Abstract
Key message
The durable late blight resistance in potato plant Ma R9 is genetically characterized. A novel R -gene is mapped. The monogenic nature and map positions of R9 are negated and rectified.
Abstract
Late blight of potato (Solanum tuberosum), caused by Phytophthora infestans, can effectively be managed by genetic resistance. The MaR9 differential plant provides durable resistance to a broad spectrum of late blight strains. This resistance is brought about by at least seven genes derived from S. demissum including R1, Rpi-abpt1, R3a, R3b, R4, R8 and, so far uncharacterized resistance gene(s). Here we set out to genetically characterize this additional resistance in MaR9. Three BC1 populations derived from MaR9 were identified that segregated for IPO-C resistance but that lacked R8. One BC1 population showed a continuous scale of resistance phenotypes, suggesting that multiple quantitative resistance genes were segregating. In two other BC1 populations resistance and susceptibility were segregating in a 1:1 ratio, suggesting a single qualitative resistance gene (R9a). A chromosome IX PCR marker, 184-81, fully co-segregated with R9a. The map position of R9a on the distal end of the lower arm of chromosome IX was confirmed using PCR markers GP101 and Stm1021. Successively, cluster-directed profiling (CDP) was carried out, revealing six closely linked markers. CDPSw58, CDPSw59 and CDPSw510 flanked the R9a gene at the distal end (5.8 cM) and, as expected, were highly homologous to Sw-5. CDPTm22 flanked R9a on the proximal side (2.9 cM). CDPTm26 and CDPTm27 fully co-segregated with resistance and had high homology to Tm-2 2, showing that R9a resides in a cluster of NBS–LRR genes with homology to Tm-2 2. Besides R9a, additional resistance of quantitative nature is found in MaR9, which remains to be genetically characterized.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Late blight of potato (Solanum tuberosum) is a devastating disease caused by the oomycete pathogen Phytophthora infestans (Pi) (Fry 2008). This pathogen is a diploid, heterothallic fungus-like oomycete with two mating types (A1 and A2). Over the past several decades, with the increased incidence of late blight epidemics worldwide, the coexistence of both mating types has been observed throughout the world rather than the confinement inside of Mexico (Fry et al. 1993; Goodwin et al. 1994). Consequently, there is a possibility of sexual recombination and increased genetic diversity among progeny of the pathogen which increases the difficulty in late blight control. Furthermore, developing resistance to fungicides in Pi populations, which has been demonstrated by widespread resistance to metalaxyl, a key component of fungicides for potato production, requires more frequent applications during the season to control late blight, causing contamination harmful to both humans and environment (Deahl et al. 1993; Goodwin et al. 1994; Grünwald et al. 2001).
Genetic disease resistance has long been considered a promising method for the management of late blight as an alternative to fungicides and healthy seed tubers for late blight control. Two kinds of resistance against late blight have been described in potato: qualitative resistance and quantitative resistance (Rauscher et al. 2010). Late blight qualitative resistance is governed by resistance (R) genes that encode immunity through a hypersensitive reaction and is thought to be monogenic, specific, and often of short durability due to the high genetic variability of the pathogen. Since the early part of the last century breeding activities have mainly focused on the high level of resistance conferred by dominant major R genes from the Mexican hexaploid wild Solanum species, Solanum demissum. Presently, many cultivars with S. demissum in their pedigree are available (Umaerus et al. 1983; Šwiežynski et al. 1997).
Eleven race-specific R genes, named R1–R11, have been proposed in S. demissum and introduced into potatoes (Black 1951; Black et al. 1953; Malcolmson and Black 1966). A current international set of potato R gene differentials comprises 11 clones originating from S. demissum (Trognitz and Trognitz 2007) for the detection of late blight virulence factors. The Dutch differential set collected by Mastenbroek (1952) is also known as the Mastenbroek differential set: MaR1 to MaR11. MaR1 to MaR4 were developed by Mastenbroek and the other R gene differentials are identical to the Scottish differential set developed by Black (Huang 2005). Seven genes controlling late blight resistance within this differential set have been mapped: R1 on chromosome V (Leonards-Schippers et al. 1992), R2 on chromosome IV (Li et al. 1998), R3a, R3b, R4, R6 and R7 on chromosome XI (El-Kharbotly et al. 1996; Huang et al. 2005; Verzaux 2010) and R8 on chromosome IX (Jo et al. 2011). Although the differential set was initially thought to represent single late blight resistance factors, many exceptions have been observed: R1 was also found in the MaR5, MaR6 and MaR9 differentials (Trognitz and Trognitz 2007) and the MaR3 differential plant contained two R genes, R3a and R3b (Huang et al. 2005). Even in the differentials MaR8 and MaR9, respectively, at least four (R3a, R3b, R4 and R8) and seven (R1, Rpi-abpt1, R3a, R3b, R4, R8 and R9) R genes were present (Kim et al. 2012). In the past, R genes from MaR1, MaR2, MaR3, MaR4 and MaR10 were rapidly overcome (Wastie 1991), but S. demissum is still considered a valuable source for resistance (Niederhauser and Mills 1953; Colon et al. 1995). Especially, the MaR8 and MaR9 have been reported to show broad spectrum resistance both under laboratory and under field conditions (Fry and Goodwin 1997; Šwiežynski et al. 2000; Haynes et al. 2002; Bisognin et al. 2002; Zhang and Kim 2007). Recently, it was shown that this broad spectrum resistance is a result of R gene stacking and/or a result of individual broad spectrum R genes like R8, using a “de-stacking” approach and an “on site” Pi virulence monitoring system (Kim et al. 2012). Stacking of multiple late blight R genes in different Solanum accessions has been revealed (Verzaux 2010) and is most likely a natural defence strategy against the highly flexible late blight pathogen. Also for late blight resistance breeding, stacking of multiple R genes seems mandatory to provide sufficient durability (Jo 2013). So far, over 20 functional late blight R genes have been cloned and all belong to the CC–NB–LRR class. These include four Solanum demissum genes R1 (Ballvora et al. 2002), R2 (Lokossou et al. 2009), R3a (Huang et al. 2005), and R3b (Li et al. 2011) and R genes derived from wild Solanum species like S. bulbocastanum (Song et al. 2003; van der Vossen et al. 2003, 2005; Lokossou et al. 2009), S. stoloniferum and S. papita (Vleeshouwers et al. 2008), S. venturii (Pel et al. 2009; Foster et al. 2009) , S. mochiquense (Jones et al. 2009), S. chacoense (Vossen et al. 2010), and S. x edinense (de Vetten et al 2011).
