Abstract
In vitro regeneration of date palm (Phoenix dactylifera L.) plants through somatic embryogenesis leads to the generation of somaclonal variants. The transposition of retrotransposons and DNA transposons in a host genome can be activated by tissue culture stresses, thus these elements can be both the cause of and useful markers for genomic variation. In this study, Hordeum-specific and Phoenix-specific inter-retrotransposon amplified polymorhism (IRAP) markers together with Phoenix-specific miniature inverted-repeat transposable element (MITE) markers were used to investigate the activation of DNA transposons and retrotransposons by somaclonal variation. Ty3/gypsy-like LTR retotransposons and MITE DNA transposon sequences were extracted from P. dactylifera cv. Khalas. Phoenix-specific primers were designed from the long terminal repeat (LTR) region of Ty3/gypsy-like LTR retotransposons and MITE DNA transposons. Both IRAP and MITE markers were able to detect somaclonal variants among date palms grown in open field trials. DNA marker analyses support that the transposability of both LTR retrotransposon and MITEs in the date palm genome is activated during the tissue culture process, leading to new insertion events in somaclonal variants. This study demonstrated a simple PCR-based method for the screening of somaclonal variants in tissue cultured date palm plants and establishes the application of transposible element based DNA markers for clonal identification.
Key message
This study demonstrated a simple PCR-based method for the screening of somaclonal variants in tissue cultured date palm plants and establishes the application of transposable element-based DNA markers for clonal identification.




Similar content being viewed by others
References
Abd-Alla MM (2010) Genetic stability on Phoenix dactylifera var. Karama produced in vitro. N Y Sci J 3:70–75
Abul-Soad AA, Mahdi SM (2010) Commercial production of tissue culture date palm (Phoenix dactylifera L.) by inflorescence technique. J Genet Eng Biotechnol 8:39–44
Aitchitt M, Ainsworth CC, Thangavelu M (1993) A rapid and efficient method for the extraction of total DNA from mature leaves of the data palm (Phoenix dactylifera L.). Plant Mol Biol Rep 11:317–319
Al-Dous EK, George B, Al-Mahmoud ME, Al-Jaber MY, Wang H, Salameh YM, Al-Azwani EK, Chaluvadi S, Pontaroli AC, DeBarry J (2011) De novo genome sequencing and comparative genomics of date palm (Phoenix dactylifera). Nat Biotechnol 29:521
Al-Mssallem IS, Hu S, Zhang X, Lin Q, Liu W, Tan J, Yu X, Liu J, Pan L, Zhang T (2013) Genome sequence of the date palm Phoenix dactylifera L. Nat Commun 4:2274
Al-Wasel A (2001) Field performance of somaclonal variants of tissue culture-derived date palm (Phoenix dactylifera L.). Plant Tissue Cult Biotechnol 11:97–105
Alkhateeb A, Ali-Dinar H (2002) Date palm in Kingdom of Saudi Arabia: cultivation production and processing. Translation Authorship and Publishing Center, King Faisal University, Kingdom of Saudi Arabia 188:281–292
Alkhateeb AA (2008) A review the problems facing the use of tissue culture technique in date palm (Phoenix dactylifera L.). Sci J King Faisal Univ Basic Appl Sci 9:85–104
Alves E, Ballesteros I, Linacero R, Vazquez A (2005) RYS1, a foldback transposon, is activated by tissue culture and shows preferential insertion points into the rye genome. Theor Appl Genet 111:431–436
Alzohairy A, Yousef M, Edris S, Kerti B, Gyulai G, Bahieldin A (2012) Detection of LTR retrotransposons reactivation induced by in vitro environmental stresses in barley (Hordeum vulgare) via RT-qPCR. Life Sci J 9:5019–5026
Bairu MW, Fennell CW, van Staden J (2006) The effect of plant growth regulators on somaclonal variation in Cavendish banana (Musa AAA cv.‘Zelig’). Sci Hortic 108:347–351
Bashah M (1996) Date variety in the Kingdom of Saudi Arabia King Abdulaziz Univ Guidance booklet palms and dates. King Abdulaziz Univ Press, Riyadh, pp 1225–1319
