Abstract
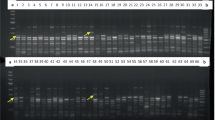
Diversity Arrays Technology (DArT) is a DNA hybridisation-based molecular marker technique that can detect simultaneously variation at numerous genomic loci without sequence information. This efficiency makes it a potential tool for a quick and powerful assessment of the structure of germplasm collections. This article demonstrates the usefulness of DArT markers for genetic diversity analyses of Musa spp. genotypes. We developed four complexity reduction methods to generate DArT genomic representations and we tested their performance using 48 reference Musa genotypes. For these four complexity reduction methods, DArT markers displayed high polymorphism information content. We selected the two methods which generated the most polymorphic genomic representations (PstI/BstNI 16.8%, PstI/TaqI 16.1%) to analyze a panel of 168 Musa genotypes from two of the most important field collections of Musa in the world: Cirad (Neufchateau, Guadeloupe), and IITA (Ibadan, Nigeria). Since most edible cultivars are derived from two wild species, Musa acuminata (A genome) and Musa balbisiana (B genome), the study is restricted mostly to accessions of these two species and those derived from them. The genomic origin of the markers can help resolving the pedigree of valuable genotypes of unknown origin. A total of 836 markers were identified and used for genotyping. Ten percent of them were specific to the A genome and enabled targeting this genome portion in relatedness analysis among diverse ploidy constitutions. DArT markers revealed genetic relationships among Musa genotype consistent with those provided by the other markers technologies, but at a significantly higher resolution and speed and reduced cost.



Similar content being viewed by others
References
Akbari M, Wenzl P, Vanessa C, Carling J, Xia L, Yang S, Uszynski G, Mohler V, Lehmensiek A, Kuchel H, Hayden MJ, Howes N, Sharp P, Rathmell B, Vaughan P, Huttner E, Kilian A (2006) Diversity Arrays Technology (DArT) for high-throughput profiling of the hexaploid wheat genome. Theor Appl Genet 113:1409–1420
Amorim EP, Vilarinhos AD, Cohen KO, Amorim VBO, Santos-Serejo JA, Silva SO, Pestana KN, Santos VJ, Paes NS, Monte DC, Rei RV (2009) Genetic diversity of carotenoid-rich bananas evaluated by Diversity Arrays Technology (DArT). Genet Mol Biol 32:96–103
Anderson JA, Churchill GA, Autrique JE, Tanskley SD, Sorrells ME (1993) Optimizing parental selection for genetic linkage maps. Genome 396:181–186
Carreel F, Fauré S, Gonzalez de Leon D, Lagoda PLJ, Perrier X, Bakry F, Tezenas du Montcel H, Lanaud C, Horry JP (1994) Evaluation de la diversité génétique chez les bananiers diploïdes (Musa sp). Genet Sel Evol 26(Suppl):125s–136s
Carreel F, Gonzalez de Leon D, Lagoda P, Lanaud C, Jenny C, Horry J, Tezenas du Montcel H (2002) Ascertaining maternal and paternal lineage within Musa by chloroplast and mitochondrial DNA RFLP analyses. Genome 45:679–692
Creste S, Tulmann Neto A, de Oliveira Silva S, Figueira A (2003) Genetic characterization of banana cultivars (Musa spp.) from Brazil using microsatellite markers. Euphytica 132:259 258
Creste S, Tulmann Neto A, Vencovsky R, de Oliveira Silva S, Figueira A (2004) Genetic diversity of Musa diploid and triploid accessions from the Brazilian banana breeding program estimated by microsatellite markers. Genet Res Crop Evol 51:723–733
Crouch JH, Crouch HK, Constandt H, Van Gysel A, Breyne P, Van Montagu M, Jarret RL, Ortiz R (1999) Comparison of PCR-based molecular marker analyses of Musa breeding populations. Mol Breeding 5:233–244
D’Hont A, Paget-Goy A, Escoute J, Carreel F (2000) The interspecific genome structure of cultivated banana, Musa spp. revealed by genomic DNA in situ hybridization. Theor Appl Genet 100:177–183