Quantitative resistance, usually due to the effect of multiple genes of minor effects, is characterized by a slower development of the disease, considered to be race non-specific and influenced by environmental conditions (Wastie 1991). Quantitative resistance locus (QRL) mapping enables to identify multiple loci with important phenotypic effects throughout the genome and characterize epistatic interactions among these loci (Kover and Caicedo 2001). In potato, late blight QRL studies have revealed that some QRLs co-localize with positions of known R genes (Stewart et al. 2003; Tan et al. 2008; Rauscher et al. 2010), R gene clusters (Gebhardt and Valkonen 2001), and defence genes (Trognitz et al. 2002). Interestingly, some R genes were mapped as both qualitative and quantitative traits like R11 or a quantitative trait like R10 (Bradshaw et al. 2006). Quantitative resistance has also been reported to be present in plants carrying defeated R genes such as R1, R2, R10, and R11 (Stewart et al. 2003; Pilet et al. 2005) and Rpi-ber (Rauscher et al. 2010) in the presence of compatible isolates.
In our previous study (Kim et al. 2012), it has been suggested that the MaR9 differential contained multiple R genes. The present study describes the genetic dissection of the uncovered, remaining resistance from potato differential set plant MaR9. Using a series of BC1 populations it was found that the remaining resistance is conferred by a qualitative resistance gene R9a as well as by an uncharacterized quantitative resistance. Using a dedicated set of candidate markers, polymorphisms were identified that were linked to, and co-segregated with, R9a. It was concluded that R9a resides at the bottom end of Chromosome IX, proximal to the R8 locus. Thereby, the previous suggestion that R9 would be an allelic variant of R3 on chromosome XI (Huang et al. 2005) was rejected.
Materials and methods
Plant materials
The MaR9 differential, corresponding to 2573(2) and LB1 (Black et al. 1953; Malcolmson and Black 1966), was crossed as female parent with the cultivar Concurrent (which contains R10). BC1 populations were generated by crossing resistant F1 progenies as female parents with susceptible cultivar Katahdin as a male parent and six populations were selected (3150, 3151, 3153, 3154, 3155 and 3247). To further characterize 3151, the BC2 population 3253 was made by crossing R8 and R9a lacking resistant seedling 3151-03 from BC1 population 3151 with cultivar Desiree as a male parent. Neither of the F1, BC1 and BC2 male parents contains R genes that cause incompatibility of the P. infestans IPO-C isolate. Seeds were sown under sterile conditions, and plants were maintained in in vitro culture and propagated for multiple field trials and for the whole plant assays in a climate cell.
Phytophthora infestans isolate and late blight resistance tests
Phytophthora infestans isolate IPO-C (race 1, 2, 3a, 3b, 4, 5, 6, 7, 10, 11) was used in both field trials and whole plant climate cell assays. Field trials were done as described by Jo et al. (2011). For whole plant climate cell assays, seedlings from population 3151 and 3154 and 3253 were planted in pots and grown in the greenhouse at 22 °C with a 10 h day/14 h night photoperiod and a relative humidity of 70–80 %. One month after growth of plants, they were transferred into a growth chamber maintained at 15 °C with a photoperiod of 16 h/8 h day/night at 70 % relative humidity. Inoculum preparation and inoculation were performed essentially as described by Vleeshouwers et al. (1999). Three leaves per plant for two plants of each genotype of the populations were inoculated with 10 µl droplets of inoculum (5 × 104 zoospores/ml). 100 % humidity was maintained for the first 3 days after inoculation, after which a humidity of 70 % was restored. Seven days after inoculation phenotypes were classified into three groups, resistant (no symptoms, hypersensitive (HR) lesions), susceptible (sporulating lesions on all inoculated leaves), or intermediate phenotypes. For 3151 and 3154 genotypes that were tested both in the climate cell and in the field trial, there was a complete agreement between field trial and climate cell late blight trials. In the field, scoring was performed by estimating the percentage of blight-affected leaf area.
Agroinfiltration assay
Two leaves per plant for three plants of each of the BC1 genotypes were infiltrated with the following constructs: Avr8, 1:1 mixture of R3a and Avr3a (Bos et al. 2006) as the positive control, and empty pK7FWG2.0 (Karimi et al. 2002) as the negative control. Agroinfiltration assay was carried out as described by Rietman et al. (2012). Agrobacterium tumefaciens strain AGL1 + pVirG was used as a carrier of the binary plasmids. A. tumefaciens strains were maintained as glycerol stocks and were grown in LB medium supplemented with appropriate antibiotics (R3a construct; 50 mg/L kanamycin and 75 mg/L chloramphenicol, Avr8 and Avr3a construct; 50 mg/L carbenicilin and 75 mg/L chloramphenicol) at 28 °C overnight. One day before agroinfiltration, the cultures were reinoculated in YEB medium (5 g beef extract, 5 g bacteriological peptone, 5 g sucrose, 1 g yeast extract, 2 ml 1 M MgSO4 in 1 litre of milli-Q water) supplemented with antibiotics, 10 µl of 200 mM acetosyringone and 1000 µl of 1 M MES pH = 5.5. On the day of agroinfiltration, the cells were harvested and resuspended in MMA solution (20 g sucrose, 5 g MS salts and 1.95 g MES in 1 litre of distilled water, adjusted to pH5.5) supplemented with 1 ml of 200 mM acetosyringone to a final OD600 of 0.3. The leaves of 4- to 5-week-old potato plants were infiltrated with this suspension. Responses were scored 3–4 days after infiltration.
