Abstract
With an unsupervised GC–MS metabolomics approach, polar metabolite changes of the microalgae Coccomyxa melkonianii SCCA 048 grown under standard conditions for seven weeks were studied. C. melkonianii was sampled at the Rio Irvi River, in the mining site of Montevecchio-Ingurtosu (Sardinia, Italy), which is severely contaminated by heavy metals and shows high concentrations of sulfates. The partial-least-square (PLS) analysis of the GC–MS data indicated that growth of C. melkonianii was characterized by an increase of the levels of threonic acid, myo-inositol, malic acid, and fumaric acid. Furthermore, at the sixth week of exponential phase the lipid fingerprint of C. melkonianii was studied by LC-QTOF-MS. C. melkonianii lipid extract characterized through an iterative MS/MS analysis showed the following percent levels: 61.34 ± 0.60% for triacylglycerols (TAG); 11.55 ± 0.09% for diacylglyceryltrimethyl homoserines (DGTS), 11.34 ± 0.10% for sulfoquinovosyldiacylglycerols (SQDG) and, 5.29 ± 0.04% for lysodiacylglyceryltrimethyl homoserines (LDGTS). Noteworthy, we were able to annotate different fatty acid ester of hydroxyl fatty acid, such as FAHFA (18:1_20:3), FAHFA (18:2_20:4), FAHFA (18:0_20:2), and FAHFA (18:1_18:0), with relevant biological activity. These approaches can be useful to study the biochemistry of this extremophile algae in the view of its potential exploitation in the phycoremediation of polluted mining areas.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Microalgae belonging to the class of Trebouxiophyceae can be found in different terrestrial and aquatic environments, such as mesophilic habitats comprising soil, salty waters and extreme environments (Büdel et al. 2009; Cavacini 2001; Fermani et al. 2007; Flechtner et al. 2013; Heesch et al. 2012; Hodač et al. 2016; Juárez et al. 2011; Malavasi et al. 2016; Tragin and Vaulot 2018).
Trebouxiophyceae show different phenotypic, physiologic, and genetic characteristics determining their ability to survive in these environments (Assunção et al. 2017; Cannell 1993). Belonging to this class, Coccomyxa spp. with more than 60 described species (Guiry et al. 2020) are morphological characterized by a parietal chloroplast without a pyrenoid and a thin three-layered cell wall, and by the lack of flagellum (Brunner and Honegger 1981; Darienko et al. 2015; Malavasi et al. 2016). In recent years, in response to the search for more biofuel sources, strains of the genus Coccomyxa have been studied for their ability to produce lipids and, given the high concentration of polyunsaturated ω-3 and ω-6 fatty acids, some strains may cover an important role in the food and feed industry (Jeong et al. 2011). Moreover, green algal photobionts of Coccomyxa are often symbiotic partners in Peltigera lichens which are relatively sensitive to environment while growing exclusively in mild moist habitats (Guschina and Harwood 2006). These species can also resist in strongly polluted environments (Kalinowska and Pawlik-Skowrońska 2010; Koechler et al. 2016; la Rocca et al. 2009).
In our study, the extremophile microalga C. melkioananii SCCA 048 was sampled in acidic mine drainage waters severely contaminated by heavy metals. This mining site of Montevecchio-Ingurtosu (Sardinia, Italy) is characterized by minerals such as galena (PbS2) containing appreciable levels of silver, sphalerite ((Zn, Fe)S) containing Cd, Ga, In, and the oxide mineral goethite (α-FeO(OH)). For this reason, this site is rich in pollutants, with levels of Zn2+ at 956 mg/L, Fe2+ at 227 mg/L, and sulfates at 3697 mg/L, released into the river by oxidation reactions involving the sulfide minerals still present in the ore bodies after flooding of galleries (De Giudici et al. 2018).
This C. melkonianii strain well adapted to this contaminated environment may be used for the development of new and sustainable phycoremediation technology strategies (De Giudici et al. 2018; Kothe et al. 2005). Furthermore, Coccomyxa actinabiotis showed high silver levels confined inside microalgae, when grown in contaminated waters containing silver ions (Leonardo et al. 2016). Additionally, this species isolated from a nuclear facility showed the ability to uptake radionuclides (Rivasseau et al. 2016; Sukla and Pradhan 2019). In this context, Coccomyxa subellipsoidea, a unicellular green acidophilic microalga isolated from the Antarctic, grows over a range of temperate climates, accumulating high levels of triglycerides under abiotic stress conditions and thus relevant for biofuel/bioproduct production, or for providing biomass for nitrogenous biofertilizers, and for the degradation of organophosphate pesticides (Allen et al. 2015; Heesch et al. 2012; Hirooka et al. 2017; Nicodemus et al. 2020). It is also known that Coccomyxa species can produce lipid-like compounds containing a dimethylarsinoyl group cultivated at high concentration of disodium hydrogen arsenate (Řezanka et al. 2019). Furthermore, this strain can be exploited for the production of isotopically labeled (non-radioactive) lipids to be used in the field of analytical chemistry (Beherens 1994).
In our previous studies, this strain was investigated for its ability of growing at different pH values. Our results demonstrated the extreme plasticity of this species, being able to cope with different environments and to produce more than 20% by weight of lipids (Soru et al. 2019a). Interestingly, a change in the profile of fatty acid methyl esters (FAMEs) and an increase in lipid content under nitrogen starvation was also observed, showing that C. melkonianii SCCA 048 can be used for biofuels production (Soru et al. 2019b).
