Abstract
Oncobiosis has emerged as a key contributor to the development, and modulator of the treatment efficacy of cancer. Hereby, we review the modalities through which the oncobiome can support the progression of tumors, and the emerging therapeutic opportunities they present. The review highlights the inherent challenges and limitations faced in sampling and accurately characterizing oncobiome. Additionally, the review underscores the critical need for the standardization of microbial analysis techniques and the consistent reporting of microbiome data. We provide a suggested metadata set that should accompany microbiome datasets from oncological settings so that studies remain comparable and decipherable.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer remains one of the leading causes of morbidity and mortality worldwide, with breast cancer accounting for a significant proportion of these cases [1, 2]. Amidst the myriad of factors contributing to cancer progression, the role of microorganisms, particularly bacteria, has emerged as a critical area of research [3,4,5,6,7,8,9,10]. Oncobiome refers to a microbiome with altered composition in cancer patients, which is implicated in supporting progression, and metastasis of various cancers, including breast cancer [11,12,13]. The oncobiome can influence cancer through direct interactions with cancer cells, modulation of the immune system, or alterations in the local tumor microenvironment [13]. The oncobiome is thought to interact with anticancer therapy as well, influencing efficacy and outcome [14]. This review aims to dissect the current understanding oncobiosis, highlighting key findings, challenges, and future directions in this evolving field and to provide a comprehensive reporting and sampling guideline for oncobiome studies.
ncobiome: an overview
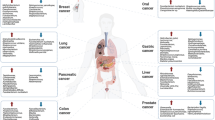
The intricate relationship between the human microbiome and cancer has garnered significant attention, leading to the identification of a phenomenon known as oncobiosis [15]. Oncobiosis refers to the dysbiosis of microbiome compartments in the presence of neoplasia, which is often associated with cancer progression, representing a shift away from the normal microbial balance [15]. This alteration in the microbial ecosystem is not restricted to a single niche but affects multiple microbiome compartments that for example in breast cancer includes, but is likely not limited to, the fecal/gut, breast tissue, milk ducts, tumor sites, oral cavity, pharyngeal region, urinary tract, and even the blood (Table 1, [3]). It is important to note that the specific pattern and impact of oncobiotic changes vary across different types of cancer, underscoring the complex and distinct nature of microbial associations with neoplastic diseases [102, 103].
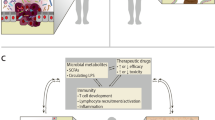
While the list of bacteria directly implicated in oncogenesis remains relatively concise, the broader concept of oncobiosis encompasses a range of microbial interactions that support cancer progression and influence disease outcomes [104]. These interactions manifest through three, somewhat overlapping primary modalities:
-
1)
Colonization of tumor tissue: Certain bacteria have been found to colonize tumor tissues, triggering chronic inflammation and/or secreting toxins that can promote tumor growth and progression or induce genomic instability [3, 13, 46]. Bacteria in the oncobiome can suppress the activity of immune cells capable of targeting cancer cells, such as natural killer (NK) cells and T-cells. The specific mechanisms of immune modulation can include the alteration of cytokine and chemokine profiles, modification of antigen presentation, and the induction of immunosuppressive cells within the tumor microenvironment [3, 13, 46].
-
2)
Production of metabolites and toxins with paracrine or hormone-like properties: Microbial metabolites and toxins can act in a paracrine fashion or mimic hormone action, influencing cancer cell behavior and tumor microenvironment dynamics [3, 105, 106].
-
3)
Immune modulation/immune evasion: The microbiome plays a crucial role in modulating the host immune response, which can influence the efficacy of immune surveillance against tumors and impact cancer progression [3, 64, 107, 108].
Significantly, the microbiome has been identified as a pivotal player in controlling metastasis formation, thereby determining the overall outcome of the disease [31, 40, 108, 109]. This suggests that interventions targeting oncobiotic changes could potentially offer novel therapeutic avenues for cancer treatment and metastasis prevention.
A handful of bacteria have been firmly established in the literature as being linked to cancer as tumor inducers. Helicobacter pylori is well-known for its association with gastric cancer [110,111,112] and mucosa-associated lymphoid tissue lymphoma [113,114,115,116]. Several bacterial species play significant roles in the initiation and progression of colorectal cancer through the induction of chronic inflammation, production of genotoxic substances, and modulation of the immune response [117,118,119]. Fusobacterium nucleatum can adhere to and invade colorectal epithelial cells using its FadA adhesin to bind to E-cadherin declutching β-catenin signaling [120,121,122], promotes a pro-inflammatory environment [123,124,125,126] and interacts with the host’s immune system to shield tumor cells from immune surveillance [127,128,129,130,131,132] conducive to cancer progression. Enterotoxigenic Bacteroides fragilis (ETBF) secretes a toxin known as Bacteroides fragilis toxin (BFT) that can disrupt the mucosal barrier and can induce oncogenic signaling pathways as STAT3 and NF-κB, and contribute to the formation of a tumorigenic environment in the colon [131, 133,134,135,136]. Strains of Escherichia coli (strains harboring the polyketide synthase gene, pks +) produce colibactin, a genotoxin that causes DNA damage and mutations, thereby contributing to colorectal carcinogenesis [137,138,139]. Streptococcus gallolyticus (formerly known as Streptococcus bovis biotype I) has been associated with an increased risk of colorectal cancer [140,141,142,143,144,145] through inducing chronic inflammation and potentially modulating the immune response within the gastrointestinal tract.
The number of directly oncogenic bacteria, as mentioned, is only a handful, and these species are not necessarily overrepresented in neoplasia patients. Oncobiosis in these cases yields a maladaptive microbiome with disproportionate, abnormal taxonomical composition coinciding with pathological functional adaptation with large interpersonal variability.
Detection and analysis of the oncobiome
The detection and analysis of oncobiome are crucial for understanding the microbiome’s role in cancer development and for exploring new diagnostic and therapeutic strategies. This section covers the contemporary techniques and methodologies employed in assessing the oncobiome, highlighting the challenges and limitations of these methods.
16S rRNA gene sequencing
16S rRNA gene sequencing targets the hypervariable regions of the 16S ribosomal RNA gene that can distinguish between different taxa. The advantage of the 16S hypervariable region sequencing is that it can provide taxonomical information from samples with low bacteria-to-host nucleic acid ratio and its relative cheapness compared to whole genome sequencing or to shotgun sequencing (see below). However, despite its widespread use and valuable insights, 16S rRNA gene sequencing comes with important limitations, particularly concerning resolution at the genus and species level. The method relies on the analysis of specific hypervariable regions within the 16S rRNA gene to distinguish between bacterial taxa. However, the degree of variability in these regions can differ among taxa; therefore, the resolution may not be sufficient to distinguish closely related genera in some cases [146]. The challenge becomes even more pronounced at the species level, where the variability in the 16S rRNA gene may not be sufficient to distinguish between closely related species. This is particularly problematic in the context of oncobiome, where different species within the same genus may have vastly different (or even opposite) effects on cancer development. Another technical issue is that 16S rRNA sequencing may skew the representation of low-abundance taxons as compared to whole genome sequencing [147]. It is possible to predict functional changes of microbial communities using algorithms as PICRUSt [148]. It is also important to note that samples with low abundance of bacterial DNA are prone for pollution from the environment (e.g., the skin in samples from surgical interventions, pollution of reagents, or pollution of tap water from formalin-fixes, paraffin-embedded blocks, or even from plasticware and kits [149]) that needs to be meticulously controlled.
Whole genome sequencing (WGS)
Shotgun sequencing is a powerful tool that revolutionized our understanding of the microbiome’s complexity and its impact on health and disease. By sequencing all the DNA (metagenomics) or RNA (metatranscriptomics) in a sample, these methods allow for the identification of not only bacterial communities, but also viruses, fungi, and other microorganisms, offering a comprehensive view of the microbial ecosystem. In addition, metagenomics and metatranscriptomics provide insights into the genetic and functional potential of these microbial communities.
Despite their considerable advantages, metagenomics and metatranscriptomics come with limitations. One such limitation is cost. Further, the large volume of data generated requires substantial computational resources for storage, processing, and analysis. Moreover, data analysis is complex and challenging; it requires sophisticated bioinformatic tools and expertise to assemble, annotate, and interpret the taxonomical composition and biochemical functions of a bacterial community. Another limitation is the signal-to-noise ratio in samples with low microbial biomass, such as human tissue. The DNA from the host or other sources (e.g., environmental contaminants) can overwhelm the microbial DNA, making it challenging to accurately profile the microbiome. Tackling this requires careful sample handling, processing, and data analysis techniques to minimize contamination and to ensure that the microbial signal can be accurately detected and analyzed as well as the use of environmental controls.
Metagenomics and metatranscriptomics of human-associated samples raise ethical and privacy concerns, particularly when human DNA is sequenced alongside microbial DNA. The potential for identifying genetic information about the host requires careful consideration of consent, data storage, and data sharing practices to protect individuals’ privacy.
