Abstract
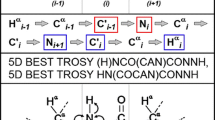
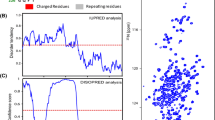
The C-terminal domain of histone H1.0 (C-H1.0) is involved in DNA binding and is a main determinant of the chromatin condensing properties of histone H1.0. Phosphorylation at the (S/T)-P-X-(K/R) motifs affects DNA binding and is crucial for regulation of C-H1.0 function. Since C-H1.0 is an intrinsically disordered domain, solution NMR is an excellent approach to characterize the effect of phosphorylation on the structural and dynamic properties of C-H1.0. However, its very repetitive, low-amino acid-diverse and Pro-rich sequence, together with the low signal dispersion observed at the 1H–15N HSQC spectra of both non- and tri-phosphorylated C-H1.0 preclude the use of standard 1H-detected assignment strategies. We have achieved an essentially complete assignment of the heavy backbone atoms (15N, 13C′ and 13Cα), as well as 1HN and 13Cβ nuclei, of non- and tri-phosphorylated C-H1.0 by applying a novel 13C-detected CON-based strategy. No C-H1.0 region with a clear secondary structure tendency was detected by chemical shift analyses, confirming at residue level that C-H1.0 is disordered in aqueous solution. Phosphorylation only affected the chemical shifts of phosphorylated Thr’s, and their adjacent residues. Heteronuclear {1H}–15N NOEs were also essentially equal in the non- and tri-phosphorylated states. Hence, structural tendencies and dynamic properties of C-H1.0 free in aqueous solution are unmodified by phosphorylation. We propose that the assignment strategy used for C-H1.0, which is based on the acquisition of only a few 3D spectra, is an excellent choice for short-lived intrinsically disordered proteins with repetitive sequences.



Similar content being viewed by others
References
Bermel W et al (2005) Complete assignment of heteronuclear protein resonances by protonless NMR spectroscopy. Angew Chem Int Ed Engl 44:3089–3092
Bermel W et al (2009) H-start for exclusively heteronuclear NMR spectroscopy: the case of intrinsically disordered proteins. J Magn Reson 198:275–281
Bermel W et al (2012) Speeding up sequence specific assignment of IDPs. J Biomol NMR 53:293–301
Bermel W et al (2013) High-dimensionality 13C direct-detected NMR experiments for the automatic assignment of intrinsically disordered proteins. J Biomol NMR 57:353–361
Bienkiewicz EA, Lumb KJ (1999) Random-coil chemical shifts of phosphorylated amino acids. J Biomol NMR 15:203–206
Borgia A et al (2018) Extreme disorder in an ultrahigh-affinity protein complex. Nature 555:61–66
Caterino TL, Fang H, Hayes JJ (2011) Nucleosome linker DNA contacts and induces specific folding of the intrinsically disordered H1 carboxyl-terminal domain. Mol Cell Biol 31:2341–2348
Delaglio F et al (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Dziekanski P, Grudziaz K, Jarvoll P, Kozminski W, Zawadzka-Kazimierczuk A (2015) (13)C-detected NMR experiments for automatic resonance assignment of IDPs and multiple-fixing SMFT processing. J Biomol NMR 62:179–190
Fang H, Clark DJ, Hayes JJ (2012) DNA and nucleosomes direct distinct folding of a linker histone H1 C-terminal domain. Nucleic Acids Res 40:1475–1484
Fang H, Wei S, Lee TH, Hayes JJ (2016) Chromatin structure-dependent conformations of the H1 CTD. Nucleic Acids Res 44:9131–9141
Grzesiek S, Anglister J, Ren H, Bax A (1993) C-13 line narrowing by H-2 decoupling in H-2/C-13/N-15-enriched proteins—application to triple-resonance 4d J-connectivity of sequential amides. J Am Chem Soc 115:4369–4370
Hafsa NE, Arndt D, Wishart DS (2015) CSI 3.0: a web server for identifying secondary and super-secondary structure in proteins using NMR chemical shifts. Nucleic Acids Res 43:W370–W377
Hendzel MJ, Lever MA, Crawford E, Th’ng JP (2004) The C-terminal domain is the primary determinant of histone H1 binding to chromatin in vivo. J Biol Chem 279:20028–20034