Branco CJ, Vieira EA, Malone G, Kopp MM, Malone E, Bernardes A, Mistura CC, Carvalho FI, Oliveira CA (2007) IRAP and REMAP assessments of genetic similarity in rice. J Appl Genet 48:107–113
Campbell BC, LeMare S, Piperidis G, Godwin ID (2011) IRAP, a retrotransposon-based marker system for the detection of somaclonal variation in barley. Mol Breed 27:193–206
Chen J, Hu Q, Zhang Y, Lu C, Kuang H (2014) P-MITE: a database for plant miniature inverted-repeat transposable elements. Nucleic Acids Res 42:D1176
Chiu L-W, Zhou X, Burke S, Wu X, Prior RL, Li L (2010) The purple cauliflower arises from activation of a MYB transcription factor. Plant Physiol 154:1470–1480
Cohen Y, Korchinsky R, Tripler E (2004) Flower abnormalities cause abnormal fruit setting in tissue culture-propagated date palm (Phoenix dactylifera L.). J Hortic Sci Biotechnol 79:1007–1013
Damasco OP, Smith MK, Adkins SW, Hetherington SE, Godwin ID (1997) Identification and characterisation of dwarf off-types from micropropagated Cavendish bananas. In: II International Symposium on Banana: I International Symposium on Banana in the Subtropics 490:79–84
Dellaporta S, Chomet P, Mottinger J, Wood J, Yu S-M, Hicks J (1984) Endogenous transposable elements associated with virus infection in maize. In: Cold Spring Harbor symposia on quantitative biology. Cold Spring Harbor Laboratory Press, pp 321–328
Diaz S, Pire C, Ferrer J, Bonete MJ (2003) Identification of Phoenix dactylifera L. varieties based on amplified fragment length polymorphism (AFLP) markers. Cell Mol Biol Lett 8:891–900
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
El-Assar A, Krueger R, Devanand P, Chao C (2005) Genetic analysis of Egyptian date (Phoenix dactylifera L.) accessions using AFLP markers. Genet Resour Crop Evol 52:601–607
Eshraghi P, Zarghami R, Ofoghi H (2005) Genetic stability of micropropagated plantlets in date palm. J Sci Islam Repub Iran 16:311–315
Grafi G, Avivi Y (2004) Stem cells: a lesson from dedifferentiation. Trends Biotechnol 22:388–389
Grandbastien M-A (1998) Activation of plant retrotransposons under stress conditions. Trends Plant Sci 3:181–187
Grandbastien M-A (2015) LTR retrotransposons, handy hitchhikers of plant regulation and stress response. Biochem Biophys Acta 1849:403–416
Gurevich V, Lavi U, Cohen Y (2005) Genetic variation in date palms propagated from offshoots and tissue culture. J Am Soc Hortic Sci 130:46–53
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In: Nucleic acids symposium series. vol 41. [London]: Information Retrieval Ltd., c1979-c2000., pp 95–98
Hirochika H (1993) Activation of tobacco retrotransposons during tissue culture. EMBO J 12:2521
Hirochika H, Sugimoto K, Otsuki Y, Tsugawa H, Kanda M (1996) Retrotransposons of rice involved in mutations induced by tissue culture. Proc Natl Acad Sci USA 93:7783–7788
Hosaka A, Kakutani T (2018) Transposable elements, genome evolution and transgenerational epigenetic variation. Curr Opin Genet Dev 49:43–48
James MG, Stadler J (1989) Molecular characterization of Mutator systems in maize embryogenic callus cultures indicates Mu element activity in vitro. Theor Appl Genet 77:383–393
Jatoi MA, Abul-Soad AA, Markhand GS, Solangi N (2015) Establishment of an efficient protocol for micropropagation of some Pakistani cultivars of date palm (Phoenix dactylifera L.) using novel inflorescence explants. Pak J Bot 47:1921–1927
Jiang N, Feschotte C, Zhang X, Wessler SR (2004) Using rice to understand the origin and amplification of miniature inverted repeat transposable elements (MITEs). Curr Opin Plant Biol 7:115–119
Kalendar R, Grob T, Regina M, Suoniemi A, Schulman A (1999) IRAP and REMAP: two new retrotransposon-based DNA fingerprinting techniques. TAG Theor Appl Genet 98:704–711
Kalendar R (2011) The use of retrotransposon-based molecular markers to analyze genetic diversity. Field Veg Crops Res/Ratarstvo i povrtarstvo 48:261–274
Kalendar R, Khassenov B, Ramankulov Y, Samuilova O, Ivanov KI (2017) FastPCR: an in silico tool for fast primer and probe design and advanced sequence analysis. Genomics 109:312–319