Fauré S, Noyer JL, Horry JP, Bakry F, Lanaud C, Gońzalez de León D (1993) A molecular marker-based linkage map of diploid bananas (Musa acuminata). Theor Appl Genet 87:517–526
Gawel NJ, Jarret RL, Whittemore AP (1992) Restriction fragment length polymorphism (RFLP)-based phylogenetic analysis of Musa. Theor Appl Genet 84:286–290
Godwin ID, Aitken EAB, Smith LW (1997) Application of inter simple sequence repeat (ISSR) markers to plant genetics. Electrophoresis 18:1524–1528
Grapin A, Noyer JL, Carreel F, Dambler D, Baurens FC, Lanaud C, Lagoda PJL (1998) Diploid Musa acuminata genetic diversity assayed with sequence tagged microsatellite sites. Electrophoresis 19:1374–1380
Horry J, Jay M (1988) Distribution of anthocyanins in wild and cultivated banana varieties. Phytochemistry 27:2667–2672
Howell EC, Newbury HJ, Swennen RL, Withers LA, Ford-Lloyd BV (1994) The use of RAPD for identifying and classifying Musa germplasm. Genome 37:328–332
Jaccoud D, Peng K, Feinstein D, Kilian A (2001) Diversity Arrays: a solid-state technology for sequence information independent genotyping. Nucleic Acids Res 29:e25
Jarret RL, Gawel NJ, Whittemore A, Sharrock S (1992) RFLP-based phylogeny of Musa species in Papua New Guinea. Theor Appl Genet 84:579–584
Jenny C, Carreel F, Kodjo T, Perrier X, Dubois C, Horry JP, Tézenas du Montcel H (2002) Banana. In: Hamon P, Seguin M, Perrier X, Glaszmann JC (eds) Genetic diversity of cultivated tropical plants. Science Publishers Inc USA and CIRAD France, pp 99–124
Lanaud C, Tezenas du Montcel H, Jolivot MP, Glaszmann JC, Gonzalez De Leon D (1992) Variation of ribosomal gene spacer length among wild and cultivated banana. Heredity 68:147–156
Lescot M, Piffanelli P, Ciampi AY, Ruiz M, Blanc G, Leebens-Mack J, da Silva FR, Santos CMR, D’Hont A, Garsmeur O, Vilarinhos AD, Kanamori H, Matsumoto T, Ronning CM, Cheung F, Haas BJ, Althoff R, Arbogast T, Hine E, Pappas GJ, Sasaki T, Souza MT, Miller RG, Glaszmann JC, Town CD (2008) Insights into the Musa genome: syntenic relationships to rice and between Musa species. BMC Genomics 9:58
Lezar S, Myburg AA, Berger DK, Wingfield MJ, Wingfield BD (2004) Assessment of microarray-based DNA fingerprinting in Eucalyptus grandis. Theor Appl Genet 109:1329–1336
Mace ES, Xia L, Jordan DR, Halloran K, Parh DK, Huttner E, Wenzl P, Kilian A (2008) DArT markers: diversity analyses and mapping in Sorghum bicolor. BMC Genomics 9:26
Nair AS, Teo CH, Schwarzacher T, Heslop-Harisson JS (2005) Genome classification of banana cultivars from South India using IRAP markers. Euphytica 114:285–290
Noyer JL, Causse S, Tomekpe K, Bouet A, Baurens FC (2004) A new image of plantain diversity assessed by SSR, AFLP and MSAP markers. Genetica 124:61–69
Perrier X, Flori A, Bonnot F (2003) Data analysis methods. In: Hamon P, Seguin M, Perrier X, Glaszmann JC (eds) Genetic diversity of cultivated tropical plants. Science Publishers Inc USA and CIRAD France, pp 43–76
Perrier X, Bakry F, Carreel F, Jenny C, Horry JP, Lebot V, Hippolyte I (in press) Combining biological approaches to highlight the evolution towards edible bananas Sixth World Archaeological Congress Dublin. Ethnobotany Research and Application
Pillay M, Ogundiwin E, Nwakanma DC, Ude G, Tenkouano A (2001) Analysis of genetic diversity and relationships in East African banana germplasm. Theor Appl Genet 102:965–970
Raboin LM, Carreel F, Noyer JL, Baurens FC, Horry JP, Bakry F, Tezenas du Montcel H, Ganry J, Lanaud C, Lagoda PJL (2005) Diploid ancestors of triploid export banana cultivars: molecular identification of 2n restitution gamete donors and n gamete donors. Mol Breed 16:333–341