Metabolomics is as a new “omics” science widely used in system biology. Metabolomics is the holistic study of the metabolome of a system cell, tissue, or organism performed using specific analytical instrumentation and statistical methods. The metabolome and lipidome are the result of the interaction of the genome of the system with its environment (Rochfort 2005) and comprise the collection of all low molecular weight compounds, such as amino acids, carbohydrates, nucleotides, organic acids, fatty acids, lipids, and cofactors responsible for maintaining cell's biological processes (Dunn and Ellis 2005). Metabolomics and lipidomics studies play an important role to understand the physiological changes in living organisms, thus offering additional insight for microalgae strain engineering consideration (Arora et al. 2018; Ito et al. 2013). Several metabolomics studies evidenced that the production of metabolites is highly dependent on the development of the microalgal cells (Blifernez-Klassen et al. 2018; Treves et al. 2017; Vidoudez and Pohnert 2012) and omics approach has been used to characterize compounds synthesized during lipids accumulation (Arora et al., 2018). Synthesis of microalgae metabolites is tighten correlated to the accumulation of triacylglycerols (TAGs) during nitrogen starvation and the presence of specific compounds. For example, an increase of Krebs cycle metabolites, such as citrate, 2-oxoglutarate, and phosphorylated sugars, with a concomitant decreases in amino acids levels indicates an accumulation of intracellular lipids (Blaby et al. 2013; Chen et al. 2017; Ito et al. 2013; Wase et al. 2014). Similarly, cell metabolites, such as glycerol, 3-phosphoglyceric acid and 2-ketoglutaric acid, play a crucial role in the increase of TAGs in response to environmental stress factors, such as high salinity, high intensity of light or to chemical challenge with phytohormones (Ho et al. 2015, 2014; Malavasi and Cao 2015; Yu et al. 2016). Furthermore, polar lipids, i.e. phosphatidic acid (PA), phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylglycerol (PG), digalactosyldiacylglycerol (DGDG), monogalactosyldiacylglycerol (MGDG), sulphoquinovosyl diacylglycerols (SQDG), and diacylglyceryltrimethylhomoserine (DGTS), play an important role in membranes structure, photosynthesis, energy storage and cellular signaling (Darienko et al., 2015; Van Meer et al., 2008).
Despite its physiological and technological importance, little is known on the metabolome and lipidome of C. melkonianii SCCA 048 (Pasqualetti et al. 2015; Soru et al. 2019a, b).
Microalgae contain large amounts of lipids, proteins, and carbohydrates, while one of the main bottlenecks for their possible application is that the composition of such compounds dramatically changes during growth depending upon the relevant species and strains (Guedes et al. 2011; Cannell 1993; De Morais et al. 2015). It is also highly desirable to take advantage of suitable microalgal species which can be exploited for the isolation of high-value metabolites (Lee et al. 2018) and lipids as well as potent and ecofriendly tools for bioremediation of polluted waters.
To understand the molecular profiling and to evaluate the changes occurring during the growth of this microalga, we employed an untargeted metabolomic approach.
In this longitudinal study, the changes of cellular metabolites of Coccomyxa melkonianii strain SCCA 048 during seven weeks of cultivation, were studied using a gas chromatography mass spectrometry (GC–MS) approach followed by a partial-least-square analysis (PLS). Furthermore, using a UHPLC-QTOF-MS iterative approach we studied the lipidome fingerprint of C. melkonianii. The aim of this study was to explore the metabolic profiles of C. melkonianii SCCA 048 grown under standard conditions, and to obtain basic lipidome profile information for the potential use of this microalga in the bioremediation of the drainage waters of the Rio Irvi insisting in the mining area of Ingurtosu.
Materials and methods
Strains, culture conditions and growth measurement
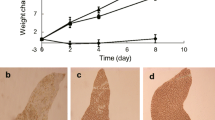
In this work, freshwater strains of C. melkonianii SCCA 048, sampled at the Rio Irvi river (West-Sardinia, Italy), was investigated (Fig. 1b). The strain was maintained under axenic conditions at the Sardinian Culture Collection of Algae (SCCA) (Malavasi and Cao 2015). The alga strain was phototrophically cultivated at 25 °C under 12:12 light–dark illumination of 60–80 μmol photons/m2/s (Light meter Delta) white light and continuous agitation at 100 rpm (Stuart SSM1, Biosigma orbital shaker) in BBM medium for 49 days. The cultivation was performed in cylindrical Erlenmeyer flasks (with maximal capacity of 250 mL) with a total starting volume of 150 mL per replicate. Flasks as well as the culture media were autoclaved at 121 °C for 15 min prior to microalgae inoculation. Flasks were stoppered by cotton plugs wrapped in cotton gauze during cultivation. All operations were conducted under a microbiological safety cabinet. A pre-culture of 5 days was prepared as inoculum for the experiments and used when cells started their exponential growth. Such pre-culture was incubated and maintained under the same conditions described above. The microalgae growth was monitored in vivo by detecting the chlorophyll-a optical density (OD) of the culture at 663 nm (Genesys 20. Thermo Scientific. Waltham. USA). Cells morphology was investigated using an inverted light microscope (Olympus, Tokyo, Japan).
Chemicals
Chemicals and solvents were purchased from (Sigma Aldrich, Milano, Italy). Bi-distilled water was obtained with a MilliQ purification system (Millipore, Milan, Italy). Methoxyamine hydrochloride, NO-bis (trimethylsilyl) trifluoroacetamide (BSTFA) were used for the derivatization process. A SPLASH® LIPIDOMIX® standard component mixture was purchased from Sigma Aldrich (Milan, Italy) PC (15:0–18:1) (d7), PE(15:0–18:1) (d7), PS (15:0–18:1) (d7), PG (15:0–18:1) (d7), PI (15:0–18:1) (d7), PA (15:0–18:1) (d7), LPC (18:1) (d7) LPC 25, LPE (18:1) (d7), Chol Ester (18:1) (d7), MG (18:1) (d7), DAG (15:0–18:1) (d7), TAG ((15:0–18:1) (d7)-15:0)), SM (18:1) (d9), Cholesterol (d7).