DNA isolation and marker analysis
Total genomic DNA was isolated from young leaves as described by Fulton et al. (1995). The Retsch machine (RETSCH Inc., Hannover, Germany) was used to grind young plant materials frozen in liquid nitrogen. For the identification of R gene-specific clusters, a modification of the NBS profiling protocol of van der Linden et al. (2004) was carried out as described by Jo et al. (2011). The restriction ligation reaction was done using MseI restriction enzyme and eight Tm-2 2 primers (Tm1R, Tm2F, Tm3F, Tm3R, Tm6F, Tm15F, Tm15R, and Tm19F) described by Verzaux (2010) were used for the successive PCRs. For Sw-5-CDP, seven specific primers were designed on cluster-specific conserved domains encoding CC and LRR after the alignment of Sw-5 sequences available from NCBI (http://www.ncbi.nlm.nih.gov/). The Sw-5-specific primers described by Dianese et al. (2010) were also used for Sw-5-CDP. The CDP primers were used in combination with a labelled adapter primer (fluorescent dye IRD700) and labelled R gene-targeted PCR products were separated on a denaturing polyacrylamide gel using a NEN® IR2 DNA analyser (LI-COR® Biosciences, Lincoln, NE, USA). A set of the ten resistant and the ten susceptible BC1 plants, including parents, was used to obtain CDP markers linked to resistance in mapping population (Vossen et al. 2013). If linked CDP markers were found, a second round of CDP was applied to all the remaining individuals of a segregating population. Fragments were excised as described in the Odyssey® manual for band extraction (Westburg, The Netherlands) and re-amplified with the specific profiling primer and the adaptor primer. The sizes of PCR products were checked on polyacrylamide gels and fragments were cloned into the pGEM-T Easy vector (Promega, USA) prior to sequencing with M13 primers. Sequencing was carried out with the BigDye Terminator kit and an ABI 3700 automated sequencer from Applied Biosystems (USA). The marker nomenclature followed as described by Jo et al. (2011). PCRs for GP101, 184-81, and Stm1021 were performed using DreamTaqTM polymerase (Fermentas) in a standard PCR program (94 °C for 60 s followed by 30 cycles of 94 °C for 30 s, 58 °C for 60 s, 72 °C for 90 s and a final extension time of 5 min at 72 °C). To screen for cleaved amplified polymorphic sequences (CAPS), PCR was done using primers listed in Table 1 and subsequently PCR products were digested using the restriction enzymes listed in Table 1. For SSR, a labelled forward primer (fluorescent dye IRD800) was used to enable visualization on a denaturing polyacrylamide gel using a NEN® IR2 DNA analyser (LI-COR® Biosciences, Lincoln, NE, USA). Tm-2 2-like or Sw-5-like sequences available from NCBI (http://www.ncbi.nlm.nih.gov/) and S. tuberosum Group Phureja DM1-3 516R44 (CIP801092) Genome Annotation v3.4 (based on v3 superscaffolds) PGSC_DM_v3.4_gene.fasta (http://potatogenomics.plantbiology.msu.edu/blast.html) available from the Potato Genome Sequencing Consortium (PGSC), were collected and aligned using DNASTAR SeqMan Pro™ (DNASTAR. Inc.).
Map construction and comparison
Co-segregating, simplex-inherited CDP markers from the tetraploid female parent (MaR9) were scored as dominant markers (Wu et al. 1992). The marker order was determined by TetraploidMap (Hackett and Luo 2003). The map distance was calculated based on the frequency of the recombination between markers. Publicly available potato and tomato genetic maps from SGN (http://sgn.cornell.edu/cview/map.pl?map_id=9&show_offsets=1&show_ruler=1) and GABI (http://www.gabipd.org/database/) databases, and physical map (Jupe et al. 2012) were included for comparison of marker positions and synteny.
Results
Development of populations lacking R8
Six IPO-C-resistant F1 plants derived from a cross between MaR9 and cv. Concurrent (population 3025) were selected in late blight field trials in 2009. To generate BC1 populations with a reduced R gene content, the selected resistant F1 plants were crossed with susceptible cultivar Katahdin. The BC1 populations were tested in field trials inoculated with IPO-C in the potato-growing seasons of 2010, 2011 and 2013. Four BC1 populations showed a clear segregation into resistant and susceptible groups, while one BC1 population (3247) displayed a continuous distribution of late blight severity (Supplementary Figure 1) and could be roughly grouped into three resistant, 37 susceptible and 10 intermediate-resistant individuals (Table 2). To select BC1 populations without R8 the populations were tested for the presence of a R8 PCR marker (CDP3, which fully co-segregated with resistance; Jo et al. 2011). The progeny of three populations (3247, 3151 and 3154) lacked the CDP3 marker. To rule out that the absence of the CDP3 marker was caused by genetic recombination between R8 and CDP3, the absence of R8 was further confirmed by testing the response of the BC1 plants to AVR8, the cognate effector of R8 (Jo 2013). As expected, none of the BC1 progeny produced a hypersensitive response (HR) upon AVR8 infiltration (Table 2). According to the marker analysis and effector responsiveness, it was concluded that these BC1 populations (3247, 3151 and 3154) did not contain R8 and that the resistance in these plants must be conferred by additional resistance gene(s). Since the segregation of resistance in population 3247 was of quantitative nature, and because the resistance in populations 3151 and 3154 was of qualitative nature, it was postulated that the remaining resistance in MaR9 was constituted by different types of resistances. To avoid suggestions about a potential monogenic nature, we will no longer use the name “R9” but refer to the individual genetic components as R9a, R9b, etc.
Characterisation of the qualitative R9a resistance
Seventeen and ten individuals, respectively, of populations 3151 and 3154 were tested in field trials in 2010 and 2011 and a clear segregation into resistant and susceptible groups was found (Table 2). For mapping the IPO-C resistance a larger population needed to be phenotyped. Since field trials can only be performed in summer, we developed a whole plant late blight assay in a climate cell. The initial small populations that were tested in field trials in 2010 and 2011 were now tested in this climate cell assay and the results fully matched the results of the field assay. In the same climate cell assay 69 additional seedlings of the 3154 population were phenotyped and again a clear segregation into resistant and susceptible groups was observed. In population 3154 resistance segregated in a 1:1 fashion (χ 2 = 0.5, p > 0.05), indicating the expected simplex-based inheritance (Table 3). The causal resistance gene was designated as R9a.
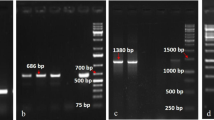
As described, R8 (CDPHero3 marker and AVR8 response) was absent from the 3154 population. Interestingly, when the 184-81 marker that flanked the R8 gene at 1 cM distance was tested in the 3154 population we found that this marker fully co-segregated with the resistance. This suggests that, like R8, R9a locates on chromosome IX. To verify this finding, we set out to develop additional commonly used markers (GP101, S2g3, TG591A, GP41, CT220, T0521, S1d11, S1d5-a, T1065, TG328, TG424, St_At3g23400) from the SGN and GABI databases on the long arm of chromosome IX. A cleaved amplified polymorphism (CAPS) in GP101 was found and located 2.9 cM proximal (two recombinants) relative to R9a in population 3154 (Fig. 1). A polymorphism in SSR marker Stm1021, which is present in RH9 BIN65 of the SH x RH map (Van Os et al. 2006), mapped at 20.3 cM (14 recombinants) proximal to R9a. In this interval of chromosome IX, two R gene clusters (C42 and C43) are known (Jupe et al. 2012). These clusters were targeted for R gene cluster-directed profiling (CDP; Vossen et al. 2013). Using eight Tm-2 2 primers, population 3154 was screened for linked markers. Three markers, CDPTm22 (120 bp), CDPTm26 (375 bp) and CDPTm27 (430 bp) were identified (Fig. 2) that mapped in close proximity to R9a. CDPTm22 marker is at 1.5 cM distance (one recombinant), proximal from R9a and the other two markers fully co-segregated with the resistance in population 3154 (Figs. 1, 2). Using Sw-5-CDP, three linked markers were found; CDPSw58, CDPSw59 and CDPSw510 (Fig. 2). All CDPSw5 markers were located at 5.8 cM (4 recombinants) to the opposite side (distal) of the CDPTm2 markers from the R9a gene (Fig. 1). The CDP markers were excised from the gel and subjected to sequence analysis. The sequence of CDPTm22 was identical to CDPTm22 found in mapping R8 gene (Jo et al. 2011; Genbank accession number JF317285.1). All three CDPTm2 markers identified showed similarity to Tm-2 2 and showed 90–92 % identity with PGSC0003DMG402020585. This is an NB-LRR gene which locates in the Tm-2 2-like cluster C42 (Jupe et al. 2012). CDPSw58, CDPSw59 and CDPSw510 were confirmed to be similar to Sw-5, an S. lycopersicon tospovirus resistance gene (Brommonschenkel and Tanksley 1997). When the CDPSw5 markers from the R9a and R8 maps were compared with the physical map of unique DMGs encoding NB–LRR-type proteins (Jupe et al. 2012), they were found in cluster C43. Marker CDPSw510 in the R9a map and CDPSw54 in R8 map had 70 and 85 % of identity to DMG400016601, respectively. There is a good agreement between the relative positions of the Tm-2 2 and Sw-5 homologous markers identified in the MaR9-derived BC1 population and the R8 map (Jo et al. 2011) and DMG maps (Fig. 1). In conclusion, R9a resides on the telomeric end of the southern arm of chromosome IX and locates in or near a Tm-2 2 cluster.