Hyberts SG, Frueh DP, Arthanari H, Wagner G (2009) FM reconstruction of non-uniformly sampled protein NMR data at higher dimensions and optimization by distillation. J Biomol NMR 45:283–294
Johnson BA, Blevins RA, NMR, View (1994) A computer program for the visualization and analysis of NMR data. J Biomol NMR 4:603–614
Kalashnikova AA et al (2013) Linker histone H1.0 interacts with an extensive network of proteins found in the nucleolus. Nucleic Acids Res 41:4026–4035
Kalashnikova AA, Rogge RA, Hansen JC (2016) Linker histone H1 and protein-protein interactions. Biochim Biophys Acta 1859:455–461
Kjaergaard M, Poulsen FM (2011) Sequence correction of random coil chemical shifts: correlation between neighbor correction factors and changes in the Ramachandran distribution. J Biomol NMR 50:157–165
Kjaergaard M, Brander S, Poulsen FM (2011) Random coil chemical shift for intrinsically disordered proteins: effects of temperature and pH. J Biomol NMR 49:139–149
Liao R, Mizzen CA (2016) Interphase H1 phosphorylation: regulation and functions in chromatin. Biochim Biophys Acta 1859:476–485
Lopez R et al (2015) Linker histone partial phosphorylation: effects on secondary structure and chromatin condensation. Nucleic Acids Res 43:4463–4476
Mantylahti S, Aitio O, Hellman M, Permi P (2010) HA-detected experiments for the backbone assignment of intrinsically disordered proteins. J Biomol NMR 47:171–181
Mantylahti S, Hellman M, Permi P (2011) Extension of the HA-detection based approach: (HCA)CON(CA)H and (HCA)NCO(CA)H experiments for the main-chain assignment of intrinsically disordered proteins. J Biomol NMR 49:99–109
Marley J, Lu M, Bracken C (2001) A method for efficient isotopic labeling of recombinant proteins. J Biomol NMR 20:71–75
Motackova V et al (2010) Strategy for complete NMR assignment of disordered proteins with highly repetitive sequences based on resolution-enhanced 5D experiments. J Biomol NMR 48:169–177
Novacek J et al (2011) 5D 13C-detected experiments for backbone assignment of unstructured proteins with a very low signal dispersion. J Biomol NMR 50:1–11
Novacek J, Haba NY, Chill JH, Zidek L, Sklenar V (2012) 4D non-uniformly sampled HCBCACON and (1)J(NCalpha)-selective HCBCANCO experiments for the sequential assignment and chemical shift analysis of intrinsically disordered proteins. J Biomol NMR 53:139–148
Novacek J, Janda L, Dopitova R, Zidek L, Sklenar V (2013) Efficient protocol for backbone and side-chain assignments of large, intrinsically disordered proteins: transient secondary structure analysis of 49.2 kDa microtubule associated protein 2c. J Biomol NMR 56:291–301
Panchal SC, Bhavesh NS, Hosur RV (2001) Improved 3D triple resonance experiments, HNN and HN(C)N, for H-N and N-15 sequential correlations in (C-13, N-15) labeled proteins: application to unfolded proteins. J Biomol NMR 20:135–147
Pantoja-Uceda D, Santoro J (2009) Aliasing in reduced dimensionality NMR spectra: (3,2)D HNHA and (4,2)D HN(COCA)NH experiments as examples. J Biomol NMR 45:351–356
Pantoja-Uceda D, Santoro J (2014) New 13C-detected experiments for the assignment of intrinsically disordered proteins. J Biomol NMR 59:43–50
Permi P, Annila A (2004) Coherence transfer in proteins. Prog Nucl Mag Res Sp 44:97–137
Piai A et al (2014) “CON-CON” assignment strategy for highly flexible intrinsically disordered proteins. J Biomol NMR 60:209–218
Raghuram N et al (2013) Pin1 promotes histone H1 dephosphorylation and stabilizes its binding to chromatin. J Cell Biol 203:57–71
Renner C, Schleicher M, Moroder L, Holak TA (2002) Practical aspects of the 2D 15N-[1h]-NOE experiment. J Biomol NMR 23:23–33
Roque A, Orrego M, Ponte I, Suau P (2004) The preferential binding of histone H1 to DNA scaffold-associated regions is determined by its C-terminal domain. Nucleic Acids Res 32:6111–6119