GC–MS analysis
For GC–MS analysis, 10 mL of culture was taken at seven different time points (1, 7, 15, 22, 29, 36, and 43 days) of the cultivation and transferred into a sterile 15 mL Falcon tube. Samples were then stored at -20 °C. After thawing, microalgae samples were sonicated for 15 min at 4 °C at 13,000 rpm. Samples were vortexed for 30 s and 250 μL from each sample was transferred into Eppendorf tubes, to which 250 μL of methanol and 125 μL of chloroform (2:1 v/v) were subsequently added (Folch et al. 1987). Samples were stored at room temperature, vortexed every 15 min and then centrifuged for 10 min at 13,000 rpm at 4 °C. The supernatant was separated, dried under nitrogen flow and derivatized with 50 μL of methoxyamine/pyridine solution (10 mg/mL). After 17 h, 50 μL of BSTFA were added for 40 min at 50 °C. Samples were re-suspended with 50 μL of hexane. After derivatization, samples were injected in a Hewlett Packard 6850 Gas Chromatograph, 5973 mass selective detector (Agilent Technologies, Palo Alto, CA), using helium as carrier gas at 1.0 mL/min flow. 1 μL of each sample was injected in the split-less mode and resolved on a 30 m × 0.25 mm × 0.25 μm DB-5MS column (Agilent Technologies, Palo Alto, CA). Inlet, interface, and ion source temperatures were 250, 250 and 230 °C, respectively. Oven starting temperature was set to 50 °C, final temperature to 230 °C with a heating rate of 5 °C/min for 36 min and then for 2 min at a constant temperature. Electron impact mass spectra were recorded from m/z 50 to 550 at 70 eV. Chromatograms in the AIA format were then uploaded to the XCMS Online platform (Tautenhahn et al. 2012). The outputs of XCMS consisted of a list of features corresponding to the intensity value of each m/z ion at a specific retention time value. The identification of metabolites was performed by mass spectra comparison with analytical standards, using the NIST14 library database of the National Institute of Standards and Technology (Gaithersburg, MD), Golm library (http://gmd.mpimp-golm.mpg.de/), and an in-house library of metabolites.
Multivariate analysis (MVA)
The GC–MS data were submitted to MVA as implemented in SIMCA-P + software (version 14.1. Umetrics, Umeå, Sweden). Prior to MVA, GC features were mean centered and scaled to unit variance column-wise. Principal component analysis (PCA) was performed to investigate sample distributions, deviating features and prevailing trends. GC–MS data were correlated to the calculated biomass concentration (g/L) by a single-Y Partial Least Squares Projections to Latent Structures (PLS) analysis and to highlight the metabolites mostly correlated to the time evolution of biomass by its Orthogonal variant (OPLS). The quality of the model was evaluated based on the cumulative parameters R2Y and Q2Y, being the latter estimated by the default leave-1/7th-out cross-validation. The variable influence on projection (VIP) scores, that summarize the contribution of each variable to the model, were analyzed (Scano et al. 2014). GC–MS features showing VIP values > 1 underwent a manual annotation using GC–MS library mass spectral databases. A metabolite was considered significant only when at least two of its most abundant mass fragments and a retention index deviation < 0.05 min were found in the list of VIP having a score greater than 1. For quantification purposes for each metabolite, we considered the intensity of the most abundant mass fragment.
UHPLC-QTOF-MS/MS analysis
The chloroform layer of the sample at the sixth week of the exponential phase was obtained by Folch extraction (Folch et al., 1987) was evaporated under a gentle nitrogen stream, and dissolved in 100 µL of a mixture of acetonitrile/water (1:1 v/v) and 10 µL of a mixture of methanol/chloroform (1:1 v/v), and finally added of 10 µL of the internal lipid standard SPLASH solution. Samples were then analyzed with a LC-QTOF-MS coupled with an Agilent 1290 Infinity II LC system. An aliquot of 1.0 μL from each sample was injected in a Kinetex 5 µm EVO C18 100 A, 150 mm × 2.1 μm column (Agilent Technologies, Palo Alto, CA). The column was maintained at 50 °C at a flow rate of 0.4 mL/min. The mobile phase for positive ionization mode consisted of (A) 10 mM ammonium formate solution in 60% of milliQ water and 40% of acetonitrile and (B) 10 mM ammonium formate solution containing 90% of isopropanol, 10% of acetonitrile. In positive ionization mode, the chromatographic separation was obtained with the following gradient: initially 60% of A, then a linear decrease from 60 to 50% of A in 2 min then at 1% in 5 min staying at this percentage for 1.9 min and then brought back to the initial conditions in 1 min. The mobile phase for negative ionization mode differed only for the use of 10 mM ammonium acetate instead of ammonium formate. We used an Agilent jet stream technology source which was operated in both positive and negative ion modes with the following parameters: gas temperature, 200 °C; gas flow (nitrogen) 10 L/min; nebulizer gas (nitrogen), 50 psig; sheath gas temperature, 300 °C; sheath gas flow, 12 L/min; capillary voltage 3500 V for positive and 3000 V for negative; nozzle voltage 0 V; fragmentor 150 V; skimmer 65 V, octapole RF 7550 V; mass range, 50 − 1700 m/z; capillary voltage, 3,5 kV; collision energy 20 eV in positive and 25 eV in negative mode, mass precursor per cycle = 3; threshold for MS/MS 5000 counts. Before the analysis, the instrument was calibrated using an Agilent tuning solution at the mass range of m/z 50–1700. Samples were acquired in an auto MS/MS method in the iterative mode with a mass error tolerance of 20 ppm with a retention exclusion tolerance of 0.2 min. The Agilent MassHunter LC/MS Acquisition console (revision B.09.00) and Lipid annotator from the MassHunter suite was used for data acquisition and data processing. This method consists in injecting the same sample multiple times, while precursors previously selected for MS/MS fragmentation are excluded on a rolling basis. Five different iterative analyses were performed for maximizing the maximum number of lipid species detected. In the positive mode, lipids were quantified using the following standards TAG 15:0–18:1 (d7)-15:0, PC 15:0–18:1 (d7), PG 15:0–18:1 (d7), and PE 15:0–18:1 (d7). Microalgae lipid analysis was conducted using an innovative iterative auto MS/MS mode.