Hybridization-based techniques
The PathoChip or the GeoChip technologies are based on the hybridization of nucleic acid samples from biomaterials to probes attached to the chip [150, 151]. The probes on the GeoChip surface recognize bacteria, archaea, fungi, protists, and viruses [150]. In addition, the PathoChip contains probes for virulence factors, toxin, and siderophore genes; hence, the system provides besides taxonomical, microbial function information [151]. This methodology is an alternative for low bacterial-to-host nucleic acid ratio samples. The technology has similar advantages and drawbacks as the hybridization-based transcriptomic and genomic experimental techniques [152]. Multiple studies were conducted using the technology on multiple neoplasias [34, 153,154,155,156].
Quantitative polymerase chain reaction (qPCR)
qPCR is employed to quantify specific bacterial species or genes of interest in tissue samples. This technique is valuable for confirming the presence of a specific bacterial species or a bacterial gene identified through sequencing methods.
Imaging techniques
Fluorescence in situ hybridization (FISH) and immunohistochemistry (IHC) are used to visualize bacteria within tissue samples, allowing researchers to examine the spatial distribution of the oncobiome in relation to cancer cells and the tumor microenvironment (as an example see [18]).
Classical bacterial culture techniques
Culturable bacteria were isolated from human tumors [18]. This observation points towards the possible applicability of classical bacterial culture in characterizing bacterial communities (hereby we cite an example of a non-neoplastic disease [157]). The advantage of this approach is that the bacteria can be actually characterized in terms of immunogenic and biochemical properties or their effects on cancer (e.g., can be fed to an animal with cancer). The flip side of this approach is that unculturable bacteria cannot be assessed.
Optimizing oncobiome research in cancer: guidelines and standards
The incidence of neoplastic diseases escalates with advancing age [158], a phenomenon paralleled by aging-related shifts in the microbiome [159,160,161,162,163,164,165,166,167]. These microbiome changes are implicated in a range of aging-associated conditions, from cognitive decline to systemic inflammation and metabolic diseases linked to aging [168, 169]. Moreover, aging influences the microbiome’s composition, notably diminishing its diversity across various compartments [159,160,161,162,163,164,165,166,167]. This observation hints at a potential connection between aging-associated dysbiosis and heightened neoplastic disease risk in the elderly.
Amidst the surge in microbiome-related studies, research on oncobiosis has been expanding at a similar pace. However, not all studies meet the high standards required for impactful scientific contributions, as highlighted in select critiques [170]. This discrepancy underscores the urgent need for standardized guidelines to aid in the structuring of microbiome-related oncological research. Our objective is to contribute towards the standardization of patient cohort characterization, using breast cancer-related studies as a benchmark. Recent general guidelines for microbiome research have been set forth by Mirzayi et al. [171] and Bharti et al. [172], providing a foundational technical checklist.
In reviewing the literature on microbiome studies related to breast cancer (Table 1)—chosen as a model due to the correlation between breast cancer risk and age [173]—we screened PubMed using “microbiome” and “breast cancer” as search terms. Our analysis covered 90 papers focusing on human breast cancer, with the majority examining changes in the tumor tissue microbiome (39) and the GI tract microbiome (36). A smaller fraction addressed other compartments. Notably, approximately 40% of all studies, and a worrying 55% of cross-sectional studies, neglected to report participant ages or ensure age-matched cohorts, failing to account for age-related microbiome composition shifts.
The consistency in clinical characterization of study participants often falls short, hampering the comparability of research findings. Common characterization criteria include cancer stage, grade, and molecular subtype [174,175,176,177,178], with reporting frequencies of 57%, 34%, and 71%, respectively. Molecular subtype classification typically hinges on the expression profiles of the estrogen receptor, progesterone receptor, and HER2 receptor. However, detailed reporting on luminal A and B subtypes separately is rare. When documenting these variables, specifying the version of the classification system used is crucial. These details should accompany the sequencing data as metadata (refer to Fig. 1 for a recommended panel of patient data). For stratified cohorts, reporting should extend to sex distribution, average age among subgroups, and, for non-incident patients, medication and chemoradiotherapy regimens.
Control group selection criteria, including disease-free status assessment and duration of being disease-free (i.e., were the controls checked once before sampling for being disease-free or is the disease-free status maintained over extended periods), should be transparent. Furthermore, the study should account for potential confounders such as smoking habits, nutrition, BMI, menarche, and ECOG status. Sampling conditions, including the time lapse between sample collection and biobanking and the use of nucleic acid preservatives, must also be disclosed. Power calculations are advisable for experimental cohort setup, enhancing study robustness and reliability.
By adhering to these comprehensive guidelines, the microbiome research community can ensure greater consistency and comparability across studies, paving the way for advancements in understanding the oncobiome’s role in aging and neoplastic diseases.
Therapeutic implications and future direction
The emerging understanding of oncobiome’s role opens new avenues for therapeutic interventions and highlights the potential for innovative approaches to treatment and prevention.
The identification of specific bacterial species associated with cancer progression presents a unique opportunity to explore antibiotics and antimicrobial strategies as potential therapeutic interventions. Targeted antibiotics could be employed to disrupt harmful bacterial populations, potentially slowing or reversing tumor growth. However, this approach requires careful consideration to avoid disrupting the beneficial components of the microbiome [179,180,181], unwanted drug-drug interaction during chemotherapy [182, 183], and to prevent the development of antibiotic resistance. Antimicrobial peptides (AMPs) represent another promising strategy, offering the possibility of selectively targeting cancer-associated bacteria with reduced risk of disturbing the overall microbiome balance [184].
Probiotics, live microorganisms that confer health benefits when administered in adequate amounts, and prebiotics, non-digestible fibers that promote the growth of beneficial bacteria, offer another strategy for modulating the microbiome in favor of cancer prevention and treatment [10, 185,186,187,188,189,190]. These interventions could help restore a healthy microbial balance, potentially reducing inflammation and inhibiting the growth of species with negative effects [185,186,187,188]. Clinical trials exploring the efficacy of specific probiotic strains and prebiotic compounds in modulating the microbiome and their impact on cancer outcomes are needed to validate this approach.
The field of oncobiome and oncobiosis is ripe with opportunities for groundbreaking research that could transform our understanding and treatment of the disease. Key future directions include leveraging detailed microbiome profiles to tailor prevention and treatment strategies to the individual’s unique microbial and genetic landscape, potentially improving the efficacy of therapies and reducing side effects. Further, investigating the potential for vaccines targeting specific species of the oncobiome associated with cancer may offer a proactive approach to prevention and treatment. Exploring the combination of microbiome-targeted therapies with conventional treatments like chemotherapy, radiation, and immunotherapy may lead to enhance overall treatment efficacy and potentially reduce side effects. Further research is also needed to elucidate the complex interactions between the microbiome/oncobiome, the immune system, and cancer cells, including the mechanisms by which bacteria influence cancer development and progression [3, 102, 103, 105, 191].
Conclusion
This review aims at illuminating the pivotal role of oncobiosis, highlighting how microorganisms contribute to the disease’s initiation, progression, and possibly its response to treatment. The intricate interactions between the oncobiome and the host’s immune system, along with their influence on the tumor microenvironment, underscore the complexity of cancer’s etiology and the potential for microbial involvement in its pathology. Reflecting on the insights garnered, it becomes evident that the study of the oncobiome opens new avenues for the development of innovative diagnostics and therapeutic strategies. The identification of specific bacterial signatures associated with cancer offers the promise of novel biomarkers for early detection and prognosis, enhancing our ability to tackle the disease at its onset. Furthermore, understanding the mechanisms through which the oncobiome influences cancer initiation or progression provides a foundation for exploring antimicrobial treatments, probiotics, and prebiotics as potential adjuncts to conventional therapies, potentially improving outcomes and reducing side effects. However, the path to integrating microbiome research into breast cancer management is fraught with challenges, including the need for standardized methodologies in microbial analysis. Therefore, this review serves as a call to action for continued and expanded research in this emerging field. There is a pressing need for interdisciplinary collaboration among microbiologists, oncologists, geroscientists, and bioinformaticians to further elucidate the role of oncobiosis, refine diagnostic tools, and develop microbial-based interventions.
Data Availability
Not applicable.
References
Xu S, Liu Y, Zhang T, Zheng J, Lin W, Cai J, et al. The global, regional, and national burden and trends of breast cancer from 1990 to 2019: results from the Global Burden Of Disease Study 2019. Front Oncol. 2021;11: 689562. https://doi.org/10.3389/fonc.2021.689562.
Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73(1):17–48. https://doi.org/10.3322/caac.21763.
Kovács T, Mikó E, Ujlaki G, Yousef H, Csontos V, Uray K, et al. The involvement of oncobiosis and bacterial metabolite signaling in metastasis formation in breast cancer. Cancer Metastasis Rev. 2021;40(4):1223–49.