CDP profiles for ten resistant clones and ten susceptible clones of BC1 population 3154. Pr-resistant parent (MaR9), Ps-susceptible parent (Katahdin), M molecular weight marker. The arrows in blue and in black on the right side indicate linked markers and the bands size of molecular weight marker, respectively (color figure online)
Comparison of R9a map with R8 map and potato DM genome in the distal end of potato chromosome IX. Blue dotted arrows indicate similar or identical markers/sequences in different maps. Genetic distances in centimorgan are indicated by black arrows. The R9a gene maps proximal to the R8 locus (color figure online)
Additional resistance in MaR9, besides R9a, is of quantitative nature
Now that R9a markers were available, it was possible to better characterise potential additional resistance from MaR9. In BC1 population 3153 there was additional resistance besides R8 (Table 2) since there were four plants without R8 that were resistant to IPO-C. Using marker CDPTm27 it was shown that seven plants, including the four R8-free plants from the 3153 populations carried the R9a gene. This result showed that R8 and R9a are in similar chromosomal location, but they are not linked on the same haplotype. Besides, it was concluded that no additional resistance was present in 3153.
In population 3151 (n = 104), the vast majority of the IPO-C-resistant plants carried the 184-81 marker, showing that R9a was the major constituent of resistance in population 3151 (χ 2 = 0.7, p > 0.05). However, six plants were observed which were resistant but had neither R8, nor R9a marker (Tables 2, 3). To further investigate the additional resistance in population 3151, R9a- and R8-free plants from this population were crossed with susceptible cultivar Desiree to produce BC2 population 3253. Two batches of 100 3253 seedlings were inoculated with IPO-C in climate cells. Like in BC1 population 3247 no clear segregation into resistant and susceptible groups was observed (Supplementary Figures 1, 2). It was concluded that the additional resistance in BC1 population 3151 was of quantitative nature. As described above, in population 3247 a continuous distribution of late blight severity was observed. Using marker analysis, we indeed found that besides R8, R9a was absent from this population (Table 2). Quantitative resistance may have been caused by residual effects of defeated R genes, like the R10 gene that was introduced through the susceptible parent Concurrent. In our assays we do not expect that R10 contributes to resistance since MaR10 plants are fully susceptible to IPO-C (data not shown).
Based on the currently tested populations we can conclude that the additional resistance in MaR9 is constituted of a qualitative resistance gene R9a and quantitative resistance that remain to be characterized.
Discussion
Characterization of the MaR9 resistance
In this study, we genetically characterized the additional resistance to P. infestans isolate IPO-C from the late blight differential plant MaR9. Both qualitative and quantitative resistances were encountered. The qualitatively inherited resistance gene R9a resides on the distal end of chromosome IX like R8. Both R8 and R9 genes have previously been suggested to locate on chromosome XI as allelic variants of R3 (Huang et al. 2005) because of the presence of the R3a haplotype in the R8 and R9 plants. In retrospect, the presence of the haplotype was associated with the presence of the R3a gene itself that was a contaminant of the MaR8 and MaR9 differentials (Kim et al. 2012) rather than with the location of the R8 and R9a genes in the R3 haplotype. We employed an R gene “de-stacking” (making offspring plants containing different R gene combinations) approach using marker analysis and effector response to eliminate the effects of other known R genes for revealing the remaining resistance in MaR9. The resulting BC1 populations were analysed using R gene cluster-directed profiling (CDP) strategies for mapping R9a. It was essential to make a clear discrimination between R9a and R8 because MaR9 contained R8 as well as R9a and because isolate IPO-C was avirulent on both R8- and R9a-containing plants. Analysis of the absence of R8 in mapping populations using the R8 marker alone was not sufficient since R9a was in a similar chromosomal position as R8. Another molecular tool, in planta AVR8 expression, was used to further confirm the absence of R8. It was confirmed that populations lacking the R8 marker indeed were not responsive to AVR8, so it could be concluded that the R8 gene could not be responsible for the resistance to IPO-C in the selected populations. By investigating BC1 and BC2 populations, it could also be revealed that quantitative resistance contributes to the resistance to IPO-C in MaR9. Regarding the quantitative resistance, there would be two possibilities: (1) QRL as a new genetic entities and/or (2) residual effects of resistance (Stewart et al. 2003; Rauscher et al. 2010) from different combinations of the defeated R genes from MaR9, as well as R10 from cv. Concurrent when progeny plants were infected with a compatible isolate IPO-C. Further analysis regarding insights into the genetic nature and stability of the quantitative resistance in MaR9 through isolates and environments remains to be conducted.