Results and discussion
Growth evaluation
As a first step towards the metabolic profiling of C. melkonianii SCCA 048, we monitored the growth of the microalgal strain along seven weeks. The experiments started (day 1) from biomass concentration lower than 0.1 g/L and showed a continuous increase in the time course of the cultivation with an almost linear growth pattern during the investigated period. At the end of cultivation, C. melkonianii achieved a final biomass concentration of approximately 1.05 g/L. The time evolution of biomass during cultivation is reported in Fig. 1e.
GC–MS metabolomics
GC–MS chromatograms of these alga were analyzed, 53 polar metabolites were detected, 39 of which were identified (Table 1). The GC–MS data, obtained by the XCMS pipeline, underwent multivariate analysis. To identify those metabolites mostly correlated with the growth of the culture of C. melkonianii, the OPLS predictive statistical analysis was applied. Results of the analysis, shown as correlation plot in Fig. 2, indicated that for this alga, the whole metabolite profile linearly changes with the biomass increase. Those metabolites with a variable influence on projection (VIP) values > 1 that increased during growth are reported in Table 2. Among these, threonic acid was found correlated with algae growth. Threonic acid is a storage metabolite of the ascorbate metabolism pathway. This metabolite is strongly elevated under stress conditions, playing a part as a stress-responsive factor with a bio-protective role, and thus helping the cells to retain cellular integrity and stability (Blifernez-Klassen et al. 2018). Several studies suggest that ascorbate efflux plays a role in Fe3+ reduction in plants and algae (Kobayashi and Nishizawa 2012; Urzica et al. 2012; Grillet et al. 2014; Smirnoff 2018). Levels of threonic acid as housekeeping metabolite in C. melkonianii might correlate with its adaptation to harsh habitats (Malavasi et al. 2016). Moreover, malic acid and citric acid that are time-correlated with the growth of C. melkonianii are also known to be able to chelate metals (Perpetuo et al., 2011; Asemave 2018) Furthermore, myo-inositol is required for the cell growth and development, being involved in the phosphatidylinositol signaling pathway that takes part in auxin transport, cell wall biosynthesis, phytic acid biosynthesis, and the production of stress-related compounds (Cho et al. 2015).
Consistently, malic acid, succinic acid, citric acid, and fumaric acid, all involved in TCA cycle, increased during algal growth. It is well recognized that endogenous organic acids are the source of both carbon skeleton and energy for cells and are used in the respiratory cycle and other biochemical pathways. Previous studies demonstrated that the malic acid pathway is critical for lipid accumulation (Xue et al. 2016; Red et al. 2016), γ-aminobutyric acid (GABA) was also found upregulated during growth suggesting this metabolite may play a molecular signaling role in response to various environmental stress, including oxidative stress (Bouché and Fromm 2004).
To sum up the GC–MS metabolomics data, this study provides an overview of the dynamic metabolic changes during 7 weeks on the SCCA strain C. melkonianii, which potentially provides screening for the selection of their biologically active natural products. Moreover, the results of the metabolite analysis in this work can be used to further explore the uses and product synthesis of this microalga, which might be relevant for different biotechnological and bioremediation applications.
UHPLC-QTOF-MS/MS analysis
In this work, we also investigated the lipid profile of C. melkonianii, in the last stages of growth, using a UHPLC-QTOF-MS/MS analytical platform after lipid extraction with chloroform. Total percent composition, based on ionic abundance, of different lipids species is reported in Table 3. TAG (61.34 ± 0.60%) were the prevalent lipids followed by LDGTS, GlcADG, DGTS and SQDG. In particular, in the ESI + mode, the lipid profile, expressed as µg/mL and percent composition, is reported in Table 4. The most abundant triacylglycerols were: TAG (16:0_18:1_18:1), TAG (16:0_18:1_18:2), TAG (18:1_18:1_18:1), and TAG (18:1_18:1_18:2) with a carbon number of annotated TAG comprised between 48 and 56 with 9 as the maximum number of insaturation. We were also able to annotate different lysobetaine lipids, such as lysodiacylglyceryltrimethylhomoeserine 16:0 (LDGTS 16:0) and different betaine lipids diacylglyceryltrimethylhomoeserine with carbon number, comprised between 18 and 34 and 3 as the maximum number of insaturations. Interestingly, based on their abundance, we measured the percent levels of different glucoronosyl diacylglycerol 34:1 and 34:2 and sulfoquinovosyldiacylglycerols (SQDG): SQDG (16:0_16:0), SQDG (16:0_18:2), SQDG (16:0_18:3) and SQDG (16:0_18:1).