Wang N, Sun T, Xu J. Tumor-related microbiome in the breast microenvironment and breast cancer. J Cancer. 2021;12(16):4841–8. https://doi.org/10.7150/jca.58986.
Sohail S, Burns MB. Integrating current analyses of the breast cancer microbiome. PLoS ONE. 2023;18(9): e0291320. https://doi.org/10.1371/journal.pone.0291320.
Peters BA, Kelly L, Wang T, Loudig O, Rohan TE. The breast microbiome in breast cancer risk and progression: a narrative review. Cancer Epidemiol Biomarkers Prev. 2024;33(1):9–19. https://doi.org/10.1158/1055-9965.Epi-23-0965.
Hong W, Huang G, Wang D, Xu Y, Qiu J, Pei B, et al. Gut microbiome causal impacts on the prognosis of breast cancer: a Mendelian randomization study. BMC Genomics. 2023;24(1):497. https://doi.org/10.1186/s12864-023-09608-7.
Hoang J, Gilbertson-White S, Cady N, Yadav M, Shahi S, Aguilar L, et al. Preliminary analysis of gut microbiome and gastrointestinal symptom burden in breast cancer patients receiving chemotherapy compared to healthy controls. Biol Res Nurs. 2024;26(2):219–30. https://doi.org/10.1177/10998004231205277.
Chen F, Yang J, Guo Y, Su D, Sheng Y, Wu Y. Integrating bulk and single-cell RNA sequencing data reveals the relationship between intratumor microbiome signature and host metabolic heterogeneity in breast cancer. Front Immunol. 2023;14:1140995. https://doi.org/10.3389/fimmu.2023.1140995.
Avtanski D, Reddy V, Stojchevski R, Hadzi-Petrushev N, Mladenov M. The microbiome in the obesity-breast cancer axis: diagnostic and therapeutic potential. Pathogens. 2023;12(12):1402. https://doi.org/10.3390/pathogens12121402.
Brennan CA, Garrett WS. Fusobacterium nucleatum - symbiont, opportunist and oncobacterium. Nat Rev Microbiol. 2019;17(3):156–66. https://doi.org/10.1038/s41579-018-0129-6.
Saenz JB. Follow the metaplasia: characteristics and oncogenic implications of metaplasia’s pattern of spread throughout the stomach. Front Cell Dev Biol. 2021;9: 741574. https://doi.org/10.3389/fcell.2021.741574.
El Tekle G, Garrett WS. Bacteria in cancer initiation, promotion and progression. Nat Rev Cancer. 2023;23(9):600–18. https://doi.org/10.1038/s41568-023-00594-2.
Oh B, Boyle F, Pavlakis N, Clarke S, Guminski A, Eade T, et al. Emerging Evidence of the Gut Microbiome in Chemotherapy: A Clinical Review. Front Oncol. 2021;11:706331. https://doi.org/10.3389/fonc.2021.706331.
Thomas RM, Jobin C. The microbiome and cancer: is the ‘oncobiome’ mirage real? Trends Cancer. 2015;1(1):24–35. https://doi.org/10.1016/j.trecan.2015.07.005.
Li X, Sun X, Zhang A, Pang J, Li Y, Yan M, et al. Breast microbiome associations with breast tumor characteristics and neoadjuvant chemotherapy: a case-control study. Front Oncol. 2022;12: 926920. https://doi.org/10.3389/fonc.2022.926920.
Chan AA, Bashir M, Rivas MN, Duvall K, Sieling PA, Pieber TR, et al. Characterization of the microbiome of nipple aspirate fluid of breast cancer survivors. Sci Rep. 2016;6:28061. https://doi.org/10.1038/srep28061.
Nejman D, Livyatan I, Fuks G, Gavert N, Zwang Y, Geller LT, et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science. 2020;368(6494):973–80. https://doi.org/10.1126/science.aay9189.
Feng K, Ren F, Wang X. Relationships among breast, gut, and oral microbiota across diverse pathological types of breast cancer, a Chinese cohort study. Front Mol Biosci. 2023;10:1325552. https://doi.org/10.3389/fmolb.2023.1325552.
Xuan C, Shamonki JM, Chung A, Dinome ML, Chung M, Sieling PA, et al. Microbial dysbiosis is associated with human breast cancer. PLoS ONE. 2014;9(1): e83744.
Smith A, Cao X, Gu Q, Kubi Amos-Abanyie E, Tolley EA, Vidal G, et al. Characterization of the metabolome of breast tissues from non-Hispanic Black and non-Hispanic White women reveals correlations between microbial dysbiosis and enhanced lipid metabolism pathways in triple-negative breast tumors. Cancers (Basel). 2022;14(17). https://doi.org/10.3390/cancers14174075.
Duan Y, Ma F, Guo B, Chen Z, Liu Y, Jiang X, et al. Listeria monocytogenes promotes breast cancer proliferation and enhances the survival rate of circulating breast cancer cells. Research Square; 2023.
Lee CC, Yang HW, Liu CJ, Lee F, Ko WC, Chang YC, et al. Unraveling the connections between gut microbiota, stress, and quality of life for holistic care in newly diagnosed breast cancer patients. Sci Rep. 2023;13(1):17916. https://doi.org/10.1038/s41598-023-45123-1.
Liu E, Zhang F, Xu T, Ye L, Ma SSQ, Ji Z-S. Relationship between tumor microbiota transcriptional activity and gene expression in breast cancer. BMC Cancer. 2023;23(1):252. https://doi.org/10.1186/s12885-023-10726-4.
Desalegn Z, Smith A, Yohannes M, Cao X, Anberber E, Bekuretsion Y, et al. Human breast tissue microbiota reveals unique microbial signatures that correlate with prognostic features in adult Ethiopian women with breast cancer. Cancers (Basel). 2023;15(19). https://doi.org/10.3390/cancers15194893.
Kim HE, Kim J, Maeng S, Oh B, Hwang KT, Kim BS. Microbiota of breast tissue and its potential association with regional recurrence of breast cancer in Korean women. J Microbiol Biotechnol. 2021;31(11). https://doi.org/10.4014/jmb.2106.06039.
Kartti S, Bendani H, Boumajdi N, Bouricha EM, Zarrik O, El Agouri H, et al. Metagenomics analysis of breast microbiome highlights the abundance of Rothia genus in tumor tissues. J Pers Med. 2023;13(3). https://doi.org/10.3390/jpm13030450.
Hogan G, Eckenberger J, Narayanen N, Walker SP, Claesson MJ, Corrigan M, et al. Biopsy bacterial signature can predict patient tissue malignancy. Sci Rep. 2021;11(1):18535. https://doi.org/10.1038/s41598-021-98089-3.
Costantini L, Magno S, Albanese D, Donati C, Molinari R, Filippone A, et al. Characterization of human breast tissue microbiota from core needle biopsies through the analysis of multi hypervariable 16S-rRNA gene regions. Sci Rep. 2018;8(1):16893.
Thyagarajan S, Zhang Y, Thapa S, Allen MS, Phillips N, Chaudhary P, et al. Comparative analysis of racial differences in breast tumor microbiome. Sci Rep. 2020;10(1):14116. https://doi.org/10.1038/s41598-020-71102-x.
Fu A, Yao B, Dong T, Chen Y, Yao J, Liu Y, et al. Tumor-resident intracellular microbiota promotes metastatic colonization in breast cancer. Cell. 2022. https://doi.org/10.1016/j.cell.2022.02.027.
Banerjee S, Wei Z, Tan F, Peck KN, Shih N, Feldman M, et al. Distinct microbiological signatures associated with triple negative breast cancer. Sci Rep. 2015;5:15162.
Banerjee S, Wei Z, Tian T, Bose D, Shih NNC, Feldman MD, et al. Prognostic correlations with the microbiome of breast cancer subtypes. Cell Death Dis. 2021;12(9):831. https://doi.org/10.1038/s41419-021-04092-x.
Banerjee S, Tian T, Wei Z, Shih N, Feldman MD, Peck KN, et al. Distinct microbial signatures associated with different breast cancer types. Front Microbiol. 2018;9:951.
Hieken TJ, Chen J, Hoskin TL, Walther-Antonio M, Johnson S, Ramaker S, et al. The microbiome of aseptically collected human breast tissue in benign and malignant disease. Sci Rep. 2016;6:30751.
Hoskinson C, Zheng K, Gabel J, Kump A, German R, Podicheti R, et al. Composition and functional potential of the human mammary microbiota prior to and following breast tumor diagnosis. mSystems. 2022;7(3):e0148921. https://doi.org/10.1128/msystems.01489-21.
Esposito MV, Fosso B, Nunziato M, Casaburi G, D’Argenio V, Calabrese A, et al. Microbiome composition indicate dysbiosis and lower richness in tumor breast tissues compared to healthy adjacent paired tissue, within the same women. BMC Cancer. 2022;22(1):30. https://doi.org/10.1186/s12885-021-09074-y.
