Abstract
Crop wild relatives are considered as important genetic resources of allelic diversity for domesticated crop species. Their utilization in breeding programs, however, is often limited due to crossing barriers and genome incompatibilities. Wild relatives of barley possess attractive properties and hence allelic diversity for adapting barley better to changing environmental conditions. Therefore, gaining a better knowledge about genomic synteny between cultivated barley and wild relatives of the same genus is an important task. To visualize genomic collinearity in related species, 22 genomic single-copy and 14 complementary DNA (cDNA) chromosome 3H-specific probes were mapped to the chromosomes of Hordeum bulbosum, Hordeum marinum, Hordeum pubiflorum, Hordeum murinum, and Secale cereale by fluorescent in situ hybridization (FISH). Most probes showed reliable signals confirming homoeology between cultivated barley and related species. Differences in order and position of FISH markers demonstrated sequence movements or small-scale chromosomal rearrangements within genus Hordeum and confirmed interchromosomal rearrangements between barley and rye. Comparison between repeat-free genomic and cDNA probes showed that gene-containing single-copy genomic DNA (gDNA) probes are performing more reliably for FISH-based analysis of synteny.




Similar content being viewed by others
Abbreviations
- Alexa 488:
-
Green-fluorescent dye
- cDNA:
-
Complementary DNA
- Cy3:
-
Cyanine dye
- dUTP:
-
Deoxyuridine triphosphate
- DAPI:
-
4′,6′-Diamidino-2-phenylindole
- fl-cDNA:
-
Full-length complementary DNA
- FISH:
-
Fluorescence in situ hybridization
- FPcontig:
-
FingerPrinted Contig
- gDNA:
-
Genomic DNA
- NOR:
-
Nucleolus organizer region
- rDNA:
-
Ribosomal DNA
References
Akhunov ED, Akhunova AR, Linkiewicz AM, Dubcovsky J, Hummel D, Lazo G, Chao S, Anderson OD, David J, Qi L, Echalier B, Gill BS, Miftahudin, Gustafson JP, La Rota M, Sorrells ME, Zhang D, Nguyen HT, Kalavacharla V, Hossain K, Kianian SF, Peng J, Lapitan NL, Wennerlind EJ, Nduati V, Anderson JA, Sidhu D, Gill KS, McGuire PE, Qualset CO, Dvorak J (2003) Synteny perturbations between wheat homoeologous chromosomes caused by locus duplications and deletions correlate with recombination rates. Proc Natl Acad Sci U S A 100:10836–10841
Aliyeva-Schnorr L, Beier S, Karafiatova M, Schmutzer T, Scholz U, Dolezel J, Stein N, Houben A (2015a) Cytogenetic mapping with centromeric bacterial artificial chromosomes contigs shows that this recombination-poor region comprises more than half of barley chromosome 3H. Plant J 84:385–394
Aliyeva-Schnorr L, Ma L, Houben A (2015b) A fast air-dry dropping chromosome preparation method suitable for FISH in plants. J Vis Exp
Bedbrook JR, Jones J, O’Dell M, Thompson RD, Flavell RB (1980) A molecular description of telomeric heterochromatin in Secale species. Cell 19:545–560
Blattner FR (2004) Phylogenetic analysis of Hordeum (Poaceae) as inferred by nuclear rDNA ITS sequences. Mol Phylogenet Evol 33:289–299
Blattner FR (2009) Progress in phylogenetic analysis and a new infrageneric classification of the barley genus Hordeum (Poaceae: Triticeae). Breed Sci 59:471–480
Brassac J, Blattner FR (2015) Species-level phylogeny and polyploid relationships in Hordeum (Poaceae) inferred by next-generation sequencing and in silico cloning of multiple nuclear loci. Syst Biol 64:792–808
Carmona A, Friero E, de Bustos A, Jouve N, Cuadrado A (2013a) Cytogenetic diversity of SSR motifs within and between Hordeum species carrying the H genome: H. vulgare L. and H. bulbosum L. Theor Appl Genet 126:949–961
Carmona A, Friero E, de Bustos A, Jouve N, Cuadrado A (2013b) The evolutionary history of sea barley (Hordeum marinum) revealed by comparative physical mapping of repetitive DNA. Ann Bot 112:1845–1855
Chalmers KJ, Waugh R, Watters J, Forster BP, Nevo E, Abbott RJ, Powell W (1992) Grain isozyme and ribosomal DNA variability in Hordeum spontaneum populations from Israel. Theor Appl Genet 84:313–322
Chloupek O, Forster BP, Thomas WT (2006) The effect of semi-dwarf genes on root system size in field-grown barley. Theor Appl Genet 112:779–786