Risterucci AM, Grivet L, N’Goran JAK, Pieretti I, Flament MH, Lanaud C (2000) A high density linkage map of Theobroma cacao L. Theor Appl Genet 101:948–955
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Semagn K, Bjornstad A, Skinnes H, Maroy AG, Tarkegne Y, William M (2006) Distribution of DArT, AFLP, and SSR markers in a genetic linkage map of a doubled-haploid hexaploid wheat population. Genome 49:545–555
Shepherd K (1999) Cytogenetics of the genus Musa. Ipgri-Inibap, Rome
Simmonds NW (1962) The evolution of the bananas. Longmans, London
Simmonds NW (1995) Bananas Musa (Musaceae). In: Smartt J, Simmonds NW (eds) Evolution of crop plants. Longman Scientific and Technical, London, pp 370–375
Sokal RR, Michener CD (1958) A statistical method for evaluating systematic relationships. Univ Kansas Sci Bull 28:1409–1438
Storey J, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Nat Acad Sci USA 100:9440–9945
Tenkouano A, Crouch JH, Crouch HK, Vuylsteke D, Ortiz R (1999) Comparison of DNA marker and pedigree-based methods of genetic analysis of plantain and banana (Musa spp.) clones. I. Estimation of genetic relationships. Theor Appl Genet 98:62–68
Teo CH, Tan SH, Ho CH, Faridah QZ, Othman YR, Heslop-Harrison JS, Kalendar R, Schulman AH (2005) Genome constitution and classification using retrotransposon based markers in the orphan crop banana. J Plant Biol 48:96–105
Ude G, Pillay M, Ogundiwin E, Tenkouano A (2003) Genetic diversity in an African plantain core collection using AFLP and RAPD markers. Theor Appl Genet 107:248–255
Wenzl P, Carling J, Kudrna D, Jaccoud D, Huttner E, Kleinhofs A, Kilian A (2004) Diversity Arrays Technology (DArT) for whole-genome profiling of barley. Proc Nat Acad Sci USA 101:9915–9920
Wenzl P, Li H, Carling J, Zhou M, Raman H, Paul E, Hearnden P, Maier C, Xia L, Caig V, Ovesna J, Cakir M, Poulsen D, Wang J, Raman R, Smith KP, Muehlbauer GJ, Chalmers KJ, Kleinhofs A, Huttner E, Kilian A (2006) A high-density consensus map of barley linking DArT markers to SSR, RFLP and STS loci and phenotypic traits. BMC Genomics 7:206
White J, Law JR, Mackay I, Chalmers KJ, Smith JSC, Kilian A, Powell W (2008) The genetic diversity of UK, US and Australian cultivars of Triticum aestivum measured by DArT markers and considered by genome. Theo Appl Genet 116:439–453
Wittenberg A, van der Lee T, Cayla C, Kilian A, Visser R, Schouten H (2005) Quality assessment of the Diversity Array Technology (DArT) using the model plant Arabidopsis thaliana. Mol Gen Genomics 274:30–39
Wong C, Kiew R, Loh JP, Gan LH, Lee SK, Ohn S, Lum S, Gan YY (2001) Genetic diversity of the wild banana Musa acuminata Colla in Malaysia as evidenced by AFLP. Ann Bot 88:1017–1025
Wong C, Kiew R, Argent G, Set O, Lee SK, Ohn S, Gan YY (2002) Assessment of the validity of section in Musa (Musaceae) using AFLP. Ann Bot 90:231–238
Xia L, Peng K, Yang S, Wenzl P, de Vicente MC, Fregene M, Kilian A (2005) DArT for high-throughput genotyping of cassava (Manihot esculenta) and its wild relatives. Theor Appl Genet 110:1092–1098
Yang S, Pang W, Ash G, Harper J, Carling J, Wenzl P, Huttner E, Zong X, Kilian A (2006) Low level of genetic diversity in cultivated Pigeonpea compared to its relatives is revealed by diversity arrays technology. Theor Appl Genet 113:585–595
Acknowledgment
We thank the Generation Challenge Programme for financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Nybom.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Risterucci, AM., Hippolyte, I., Perrier, X. et al. Development and assessment of Diversity Arrays Technology for high-throughput DNA analyses in Musa . Theor Appl Genet 119, 1093–1103 (2009). https://doi.org/10.1007/s00122-009-1111-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1111-5