Soto-Pantoja DR, Gaber M, Arnone AA, Bronson SM, Cruz-Diaz N, Wilson AS, et al. Diet alters entero-mammary signaling to regulate the breast microbiome and tumorigenesis. Cancer Res. 2021;81(14):3890–904. https://doi.org/10.1158/0008-5472.Can-20-2983.
Feng Z, Hu Y, Wang X, Li Y, Yu Y, He J, et al. In situ imaging for tumor microbiome interactions via imaging mass cytometry on single-cell level. Cytometry A. 2022;101(8):617–29. https://doi.org/10.1002/cyto.a.24550.
Hilmi M, Kamal M, Vacher S, Dupain C, Ibadioune S, Halladjian M, et al. Intratumoral microbiome is driven by metastatic site and associated with immune histopathological parameters: an ancillary study of the SHIVA clinical trial. Eur J Cancer. 2023;183:152–61. https://doi.org/10.1016/j.ejca.2023.01.024.
Hadzega D, Minarik G, Karaba M, Kalavska K, Benca J, Ciernikova S, et al. Uncovering microbial composition in human breast cancer primary tumour tissue using transcriptomic RNA-seq. Int J Mol Sci. 2021;22(16):9058. https://doi.org/10.3390/ijms22169058.
Wang Y, Qu D, Zhang Y, Jin Y, Feng Y, Zhang H, et al. Intra-tumoral microbial community profiling and associated metabolites alterations of TNBC. Front Oncol. 2023;13:1143163. https://doi.org/10.3389/fonc.2023.1143163.
Klann E, Williamson JM, Tagliamonte MS, Ukhanova M, Asirvatham JR, Chim H, et al. Microbiota composition in bilateral healthy breast tissue and breast tumors. Cancer Causes Control. 2020;31(11):1027–38. https://doi.org/10.1007/s10552-020-01338-5.
Tzeng A, Sangwan N, Jia M, Liu C-C, Keslar KS, Downs-Kelly E, et al. Human breast microbiome correlates with prognostic features and immunological signatures in breast cancer. Genome Med. 2021;13(1):60. https://doi.org/10.1186/s13073-021-00874-2.
Meng S, Chen B, Yang J, Wang J, Zhu D, Meng Q, et al. Study of microbiomes in aseptically collected samples of human breast tissue using needle biopsy and the potential role of in situ tissue microbiomes for promoting malignancy. Front Oncol. 2018;8:318.
Urbaniak C, Gloor GB, Brackstone M, Scott L, Tangney M, Reid G. The microbiota of breast tissue and its association with breast cancer. Appl Environ Microbiol. 2016;82(16):5039–48. https://doi.org/10.1128/AEM.01235-16.
Smith A, Pierre JF, Makowski L, Tolley E, Lyn-Cook B, Lu L, et al. Distinct microbial communities that differ by race, stage, or breast-tumor subtype in breast tissues of non-Hispanic Black and non-Hispanic White women. Sci Rep. 2019;9(1):11940.
Wang H, Altemus J, Niazi F, Green H, Calhoun BC, Sturgis C, et al. Breast tissue, oral and urinary microbiomes in breast cancer. Oncotarget. 2017;8(50):88122–38.
Giallourou N, Urbaniak C, Puebla-Barragan S, Vorkas PA, Swann JR, Reid G. Characterizing the breast cancer lipidome and its interaction with the tissue microbiota. Commun Biol. 2021;4(1):1229. https://doi.org/10.1038/s42003-021-02710-0.
Luo L, Fu A, Shi M, Hu J, Kong D, Liu T, et al. Species-level characterization of the microbiome in breast tissues with different malignancy and hormone-receptor statuses using nanopore sequencing. J Pers Med. 2023;13(2). https://doi.org/10.3390/jpm13020174.
Thompson KJ, Ingle JN, Tang X, Chia N, Jeraldo PR, Walther-Antonio MR, et al. A comprehensive analysis of breast cancer microbiota and host gene expression. PLoS ONE. 2017;12(11): e0188873.
Poore GD, Kopylova E, Zhu Q, Carpenter C, Fraraccio S, Wandro S, et al. Microbiome analyses of blood and tissues suggest cancer diagnostic approach. Nature. 2020;579(7800):567–74. https://doi.org/10.1038/s41586-020-2095-1.
Parida S, Siddharth S, Xia Y, Sharma D. Concomitant analyses of intratumoral microbiota and genomic features reveal distinct racial differences in breast cancer. npj Breast Cancer. 2023;9(1):4. https://doi.org/10.1038/s41523-023-00505-6.
Gao X, Yang H, Chu Y, Zhang W, Wang Z, Ji L. The specific viral composition in triple-negative breast cancer tissue shapes the specific tumor microenvironment characterized on pathological images. Microb Pathog. 2023;184: 106385. https://doi.org/10.1016/j.micpath.2023.106385.
An J, Yang J, Kwon H, Lim W, Kim YK, Moon BI. Prediction of breast cancer using blood microbiome and identification of foods for breast cancer prevention. Sci Rep. 2023;13(1):5110. https://doi.org/10.1038/s41598-023-32227-x.
An J, Kwon H, Kim YJ. The firmicutes/bacteroidetes ratio as a risk factor of breast cancer. J Clin Med. 2023;12(6). https://doi.org/10.3390/jcm12062216.
An J, Kil BJ, Kwon H, Kim YJ. Analysis of the impact of the presence of phylum Cyanobacteria in the microbiome of patients with breast cancer on their prognosis. J Clin Med. 2022;11(24). https://doi.org/10.3390/jcm11247272.
An J, Kwon H, Kim YJ. The role of blood microbiome in the development of thyroid cancer in breast cancer survivors. Cancers (Basel). 2023;15(18). https://doi.org/10.3390/cancers15184492.
Hou MF, Ou-Yang F, Li CL, Chen FM, Chuang CH, Kan JY, et al. Comprehensive profiles and diagnostic value of menopausal-specific gut microbiota in premenopausal breast cancer. Exp Mol Med. 2021;53(10):1636–46. https://doi.org/10.1038/s12276-021-00686-9.
Yang P, Wang Z, Peng Q, Lian W, Chen D. Comparison of the gut microbiota in patients with benign and malignant breast tumors: a pilot study. Evol Bioinformatics Online. 2021;17:11769343211057572. https://doi.org/10.1177/11769343211057573.
Yao ZW, Yang X, Zhao BC, Deng F, Jiang YM, Pan WY, et al. Predictive and preventive potential of preoperative gut microbiota in chronic postoperative pain in breast cancer survivors. Anesth Analg. 2022;134(4):699–709. https://doi.org/10.1213/ane.0000000000005713.
Ubachs J, Ziemons J, Soons Z, Aarnoutse R, van Dijk DPJ, Penders J, et al. Gut microbiota and short-chain fatty acid alterations in cachectic cancer patients. J Cachexia Sarcopenia Muscle. 2021;12(6):2007–21. https://doi.org/10.1002/jcsm.12804.
Goedert JJ, Jones G, Hua X, Xu X, Yu G, Flores R, et al. Investigation of the association between the fecal microbiota and breast cancer in postmenopausal women: a population-based case-control pilot study. J Natl Cancer Inst. 2015;107(8):djv147. https://doi.org/10.1093/jnci/djv147.
Miko E, Vida A, Kovacs T, Ujlaki G, Trencsenyi G, Marton J, et al. Lithocholic acid, a bacterial metabolite reduces breast cancer cell proliferation and aggressiveness. Biochim Biophys Acta - Bioenerg. 2018;1859(9):958–74.
Kovács T, Mikó E, Vida A, Sebő É, Toth J, Csonka T, et al. Cadaverine, a metabolite of the microbiome, reduces breast cancer aggressiveness through trace amino acid receptors. Sci Rep. 2019;9(1):1300. https://doi.org/10.1038/s41598-018-37664-7.
Sári Z, Kovács T, Csonka T, Török M, Sebő É, Toth J, et al. Fecal expression of E. coli lysine decarboxylase (LdcC) is downregulated in E-cadherin negative lobular breast carcinoma. Physiol Int. 2020;107(2):349–58. https://doi.org/10.1556/2060.2020.00016.
Goedert JJ, Hua X, Bielecka A, Okayasu I, Milne GL, Jones GS, et al. Postmenopausal breast cancer and oestrogen associations with the IgA-coated and IgA-noncoated faecal microbiota. Br J Cancer. 2018;23(10):435.
Zhu J, Liao M, Yao Z, Liang W, Li Q, Liu J, et al. Breast cancer in postmenopausal women is associated with an altered gut metagenome. Microbiome. 2018;6(1):136.
Wenhui Y, Zhongyu X, Kai C, Zhaopeng C, Jinteng L, Mengjun M, et al. Variations in the gut microbiota in breast cancer occurrence and bone metastasis. Front Microbiol. 2022;13: 894283. https://doi.org/10.3389/fmicb.2022.894283.