Cuadrado A, Jouve N (2007) The nonrandom distribution of long clusters of all possible classes of trinucleotide repeats in barley chromosomes. Chromosome Res 15:711–720
Cuadrado A, Jouve N, Ceoloni C (1995) Variation in highly repetitive DNA composition of heterochromatin in rye studied by fluorescence in situ hybridization. Genome 38:1061–1069
Cuadrado A, Cardoso M, Jouve N (2008) Increasing the physical markers of wheat chromosomes using SSRs as FISH probes. Genome 51:809–815
Cuadrado A, Carmona A, Jouve N (2013) Chromosomal characterization of the three subgenomes in the polyploids of Hordeum murinum L.: new insight into the evolution of this complex. PLoS One 8:e81385
Danilova TV, Birchler JA (2008) Integrated cytogenetic map of mitotic metaphase chromosome 9 of maize: resolution, sensitivity, and banding paint development. Chromosoma 117:345–356
Danilova TV, Friebe B, Gill BS (2014) Development of a wheat single gene FISH map for analyzing homoeologous relationship and chromosomal rearrangements within the Triticeae. Theor Appl Genet 127:715–730
Fukui K, Kamisugi Y, Sakai F (1994) Physical mapping of 5S rDNA loci by direct-cloned biotinylated probes in barley chromosomes. Genome 37:105–111
Hasterok R, Marasek A, Donnison IS, Armstead I, Thomas A, King IP, Wolny E, Idziak D, Draper J, Jenkins G (2006) Alignment of the genomes of Brachypodium distachyon and temperate cereals and grasses using bacterial artificial chromosome landing with fluorescence in situ hybridization. Genetics 173:349–362
Huang S, Sirikhachornkit A, Su X, Faris J, Gill B, Haselkorn R, Gornicki P (2002) Genes encoding plastid acetyl-CoA carboxylase and 3-phosphoglycerate kinase of the Triticum/Aegilops complex and the evolutionary history of polyploid wheat. Proc Natl Acad Sci U S A 99:8133–8138
International Barley Genome Sequencing, Mayer CKF, Waugh R, Brown JW, Schulman A, Langridge P, Platzer M, Fincher GB, Sato K, Close TJ, Wise RP, Stein N (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–716
Jacobsen N, von Bothmer R (1995) Taxonomy in the Hordeum murinum complex (Poaceae). Nord J Bot 15:449–458
Jakob SS, Blattner FR (2010) Two extinct diploid progenitors were involved in allopolyploid formation in the Hordeum murinum (Poaceae: Triticeae) taxon complex. Mol Phylogenet Evol 55:650–659
Jakob SS, Meister A, Blattner FR (2004) The considerable genome size variation of Hordeum species (Poaceae) is linked to phylogeny, life form, ecology, and speciation rates. Mol Biol Evol 21:860–869
Kato A, Lamb JC, Birchler JA (2004) Chromosome painting using repetitive DNA sequences as probes for somatic chromosome identification in maize. Proc Natl Acad Sci U S A 101:13554–13559
Kato A, Albert PS, Vega JM, Birchler JA (2006) Sensitive fluorescence in situ hybridization signal detection in maize using directly labeled probes produced by high concentration DNA polymerase nick translation. Biotech Histochem 81:71–78
Kim MJ, Kim H, Shin JS, Chung CH, Ohlrogge JB, Suh MC (2006) Seed-specific expression of sesame microsomal oleic acid desaturase is controlled by combinatorial properties between negative cis-regulatory elements in the SeFAD2 promoter and enhancers in the 5′-UTR intron. Mol Genet Genomics 276:351–368
Komatsuda T, Mano Y (2002) Molecular mapping of the intermedium spike-c (int-c) and non-brittle rachis 1 (btr1) loci in barley (Hordeum vulgare L.). Theor Appl Genet 105:85–90
Komuro S, Endo R, Shikata K, Kato A (2013) Genomic and chromosomal distribution patterns of various repeated DNA sequences in wheat revealed by a fluorescence in situ hybridization procedure. Genome 56:131–137
Lange W (1971) Crosses between Hordeum vulgare L and H. bulbosum L. 2. Elimination of chromosomes in hybrid tissues. Euphytica 20:181
Linde-Laursen I, von Bothmer R, Jacobsen N (1990) Giemsa C-banded karyotypes of diploid and tetraploid Hordeum bulbosum (Poaceae). Pl Syst Evol 172:141–150
Ma J, Wing RA, Bennetzen JL, Jackson SA (2007) Evolutionary history and positional shift of a rice centromere. Genetics 177:1217–1220
