologous host Escherichia coli, and the recombinant protein was purified in a two-step purification procedure to homogeneity indicating an obligate dimerization of monomers. Four conserved amino acid residues involved in metal ion coordination were substituted with alanine, and their importance for catalytic activity was confirmed by circular dichroism spectrum determination, in vitro, and growth experiments. PaYqhD exhibits optimal activity at 40 °C with short carbon chain aldehyde compounds and NADPH as cofactor indicating the enzyme to be an aldehyde reductase. No oxidative activities towards alcoholic compounds were detectable. EDTA completely inhibited catalytic activity and was fully restored by the addition of Co2+. Activity measurements together with sequence alignments and structure analysis confirmed that PaYqhD belongs to the butanol dehydrogenase-like enzymes within group III of ADHs.








Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Atsumi S, Li Z, Liao JC (2009) Acetolactate synthase from Bacillus subtilis serves as a 2-ketoisovalerate decarboxylase for isobutanol biosynthesis in Escherichia coli. Appl Environ Microbiol 75:6306–6311
Atsumi S, Wu TY, Eckl EM, Hawkins SD, Buelter T, Liao JC (2010) Engineering the isobutanol biosynthetic pathway in Escherichia coli by comparison of three aldehyde reductase/alcohol dehydrogenase genes. Appl Microbiol Biotechnol 85:651–657
Bell KS, Sebaihia M, Pritchard L, Holden MT, Hyman LJ, Holeva MC, Thomson NR, Bentley SD, Churcher LJ, Mungall K, Atkin R, Bason N, Brooks K, Chillingworth T, Clark K, Doggett J, Fraser A, Hance Z, Hauser H, Jagels K, Moule S, Norbertczak H, Ormond D, Price C, Quail MA, Sanders M, Walker D, Whitehead S, Salmond GP, Birch PR, Parkhill J, Toth IK (2004) Genome sequence of the enterobacterial phytopathogen Erwinia carotovora subsp. atroseptica and characterization of virulence factors. Proc Natl Acad Sci U S A 101:11105–11110
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Conway T, Sewell GW, Osman YA, Ingram LO (1987) Cloning and sequencing of the alcohol dehydrogenase II gene from Zymomonas mobilis. J Bacteriol 169:2591–2597
Deng Y, Wang Z, Gu S, Ji C, Ying K, Xie Y, Mao Y (2002) Cloning and characterization of a novel human alcohol dehydrogenase gene (ADHFe1). DNA Seq 13:301–306
Elleuche S, Antranikian G (2013) Bacterial group III alcohol dehydrogenases—function, evolution and biotechnological applications. OA Alcohol 1:3
Elleuche S, Döring K, Pöggeler S (2008) Minimization of a eukaryotic mini-intein. Biochem Biophys Res Commun 366:239–243
Elleuche S, Fodor K, Klippel B, von der Heyde A, Wilmanns M, Antranikian G (2013a) Structural and biochemical characterisation of a NAD+-dependent alcohol dehydrogenase from Oenococcus oeni as a new model molecule for industrial biotechnology applications. Appl Microbiol Biotechnol 97:8963–8975
Elleuche S, Klippel B, von der Heyde A, Antranikian G (2013b) Comparative analysis of two members of the metal ion-containing group III-alcohol dehydrogenases from Dickeya zeae. Biotechnol Lett 35:725–733
Fenghuan W, Huijin Q, He H, Tan T (2005) High-level expression of the 1,3-propanediol oxidoreductase from Klebsiella pneumoniae in Escherichia coli. Mol Biotechnol 31:211–219
Hernandez-Tobias A, Julian-Sanchez A, Pina E, Riveros-Rosas H (2011) Natural alcohol exposure: is ethanol the main substrate for alcohol dehydrogenases in animals? Chem Biol Interact 191:14–25
Horn U, Strittmatter W, Krebber A, Knupfer U, Kujau M, Wenderoth R, Muller K, Matzku S, Pluckthun A, Riesenberg D (1996) High volumetric yields of functional dimeric miniantibodies in Escherichia coli, using an optimized expression vector and high-cell-density fermentation under non-limited growth conditions. Appl Microbiol Biotechnol 46:524–532
Jarboe LR (2011) YqhD: a broad-substrate range aldehyde reductase with various applications in production of biorenewable fuels and chemicals. Appl Microbiol Biotechnol 89:249–257
Lan EI, Liao JC (2012) ATP drives direct photosynthetic production of 1-butanol in cyanobacteria. Proc Natl Acad Sci U S A 109:6018–6023
Lee C, Kim I, Lee J, Lee KL, Min B, Park C (2010) Transcriptional activation of the aldehyde reductase YqhD by YqhC and its implication in glyoxal metabolism of Escherichia coli K-12. J Bacteriol 192:4205–4214