Ma Z, Qu M, Wang X. Analysis of gut microbiota in patients with breast cancer and benign breast lesions. Pol J Microbiol. 2022;71(2):217–26. https://doi.org/10.33073/pjm-2022-019.
Hoang J, Gilbertson-White S, Cady N, Yadav M, Shahi S, Aguilar L, et al. Preliminary analysis of gut microbiome and gastrointestinal symptom burden in breast cancer patients receiving chemotherapy compared to healthy controls. Biol Res Nurs. 2023:10998004231205277. https://doi.org/10.1177/10998004231205277.
Maitiniyazi G, Cao X, Chen Y, Zhang R, Liu Y, Li Z, et al. Impact of gut microbiota on the association between diet and depressive symptoms in breast cancer. Nutrients. 2022;14(6). https://doi.org/10.3390/nu14061186.
Horigome A, Okubo R, Hamazaki K, Kinoshita T, Katsumata N, Uezono Y, et al. Association between blood omega-3 polyunsaturated fatty acids and the gut microbiota among breast cancer survivors. Beneficial microbes. 2019;10(7):751–8. https://doi.org/10.3920/bm2019.0034.
Fruge AD, Van der Pol W, Rogers LQ, Morrow CD, Tsuruta Y, Demark-Wahnefried W. Fecal Akkermansia muciniphila is associated with body composition and microbiota diversity in overweight and obese women with breast cancer participating in a presurgical weight loss trial. J Acad Nutr Diet. 2018;9(18):164.
Wu AH, Tseng C, Vigen C, Yu Y, Cozen W, Garcia AA, et al. Gut microbiome associations with breast cancer risk factors and tumor characteristics: a pilot study. Breast Cancer Res Treat. 2020. https://doi.org/10.1007/s10549-020-05702-6.
Vernaci G, Savarino EV, Patuzzi I, Facchin S, Zingone F, Massa D, et al. Characterization of gut microbiome composition in patients with triple-negative breast cancer treated with neoadjuvant chemotherapy. Oncologist. 2023. https://doi.org/10.1093/oncolo/oyad060.
Terrisse S, Derosa L, Iebba V, Ghiringhelli F, Vaz-Luis I, Kroemer G, et al. Intestinal microbiota influences clinical outcome and side effects of early breast cancer treatment. Cell Death Differ. 2021. https://doi.org/10.1038/s41418-021-00784-1.
Walker J, Joy AA, Vos LJ, Stenson TH, Mackey JR, Jovel J, et al. Chemotherapy-induced weight gain in early-stage breast cancer: a prospective matched cohort study reveals associations with inflammation and gut dysbiosis. BMC Med. 2023;21(1):178. https://doi.org/10.1186/s12916-023-02751-8.
Wu AH, Vigen C, Tseng C, Garcia AA, Spicer D. Effect of chemotherapy and weight change on the gut microbiome of breast cancer patients during the first year of treatment. Research Square. 2021. https://www.researchsquare.com/article/rs-970564/v1. Accessed 2021 11 09.
Smith KS, Tissier A, Bail JR, Novak JR, Morrow CD, Demark-Wahnefried W, et al. Health-related quality of life is associated with fecal microbial composition in breast cancer survivors. Support Care Cancer. 2022;31(1):10. https://doi.org/10.1007/s00520-022-07496-3.
Aarnoutse R, Ziemons J, Hillege LE, de Vos-Geelen J, de Boer M, Bisschop SMP, et al. Changes in intestinal microbiota in postmenopausal oestrogen receptor-positive breast cancer patients treated with (neo)adjuvant chemotherapy. NPJ Breast Cancer. 2022;8(1):89. https://doi.org/10.1038/s41523-022-00455-5.
Sári Z, Mikó E, Kovács T, Jankó L, Csonka T, Sebő E, et al. Indolepropionic acid, a metabolite of the microbiome, has cytostatic properties in breast cancer by activating AHR and PXR receptors and inducing oxidative stress. Cancers (Basel). 2020;12(9):2411. https://doi.org/10.3390/cancers12092411.
Luu TH, Michel C, Bard JM, Dravet F, Nazih H, Bobin-Dubigeon C. Intestinal proportion of Blautia sp. is associated with clinical stage and histoprognostic grade in patients with early-stage breast cancer. Nutr Cancer. 2017;69(2):267–75.
Robinson S, Teng N, Malfettone A, Dalby M, Kiu R, Seki D, et al. Profiling the gut and oral microbiota of hormone-receptor positive, HER2-negative metastatic breast cancer patients receiving pembrolizumab and eribulin. Research Square. 2024. https://doi.org/10.21203/rs.3.rs-3787741/v1.
Li Y, Dong B, Wu W, Wang J, Jin H, Chen K, et al. Metagenomic analyses reveal distinct gut microbiota signature for predicting the neoadjuvant chemotherapy responsiveness in breast cancer patients. Front Oncol. 2022;12: 865121. https://doi.org/10.3389/fonc.2022.865121.
Di Modica M, Gargari G, Regondi V, Bonizzi A, Arioli S, Belmonte B, et al. Gut microbiota condition the therapeutic efficacy of trastuzumab in HER2-positive breast cancer. Cancer Res. 2021;81(8):2195–206. https://doi.org/10.1158/0008-5472.Can-20-1659.
Wong CW, Yost SE, Lee JS, Gillece JD, Folkerts M, Reining L, et al. Analysis of gut microbiome using explainable machine learning predicts risk of diarrhea associated with tyrosine kinase inhibitor neratinib: a pilot study. Front Oncol. 2021;11: 604584. https://doi.org/10.3389/fonc.2021.604584.
Wu Z, Byrd DA, Wan Y, Ansong D, Clegg-Lamptey J-N, Wiafe-Addai B, et al. The oral microbiome and breast cancer and nonmalignant breast disease, and its relationship with the fecal microbiome in the Ghana Breast Health Study. Int J Cancer. 2022;151(8):1248–60. https://doi.org/10.1002/ijc.34145.
Murray WR, Blackwood A, Calman KC, MacKay C. Faecal bile acids and clostridia in patients with breast cancer. Br J Cancer. 1980;42(6):856–60. https://doi.org/10.1038/bjc.1980.333.
Lasagna A, De Amici M, Rossi C, Zuccaro V, Corbella M, Petazzoni G, et al. The bio-diversity and the role of gut microbiota in postmenopausal women with luminal breast cancer treated with aromatase inhibitors: an observational cohort study. Pathogens. 2022;11(12). https://doi.org/10.3390/pathogens11121421.
Byrd DA, Vogtmann E, Wu Z, Han Y, Wan Y, Clegg-Lamptey J-N, et al. Associations of fecal microbial profiles with breast cancer and non-malignant breast disease in the Ghana Breast Health Study. Int J Cancer. 2021;148(11):2712–23. https://doi.org/10.1002/ijc.33473.
Bobin-Dubigeon C, Luu HT, Leuillet S, Lavergne SN, Carton T, Le Vacon F, et al. Faecal microbiota composition varies between patients with breast cancer and healthy women: a comparative case-control study. Nutrients. 2021;13(8). https://doi.org/10.3390/nu13082705.
Shrode RL, Knobbe JE, Cady N, Yadav M, Hoang J, Cherwin C, et al. Breast cancer patients from the Midwest region of the United States have reduced levels of short-chain fatty acid-producing gut bacteria. Sci Rep. 2023;13(1):526. https://doi.org/10.1038/s41598-023-27436-3.
Caleça T, Ribeiro P, Vitorino M, Menezes M, Sampaio-Alves M, Mendes AD, et al. Breast cancer survivors and healthy women: could gut microbiota make a difference?-“BiotaCancerSurvivors”: a case-control study. Cancers (Basel). 2023;15(3). https://doi.org/10.3390/cancers15030594.
Bilenduke E, Sterrett JD, Ranby KW, Borges VF, Grigsby J, Carr AL, et al. Impacts of breast cancer and chemotherapy on gut microbiome, cognitive functioning, and mood relative to healthy controls. Sci Rep. 2022;12(1):19547. https://doi.org/10.1038/s41598-022-23793-7.
Wu R, Yu I, Tokumaru Y, Asaoka M, Oshi M, Yan L, et al. Elevated bile acid metabolism and microbiome are associated with suppressed cell proliferation and better survival in breast cancer. Am J Cancer Res. 2022;12(11):5271–85.
An J, Kim JB, Yang EY, Kim HO, Lee W-H, Yang J, et al. Bacterial extracellular vesicles affect endocrine therapy in MCF7 cells. Medicine. 2021;100(18).
Kim HY, Kim TH, Shin JH, Cho K, Ha HK, Lee A, et al. Navigating the microbial community in the trachea-oropharynx of breast cancer patients with or without neoadjuvant chemotherapy (NAC) via endotracheal tube: has NAC caused any change? PeerJ. 2023;11: e16366. https://doi.org/10.7717/peerj.16366.
