Abstract
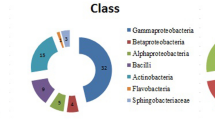
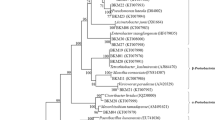
Molecular approaches [PCR-denaturing gradient gel electrophoresis (DGGE)] were used to determine whether three different vetiver (Chrysopogon zizanioides) genotypes, commercially used in Brazil and considered economically important over the world, select specific bacterial populations to coexist in their rhizospheres. DGGE profiles revealed that the predominant rhizospheric bacterial community hardly varies regarding the vetiver genotype. Moreover, using traditional cultivation methods, bacterial strains were isolated from the different rhizospheres. Colonies presenting different morphologies (83) were selected for determining their potential for plant growth promotion. More than half of the strains tested (57.8%) were amplified by PCR using nifH-based primers, specific for the enzyme nitrogenase reductase. The production of siderophores was observed in 88% of the strains, while the production of antimicrobial substances was detected in only 14.5% of the isolates when Micrococcus sp. was used as the indicator strain. Production of indole-3-acetic acid and the solubilization of phosphate were observed in 55.4% and 59% of the isolates, respectively. In total, 44 strains (53%) presented at least three characteristics of plant growth promotion and were submitted to amplified ribosomal DNA restriction analysis. Twenty-four genetic groups were formed at 100% similarity and one representative of each group was selected for their identification by partial 16S rRNA gene sequencing. They were affiliated with the genera Acinetobacter, Comamonas, Chryseobacterium, Klebsiella, Enterobacter, Pantoea, Dyella, Burkholderia, or Pseudomonas. These strains can be considered of great importance as possible biofertilizers in vetiver.
Similar content being viewed by others
References
Adams, R.P., M. Habte, S. Park, and M.R. Dafforn. 2004. Preliminary comparison of vetiver root essential oils from cleansed (bacteria- and fungus-free) versus non-cleansed (normal) vetiver plants. Biochem. Syst. Ecol. 32, 1137–1144.
Adams, R.P., R.N. Pandey, M.R. Dafforn, and S.A. James. 2003. Vetiver DNA-fingerprinted cultivars: effects of environment on growth, oil yields and composition. J. Ess. Oil Res. 15, 363–371.
Akhila, A. and M. Rani. 2002. Chemical constituents and essential oil biogenesis in Vetiveria zizanioides, p. 73–109. In M. Maffei (ed.), The Genus Vetiveria. Taylor & Francis, New York, N.Y., USA.
Albuquerque, J.P., F.F. Mota, I. von der Weid, and L. Seldin. 2006. Diversity of Paenibacillus durus strains isolated from soil and different plant rhizospheres evaluated by ARDRA and gyrB-RFLP analysis. Eur. J. Soil Biol. 42, 200–207.
Bent, E., S. Tuzun, C.P. Chanway, and S. Eneback. 2001. Alterations in plant growth and in root hormone levels of lodgepole pines inoculated with rhizobacteria. Can. J. Microbiol. 47, 793–800.
Brimecombe, M.J., F.A. De Leij, and J.M. Lynch. 2001. The effect of root exudates on rhizosphere microbial populations, p. 95–140. In R. Pinton, Z. Varanini, and P. Nannipieri (eds.), The Rhizosphere — biochemistry and organic substances at soil-plant interface. Marcel Dekker, New York, N.Y., USA.
Champagnat, P., G. Figueredo, J.C. Chalchat, A.P. Carnat, and J.M. Bessiere. 2006. A study on the composition of commercial Vetiveria zizanioides oils from different geographical origins. J. Ess. Oil Res. 18, 416–422.
Chomchalow, N. 2001. The utilization of vetiver as medicinal and aromatic plants with special reference to Thailand. PRVN Tech. Bull. No. 200/1, ORDPB, Bangkok.
Chomchalow, N. and K. Chapman. 2003. Other uses and utilization of vetiver, p. 474–485. Proceedings of the Third International Vetiver Conference. China.
da Silva, K.R.A., J.F. Salles, L. Seldin, and J.D. van Elsas. 2003. Assessment of the diversity of Paenibacillus spp. in the rhizosphere of different maize cultivars in two soils by Paenibacillus-specific PCR-DGGE and sequence analysis. J. Microbiol. Methods 54, 213–231.