The long arm of chromosome IX is a hot spot for resistance
The long arm of chromosomes IX is a hot spot for resistance in Solanaceous genomes (Sliwka et al. 2006; Pel et al. 2009). The R gene clusters on the long arm of chromosome IX contain the late blight resistance genes Rpi-moc1 of S. mochiquense (Smilde et al. 2005) Rpi-phu1 which is identical to Rpi-vnt1, most likely from S. phureja (Sliwka et al. 2006), Ph-3, a major QRL for late blight resistance from S. pimpenelifolium (Zhang et al. 2014) the R8 late blight resistance gene from S. demissum (Jo et al. 2011), Rpi-dlc1, from S. dulcamara, a Solanum species native to Europe (Golas et al. 2010), and Rpi-edn2 from S.x edinense, a natural pentaploid hybrid between S. demissum and the South American cultivated potato S. tuberosum spp. Andigena (Verzaux 2010), mapped in a similar region as R9a. Also resistances to other diseases like Sw-5, a tomato gene for tospoviruses resistance (Brommonschenkel and Tanksley 1997), Nx encoding hypersensitive resistance to Potato virus X of S. phureja (Tommiska et al. 1998), Gpa6, a QRL for resistance to Globodera pallida in potato (Rouppe van der Voort et al. 2000), the gene Gm conferring resistance to Potato virus M (Marczewski et al. 2006), the Ry chc and Ny-1 potato virus Y resistance genes (Sato et al. 2006; Szajko et al. 2008), as well as a QRL for resistance to Erwinia carotovora ssp. atroseptica (Zimnoch-Guzowska et al. 2000) have been identified in similar genomic regions. The long arm of chromosome IX features two large heterogeneous clusters (Jupe et al. 2012). Cluster 42 harbours eight TIR–NB–LRR genes that are separated by eight paralogs of Tm-2 2, whereas the distal cluster C43 contains 15 paralogs of the Tospovirus resistance gene Sw-5 (Jupe et al. 2012). When compared to tomato, these clusters seem to have drastically expanded in tomato as both the Tm-2 2 and Sw-5 cluster in the Heinz tomato genome count only four members (Andolfo et al. 2014). Alternatively, tomato clusters may have been subject to shrinking as the S. pimpenellifolium genome only counts two Tm-2 2 paralogs (Zhang et al. 2014). Therefore, R genes on the long arm of chromosomes IX could be members of either the Tm-2 2 cluster or the Sw-5 cluster so that R gene cloning by paralog mining (Vossen et al. 2014) or other homology-based approaches (Jupe et al. 2012 would be encouraged, although it will require sophisticated bioinformatics for efficient assembly of paralogous sequences.
Implications for late blight resistance breeding
The utility of R9a in combination with the R8 gene would be of particular interest in the context of late blight resistance breeding of potato. The co-introgression of multiple R gene(s) is essential to provide durability to new potato varieties. Another candidate for co-introgression is R2 that is known to be defeated but remains providing quantitative resistance in many potato-growing regions (Pilet et al. 2005; Wang et al. 2012; Li 2012), could most likely contribute to durability. The combination of R2, R8 and R9a was correlated with durable resistance observed in MaR9 (Kim et al. 2012; Jo 2013). However, R9 QRL were most likely unevenly distributed over MaR9 BC1 offspring in these studies. The presence of the R gene combinations in dissimilar genetic backgrounds may have blurred this on-site durability monitoring approach. The study of R genes and R gene combinations in isogenic genetic backgrounds is essential for future durability studies of R gene combinations (Zhu et al. 2014).
Two approaches can be used to produce potato clones with late blight R gene combinations; marker-assisted selection or GM breeding strategies (Zhu et al. 2013). Marker-assisted selection allows transfer of multiple R genes into potato using traditional breeding methods, an alternative to deploying the R gene through genetic transformation which has been a non-stop controversial issue. The fact that R9a and R8 genes reside in the adjacent Tm-2 2- and/or Sw-5 clusters which are physically separated by a maximum distance of 1120 kb (Fig. 2), and that they are naturally located on different chromatids (this study), provide a potential challenge to recombining them into coupling phase, as demonstrated by Robbins et al. (2010). This would result in a chromosome fragment containing R8 and R9a. As a consequence, possible genes with a negative effect (linkage drag) located between R8 and R9a will not be lost by backcrossing. Cisgenic breeding based on the introduction of cloned natural R genes from crossable species using A. tumefaciens-mediated marker-free transformation, would not encounter the problem of linkage drag that could not be removed (Jo et al. 2014).
Author contribution statement
JKR performed the molecular experiments and manuscript writing. RV and EJ were involved in manuscript writing and revisions. JV performed phenotyping experiments and manuscript writing.
References
Andolfo G, Jupe F, Witek Etherington GJ, Ercolano MR, Jones JD (2014) Defining the full tomato NB-LRR resistance gene repertoire using genomic and cDNA RenSeq. BMC Plant Biol 14:120
Ballvora A, Ercolano MR, Weiss J, Meksem K, Bormann CA, Oberhagemann P, Salamini F, Gebhardt C (2002) The R1 gene for potato resistance to late blight (Phytophthora infestans) belongs to the leucine zipper/NBS/LRR class of plant resistant genes. Plant J 30:361–371
Bisognin DA, Douches DS, Jastrzebski K, Kirk WW (2002) Half-sib progeny evaluation and selection of potatoes resistant to the US8 genotype of Phytophthora infestans from crosses between resistant and susceptible parents. Euphytica 125:129–138
Black W (1951) Inheritance of resistance to blight (Phytophthora infestans) in potatoes: inter-relationships of genes and strains. Proc R Soc Edinburgh 64:312–352
Black W, Mastenbroek C, Mills WR, Peterson LC (1953) A proposal for an international nomenclature of races of Phytophthora infestans and of genes controlling immunity in Solanum demissum derivatives. Euphytica 2:173–178
Bos JIB, Kanneganti TD, Young C, Cakir C, Huitema E, Win J, Armstrong MR, Birch RJ, Kamoun S (2006) The C-terminal half of Phytophthora infestans RXLR effector AVR3a is sufficient to trigger R3a-mediated hypersensitivity and suppress INF1-induced cell death in Nicotiana benthamiana. Plant J 48:165–176