Karim R, Nuruzzaman M, Khalid N, Harikrishna JA (2016) Importance of DNA and histone methylation in in vitro plant propagation for crop improvement: A review. Ann Appl Biol 169(1):116
Kashkush K, Feldman M, Levy AA (2002) Gene loss, silencing and activation in a newly synthesized wheat allotetraploid. Genetics 160:1651–1659
Kikuchi K, Terauchi K, Wada M, Hirano H-Y (2003) The plant MITE mPing is mobilized in anther culture. Nature 421:167–170
Kılınç FM, Süzerer V, Çiftçi Y, Onay A, Yıldırım H, Uncuoğlu AA, Tilkat E, Koç I, Akdemir ÖF, Metin ÖK (2015) Clonal micropropagation of Pistacia lentiscus L. and assessment of genetic stability using IRAP markers. Plant Growth Regul 75:75–88
Kumar N, Modi AR, Singh AS, Gajera BB, Patel AR, Patel MP, Subhash N (2010) Assessment of genetic fidelity of micropropagated date palm (Phoenix dactylifera L.) plants by RAPD and ISSR markers assay. Physiol Mol Biol Plants 16:207–213
Kumar N, Singh AS, Modi AR, Patel AR, Gajera BB, Subhash N (2010) Genetic stability studies in micropropagated date palm (Phoenix dactylifera L.) Plants using microsatellite marker. J For Environ Sci 26:31–36
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lisch D, Bennetzen JL (2011) Transposable element origins of epigenetic gene regulation. Curr Opin Plant Biol 14:156–161
McClintock B (1951) Chromosome organization and genic expression. In: Cold Spring Harbor symposia on quantitative biology. Cold Spring Harbor Laboratory Press pp 13–47
McClintock B (1978) Mechanisms that rapidly reorganize the genome. In: Stadler genetics symposia, University of Missouri pp 25–48
McCubbin M, Zaid A, Van Stade J (2004) A southern African survey conducted for off-types on date palms produced using somatic embryogenesis. Emir J Food Agric 8–14
Mirani A, Teo CH, Abul-Soad A, Markhand G, Jatt T, Mirbahar A, Solangi N (2019) Phenotypic reversion of somaclonal variants derived from inflorescence of date palm (Phoenix dactylifera L.) in the open field trials. Sarhad J Agric 35:719–726
Muhammad AJ, Othman FY (2005) Characterization of fusarium wilt-resistant and fusarium wilt-susceptible somaclones of banana cultivar rastali (Musa AAB) by random amplified polymorphic DNA and retrotransposon markers. Plant Mol Biol Rep 23:241–249
Nei M, Li W-H (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Neuffer M (1966) Stability of the suppressor element in two mutator systems at the A1 locus in maize. Genetics 53:541–549
Ong-Abdullah M, Ordway JM, Jiang N, Ooi S-E, Kok S-Y, Sarpan N, Azimi N, Hashim AT, Ishak Z, Rosli SK (2015) Loss of Karma transposon methylation underlies the mantled somaclonal variant of oil palm. Nature 525:533
Othmani A, Rhouma S, Bayoudh C, Mzid R, Drira N, Trifi M (2010) Regeneration and analysis of genetic stability of plantlets as revealed by RAPD and AFLP markers in date palm (Phoenix dactylifera L.) cv. Deglet Nour. Int Res J Plant Sci 1:48–55
Pearce S, Kumar A, Flavell A (1996) Activation of the Ty1-copia group retrotransposons of potato (Solatium tuberosum) during protoplast isolation. Plant Cell Rep 15:949–953
Peschke V, Phillips R, Gengenbach B (1991) Genetic and molecular analysis of tissue-culture-derived Ac elements. Theor Appl Genet 82:121–129
Peschke VM, Phillips RL, Gengenbach BG (1987) Discovery of transposable element activity among progeny of tissue culture-derived maize plants. Science 238:804–808
Peterson PA (1960) The pale green mutable system in maize. Genetics 45:115–133
Racchi M, Bove A, Turchi A, Bashir G, Battaglia M, Camussi A (2014) Genetic characterization of Libyan date palm resources by microsatellite markers. 3 Biotech 4:21–32
Rhee Y, Lin H, Buell R, Childs K, Kaeppler S (2009) Ac2 allele of maize identified in regenerant-derived progeny from tissue culture results from insertion of a novel transposon. Maydica 54:429–437
Rodrigues P, Tulmann Neto A, Cassieri Neto P, Mendes B (1997) Influence of the number of subcultures on somaclonal variation in micropropagated nanicao (Musa spp., AAA Group). In: II International Symposium on Banana: I International Symposium on Banana in the Subtropics 490 pp 469–474