Lee C, Park C (2012) Development of a suicidal vector-cloning system based on butanal susceptibility due to an expression of YqhD aldehyde reductase. J Microbiol 50:249–255
Mansfield J, Genin S, Magori S, Citovsky V, Sriariyanum M, Ronald P, Dow M, Verdier V, Beer SV, Machado MA, Toth I, Salmond G, Foster GD (2012). Top 10 plant pathogenic bacteria in molecular plant pathology. Mol Plant Pathol 13(6):614-629
Marcal D, Rego AT, Carrondo MA, Enguita FJ (2009) 1,3-Propanediol dehydrogenase from Klebsiella pneumoniae: decameric quaternary structure and possible subunit cooperativity. J Bacteriol 191:1143–1151
Miller EN, Jarboe LR, Turner PC, Pharkya P, Yomano LP, York SW, Nunn D, Shanmugam KT, Ingram LO (2009) Furfural inhibits growth by limiting sulfur assimilation in ethanologenic Escherichia coli strain LY180. Appl Environ Microbiol 75:6132–6141
Montella C, Bellsolell L, Perez-Luque R, Badia J, Baldoma L, Coll M, Aguilar J (2005) Crystal structure of an iron-dependent group III dehydrogenase that interconverts L-lactaldehyde and L-1,2-propanediol in Escherichia coli. J Bacteriol 187:4957–4966
Moon JH, Lee HJ, Park SY, Song JM, Park MY, Park HM, Sun J, Park JH, Kim BY, Kim JS (2011) Structures of iron-dependent alcohol dehydrogenase 2 from Zymomonas mobilis ZM4 with and without NAD+ cofactor. J Mol Biol 407:413–424
Perez JM, Arenas FA, Pradenas GA, Sandoval JM, Vasquez CC (2008) Escherichia coli YqhD exhibits aldehyde reductase activity and protects from the harmful effect of lipid peroxidation-derived aldehydes. J Biol Chem 283:7346–7353
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Radianingtyas H, Wright PC (2003) Alcohol dehydrogenases from thermophilic and hyperthermophilic archaea and bacteria. FEMS Microbiol Rev 27:593–616
Reid MF, Fewson CA (1994) Molecular characterization of microbial alcohol dehydrogenases. Crit Rev Microbiol 20:13–56
Ruzheinikov SN, Burke J, Sedelnikova S, Baker PJ, Taylor R, Bullough PA, Muir NM, Gore MG, Rice DW (2001) Glycerol dehydrogenase: structure, specificity, and mechanism of a family III polyol dehydrogenase. Structure 9:789–802
Seo JW, Seo MY, Oh BR, Heo SY, Baek JO, Rairakhwada D, Luo LH, Hong WK, Kim CH (2010) Identification and utilization of a 1,3-propanediol oxidoreductase isoenzyme for production of 1,3-propanediol from glycerol in Klebsiella pneumoniae. Appl Microbiol Biotechnol 85:659–666
Sulzenbacher G, Alvarez K, Van Den Heuvel RH, Versluis C, Spinelli S, Campanacci V, Valencia C, Cambillau C, Eklund H, Tegoni M (2004) Crystal structure of E. coli alcohol dehydrogenase YqhD: evidence of a covalently modified NADP coenzyme. J Mol Biol 342:489–502
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Vaidyanathan H, Kandasamy V, Gopal Ramakrishnan G, Ramachandran K, Jayaraman G, Ramalingam S (2011) Glycerol conversion to 1, 3-propanediol is enhanced by the expression of a heterologous alcohol dehydrogenase gene in Lactobacillus reuteri. AMB Express 1:37
Walter KA, Bennett GN, Papoutsakis ET (1992) Molecular characterization of two Clostridium acetobutylicum ATCC 824 butanol dehydrogenase isozyme genes. J Bacteriol 174:7149–7158
Zhang G, Yang G, Li J (2011) Purification and characterization of a novel 1,3-propanediol oxidoreductase from Klebsiella oxytoca. Afr J Biotechnol 10:14913–14919
Acknowledgments
The authors would like to express their gratitude to Henning Piascheck for his help with the fermentations. Stéphane Boivin and Marvin Boothby at the “Sample Preparation and Characterization” facility of the EMBL at Hamburg are thanked for their help with the CD measurements. This work was funded by the Excellence Cluster in the Excellence Initiative by the State of Hamburg “Fundamentals of Synthetic Biological Systems (SynBio).”
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 596 kb)
Rights and permissions
About this article
Cite this article
Elleuche, S., Fodor, K., von der Heyde, A. et al. Group III alcohol dehydrogenase from Pectobacterium atrosepticum: insights into enzymatic activity and organization of the metal ion-containing region. Appl Microbiol Biotechnol 98, 4041–4051 (2014). https://doi.org/10.1007/s00253-013-5374-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-5374-z