Roque A, Iloro I, Ponte I, Arrondo JL, Suau P (2005) DNA-induced secondary structure of the carboxyl-terminal domain of histone H1. J Biol Chem 280:32141–32147
Roque A, Ponte I, Arrondo JL, Suau P (2008) Phosphorylation of the carboxy-terminal domain of histone H1: effects on secondary structure and DNA condensation. Nucleic Acids Res 36:4719–4726
Roque A, Ponte I, Suau P (2016) Interplay between histone H1 structure and function. Biochim Biophys Acta 1859:444–454
Roque A, Ponte I, Suau P (2017) Post-translational modifications of the intrinsically disordered terminal domains of histone H1: effects on secondary structure and chromatin dynamics. Chromosoma 126:83–91
Sattler M, Schleucher J, Griesinger C (1999) Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog Nucl Mag Res Sp 34:93–158
Schubert M, Labudde D, Oschkinat H, Schmieder P (2002) A software tool for the prediction of Xaa-Pro peptide bond conformations in proteins based on 13C chemical shift statistics. J Biomol NMR 24:149–154
Shen Y, Bax A (2010) Prediction of Xaa-Pro peptide bond conformation from sequence and chemical shifts. J Biomol NMR 46:199–204
Sun ZY, Frueh DP, Selenko P, Hoch JC, Wagner G (2005) Fast assignment of 15N-HSQC peaks using high-resolution 3D HNcocaNH experiments with non-uniform sampling. J Biomol NMR 33:43–50
Talbert PB et al (2012) A unified phylogeny-based nomenclature for histone variants. Epigenetics Chromatin 5:7
Th’ng JP, Sung R, Ye M, Hendzel MJ (2005) H1 family histones in the nucleus. Control of binding and localization by the C-terminal domain. J Biol Chem 280:27809–27814
Vila R, Ponte I, Jimenez MA, Rico M, Suau P (2000) A helix-turn motif in the C-terminal domain of histone H1. Protein Sci 9:627–636
Vila R et al (2001a) DNA-induced alpha-helical structure in the NH2-terminal domain of histone H1. J Biol Chem 276:46429–46435
Vila R, Ponte I, Collado M, Arrondo JL, Suau P (2001b) Induction of secondary structure in a COOH-terminal peptide of histone H1 by interaction with the DNA: an infrared spectroscopy study. J Biol Chem 276:30898–30903
Vila R, Ponte I, Jimenez MA, Rico M, Suau P (2002) An inducible helix-Gly-Gly-helix motif in the N-terminal domain of histone H1e: a CD and NMR study. Protein Sci 11:214–220
Widlak P et al (2005) The histone H1 C-terminal domain binds to the apoptotic nuclease, DNA fragmentation factor (DFF40/CAD) and stimulates DNA cleavage. Biochemistry 44:7871–7878
Williamson MP (2013) Using chemical shift perturbation to characterise ligand binding. Prog Nucl Magn Reson Spectrosc 73:1–16
Zakharova NI et al (2011) [Prothymosin alpha interacts with C-terminal domain of histone H1 and dissociates p53-histone H1 complex]. Mol Biol 45:679–688
Zerko S et al (2016) Five and four dimensional experiments for robust backbone resonance assignment of large intrinsically disordered proteins: application to Tau3x protein. J Biomol NMR 65:193–203
Acknowledgements
This work has been supported projects CTQ2014-52633-P and CTQ2017-84371-P from Spanish MINECO and MCIU, respectively (both co-financed by European FEDER funds). BB was a recipient of pre-doctoral FPI scholarship BES-2015-073383 from Spanish MINECO. The NMR experiments were performed in the “Manuel Rico” NMR laboratory (LMR) of the Spanish National Research Council (CSIC).
Author information
Authors and Affiliations
Corresponding author
Additional information
Dedicated to Prof. Jorge Santoro on the occasion of his retirement.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chaves-Arquero, B., Pantoja-Uceda, D., Roque, A. et al. A CON-based NMR assignment strategy for pro-rich intrinsically disordered proteins with low signal dispersion: the C-terminal domain of histone H1.0 as a case study. J Biomol NMR 72, 139–148 (2018). https://doi.org/10.1007/s10858-018-0213-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-018-0213-2