Bradshaw JE, Bryan GJ, Lees AK, McLean K, Solomon-Blackburn RM (2006) Mapping the R10 and R11 genes for resistance to late blight (Phytophthora infestans) present in the potato (Solanum tuberosum) R gene differentials of Black. Theor Appl Genet 112:744–751
Brommonschenkel SH, Tanksley SD (1997) Map-based cloning of the tomato genomic region that spans the Sw-5 tospovirus resistance gene in tomato. Mol Gen Genet 256:121–126
Collins A, Milbourne D, Ramsay L, Meyer R, Chatot-Balandras C, Oberhagemann P, De Jong W, Gebhardt C, Bonnel E, Waugh R (1999) QTL for field resistance to late blight in potato are strongly correlated with maturity and vigour. Mol Breeding 5:387–398
Colon LT, Jansen RC, Budding DJ (1995) Partial resistance to late blight (Phytophthora infestans) in hybrid progenies of four South American Solanum species crossed with diploid S. tuberosum. Theor Appl Genet 90:691–698
Deahl KL, Inglis DA, DeMuth SP (1993) Testing for resistance to metalaxyl in Phytophthora infestans isolates from Northwestern Washington. Am Potato J 70:779–795
de Vetten NCMH, Verzaux EC, Vossen JH, Rietman H, Vleeshouwers VGAA, Jacobsen E, Visser RGF (2011) Cloning and exploitation of a functional R-gene from Solanum x edinense. Patent application: published by: IPO: PCT/NL2011/050386
Dianese EC, Fonseca MEN, Goldbach R, Kormelink R, Inoue-Nagata AK, Resende RO, Boiteux LS (2010) Development of a locus specific, co-dominant SCAR marker for assisted-selection of the Sw-5 (Tospovirus resistance) gene cluster in a wide range of tomato accessions. Mol Breed 25:133–142
El-Kharbotly A, Palomino-Sánchez C, Salamini F, Jacobsen E, Gebhardt C (1996) R6 and R7 alleles of potato conferring race specific resistance to Phytophthora infestans (Mont.) de Bary identified genetic loci clustering with the R3 locus on chromosome 9. Theor Appl Genet 92:880–884
Foster SJ, Park TH, Pel MA, Brigneti G, Sliwka J, Jagger L, Van der Vossen EAG, Jones JDG (2009) Rpi-vnt1.1, a Tm-2 2 homolog from Solanum venturii confers resistance to potato late blight. Mol Plant Microbe Interact 22:589–600
Fry W (2008) Phytophthora infestans: the plant (and R-gene) destroyer. Mol Plant Pathol 9:385–402
Fry WE, Goodwin SB (1997) Resurgence of the Irish potato famine fungus. Bioscience 47:363–367
Fry WE, Goodwin SB, Dyer AT, Matuszak JM, Drenth A, Tooley PW, Sujkowski LS, Koh YJ, Cohen BA, Spielman LJ, Deahl KL, Inglis DA, Sandlan KP (1993) Historical and recent migrations of Phytophthora infestans: chronology, pathways, and implications. Plant Dis 77:653–661
Fulton T, Chunwongse J, Tanksley S (1995) Microprep protocol for extraction of DNA from tomato and other herbaceous plants. Plant Mol Biol Rep 13:207–209
Gebhardt C, Valkonen JP (2001) Organization of genes controlling disease resistance in the potato genome. Annu Rev Phytopathol 39:79–102
Golas TM, Sikkema A, Gros J, Feron RMC, van den Berg RG, van der Weerden GM, Mariani C, Allefs JJHM (2010) Identification of a resistance gene Rpi-dlc1 to hytophthora infestans in European accessions of Solanum dulcamara. Theor Appl Genet 120:797–808
Goodwin SB, Cohen BA, Fry WE (1994) Panglobal distribution of a single clonal lineage of the Irish potato famine fungus. Proc Natl Acad Sci USA 91:11591–11595
Grünwald NJ, Flier WG, Sturbaum AK, Garay-Serrano E, Van den Bosh TBM, Smart CD, Matuszak JM, Lozoya-Saldaňa H, Turkensteen LJ, Fry WE (2001) Population structure of Phytophthora infestans in the Toluca valley region of central Mexico. Phytopathology 91:882–890
Hackett CA, Luo ZW (2003) TetraploidMap: construction of a linkage map in autotetraploid species. J Hered 94:358–359
Haynes KG, Christ BJ, Weingartner DP, Douches DS, Thill CA, Secor G, Fry WE, Lambert DH (2002) Foliar resistance to late blight in potato clones evaluated in national trials in 1997. Am J Potato Res 79:451–457
Huang S (2005) Discovery and characterization of the major late blight resistance complex in potato. PhD thesis. Wageningen University, p 135
Huang S, Van der Vossen EAG, Huang H, Vleeshouwers VGAA, Zhang N, Borm TJA, van Eck HJ, Baker B, Jacobsen E, Visser RGF (2005) Comparative genomics enabled the isolation of the R3a late blight resistance gene in potato. Plant J 42:251–261
Jo KR (2013) Unveiling and deploying durability of late blight resistance in potato from natural stacking to cisgenic stacking. PhD Thesis, Wageningen University, Wageningen, The Netherlands, p 168
Jo KR, Arens M, Kim TY, Jongsma MA, Visser RGF, Jacobsen E, Vossen JH (2011) Mapping of the S. demissum late blight resistance gene R8 to a new locus on chromosome IX. Theor Appl Genet 123:1331–1340
Jo KR, Kim CJ, Kim SJ, Kim TY, Bergervoet M, Jongsma MA, Visser RGF, Jacobsen E, Vossen JH (2014) Development of late blight resistant potatoes by cisgene stacking. BMC Biotechnol 14:50
Jones JDG, Foster S, Chu Z, Park T, van der Vossen EAG, Pel M, Visser RGF (2009) Late blight resistance genes and methods. Patent application: WO2009013468
Jupe F, Pritchard L, Etherington GJ, MacKenzie K, Cock PJA, Wright F, Sharma SK, Bolser D, Bryan GJ, Jones JDG, Hein I (2012) Identification and localisation of the NB-LRR gene family within the potato genome. BMC Genom 13:75
Karimi M, Inze D, Depicker A (2002) GATEWAY(TM) vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7:193–195
Kim HJ, Lee HR, Jo KR, Mortazavian SMM, Huigen DJ, Evenhuis B, Kessel G, Visser RGF, Jacobsen E, Vossen JH (2012) Broad spectrum late blight resistance in potato differential set plants MaR8 and MaR9 is conferred by multiple stacked R genes. Theor Appl Genet 124:923–935
Kover PX, Caicedo AL (2001) The genetic architecture of disease resistance in plants and the maintenance of recombination by parasites. Mol Ecol 10:1–16
Leonards-Schippers C, Gieffers W, Salamini F, Gebhardt C (1992) The R1 gene conferring race specific resistance to Phytophthora infestans in potato is located on potato chromosome 5. Mol Gen Genet 233:278–283
Li Y (2012) Multiplex SSR analysis of Phytophthora infestans in different countries and the importance for potato breeding. PhD thesis. Wageningen University, p 206