Viana MC, Curty G, Furtado C, Singh B, Bendall ML, Viola JPB, et al. Naso-oropharyngeal microbiome from breast cancer patients diagnosed with COVID-19. Front Microbiol. 2022;13:1074382. https://doi.org/10.3389/fmicb.2022.1074382.
Bilgilier C, Füreder T, Kastner MT, Vass Z, Brandl I, Sahbegovic H, et al. Oral abundance of Actinomyces spp. in breast cancer patients. Oncology. 2022. https://doi.org/10.1159/000522070.
Klymiuk I, Bilgilier C, Mahnert A, Prokesch A, Heininger C, Brandl I, et al. Chemotherapy-associated oral microbiome changes in breast cancer patients. Front Oncol. 2022;12: 949071. https://doi.org/10.3389/fonc.2022.949071.
Sipos A, Ujlaki G, Mikó E, Maka E, Szabó J, Uray K, et al. The role of the microbiome in ovarian cancer: mechanistic insights into oncobiosis and to bacterial metabolite signaling. Mol Med. 2021;27(1):33. https://doi.org/10.1186/s10020-021-00295-2.
Kiss B, Mikó E, Sebő É, Toth J, Ujlaki G, Szabó J, et al. Oncobiosis and microbial metabolite signaling in pancreatic adenocarcinoma. Cancers (Basel). 2020;12(5):E1068. https://doi.org/10.3390/cancers12051068.
van Elsland D, Neefjes J. Bacterial infections and cancer. EMBO Rep. 2018;19(11):e46632. https://doi.org/10.15252/embr.201846632.
Miko E, Kovacs T, Sebo E, Toth J, Csonka T, Ujlaki G, et al. Microbiome-microbial metabolome-cancer cell interactions in breast cancer-familiar, but unexplored. Cells. 2019;8(4):E293.
Režen T, Rozman D, Kovács T, Kovács P, Sipos A, Bai P, et al. The role of bile acids in carcinogenesis. Cell Mol Life Sci. 2022;79(5):243. https://doi.org/10.1007/s00018-022-04278-2.
Buchta Rosean CM, Rutkowski MR. The influence of the commensal microbiota on distal tumor-promoting inflammation. Semin Immunol. 2017;32:62–73. https://doi.org/10.1016/j.smim.2017.06.002.
Buchta Rosean C, Bostic RR, Ferey JCM, Feng TY, Azar FN, Tung KS, et al. Preexisting commensal dysbiosis is a host-intrinsic regulator of tissue inflammation and tumor cell dissemination in hormone receptor-positive breast cancer. Can Res. 2019;79(14):3662–75.
Chiba A, Bawaneh A, Velazquez C, Clear KYJ, Wilson AS, Howard-McNatt M, et al. Neoadjuvant chemotherapy shifts breast tumor microbiota populations to regulate drug responsiveness and the development of metastasis. Mol Cancer Res. 2019;18:1541–7786.
Hand TW, Overacre-Delgoffe AE. The complex immunological role of Helicobacter in modulating cancer. Trends Immunol. 2022;43(10):826–32. https://doi.org/10.1016/j.it.2022.08.002.
Shirani M, Pakzad R, Haddadi MH, Akrami S, Asadi A, Kazemian H, et al. The global prevalence of gastric cancer in Helicobacter pylori-infected individuals: a systematic review and meta-analysis. BMC Infect Dis. 2023;23(1):543. https://doi.org/10.1186/s12879-023-08504-5.
Reyes VE. Helicobacter pylori and its role in gastric cancer. Microorganisms. 2023;11(5):1312. https://doi.org/10.3390/microorganisms11051312.
Suzuki H, Saito Y, Hibi T. Helicobacter pylori and gastric mucosa-associated lymphoid tissue (MALT) lymphoma: updated review of clinical outcomes and the molecular pathogenesis. Gut Liver. 2009;3(2):81–7. https://doi.org/10.5009/gnl.2009.3.2.81.
Kuo SH, Wu MS, Yeh KH, Lin CW, Hsu PN, Chen LT, et al. Novel insights of lymphomagenesis of Helicobacter pylori-dependent gastric mucosa-associated lymphoid tissue lymphoma. Cancers (Basel). 2019;11(4):547. https://doi.org/10.3390/cancers11040547.
Raderer M, Wrba F, Kornek G, Maca T, Koller DY, Weinlaender G, et al. Association between Helicobacter pylori infection and pancreatic cancer. Oncology. 1998;55(1):16–9. https://doi.org/10.1159/000011830.
Epplein M, Nomura AM, Hankin JH, Blaser MJ, Perez-Perez G, Stemmermann GN, et al. Association of Helicobacter pylori infection and diet on the risk of gastric cancer: a case-control study in Hawaii. Cancer Causes Control. 2008;19(8):869–77. https://doi.org/10.1007/s10552-008-9149-2.
Wang X, Huycke MM. Colorectal cancer: role of commensal bacteria and bystander effects. Gut Microbes. 2015;6(6):370–6. https://doi.org/10.1080/19490976.2015.1103426.
Thomas RM. Role of bacteria in the development of colorectal cancer. Clin Colon Rectal Surg. 2023;36(2):105–11. https://doi.org/10.1055/s-0042-1760679.
Koliarakis I, Messaritakis I, Nikolouzakis TK, Hamilos G, Souglakos J, Tsiaoussis J. Oral Bacteria and intestinal dysbiosis in colorectal cancer. Int J Mol Sci. 2019;20(17):4146. https://doi.org/10.3390/ijms20174146.
Rubinstein MR, Baik JE, Lagana SM, Han RP, Raab WJ, Sahoo D, et al. Fusobacterium nucleatum promotes colorectal cancer by inducing Wnt/beta-catenin modulator Annexin A1. EMBO Rep. 2019;20(4). https://doi.org/10.15252/embr.201847638.
Hashemi Goradel N, Heidarzadeh S, Jahangiri S, Farhood B, Mortezaee K, Khanlarkhani N, et al. Fusobacterium nucleatum and colorectal cancer: a mechanistic overview. J Cell Physiol. 2019;234(3):2337–44. https://doi.org/10.1002/jcp.27250.
Guo P, Tian Z, Kong X, Yang L, Shan X, Dong B, et al. FadA promotes DNA damage and progression of Fusobacterium nucleatum-induced colorectal cancer through up-regulation of chk2. J Exp Clin Cancer Res. 2020;39(1):202. https://doi.org/10.1186/s13046-020-01677-w.
Wu Z, Ma Q, Guo Y, You F. The role of Fusobacterium nucleatum in colorectal cancer cell proliferation and migration. Cancers (Basel). 2022;14(21):5350. https://doi.org/10.3390/cancers14215350.
Wang S, Liu Y, Li J, Zhao L, Yan W, Lin B, et al. Fusobacterium nucleatum acts as a pro-carcinogenic bacterium in colorectal cancer: from association to causality. Front Cell Dev Biol. 2021;9: 710165. https://doi.org/10.3389/fcell.2021.710165.
Pignatelli P, Nuccio F, Piattelli A, Curia MC. The role of Fusobacterium nucleatum in oral and colorectal carcinogenesis. Microorganisms. 2023;11(9):2358. https://doi.org/10.3390/microorganisms11092358.
Bostanghadiri N, Razavi S, Shariati A, Talebi M, Mirkalantari S, Emami Razavi A, et al. Exploring the interplay between Fusobacterium nucleatum with the expression of microRNA, and inflammatory mediators in colorectal cancer. Front Microbiol. 2023;14:1302719. https://doi.org/10.3389/fmicb.2023.1302719.
Sakamoto Y, Mima K, Ishimoto T, Ogata Y, Imai K, Miyamoto Y, et al. Relationship between Fusobacterium nucleatum and antitumor immunity in colorectal cancer liver metastasis. Cancer Sci. 2021;112(11):4470–7. https://doi.org/10.1111/cas.15126.
Mima K, Sukawa Y, Nishihara R, Qian ZR, Yamauchi M, Inamura K, et al. Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol. 2015;1(5):653–61. https://doi.org/10.1001/jamaoncol.2015.1377.
Mima K, Nishihara R, Qian ZR, Cao Y, Sukawa Y, Nowak JA, et al. Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut. 2016;65(12):1973–80. https://doi.org/10.1136/gutjnl-2015-310101.
Mehta RS, Nishihara R, Cao Y, Song M, Mima K, Qian ZR, et al. Association of dietary patterns with risk of colorectal cancer subtypes classified by Fusobacterium nucleatum in tumor tissue. JAMA Oncol. 2017;3(7):921–7. https://doi.org/10.1001/jamaoncol.2016.6374.
Joo JE, Chu YL, Georgeson P, Walker R, Mahmood K, Clendenning M, et al. Intratumoral presence of the genotoxic gut bacteria pks(+) E. coli, Enterotoxigenic Bacteroides fragilis, and Fusobacterium nucleatum and their association with clinicopathological and molecular features of colorectal cancer. Br J Cancer. 2024;130(5):728–40. https://doi.org/10.1038/s41416-023-02554-x.