In the ESI mode, microalga samples showed the most abundant fatty acids (Table 5) were the oleic acid (FA 18:1, C18:1omega9), linoleic acid (FA 18:2, C18:2omega6), palmitic acid (FA 16:0), and stearic acid (FA 18:0) at 29.11, 20.57, 19.38 and 17.13%, respectively. Complex lipids determined in the negative ionization mode were dominated by phosphatidylglycerol (16:1_18:2) with a percent level of 73.29% (Table 6). Noteworthy, in C. melkonianii samples, we were able to annotate in branched fatty acid esters of hydroxy fatty acids (FAHFA). These compounds represent a class of functional lipids called lipokines. The comparison of annotated FAHFA showed that C. melkonianii samples are characterized by a unique fingerprint of FAHFA, such as FAHFA (18:1_20:3), FAHFA (18:2_20:4), FAHFA (18:0_20:2), and FAHFA (18:1_18:0). FAHFAs were recently identified as a class of bioactive lipids with anti-diabetic and anti-inflammatory activities (Yore et al. 2014). Lower levels of mono- and di-galactosyldiacylglycerols (MGDG and DGDG, respectively), ceramide alpha-hydroxy fatty acid-dihydrosphingosines (Cer_ADS), ceramide non-hydroxyfatty acid-hihydrosphingosine (Cer_NDS), ceramide alpha-hydroxy fatty acid-phytospingosine (Cer_AP) and lysophatidylcholine LPC (18:2_0:0) were also found. The ratio between the two major galactolipids, MGDG and DGDG, is variable in microalgae since it is strongly influenced by responses to environmental and nutritional cues (Khozin-goldberg 2016). Higher-plant and microalgae chloroplast are made of four characteristic lipids, PG, MGDG, DGDG, and SQDG (Shimojima 2011; Boudière et al. 2014; Da Costa et al. 2016). The latter is sulfur-containing anionic glycerolipid component of photosynthetic membrane lipids, UDP-sulfoquinovose synthase (SQD1) and SQDG synthase (SQD2) which is responsible of the SQDG biosynthesis (Shimojima 2011). The synthesis of SQDG start from the entering of sulfates into the microalgae chloroplast. Sulfates are first transformed to adenosine 5’-phosphosulfate by iron − sulfur flavoenzyme adenosine-5 ‘-phosphosulfate (APS) and then to sulphites. The enzymatic complex UDP-sulfoquinovose synthase and ferredoxin-dependent glutamate synthase catalyze the reaction of sulphites with UDP-glucose to form UDP-sulfoquinovose (UDP-SQ). SQDG synthase (SQD2) catalyzes the reaction of UDP-SQ with DAG to form SQDG (Shimojima 2011). In the environment, this pathway may allow the microalgae C. melkonianii to efficiently cope with high levels of sulfates measured in the Rio Irvi, i.e., 3694 mg/L. In summary, in this work, we reported the primary metabolic changes during growth and the lipidomic characterization of the extremophile C. melkionanii SCCA48.
The changes of cellular polar metabolites during growth reported in this work may be potentially useful to control and adapt biofuel synthesis in Trebouxiophyceae. Besides, FA (i.e., oleic acid, linoleic acid, palmitic acid and stearic acid), this strain showed high levels of the following lipid classes: TAGs, LSGTSs, SQSGs and DGTS. Our results give a basic biochemical, metabolomic and lipidomic, snapshot of this algae to be used for the engineering of biological phycoremediation filters able to reduce freshwaters pollution as open raceway pond or closed photobioreactor. Furthermore, the in-depth lipidomic characterization of this strains opens the road to produce isotopically labeled standards of lipids to be used in the field of biochemistry.
Abbreviations
- TAG:
-
Triacylglycerol
- DAG:
-
Diacylglycerol
- PG:
-
Phosphatidylglycerol
- PC:
-
Phosphatidylcholine
- PE:
-
Phosphatidylethanolamine
- LPC:
-
Lysophophatidylcholine
- SM:
-
Sphingomyelin
- DGDG:
-
Digalactosyldiacylglycerol
- DGTS:
-
Diacylgyceryltrimethylhomoserine
- LDGTS:
-
Lysodiacylglyceryltrimethylhomoserine
- MGDG:
-
Monogalactosyldiacylglycerol
- SQDG:
-
Sulfoquinovosyl diacylglycerol
- GlcADG:
-
Glucoronosyldiacylglycerol
- Cer_ADS:
-
Ceramide alpha-hydroxy fatty acid-dihydrosphingosine
- Cer_AP:
-
Ceramide alpha-hydroxy fatty acid-phytospingosine
- Cer_NDS:
-
Ceramide non-hydroxyfatty acid-dihydrosphingosine
- FA:
-
Free fatty acid
- FAHFA:
-
Fatty acid ester of hydroxyl fatty acid
References
Allen JW, DiRusso CC, Black PN (2015) Triacylglycerol synthesis during nitrogen stress involves the prokaryotic lipid synthesis pathway and acyl chain remodeling in the microalgae Coccomyxa subellipsoidea. Algal Res 10:110–120. https://doi.org/10.1016/j.algal.2015.04.019
Arora N, Pienkos PT, Pruthi V, Poluri KM, Guarnieri MT (2018) Leveraging algal omics to reveal potential targets for augmenting TAG accumulation. Biotechnol Adv 36:1274–1292. https://doi.org/10.1016/j.biotechadv.2018.04.005