Li X, Van Eck HJ, Rouppe van der Voort JNAM, Huigen DJ, Stam P, Jacobsen E (1998) Autotetraploids and genetic mapping using common AFLP markers: the R2 allele conferring resistance to Phytophthora infestans mapped on potato chromosome 4. Theor Appl Genet 96:1121–1128
Li G, Huang S, Guo X, Li Y, Yang Y, Guo Z, Kuang H, Rietman H, Bergervoet M, Vleeshouwers VGAA, van der Vossen E, Qu D, Visser RGF, Jacobsen E, Vossen JH (2011) Cloning and characterization of R3b; members of the R3 superfamily of late blight resistance genes show sequence and functional divergence. Mol Plant Microbe Interact 24(10):1132–1142
Lokossou AA, Park TH, van Arkel G, Arens GM, Ruyter-Spira C, Morales J, Whisson SC, Birch PRJ, Visser RGF, Jacobsen E, van der Vossen EAG (2009) Exploiting knowledge of R/Avr genes to rapidly clone a new LZ-NBS-LRR family of late blight resistance genes from potato linkage group IV. Mol Plant Microbe Interact 22:630–641
Malcomson JF, Black W (1966) New R genes in Solanum demissum Lindl. and their complementary races of Phytophthora infestans (Mont.) de Bary. Euphytica 15:187–198
Marczewski W, Strzelczyk-ôyta D, Hennig J, Witek K, Gebhardt C (2006) Potato chromosomes IX and XI carry genes for resistance to potato virus M. Theor Appl Genet 112:1232–1238
Mastenbroek C (1952) Investigations into the inheritance of the immunity from Phytophthora infestans de B. of Solanum demissum Lindl. Euphytica 1:187–198
Niederhauser JS, Mills MR (1953) Resistance of Solanum species to Phytophthora infestans in Mexico. Phytopathol 43:303–313
Pel MA (2010) Mapping, isolation and characterization of genes responsible for late blight resistance in potato. PhD thesis. Wageningen University, p 210
Pel MA, Foster SJ, Park TH, Rietman H, Van Arkel G, Jones JDG, Van Eck HJ, Jacobsen E, Visser RGF, Van der Vossen EAG (2009) Mapping and cloning of late blight resistance genes from Solanum venturii using an interspecific candidate gene approach. Mol Plant Microbe Interact 22:601–615
Pilet F, Pelle R, Ellisseche D, Andrivon D (2005) Efficacy of the R2 resistance gene as a component for the durable management of potato late blight in France. Plant Pathol 54:723–732
Rauscher G, Simko I, Mayton H, Bonierbale M, Smart CD, Grünwald NJ, Greenland A, Fry WE (2010) Quantitative resistance to late blight from Solanum berthaultii cosegregates with RPi-ber: insights in stability through isolates and environment. Theor Appl Genet 121:1553–1567
Rietman H, Bijsterbosch G, Cano LM, Lee HR, Vossen JH, Jacobsen E, Visser RGF, Kamoun S, Vleeshouwers VGAA (2012) Qualitative and quantitative late blight resistance in the potato cultivar Sarpo Mira is determined by the perception of five distinct RXLR effectors. Mol Plant Microbe Interact 25:910–919
Robbins MD, Masud MAT, Panthee DR, Gardner RG, Francis DM, Stevens MR (2010) Marker-assisted selection for coupling phase resistance to tomato spotted wilt virus and Phytophthora infestans (Late Blight) in tomato. Hort Science 45:1424–1428
Rouppe van der Voort J, van der Vossen EAG, Bakker E, Overmars H, van Zandvoort P, Hutten RCB, Klein Lankhorst R, Bakker J (2000) Two additive QTLs conferring broad-spectrum resistance in potato to Globodera pallida are localized on resistance gene clusters. Theor Appl Genet 101:1122–1130
Sato M, Nishikawa K, Komura K, Kosaka K (2006) Potato virus Y resistance gene, Ry chc, mapped to the distal end of potato chromosome 9. Euphytica 149:367–372
Sliwka J, Jakuczun H, Lebecka R, Marczewski W, Gebhardt C, Zimnoch-Guzowska E (2006) The novel, major locus Rpi-phu1 for late blight resistance maps to potato chromosome 9 and is not correlated with long vegetation period. Theor Appl Genet 113:685–695
Smilde WD, Brigneti G, Jagger L, Perkins S, Jones JDG (2005) Solanum mochiquense chromosome IX carries a novel late blight resistance gene Rpi-moc1. Theor Appl Genet 110:252–258
Song J, Bradeen JM, Naess SK, Raasch JA, Wielgus SW, Haberlach GT, Liu J, Kuang H, Austin-Phillips S, Buell CR, Helgeson JP, Jiang J (2003) Gene RB cloned from Solanum bulbocastanum confers broad spectrum resistance to potato late blight. Proc Natl Acad Sci USA 100:9128–9133
Stewart HE, Bradshaw JE, Pande B (2003) The effect of the presence of R-genes for resistance to late blight (Phytophthora infestans) of potato (Solanum tuberosum) on the underlying level of field resistance. Plant Pathol 52:193–198
Šwiežynski KM, Haynes KG, Hutten RCB, Sieczka MT, Watts P, Zimnoch-Guzowska E (1997) Pedigree of European and North-American potato varieties. Plant Breed Seed Sci 41:3–149
Šwiežynski KM, Domanski L, Zarzycka H, Zimnoch-Guzowska E (2000) The reaction of potato differentials to Phytophthora infestans isolates collected in nature. Plant Breed 119:119–126
Szajko K, Chrzanowska M, Witek K, Strzelczyk-Żyta D, Zagórska H, Gebhardt C, Hennig J, Marczewski W (2008) The novel gene Ny-1 on potato chromosome IX confers hypersensitive resistance to Potato virus Y and is an alternative to Ry genes in potato breeding for PVY resistance. Theor Appl Genet 116:297–303
Tan MYA, Hutten RCB, Celis C, Park TH, Niks RE, Visser RGF, van Eck HJ (2008) The RPi-mcd1 locus from Solanum microdontum involved in resistance to Phytophthora infestans, causing a delay in infection, maps on potato chromosome 4 in a cluster of NBS–LRR genes. Mol Plant Microbe Interact 21:909–918
Tommiska TJ, Hämäläinen JH, Watanabe KN, Valkonen JPT (1998) Mapping of the gene Nxphu that controls hypersensitive resistance to potato virus X in Solanum phureja IvP35. Theor Appl Genet 96:840–843
Trognitz BR, Trognitz FC (2007) Occurrence of the R1 allele conferring resistance to late blight in potato R-gene differentials and commercial cultivars. Plant Pathol 56:150–155
Trognitz F, Manosalva P, Gysin R, Nino-Liu D, Simon R, Herrera MD, Trognitz B, Ghislain M, Nelson R (2002) Plant defense genes associated with quantitative resistance to potato late blight in Solanum phureja X dihaploid S. tuberosum hybrids. Mol Plant Microbe Interact 15:587–597