De Los Santos, P.E., R. Bustillos-Cristales, and J. Caballero-Mellado. 2001. Burkholderia, a genus rich in plant-associated nitrogen fixers with wide environmental and geographic distribution. Appl. Environ. Microbiol. 67, 2790–2798.
Del Giudice, L., D.R. Massardo, P. Pontieri, C.M. Bertea, D. Mombello, E. Carata, S.M. Tredici, A. Talà, M. Mucciarelli, V.I. Groudeva, M. De Stefano, G. Vigliotta, M.E. Maffei, and P. Alifano. 2008. The microbial community of Vetiver root and its involvement into essential oil biogenesis. Environ. Microbiol. 10, 2824–2841.
Défago, G., B.K. Duffy, and C. Keel. 1994. Risk assessment for the release of plant growth-promoting rhizobacteria, p. 254. In Proceedings of the Third International Workshop on Plant Growth-Promoting Rhizobacteria. CSIRO, Adelaide, Australia.
Dong, Z., X. Xie, X. Lu, H. Guo, and X. Sun. 2003. Study on the genetic diversity of vetiver grass (Vetiveria zizanioides), p. 524–531. Proceedings of the Third International Vetiver Conference, China.
Duineveld, B.M., G.A. Kowalchuk, A. Keijzer, J.D. van Elsas, and J.A. van Veen. 2001. Analysis of bacterial communities in the rhizosphere of chrysanthemum via denaturing gradient gel electrophoresis of PCR-amplified 16S rRNA as well as DNA fragments coding for 16S rRNA. Appl. Environ. Microbiol. 67, 172–178.
Garbeva, P., J.A. van Veen, and J.D. van Elsas. 2004. Microbial diversity in soil-selection of microbial populations by plant and soil type and implications for disease suppression. Annu. Rev. Phytopathol. 42, 243–270.
Glick, B.R. 1995. The enhancement of plant growth by free-living bacteria. Can. J. Microbiol. 41, 109–117.
Gray, E.J. and D.L. Smith. 2005. Intracellular and extracellular PGPR: Commonalities and distinctions in the plant-bacterium signaling processes. Soil Biol. Biochem. 37, 395–412.
Kang, S., G. Joo, M. Hamayun, C. Na, D. Shin, H.Y. Kim, J. Hong, and I. Lee. 2009. Gibberellin production and phosphate solubilization by newly isolated strain of Acinetobacter calcoaceticus and its effect on plant growth. Biotechnol. Lett. 31, 277–281.
Kloepper, J.W. and M.N. Schroth. 1978. Plant growth promoting rhizobacteria on radishes, p. 879–882. In Proceedings of the Fourth International Conference on Plant Pathogenic Bacteria, Angers, France.
Kumar, S., K. Tamura, and M. Nei. 2004. MEGA3: Integrated software for molecular evolutionary analysis and sequence alignment. Brief Bioinform. 5, 150–163.
Lavania, U.C. 2003. Primary and secondary centers of origin of vetiver and its dispersion, p. 424–426. Proceedings of the Third International Vetiver Conference, China.
Massol-Deya, A.A., D.A. Odelson, R.F. Hickey, and J.M. Tiedje. 1995. Bacterial community fingerprinting of amplified 16S and 16–23S ribosomal DNA gene sequences and restriction endonuclease analysis (ARDRA). In A.D.L. Akkermans, J.D. van Elsas, and F.J. de Bruijn (eds.), Molecular Microbial Ecology, Kluwer Academic Publishers, Dordrecht, The Netherlands.
Mota, F.F., E.A. Gomes, I.E. Marriel, E. Paiva, and L. Seldin. 2008. Effect of liming on the structure of bacterial and fungal communities in bulk soil and rhizospheres of aluminum-tolerant and aluminum-sensitive maize (Zea mays L.) lines cultivated in Cerrado soil. J. Microbiol. Biotechnol. 18, 805–814.
Mota, F.F., A. Nóbrega, I.E. Marriel, E. Paiva, and L. Seldin. 2002. Genetic diversity of Paenibacillus polymyxa populations isolated from the rhizosphere of four cultivars of maize (Zea mays) planted in Cerrado soil. Appl. Soil Ecol. 20, 119–132.