Asemave K (2018) Greener chelators for recovery of metals and other applications. Org Med Chem Int J. https://doi.org/10.19080/omcij.2018.06.555694
Assunção MFG, Amaral R, Martins CB, Ferreira JD, Ressurreição S, Santos SD, Varejão JMTB, Santos LMA (2017) Screening microalgae as potential sources of antioxidants. J Appl Phycol 29:865–877. https://doi.org/10.1007/s10811-016-0980-7
Behrens PW, Sicotte VJ, Delente J (1994) Microalgae as a source of stable isotopically labeled compounds. J Appl Phycol 6:113–121. https://doi.org/10.1007/BF02186065
Blaby IK, Glaesener AG, Mettler T, Fitz-Gibbon ST, Gallaher SD, Liu B, Boyle NR, Kropat J, Stitt M, Johnson S, Benning C, Pellegrini M, Casero D, Merchant SS (2013) Systems-level analysis of nitrogen starvation-induced modifications of carbon metabolism in a Chlamydomonas reinhardtii starchless mutant. Plant Cell 25:4305–4323. https://doi.org/10.1105/tpc.113.117580
Blifernez-Klassen O, Chaudhari S, Klassen V, Wördenweber R, Steffens T, Cholewa D, Niehaus K, Kalinowski J, Kruse O (2018) Metabolic survey of Botryococcus braunii: Impact of the physiological state on product formation. PLoS ONE 13:1–23. https://doi.org/10.1371/journal.pone.0198976
Bouché N, Fromm H (2004) GABA in plants: just a metabolite? Trends Plant Sci 9:110–115. https://doi.org/10.1016/j.tplants.2004.01.006
Boudière L, Michaud M, Petroutsos D, Rébeillé F, Falconet D, Bastien O, Roy S, Finazzi G, Rolland N, Jouhet J, Block MA, Maréchal E (2014) Glycerolipids in photosynthesis: composition, synthesis and trafficking. Biochim Biophys Acta Bioenerg 1837:470–480. https://doi.org/10.1016/j.bbabio.2013.09.007
Brunner U, Honegger R (1981) Sporopollenin in the cell walls of Coccomyxa and Myrmecia phycobionts of various lichens: an ultrastructural and chemical investigation. Can J Bot 59:2713–2734
Büdel B, Darienko T, Deutschewitz K, Dojani S, Friedl T, Mohr KI, Salisch M, Reisser W, Weber B (2009) Southern african biological soil crusts are ubiquitous and highly diverse in drylands, being restricted by rainfall frequency. Microb Ecol 57:229–247. https://doi.org/10.1007/s00248-008-9449-9
Cannell RJP (1993) Algae as a source of biologically active products. Pestic Sci 39:147–153. https://doi.org/10.1002/ps.2780390208
Cavacini P (2001) Soil algae from northern Victoria Land (Antarctica). Polar Biosci 14:45–60
Chen H, Zheng Y, Zhan J, He C, Wang Q (2017) Comparative metabolic profiling of the lipid-producing green microalga Chlorella reveals that nitrogen and carbon metabolic pathways contribute to lipid metabolism. Biotechnol Biofuels 1:1–20. https://doi.org/10.1186/s13068-017-0839-4
Cho K, Kim KN, Lim NL, Kim MS, Ha JC, Shin HH, Kim MK, Roh SW, Kim D, Oda T (2015) Enhanced biomass and lipid production by supplement of myo-inositol with oceanic microalga Dunaliella salina. Biomass Bioenerg 72:1–7. https://doi.org/10.1016/j.biombioe.2014.11.014
Da Costa E, Silva J, Mendonça SH, Abreu MH, Domingues MR (2016) Lipidomic approaches towards deciphering glycolipids from microalgae as a reservoir of bioactive lipids. Mar Drugs. https://doi.org/10.3390/md14050101
Darienko T, Gustavs L, Eggert A, Wolf W, Pröschold T (2015) Evaluating the species boundaries of green microalgae (Coccomyxa, Trebouxiophyceae, Chlorophyta) using integrative taxonomy and DNA barcoding with further implications for the species identification in environmental samples. PLoS ONE 10:1–31. https://doi.org/10.1371/journal.pone.0127838
De Giudici G, Medas D, Cidu R, Lattanzi P, Podda F, Frau F, Rigonat N, Pusceddu C, Da Pelo S, Onnis P, Marras PA, Wanty RB, Kimball B (2018) Application of hydrologic-tracer techniques to the Casargiu adit and Rio Irvi (SW-Sardinia, Italy): using enhanced natural attenuation to reduce extreme metal loads. Appl Geochemistry 96:42–54. https://doi.org/10.1016/j.apgeochem.2018.06.004
De Morais MG, Vaz BDS, De Morais EG, Costa JAV (2015) Biologically active metabolites synthesized by microalgae. Biomed Res Int. https://doi.org/10.1155/2015/835761
Dunn WB, Ellis DI (2005) Metabolomics: current analytical platforms and methodologies. TrAC Trends Anal Chem 24:285–294. https://doi.org/10.1016/j.trac.2004.11.021
Fermani P, Mataloni G, Van De Vijver B (2007) Soil microalgal communities on an antarctic active volcano (Deception Island, South Shetlands). Polar Biol 30:1381–1393. https://doi.org/10.1007/s00300-007-0299-6
Flechtner VR, Pietrasiak N, Lewis LA (2013) Newly revealed diversity of green microalgae from wilderness areas of joshua tree National Park (JTNP). Monogr West North Am Nat 6:43–63. https://doi.org/10.3398/042.006.0103
Folch J, Lees M, Sloane Stanley G (1987). A Simple 55: 999–1033.