Ma L, Vu GTH, Schubert V, Watanabe K, Stein N, Houben A, Schubert I (2010) Synteny between Brachypodium distachyon and Hordeum vulgare as revealed by FISH. Chromosome Res 18:841–850
Marcussen T, Sandve SR, Heier L, Spannagl M, Pfeifer M, International Wheat Genome Sequencing Consortium, Jakobsen KS, Wulff BB, Steuernagel B, Mayer KF, Olsen OA (2014) Ancient hybridizations among the ancestral genomes of bread wheat. Science 345:1250092
Martis MM, Zhou R, Haseneyer G, Schmutzer T, Vrana J, Kubalakova M, Konig S, Kugler KG, Scholz U, Hackauf B, Korzun V, Schon CC, Dolezel J, Bauer E, Mayer KF, Stein N (2013) Reticulate evolution of the rye genome. Plant Cell 25:3685–3698
Mukai Y, Friebe B, Gill BS (1992) Comparison of C-Banding patterns and in situ hybridization sites using highly repetitive and total genomic rye DNA probes of imperial rye chromosomes added to Chinese spring wheat. Jpn J Genet 67:71–83
Pickering RA, Hudakova S, Houben A, Johnston PA, Butler RC (2004) Reduced metaphase I associations between the short arms of homologous chromosomes in a Hordeum vulgare L. × H. bulbosum L. diploid hybrid influences the frequency of recombinant progeny. Theor Appl Genet 109:911–916
Pickering R, Klatte S, Butler RC (2006) Identification of all chromosome arms and their involvement in meiotic homoeologous associations at metaphase I in 2 Hordeum vulgare L. × Hordeum bulbosum L. hybrids. Genome 49:73–78
Rippe RA, Lorenzen SI, Brenner DA, Breindl M (1989) Regulatory elements in the 5′-flanking region and the first intron contribute to transcriptional control of the mouse alpha 1 type I collagen gene. Mol Cell Biol 9:2224–2227
Sato K, Nankaku N, Takeda K (2009) A high-density transcript linkage map of barley derived from a single population. Heredity 103:110–117
Schmutzer T, Ma L, Pousarebani N, Bull F, Stein N, Houben A, Scholz U (2014) Kmasker—a tool for in silico prediction of single-copy FISH probes for the large-genome species Hordeum vulgare. Cytogenet Genome Res 142:66–78
Suprunova T, Krugman T, Distelfeld A, Fahima T, Nevo E, Korol A (2007) Identification of a novel gene (Hsdr4) involved in water-stress tolerance in wild barley. Plant Mol Biol 64:17–34
Tadepally H, Burger G, Aubry M (2008) Evolution of C2H2-zinc finger genes and subfamilies in mammals: species-specific duplication and loss of clusters, genes and effector domains. BMC Evol Biol 8:176
Terzi V, Pecchioni N, Faccioli P, Kucera L, Stanca AM (2001) Phyletic relationships within the genus Hordeum using PCR-based markers. Genet Resour Crop Evol 48:447–458
Thomas HM, Pickering RA (1988) The cytogenetics of a triploid Hordeum bulbosum and of some of its hybrid and trisomic derivatives. Theor Appl Genet 76:93–96
von Bothmer R, Flink J, Jacobsen N, Kotimaki M, Landstrom T (1983) Interspecific hybridization with cultivated barley (Hordeum vulgare L.). Hereditas 99:219–244
von Bothmer R, Bengtsson M, Flink J, Linde-Laursen I (1988) Complex interspecific hybridization in barley (Hordeum vulgare L.) and the possible occurrence of apomixis. Theor Appl Genet 76:681–690
von Bothmer R, Claesson L, Flink J, Linde-Laursen I (1989) Triple hybridization with cultivated barley (Hordeum vulgare L.). Theor Appl Genet 78:818–824
Vos P, Hogers R, Bleeker M, Reijans M, Vandelee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP—a new technique for DNA-fingerprinting. Nucleic Acids Res 23:4407–4414
Wicker T, Mayer KF, Gundlach H, Martis M, Steuernagel B, Scholz U, Simkova H, Kubalakova M, Choulet F, Taudien S, Platzer M, Feuillet C, Fahima T, Budak H, Dolezel J, Keller B, Stein N (2011) Frequent gene movement and pseudogene evolution is common to the large and complex genomes of wheat, barley, and their relatives. Plant Cell 23:1706–1718
Yan G, Chen X (2007) Molecular mapping of the rps1.a recessive gene for resistance to stripe rust in BBA 2890 barley. Phytopathology 97:668–673
Acknowledgments