Borowsky J, Haruki K, Lau MC, Dias Costa A, Vayrynen JP, Ugai T, et al. Association of Fusobacterium nucleatum with specific T-cell subsets in the colorectal carcinoma microenvironment. Clin Cancer Res. 2021;27(10):2816–26. https://doi.org/10.1158/1078-0432.CCR-20-4009.
Yang J, Wang X, Hu T, Huang H, Chen G, Jin B, et al. Entero-toxigenic Bacteroides fragilis contributes to intestinal barrier injury and colorectal cancer progression by mediating the BFT/STAT3/ZEB2 pathway. Cell Cycle. 2024:1–13. https://doi.org/10.1080/15384101.2024.2309005.
Wu X, Yang C, Sun F, Zhang Y, Wang Y, Li X, et al. Enterotoxigenic Bacteroides fragilis (ETBF) enhances colorectal cancer cell proliferation and metastasis through HDAC3/miR-139-3p pathway. Biochem Genet. 2024. https://doi.org/10.1007/s10528-023-10621-4.
Purcell RV, Pearson J, Aitchison A, Dixon L, Frizelle FA, Keenan JI. Colonization with enterotoxigenic Bacteroides fragilis is associated with early-stage colorectal neoplasia. PLoS ONE. 2017;12(2): e0171602. https://doi.org/10.1371/journal.pone.0171602.
Liu QQ, Li CM, Fu LN, Wang HL, Tan J, Wang YQ, et al. Enterotoxigenic Bacteroides fragilis induces the stemness in colorectal cancer via upregulating histone demethylase JMJD2B. Gut Microbes. 2020;12(1):1788900. https://doi.org/10.1080/19490976.2020.1788900.
Terlouw D, Boot A, Ducarmon QR, Nooij S, Suerink M, van Leerdam ME, et al. Enrichment of colibactin-associated mutational signatures in unexplained colorectal polyposis patients. BMC Cancer. 2024;24(1):104. https://doi.org/10.1186/s12885-024-11849-y.
Rosendahl Huber A, Pleguezuelos-Manzano C, Puschhof J, Ubels J, Boot C, Saftien A, et al. Improved detection of colibactin-induced mutations by genotoxic E. coli in organoids and colorectal cancer. Cancer Cell. 2024;42(3):487-96 e6. https://doi.org/10.1016/j.ccell.2024.02.009.
de Oliveira AN, Dalmasso G, Nikitina D, Vaysse A, Ruez R, Ledoux L, et al. The colibactin-producing Escherichia coli alters the tumor microenvironment to immunosuppressive lipid overload facilitating colorectal cancer progression and chemoresistance. Gut Microbes. 2024;16(1):2320291. https://doi.org/10.1080/19490976.2024.2320291.
Romay E, Pericas JM, Garcia-Pais MJ, Hernandez-Meneses M, Ayuso B, Garcia-Gonzalez J, et al. Relationship among Streptococcus gallolyticus subsp. gallolyticus, Enterococcus faecalis and colorectal neoplasms in recurrent endocarditis: a historical case series. J Clin Med. 2022;11(8). https://doi.org/10.3390/jcm11082181.
Mohammad Aidid E, Shalihin MSE, Md Nor A, Hamzah HA, Ab Hamid NF, Saipol Bahri NAN, et al. Risk of colorectal cancer due to Streptococcus gallolyticus: a systematic review. Med J Malaysia. 2023;78(3):404–10.
Epplein M, Le Marchand L, Cover TL, Song M, Blot WJ, Peek RM, et al. Association of combined sero-positivity to Helicobacter pylori and Streptococcus gallolyticus with risk of colorectal cancer. Microorganisms. 2020;8(11). https://doi.org/10.3390/microorganisms8111698.
Butt J, Fernandez de Larrea N, Tjalsma H, Roelofs R, Kato I, Martin V, et al. Antibody responses to flagellin C and Streptococcus gallolyticus pilus proteins in colorectal cancer. Sci Rep. 2019;9(1):10847. https://doi.org/10.1038/s41598-019-47347-6.
Aymeric L, Donnadieu F, Mulet C, du Merle L, Nigro G, Saffarian A, et al. Colorectal cancer specific conditions promote Streptococcus gallolyticus gut colonization. Proc Natl Acad Sci U S A. 2018;115(2):E283–91. https://doi.org/10.1073/pnas.1715112115.
Andres-Franch M, Galiana A, Sanchez-Hellin V, Ochoa E, Hernandez-Illan E, Lopez-Garcia P, et al. Streptococcus gallolyticus infection in colorectal cancer and association with biological and clinical factors. PLoS ONE. 2017;12(3): e0174305. https://doi.org/10.1371/journal.pone.0174305.
Edgar RC. Accuracy of taxonomy prediction for 16S rRNA and fungal ITS sequences. PeerJ. 2018;6: e4652. https://doi.org/10.7717/peerj.4652.
Durazzi F, Sala C, Castellani G, Manfreda G, Remondini D, De Cesare A. Comparison between 16S rRNA and shotgun sequencing data for the taxonomic characterization of the gut microbiota. Sci Rep. 2021;11(1):3030. https://doi.org/10.1038/s41598-021-82726-y.
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol. 2013;31(9):814–21. https://doi.org/10.1038/nbt.2676.
de Goffau MC, Lager S, Salter SJ, Wagner J, Kronbichler A, Charnock-Jones DS, et al. Recognizing the reagent microbiome. Nature microbiology. 2018;3(8):851-3. doi: 10.1038/s41564-018-0202-y
Shi Z, Yin H, Van Nostrand JD, Voordeckers JW, Tu Q, Deng Y, et al. Functional gene array-based ultrasensitive and quantitative detection of microbial populations in complex communities. mSystems. 2019;4(4). https://doi.org/10.1128/mSystems.00296-19.
Lee Y-J, van Nostrand JD, Tu Q, Lu Z, Cheng L, Yuan T, et al. The PathoChip, a functional gene array for assessing pathogenic properties of diverse microbial communities. ISME J. 2013;7(10):1974–84. https://doi.org/10.1038/ismej.2013.88.
Jaksik R, Iwanaszko M, Rzeszowska-Wolny J, Kimmel M. Microarray experiments and factors which affect their reliability. Biol Direct. 2015;10:46. https://doi.org/10.1186/s13062-015-0077-2.
Grover S, Seckar T, Gao L, Bhatia R, Lin X, Zetola N, et al. Characterization of HPV subtypes in invasive cervical cancer in Botswana patients using a pan-pathogen microarray technology. Tumour Virus Res. 2023;15: 200262. https://doi.org/10.1016/j.tvr.2023.200262.
Banerjee S, Peck KN, Feldman MD, Schuster MG, Alwine JC, Robertson ES. Identification of fungal pathogens in a patient with acute myelogenic leukemia using a pathogen detection array technology. Cancer Biol Ther. 2016;17(4):339–45. https://doi.org/10.1080/15384047.2015.1121349.
Baldwin DA, Feldman M, Alwine JC, Robertson ES. Metagenomic assay for identification of microbial pathogens in tumor tissues. mBio. 2014;5(5):e01714–14. https://doi.org/10.1128/mBio.01714-14.
Banerjee S, Tian T, Wei Z, Shih N, Feldman MD, Alwine JC, et al. The ovarian cancer oncobiome. Oncotarget. 2017;8(22):36225–45. https://doi.org/10.18632/oncotarget.6717.
Antal D, Janka EA, Szabó J, Szabó IL, Szegedi A, Gáspár K, et al. Culture-based analyses of skin bacteria in lesional moist, and unaffected dry and sebaceous skin regions of hidradenitis suppurativa patients. J Eur Acad Dermatol Venereol. 2022. https://doi.org/10.1111/jdv.18254.
White MC, Holman DM, Boehm JE, Peipins LA, Grossman M, Henley SJ. Age and cancer risk: a potentially modifiable relationship. Am J Prev Med. 2014;46(3 Suppl 1):S7-15. https://doi.org/10.1016/j.amepre.2013.10.029.
Abdul-Azees PA, Wang H, Chun YP, Pizzini J, Dean DD, Reveles KR, et al. Changes in oral health during aging in a novel non-human primate model. Geroscience. 2023. https://doi.org/10.1007/s11357-023-00939-7.
Tzemah-Shahar R, Turjeman S, Sharon E, Gamliel G, Hochner H, Koren O, et al. Signs of aging in midlife: physical function and sex differences in microbiota. Geroscience. 2023. https://doi.org/10.1007/s11357-023-00905-3.
Shintani T, Shintani H, Sato M, Ashida H. Calorie restriction mimetic drugs could favorably influence gut microbiota leading to lifespan extension. GeroScience. 2023;45(6):3475–90. https://doi.org/10.1007/s11357-023-00851-0.