Grillet L, Ouerdane L, Flis P, Hoang MTT, Isaure MP, Lobinski R, Curie C, Mari S (2014) Ascorbate efflux as a new strategy for iron reduction and transport in plants. J Biol Chem 289:2515–2525. https://doi.org/10.1074/jbc.M113.514828
Guedes AC, Amaro HM, Malcata FX (2011) Microalgae as sources of carotenoids. Mar Drugs 9:625–644. https://doi.org/10.3390/md9040625
Guiry MD, Guiry GM (2020). No Title [WWW Document]. AlgaeBase. World-wide Electron. Publ. Natl. Univ. Ireland, Galw. https://www.algaebase.org
Guschina IA, Harwood JL (2006) Lipids and lipid metabolism in eukaryotic algae. Prog Lipid Res 45:160–186. https://doi.org/10.1016/j.plipres.2006.01.001
Halim R, Hosikian A, Lim S, Danquah MK (2010) Chlorophyll extraction from microalgae: a review on the process engineering aspects. Int J Chem Eng. https://doi.org/10.1155/2010/391632
Heesch S, John GD, Yamagishi T, Kawai H, Müller DG, Küpper FC (2012) Cryopreservation of the model alga ectocarpus (phaeophyceae). Cryo-Letters 33:327–336
Hirooka S, Hirose Y, Kanesaki Y, Higuchi S, Fujiwara T, Onuma R, Era A, Ohbayashi R, Uzuka A, Nozaki H, Yoshikawa H, Miyagishima SY (2017) Acidophilic green algal genome provides insights into adaptation to an acidic environment. Proc Natl Acad Sci USA 114:E8304–E8313. https://doi.org/10.1073/pnas.1707072114
Ho SH, Ye X, Hasunuma T, Chang JS, Kondo A (2014) Perspectives on engineering strategies for improving biofuel production from microalgae - A critical review. Biotechnol Adv 32:1448–1459. https://doi.org/10.1016/j.biotechadv.2014.09.002
Ho SH, Nakanishi A, Ye X, Chang JS, Chen CY, Hasunuma T, Kondo A (2015) Dynamic metabolic profiling of the marine microalga Chlamydomonas sp. JSC4 and enhancing its oil production by optimizing light intensity Luisa Gouveia. Biotechnol Biofuels 8:1–17. https://doi.org/10.1186/s13068-015-0226-y
Hodač L, Hallmann C, Spitzer K, Elster J, Faßhauer F, Brinkmann N, Lepka D, Diwan V, Friedl T (2016) Widespread green algae Chlorella and Stichococcus exhibit polar-temperate and tropical-temperate biogeography. FEMS Microbiol Ecol 92:1–16. https://doi.org/10.1093/femsec/fiw122
Ito T, Tanaka M, Shinkawa H, Nakada T, Ano Y, Kurano N, Soga T, Tomita M (2013) Metabolic and morphological changes of an oil accumulating trebouxiophycean alga in nitrogen-deficient conditions. Metabolomics 9:178–187. https://doi.org/10.1007/s11306-012-0463-z
Jeong E-Y, Seo PJ, Woo JC, Park C-M (2011) Fatty acid profiles and their distribution patterns in microalgae: a comprehensive analysis of more than 2000 strains from the SAG culture collection. BMC Plant Biol 11(1):1–6
Juárez ÁB, Vélez CG, Iñiguez AR, Martínez DE, Rodríguez MC, Vigna MS, De Molina MDCR (2011) A Parachlorella kessleri (Trebouxiophyceae, Chlorophyta) strain from an extremely acidic geothermal pond in Argentina. Phycologia 50:413–421. https://doi.org/10.2216/10-79.1
Kalinowska R, Pawlik-Skowrońska B (2010) Response of two terrestrial green microalgae (Chlorophyta, Trebouxiophyceae) isolated from Cu-rich and unpolluted soils to copper stress. Environ Pollut 158:2778–2785. https://doi.org/10.1016/j.envpol.2010.03.003
Khozin-goldberg I (2016) The physiology of microalgae. Physiol Microalgae. https://doi.org/10.1007/978-3-319-24945-2
Kobayashi T, Nishizawa NK (2012) Iron uptake, translocation, and regulation in higher plants. Annu Rev Plant Biol 63:131–152. https://doi.org/10.1146/annurev-arplant-042811-105522
Koechler S, Bertin PN, Plewniak F, Baltenweck R, Casiot C, Heipieper HJ, Bouchez O, Arsène-Ploetze F, Hugueney P, Halter D (2016) Arsenite response in Coccomyxa sp. Carn explored by transcriptomic and non-targeted metabolomic approaches. Environ Microbiol 18:1289–1300. https://doi.org/10.1111/1462-2920.13227
Kothe E, Bergmann H, Büchel G (2005) Supplement 1. Chem Erde 65:7–27. https://doi.org/10.1016/j.chemer.2005.06.005
la Rocca N, Andreoli C, Giacometti GM, Rascio N, Moro I (2009) Responses of the Antarctic microalga Koliella antarctica (Trebouxiophyceae, Chlorophyta) to cadmium contamination. Photosynthetica 47:471–479. https://doi.org/10.1007/s11099-009-0071-y
Lee N, Ko SR, Ahn CY, Oh HM (2018) Optimized co-production of lipids and carotenoids from Ettlia sp. by regulating stress conditions. Bioresour Technol 258:234–239. https://doi.org/10.1016/j.biortech.2018.03.006
Leonardo T, Farhi E, Pouget S, Motellier S, Boisson AM, Banerjee D, Rébeillé F, Den Auwer C, Rivasseau C (2016) Silver accumulation in the green microalga coccomyxa actinabiotis: toxicity, in situ speciation, and localization investigated using synchrotron XAS, XRD, and TEM. Environ Sci Technol 50:359–367. https://doi.org/10.1021/acs.est.5b03306
Malavasi V, Cao G (2015) Applications 101:273–283. https://doi.org/10.1127/nova
Malavasi V, Škaloud P, Rindi F, Tempesta S, Paoletti M, Pasqualetti M (2016) DNA-based taxonomy in ecologically versatile microalgae: A re-evaluation of the species concept within the coccoid green algal genus Coccomyxa (Trebouxiophyceae, Chlorophyta). PLoS ONE 11:1–25. https://doi.org/10.1371/journal.pone.0151137
Nicodemus TJ, DiRusso CC, Wilson M, Black PN (2020) Reactive Oxygen Species (ROS) mediated degradation of organophosphate pesticides by the green microalgae Coccomyxa subellipsoidea. Bioresour Technol Reports 11:100461. https://doi.org/10.1016/j.biteb.2020.100461
Pasqualetti M, Tempesta S, Malavasi V, Barghini P, Fenice M (2015) Lutein production by coccomyxa sp. SCCA048 isolated from a heavy metal-polluted river in Sardinia (Italy). J Environ Prot Ecol 16:1262–1272