Nübel, U., B. Engelen, A. Felske, J. Snaidr, A. Wieshuber, R.I. Amann, W. Ludwig, and H. Backhaus. 1996. Sequence heterogeneities of genes encoding 16S rDNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J. Bacteriol. 178, 5636–5643.
Poly, F., L.J. Monrozier, and R. Bally. 2001. Impovement in the RFLP procedure for studying the diversity of nifH genes in communities of nitrogen fixers in soil. Res. Microbiol. 152, 95–103.
Rosado, A.S., F.S. Azevedo, D.W.G. Cruz, and L. Seldin. 1998. Phenotypic and genetic diversity of Paenibacillus azotofixans strains isolated from rhizoplane or rhizosphere of different grasses. J. Appl. Microbiol. 84, 216–226.
Rosado, A.S. and L. Seldin. 1993. Production of a potentially novel anti-microbial substance by Bacillus polymyxa. World J. Microbiol. Biotechnol. 9, 521–528.
Sambrook, J., E.F. Fritsch, and T. Maniatis. 1989. Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press, New York, N.Y., USA.
Schwyn, B. and J.B. Neilands. 1987. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 160, 47–56.
Seldin, L. and D. Dubnau. 1985. DNA homology among Bacillus polymyxa, Bacillus azotofixans and other nitrogen fixing Bacillus strains. Int. J. Syst. Bacteriol. 35, 151–154.
Seldin, L., J.D. van Elsas, and E.G.C. Penido. 1983. Bacillus nitrogen fixers from Brazilian soils. Plant Soil 70, 243–255.
Seldin, L., J.D. van Elsas, and E.G.C. Penido. 1984. Bacillus azotofixans sp. nov., a nitrogen-fixing species from Brazilian soils and grass roots. Int. J. Syst. Bacteriol. 34, 451–456.
Sjöling, S. and D.A. Cowan. 2003. High 16S rDNA bacterial diversity in glacial meltwater lake sediment, Bratina Island, Antarctica. Extremophiles 7, 275–282.
Smalla, K., G. Wieland, A. Buchner, A. Zock, J. Parzy, S. Kaiser, N. Roskot, H. Heuer, and G. Berg. 2001. Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: Plant-dependent enrichment and seasonal shifts revealed. Appl. Environ. Microbiol. 67, 4742–4751.
Sulbarán, M., E. Pérez, M.M. Ball, A. Bahsas, and L.A. Yarzábal. 2009. Characterization of the mineral phosphate-solubilizing activity of Patoea aglomerans MMB051 isolated from an ironrich soil in southeastern Venezuela (Bolívar State). Curr. Microbiol. 58, 378–383.
Tang, Y.W. and J. Bonner. 1947. The enzymatic inactivation of indole-acetic acid. I. Some characteristics of the enzyme contained in pea seedlings. Arch. Biochem. 13, 11–25.
Thompson, J.D., T.J. Gibson, F. Plewniak, F. Jeanmougin, and D.G. Higgins. 1997. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882.
Truong, P. 2003. Vetiver grass system: Potential applications for soil and water conservation in northern California, p. 562–571. Proceedings of the Third International Vetiver Conference, China.
Tupinambá, G., C.S. Alviano, A.J.R. da Silva, T.C.B.S. Souto-Padron, L. Seldin, and D.S. Alviano. 2008. Antimicrobial activity of Paenibacillus polymyxa SCE2 against mycotoxin-producing fungi. J. Appl. Microbiol. 105, 1044–1053.
von der Weid, I., V. Artursson, L. Seldin, and J.K. Jansson. 2005. Antifungal and root surface colonization properties of GFP-tagged Paenibacillus brasilensis PB177. World J. Microbiol. Biotechnol. 21, 1591–1597.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Monteiro, J.M., Vollú, R.E., Coelho, M.R.R. et al. Comparison of the bacterial community and characterization of plant growth-promoting rhizobacteria from different genotypes of Chrysopogon zizanioides (L.) Roberty (Vetiver) rhizospheres. J Microbiol. 47, 363–370 (2009). https://doi.org/10.1007/s12275-009-0048-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-009-0048-3