Chaudhari DS, Jain S, Yata VK, Mishra SP, Kumar A, Fraser A, et al. Unique trans-kingdom microbiome structural and functional signatures predict cognitive decline in older adults. Geroscience. 2023;45(5):2819–34. https://doi.org/10.1007/s11357-023-00799-1.
Correa-Burrows P, Burrows R, Albala C, Court FA, Salech F, Sanhueza G, et al. Multiple events case-control study in a prospective cohort to identify systemic, cellular, and molecular biomarkers of obesity-induced accelerated aging in 30-years-olds: the ObAGE study protocol. BMC Geriatr. 2022;22(1):387. https://doi.org/10.1186/s12877-022-03032-4.
Du Y, Gao Y, Zeng B, Fan X, Yang D, Yang M. Effects of anti-aging interventions on intestinal microbiota. Gut Microbes. 2021;13(1):1994835. https://doi.org/10.1080/19490976.2021.1994835.
Shimizu Y, Nakamura K, Kikuchi M, Ukawa S, Nakamura K, Okada E, et al. Lower human defensin 5 in elderly people compared to middle-aged is associated with differences in the intestinal microbiota composition: the DOSANCO Health Study. Geroscience. 2022;44(2):997–1009. https://doi.org/10.1007/s11357-021-00398-y.
Singh H, Torralba MG, Moncera KJ, DiLello L, Petrini J, Nelson KE, et al. Gastro-intestinal and oral microbiome signatures associated with healthy aging. Geroscience. 2019;41(6):907–21. https://doi.org/10.1007/s11357-019-00098-8.
Buford TW, Carter CS, VanDerPol WJ, Chen D, Lefkowitz EJ, Eipers P, et al. Composition and richness of the serum microbiome differ by age and link to systemic inflammation. Geroscience. 2018;40(3):257–68. https://doi.org/10.1007/s11357-018-0026-y.
Barone M, D’Amico F, Rampelli S, Brigidi P, Turroni S. Age-related diseases, therapies and gut microbiome: A new frontier for healthy aging. Mech Ageing Dev. 2022;206: 111711. https://doi.org/10.1016/j.mad.2022.111711.
Ghosh TS, Shanahan F, O’Toole PW. The gut microbiome as a modulator of healthy ageing. Nat Rev Gastroenterol Hepatol. 2022;19(9):565–84. https://doi.org/10.1038/s41575-022-00605-x.
Thu MS, Pongpirul K. Response: Commentary: Human gut, breast, and oral microbiome in breast cancer: a systematic review and meta-analysis. Front Oncol. 2023;13. https://doi.org/10.3389/fonc.2023.1279862.
Mirzayi C, Renson A, Furlanello C, Sansone S-A, Zohra F, Elsafoury S, et al. Reporting guidelines for human microbiome research: the STORMS checklist. Nat Med. 2021;27(11):1885–92. https://doi.org/10.1038/s41591-021-01552-x.
Bharti R, Grimm DG. Current challenges and best-practice protocols for microbiome analysis. Brief Bioinform. 2019;22(1):178–93. https://doi.org/10.1093/bib/bbz155.
Giaquinto AN, Sung H, Miller KD, Kramer JL, Newman LA, Minihan A, et al. Breast Cancer Statistics, 2022. CA: Cancer J Clin. 2022;72(6):524–41. https://doi.org/10.3322/caac.21754.
TNM classification of malignant tumours. Oxford: John Wiley and Sons; 2017.
Cardoso F, Kyriakides S, Ohno S, Penault-Llorca F, Poortmans P, Rubio IT, et al. Early breast cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2019;30(8):1194–220.
Cardoso F, Paluch-Shimon S, Senkus E, Curigliano G, Aapro MS, André F, et al. 5th ESO-ESMO international consensus guidelines for advanced breast cancer (ABC 5). Ann Oncol. 2020;31(12):1623–49. https://doi.org/10.1016/j.annonc.2020.09.010.
de Azambuja E, Trapani D, Loibl S, Delaloge S, Senkus E, Criscitiello C, et al. ESMO management and treatment adapted recommendations in the COVID-19 era: Breast Cancer. ESMO Open. 2020;5(Suppl 3): e000793. https://doi.org/10.1136/esmoopen-2020-000793.
Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology. 1991;19(5):403–10.
Simin J, Tamimi RM, Engstrand L, Callens S, Brusselaers N. Antibiotic use and the risk of breast cancer: a systematic review and dose-response meta-analysis. Pharmacol Res. 2020;160: 105072. https://doi.org/10.1016/j.phrs.2020.105072.
Wirtz HS, Buist DS, Gralow JR, Barlow WE, Gray S, Chubak J, et al. Frequent antibiotic use and second breast cancer events. Cancer Epidemiol Biomarkers Prev. 2013;22(9):1588–99. https://doi.org/10.1158/1055-9965.EPI-13-0454.
Velicer CM, Heckbert SR, Lampe JW, Potter JD, Robertson CA, Taplin SH. Antibiotic use in relation to the risk of breast cancer. JAMA. 2004;291(7):827–35. https://doi.org/10.1001/jama.291.7.827.
Kiyomiya K, Tomabechi R, Saito N, Watai K, Takada T, Shirasaka Y, et al. Macrolide and ketolide antibiotics inhibit the cytotoxic effect of trastuzumab emtansine in HER2-positive breast cancer cells: implication of a potential drug-ADC interaction in cancer chemotherapy. Mol Pharm. 2023;20(12):6130–9. https://doi.org/10.1021/acs.molpharmaceut.3c00490.
Zhang X, Yu L, Shi J, Li S, Yang S, Gao W, et al. Antibiotics modulate neoadjuvant therapy efficiency in patients with breast cancer: a pilot analysis. Sci Rep. 2021;11(1):14024. https://doi.org/10.1038/s41598-021-93428-w.
Zhang QY, Yan ZB, Meng YM, Hong XY, Shao G, Ma JJ, et al. Antimicrobial peptides: mechanism of action, activity and clinical potential. Mil Med Res. 2021;8(1):48. https://doi.org/10.1186/s40779-021-00343-2.
Thu MS, Ondee T, Nopsopon T, Farzana IAK, Fothergill JL, Hirankarn N, et al. Effect of probiotics in breast cancer: a systematic review and meta-analysis. Biology (Basel). 2023;12(2):280. https://doi.org/10.3390/biology12020280.
Pellegrini M, Ippolito M, Monge T, Violi R, Cappello P, Ferrocino I, et al. Gut microbiota composition after diet and probiotics in overweight breast cancer survivors: a randomized open-label pilot intervention trial. Nutrition. 2020;74: 110749. https://doi.org/10.1016/j.nut.2020.110749.
Mendoza L. Potential effect of probiotics in the treatment of breast cancer. Oncol Rev. 2019;13(2):422. https://doi.org/10.4081/oncol.2019.422.
Duan D, Chen M, Cui W, Liu W, Chen X. Application of probiotics, prebiotics and synbiotics in patients with breast cancer: a systematic review and meta-analysis protocol for randomised controlled trials. BMJ Open. 2022;12(11): e064417. https://doi.org/10.1136/bmjopen-2022-064417.
Nguyen MR, Ma E, Wyatt D, Knight KL, Osipo C. The effect of an exopolysaccharide probiotic molecule from Bacillus subtilis on breast cancer cells. Front Oncol. 2023;13:1292635. https://doi.org/10.3389/fonc.2023.1292635.
Summer M, Sajjad A, Ali S, Hussain T. Exploring the underlying correlation between microbiota, immune system, hormones, and inflammation with breast cancer and the role of probiotics, prebiotics and postbiotics. Arch Microbiol. 2024;206(4):145. https://doi.org/10.1007/s00203-024-03868-x.
Kovacs T, Miko E, Ujlaki G, Sari Z, Bai P. The microbiome as a component of the tumor microenvironment. Adv Exp Med Biol. 2020;1225:137–53.
Funding
Open access funding provided by University of Debrecen. Our work was supported by grants from NKFIH (K142141, FK128387, FK146852, TKP2021-EGA-19, and TKP2021-EGA-20) and the Hungarian Academy of Sciences (POST-COVID2021-33, NKM2022-30). Project no. TKP2021-EGA-19 and TKP2021-EGA-20 were implemented with support provided by the National Research, Development and Innovation Fund of Hungary, financed under the TKP2021-EGA funding scheme. AS is supported by the Bolyai fellowship of the Hungarian Academy of Sciences. This project received funding from the HUN-REN Hungarian Research Network. Supported by the University of Debrecen Program for Scientific Publication.
Author information
Authors and Affiliations
Contributions
EM, AS, ET, AL, MF, ES, GK, and PB wrote and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
Péter Bai is a CEO and shareholder of Holobiont Diagnostics LTD, a developer of cancer diagnostic tests. Other authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Mikó, E., Sipos, A., Tóth, E. et al. Guideline for designing microbiome studies in neoplastic diseases. GeroScience (2024). https://doi.org/10.1007/s11357-024-01255-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11357-024-01255-4