We would like to thank Katrin Kumke and Oda Weiss for the technical assistance and Karin Lipfert for the art work. We also thank Prof. Angeles Bermejo Cuadrado for supplying the probe pSc119.2, Prof. Yasunari Ogihara for supplied plasmids containing wheat cDNA (NBRP-WHEAT, http://www.shigen.nig.ac.jp/wheat/komugi/ests/tissueBrowse.jsp), and Prof. Kazuhiro Sato for the plasmids containing barley cDNA (Barley Gene Expression Database, http://barleyflc.dna.affrc.go.jp/hvdb/index.html). Special thanks to Dr. Frank Blattner for his support and suggestions on this work. This work was supported by the DFG (HO1779/21-1) to AH and NS.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Hans de Jong
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
S. cereale chromosomes after FISH with pSc119.2 (green). Chromosomes are counterstained by DAPI (grey). Differentiation of all chromosome pairs was described earlier by Cuadrado et al. (1995) using the repetitive families of 120-, 350-480-, and 610-bp DNA. (GIF 117 kb)
Fig. S2
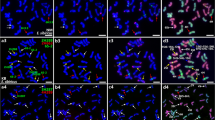
Application of barley gDNA FISH probes and barley/wheat cDNA probes on H. bulbosum and S. cereale. (a) Single-copy FISH with 22 gDNA probes (red) on metaphase chromosomes of H. bulbosum were identified by pattern of microsatellite CTT (green). (b) FISH with gDNA probes (signals in red) on metaphase chromosomes of S. cereale were identified by pattern of tandem repeat pSc119.2. (c) Single-copy FISH with cDNA probes (red) on metaphase chromosomes of H. bulbosum identified by microsatellite CTT (green). (d) FISH with cDNA probes (red) on metaphase chromosomes of S. cereale identified by the tandem repeat probe pSc119.2. All chromosomes were counterstained with DAPI (grey). (GIF 187 kb)
Fig. S3
Application of barley single-copy gene-containing (gDNA) FISH probes on H. pubiflorum, H. marinum, and H. murinum. (a) Single-copy FISH with 18 gDNA probes (red) on metaphase chromosomes of H. pubiflorum were identified by pattern of microsatellite CTT (green). (b) Single-copy FISH with 18 gDNA probes (signals in red) on metaphase chromosomes of H. marinum were identified by pattern of microsatellite CTT (green). (c) Single-copy FISH with 18 gDNA probes ( red) on metaphase chromosomes of H. murinum were identified by pattern of microsatellite CTT (green). All chromosomes were counterstained with DAPI (grey). (GIF 143 kb)
Fig. S4
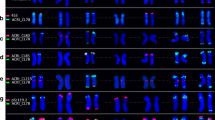
Comparative FISH mapping on metaphase chromosomes showing signals of selected cDNA probes (red) on H. vulgare and S. cereale. The insets show chromosome spreads and enlarged chromosomes with specific signals for three cDNA probes hybridized to barley (left) and rye (right). 5SrDNA (green) and pSc119.2 (green) were used as diagnostic probes for H. vulgare and for S. cereale, respectively. Chromosomes are counterstained with DAPI (grey). (GIF 198 kb)
Fig. S5
Comparison of intervals on the chromosome 3H of H. vulgare and homoeologous chromosome 3Hpub of H. pubiflorum indicated by the simultaneous hybridization of two boundary probes (1770 and 44666). The interval captures 58 % of the total physical length on H. vulgare and only 49.6 % of chromosome 3Hpub of H. pubiflorum. (GIF 107 kb)
Table S1
List of barley-specific contig derived probes with the determined cytogenetic position (CP) on (a) H. bulbosum, S. cereale (b) H. pubiflorum, H. marinum and H. murinum. Column 1 shows the first set of probes and column 2 the second set. Coding sequences are indicated in red, “na” - indicates that the corresponding probe was not tested on this species. (DOCX 22 kb)
Table S2
Subprobes and primer pairs designed for the generation of gDNA single-copy probes derived from FPcontigs (gDNA). (DOCX 29 kb)
Table S3
List of cDNAs with corresponding cytogenetic positions (CP) on chromosome 3H and its homoeologues on H. bulbosum and S. cereale. “nd” – signals were not detected on the species. (DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Aliyeva-Schnorr, L., Stein, N. & Houben, A. Collinearity of homoeologous group 3 chromosomes in the genus Hordeum and Secale cereale as revealed by 3H-derived FISH analysis. Chromosome Res 24, 231–242 (2016). https://doi.org/10.1007/s10577-016-9518-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-016-9518-8