Red E, Galdieria M, Sulphurarİa G, Araştirilmasi DA (2016) Investigation of the malic acid concentration on on extremophilic red microalga galdieria sulphuraria. Dokuz Eylül Üniversitesi Mühendislik Fakültesi Fen ve Mühendislik Dergisi, pp 101–107
Řezanka T, Nedbalová L, Barcytė D, Vítová M, Sigler K (2019) Arsenolipids in the green alga Coccomyxa (Trebouxiophyceae, Chlorophyta). Phytochemistry 164:243–251. https://doi.org/10.1016/j.phytochem.2019.05.002
Rivasseau C, Farhi E, Compagnon E, Cyr GS (2016) Coccomyxa actinabiotis sp. nov. (Trebouxiophyceae, Chlorophyta), a new green microalga living in the spent fuel cooling pool of a nuclear reactor. J Phycol 52:689–703. https://doi.org/10.1111/jpy.12442
Rochfort S (2005) Biology and implications for natural products research. J Nat Prod 68:1813–1820. https://doi.org/10.1021/np050255w
Scano P, Murgia A, Pirisi FM, Caboni P (2014) A gas chromatography-mass spectrometry-based metabolomic approach for the characterization of goat milk compared with cow milk. J Dairy Sci 97:6057–6066. https://doi.org/10.3168/jds.2014-8247
Shimojima M (2011) Biosynthesis and functions of the plant sulfolipid. Prog Lipid Res 50:234–239. https://doi.org/10.1016/j.plipres.2011.02.003
Smirnoff N (2018) Ascorbic acid metabolism and functions: a comparison of plants and mammals. Free Radic Biol Med 122:116–129. https://doi.org/10.1016/j.freeradbiomed.2018.03.033
Soru S, Malavasi V, Caboni P, Concas A, Cao G (2019a) Behavior of the extremophile green alga C. melkonianii SCCA 048 in terms of lipids production and morphology at different pH values. Extremophiles 23:79–89. https://doi.org/10.1007/s00792-018-1062-3
Soru S, Malavasi V, Concas A, Caboni P, Cao G (2019b) A novel investigation of the growth and lipid production of the extremophile microalga C. melkonianii SCCA 048 under the effect of different cultivation conditions: experiments and modeling. Chem Eng J 377:1–14. https://doi.org/10.1016/j.cej.2018.12.049
Sukla LB, Pradhan D (2019) Lala behari sukla and enketeswara subudhi and debabrata pradhan - the role of microalgae in wastewater treatment. Springer
Tautenhahn R, Patti GJ, Rinehart D, Siuzdak G (2012) XCMS online: a web-based platform to process untargeted metabolomic data. Anal Chem 84:5035–5039. https://doi.org/10.1021/ac300698c
Tragin M, Vaulot D (2018) Green microalgae in marine coastal waters: the Ocean sampling day (OSD) dataset. Sci Rep 8:1–12. https://doi.org/10.1038/s41598-018-32338-w
Treves H, Murik O, Kedem I, Eisenstadt D, Meir S, Rogachev I, Szymanski J, Keren N, Orf I, Tiburcio AF, Alcázar R, Aharoni A, Kopka J, Kaplan A (2017) Metabolic flexibility underpins growth capabilities of the fastest growing alga. Curr Biol 27:2559-2567.e3. https://doi.org/10.1016/j.cub.2017.07.014
Urzica EI, Adler LN, Page MD, Linster CL, Arbing MA, Casero D, Pellegrini M, Merchant SS, Clarke SG (2012) Impact of oxidative stress on ascorbate biosynthesis in Chlamydomonas via regulation of the VTC2 gene encoding a GDP-L-galactose phosphorylase. J Biol Chem 287:14234–14245. https://doi.org/10.1074/jbc.M112.341982
Van Meer G, Voelker DR, Feigenson GW (2008) Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol 9:112–124. https://doi.org/10.1038/nrm2330
Vidoudez C, Pohnert G (2012) Comparative metabolomics of the diatom Skeletonema marinoi in different growth phases. Metabolomics 8:654–669. https://doi.org/10.1007/s11306-011-0356-6
Wase N, Black PN, Stanley BA, Dirusso CC (2014) Integrated quantitative analysis of nitrogen stress response in Chlamydomonas reinhardtii using metabolite and protein profiling. J Proteome Res 13:1373–1396. https://doi.org/10.1021/pr400952z
Xue J, Wang L, Zhang L, Balamurugan S, Li DW, Zeng H, Yang WD, Liu JS, Li HY (2016) The pivotal role of malic enzyme in enhancing oil accumulation in green microalga Chlorella pyrenoidosa. Microb Cell Fact 15:1–9. https://doi.org/10.1186/s12934-016-0519-2
Yore MM, Syed I, Moraes-Vieira PM, Zhang T, Herman MA, Homan EA, Patel RT, Lee J, Chen S, Peroni OD, Dhaneshwar AS, Hammarstedt A, Smith U, McGraw TE, Saghatelian A, Kahn BB (2014) Discovery of a class of endogenous mammalian lipids with anti-diabetic and anti-inflammatory effects. Cell 159:318–332. https://doi.org/10.1016/j.cell.2014.09.035
Yu XJ, Sun J, Zheng JY, Sun YQ, Wang Z (2016) Metabolomics analysis reveals 6-benzylaminopurine as a stimulator for improving lipid and DHA accumulation of Aurantiochytriumsp. J Chem Technol Biotechnol 91:1199–1207. https://doi.org/10.1002/jctb.4869
Acknowledgements
The financial support of the COMISAR project (POR FESR 2014/2020 – Asse prioritario I “ricerca scientifica, sviluppo tecnologico e innovazione” Regione Autonoma della Sardegna, Italy) is gratefully acknowledged. G.F. acknowledges the financial support obtained from the University of Cagliari during his Ph.D. program in Innovation Sciences and Technologies.
Funding
Open access funding provided by Università degli Studi di Cagliari within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
GF conducted experiments, analyzed data, and edited the manuscript, VM designed research, conducted experiments, and edited the manuscript. PS analyzed data and edited the manuscript. SS conducted experiments and edited the manuscript. PC conducted experiments, analyzed data, and edited the manuscript. Fund acquisition, project administration, and supervision by GC All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by S. Albers.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fais, G., Malavasi, V., Scano, P. et al. Metabolomics and lipid profile analysis of Coccomyxa melkonianii SCCA 048. Extremophiles 25, 357–368 (2021). https://doi.org/10.1007/s00792-021-01234-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-021-01234-z