Rohlf F (1998) NTSYS-pc. Numerical taxonomy and multivariate analysis: version 2.02. Exeter Software Setauket, New York
Saker M, Adawy S, Mohamed A, El-Itriby H (2006) Monitoring of cultivar identity in tissue culture-derived date palms using RAPD and AFLP analysis. Biol Plant 50:198–204
Saker M, Bekheet S, Taha H, Fahmy A, Moursy H (2000) Detection of somaclonal variations in tissue culture-derived date palm plants using isoenzyme analysis and RAPD fingerprints. Biol Plant 43:347–351
Smulders M, De Klerk G (2011) Epigenetics in plant tissue culture. Plant Growth Regul 63:137–146
Sneath PH, Sokal RR (1973) Numerical taxonomy. The principles and practice of numerical classification. W. H. Freeman and Company, New York
Suoniemi A, Narvanto A, Schulman AH (1996) The BARE-1 retrotransposon is transcribed in barley from an LTR promoter active in transient assays. Plant Mol Biol 31:295–306
Taha RA, Hassan MM, Ibrahim EA, Abou Baker NH, Shaaban EA (2016) Carbon nanotubes impact on date palm in vitro cultures. Plant Cell Tissue Organ Cult 127:525–534
Teo CH, Tan S, Othman Y, Schwarzacher T (2002) The Cloning of Ty 1- copia-like Retrotransposons from 10 Varieties of Banana (Musa Sp.). J Biochem Mol Biol Biophys 6:193–201
Teo CH, Tan SH, Ho CL, Faridah QZ, Othman YR, Heslop-Harrison JS, Kalendar R, Schulman AH (2005) Genome constitution and classification using retrotransposon-based markers in the orphan crop banana. J Plant Biol 48:96–105
Zaid A, Al-Kaabi H (2003) Plant-off types in tissue culture-derived date palm (Phoenix dactylifera L.). Emir J Food Agric 15:17–35
Acknowledgements
This work was supported by the Higher Education Commission (HEC), Islamabad, Pakistan and CEBAR grant RU006-2017, from the University of Malaya, Malaysia.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Maria Margarida Oliveira.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1
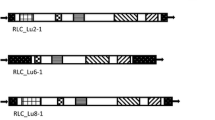
IRAP primer design. A) IRAP primers (PdGy01F and PdGy01R) designed from the 5’-end of PdGy01 Ty3/gypsy-like LTR retrotransposon; B) IRAP primers (PdGyChromo01F and PdGyChromo01R) designed from 3’-end of PdGyChromo01 Ty3/gypsy-like LTR retrotransposon (JPEG 747.5 kb)
Figure S2
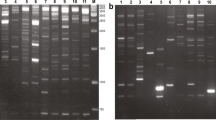
IRAP profile for date palm plants of P. dactylifera cv. Gulistan showing monomorphic bands among mother plant (M) and tissue culture regenerants (1-30; 46-47). L: GeneRuler 1 kb DNA Ladder; C: negative control (JPEG 804.9 kb)
Figure S3
MITE profile for date palm of P. dactylifera cv. Gulistan showing monomorphic bands among mother plant (M) and tissue culture regenerants (1-30; 46-47). L: GeneRuler 1 kb DNA Ladder; C: negative control (JPEG 615.3 kb)
Figure S4
IRAP profile for date palm of P. dactylifera cv. Kashuwari showing monomorphic bands among mother plant (M) and tissue culture regenerants (1-15; 31-50). L: GeneRuler 1 kb DNA Ladder; C: negative control (JPEG 770.2 kb)
Figure S5
MITE profile for date palm of P. dactylifera cv. Kashuwari showing monomorphic bands among mother plant (M) and tissue culture regenerants (1-15; 31-50). L: GeneRuler 1 kb DNA Ladder; C: negative control (JPEG 655.2 kb)
Figure S6
Dendrogram of P. dactylifera cv. Gulistan showing control (mother plant), normal tissue cultured (TC) plants and somaclonal variant tissue cultured (TC) plants. (JPEG 124.7 kb)
Figure S7
Dendrogram of P. dactylifera cv. Kashuwari showing control (mother plant), normal tissue cultured (TC) plants and somaclonal variant tissue cultured (TC) plants. (JPEG 120.4 kb)
Rights and permissions
About this article
Cite this article
Mirani, A.A., Teo, C.H., Markhand, G.S. et al. Detection of somaclonal variations in tissue cultured date palm (Phoenix dactylifera L.) using transposable element-based markers. Plant Cell Tiss Organ Cult 141, 119–130 (2020). https://doi.org/10.1007/s11240-020-01772-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-020-01772-y