Umaerus V, Umaerus M, Erjefalt L, Nilsson BA (1983) Control of Phytophthora by host resistance: problems and progress. In: Erwin CC, Bartniki-Garcia S, Tsao PH (eds) Phytophthora its biology, taxonomy, ecology and pathology. APS Press, St Paul, pp 315–326
van der Linden CG, Wouters D, Mihalka V, Kochieva EZ, Smulders MJM, Vosman B (2004) Efficient targeting of plant disease resistance loci using NBS profiling. Theor Appl Genet 109:384–393
van der Vossen EAG, Sikkema A, te Lintel Hekkert B, Gros J, Stevens P, Muskens M, Wouters D, Pereira A, Stiekema WJ, Allefs S (2003) An ancient R gene from the wild potato species Solanum bulbocastanum confers broad-spectrum resistance to Phytophthora infestans in cultivated potato and tomato. Plant J 36:867–882
van Os H, Andrzejewski S, Bakker E, Barrena I, Bryan GJ, Caromel B, Ghareeb B, Isidore E, De Jong W, Van Koert P, Lefebvre V, Milbourne D, Ritter E, Rouppe Van Der Voort JNAM, Rousselle-Bourgeois F, Van Vliet J, Waugh R, Visser RGF, Bakker J, van Eck HJ (2006) Construction of a 10,000-marker ultradense genetic recombination map of potato: providing a framework for accelerated gene isolation and a genomewide physical map. Genetics 173:1075–1087
Verzaux E (2010) Resistance and susceptibility to late blight in Solanum: gene mapping cloning and stacking. PhD Thesis, Wageningen University, The Netherlands
Vleeshouwers VGAA, van Dooijweert W, Paul Keizer LC, Sijpkes L, Govers F, Colon LT (1999) A laboratory assay for Phytophthora infestans resistance in various Solanum species reflects the field situation. Eur J Plant Pathol 105:241–250
Vleeshouwers VGAA, Rietman H, Krenek P, Champouret N, Young C, Oh SK, Wang M, Bouwmeester K, Vosman B, Visser RGF, Jacobsen E, Govers F, Kamoun S, van der Vossen EAG (2008) Effector genomics accelerates discovery and functional profiling of potato disease resistance and Phytophthora infestans avirulence genes. PLoS One 3:e2875
Vossen JH, Nijenhuis M, Arens MJB, van der Vossen EAG, Jacobsen E, Visser RGF (2010) Cloning and exploitation of a functional R-gene from Solanum chacoense. Patent application: published by: IPO: PCT/NL2010/050612
Vossen JH, Dezhsetan S, Esselink D, Arens M, Sanz MJ, Verweij W, Verzaux E, van der Linden CG (2013) Novel applications of motif-directed profiling to identify disease resistance genes in plants. Plant Methods 9:37
Vossen JH, Jo KR, Vosman B (2014) Mining the genus Solanum for increasing disease resistance. In: Genomics of Plant Genetic Resources. Springer, Netherlands, pp 27–46
Wang X, Guo M, Min F, Gao Y, Xu F, Yang S, Lu D (2012) Virulence complexity and high levels of fungicide resistance suggest population change of Phytophthora infestans in the Heilongjiang Province of China. Potato Research (Published Online)
Wastie RL (1991) Breeding for resistance. Adv Plant Pathol 7:193–224
Wu KK, Burnquist W, Sorrells ME, Tew TL, Moore PH, Tanksley SD (1992) The detection and estimation of linkage in polyploids using single-dose restriction fragments. Theor Appl Genet 83:294–300
Zhang XZ, Kim B (2007) Physiological races of Phytophthora infestans in Korea. Plant Pathol J 23:219–222
Zhang C, Liu L, Wang X, Vossen JH, Li G, Li T, Zheng Z, Gao J, Guo Y, Visser RGF, Li J, Bai Y, Du Y (2014) The Ph-3 gene from Solanum pimpinellifolium encodes CC-NBS-LRR protein conferring resistance to Phytophthora infestans. Theor Appl Genet 127:1353–1364
Zhu SX, Duwal A, Su Q, Vossen JH, Visser RGF, Jacobsen E (2013) Vector integration in triple R gene transformants and the clustered inheritance of resistance against potato late blight. Transgenic Res 22:315–325
Zhu SX, Vossen JH, Bergervoet M, Nijenhuis M, Kodde L, Kessel GJT, Vleeshouwers VGAA, Visser RGF, Jacobsen E (2014) An updated conventional- and a novel GM potato late blight R gene differential set for virulence monitoring of Phytophthora infestans. Euphytica (in press). doi:10.1007/s10681-014-1276-0
Zimnoch-Guzowska E, Marczewski W, Lebecka R, Flis B, Schäfer-Pregl R, Salamini F, Gebhardt C (2000) QTL analysis of new sources of resistance to Erwinia carotovora ssp atroseptica in potato done by AFLP, RFLP, and resistance-gene-like markers. Crop Sci 40:1156–1167
Acknowledgments
JKR was financially supported by the European Commission (EuropeAid project DCI-FOOD/2009/218-671), and the Dutch Ministry of Agriculture, Nature and Fisheries (International Cooperation project BO-10-010-112 and BO-10- 001-200). Maarten A. Jongsma is acknowledged as a project leader of those programs. JV was financially supported by the DuRPh program, granted by the Ministry of Economic Affairs, formerly the Ministry of Agriculture and Fisheries, in the Netherlands. Dr. Anton Haverkort is acknowledged as the project leader of the DuRPh program. MSc Glenn M.G. Theunissen is acknowledged for his help in the phenotypic characterisation of the BC1 and BC2 populations. We thank Dirk-Jan Huigen for making F1, BC1 and BC2 crosses and for excellent plant space management.
Conflict of interest
The authors declare to have no conflicting interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. J. Bryan.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2015_2480_MOESM1_ESM.pptx
Supplementary material 1 (PPTX 49 kb) Supplementary Fig. 1. Blight scores (mean of four plants) for 51 progeny in BC1 population in field trials. BC1 population 3247 was generated by crossing resistant F1 progeny (3025-2, originating from F1 of MaR9 x Concurrent) as female parents with susceptible cultivar Katahdin as a male parent. Blight scoring was performed by estimating the percentage of blight-affected leaf area in the field
122_2015_2480_MOESM2_ESM.pptx
Supplementary material 2 (PPTX 3383 kb) Supplementary Fig. 2. Late blight symptoms on population 3253 seedlings in climate cell assays. A continuous distribution in resistance/susceptibility levels to IPO-C was noticed
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Jo, KR., Visser, R.G.F., Jacobsen, E. et al. Characterisation of the late blight resistance in potato differential MaR9 reveals a qualitative resistance gene, R9a, residing in a cluster of Tm-2 2 homologs on chromosome IX. Theor Appl Genet 128, 931–941 (2015). https://doi.org/10.1007/s00122-015-2480-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-015-2480-6