Abstract
Glioblastoma Multiforme (GBM) is the primary brain tumor and accounts for 200,000 deaths each year worldwide. The standard therapy includes surgical resection followed by temozolomide (TMZ)-based chemotherapy and radiotherapy. The survival period of GBM patients is only 12–15 months. Therefore, novel treatment modalities for GBM treatment are urgently needed. Mounting evidence reveals that non-coding RNAs (ncRNAs) were involved in regulating gene expression, the pathophysiology of GBM, and enhancing therapeutic outcomes. The combinatory use of ncRNAs, chemotherapeutic drugs, and tumor suppressor gene expression induction might provide an innovative, alternative therapeutic approach for managing GBM. Studies have highlighted the role of Long non-coding RNAs (lncRNAs) and microRNAs (miRNAs) in prognosis and diagnosis. Dysregulation of ncRNAs is observed in virtually all tumor types, including GBMs. Studies have also indicated the blood–brain barrier (BBB) as a crucial factor that hinders chemotherapy. Although several nanoparticle-mediated drug deliveries were degrading effectively against GBM in vitro conditions. However, the potential to cross the BBB and optimum delivery of oligonucleotide RNA into GBM cells in the brain is currently under intense clinical trials. Despite several advances in molecular pathogenesis, GBM remains resistant to chemo and radiotherapy. Targeted therapies have less clinical benefit due to high genetic heterogeneity and activation of alternative pathways. Thus, identifying GBM-specific prognostic pathways, essential genes, and genomic aberrations provide several potential benefits as subtypes of GBM. Also, these approaches will provide insights into new strategies to overcome the heterogenous nature of GBM, which will eventually lead to successful therapeutic interventions toward precision medicine and precision oncology.
Similar content being viewed by others
Introduction
Brain cancer accounts for 1.2% of the various cancers, with 17,000 new cases yearly, especially in the USA. The Central Nervous System consists of the brain and spinal cord and aids in giving motor signals by processing sensory information. They control processes such as thoughts, hormone secretion, movement, heartbeats, and respiration (Controlled by the Brain stem). The various cancers found in the brain in children are Astrocytoma, Ependymoma, Diffuse Intrinsic Pontine Glioma (DIPG), Germ cell Tumors, and Medulloblastoma (Packer et al. 2008; Warren 2012). In adults, along with the above types, additional benign tumors like Oligodendroglial tumors in the temporal or frontal lobe, mixed gliomas, Pineal Parenchymal tumors in the pineal region, Craniopharyngioma, a rare type found at the anterior end of the brain just above the pituitary gland, and Meningeal tumors in the surrounding layers of the brain and spinal cord called meninges have been detected. (Farwell et al. 1977; Jeuken et al. 2004; Lutterbach et al. 2002). In GBM, Grades I-II belong to lower-grade gliomas (LGG) [angiocentric glioma and diffuse astrocytoma], and high-grade glioma (grade III-IV) include mesenchymal astrocytoma and GBM (Louis et al. 2007, 2016). The classification is based on isocitrate dehydrogenase (IDH1), alpha-thalassemia/mental retardation, X-linked (ATRX), tumor suppressor protein (TP53), and 1p/19 (Yang et al. 2016; Han et al. 2018). Most primary GBM cancers are wild-type IDH, while secondary GBM develops from lower-grade glioma and carries mutations in IDH. (Ohgaki and Kleihues 2013).
Intense research and investigations on palliative care remain essential during various stages of GBM. Emerging studies have identified the potential use of TMZ in O6-methylguanine-DNA-methyltransferase (MGMT) methylated genetic background and anaplastic glioma. Emerging studies have revealed BBB and tumor microenvironment (TME) as significant challenges for disease therapy (Tan et al. 2020). Bioinformatics analysis identified overexpression of a few genes such as Cell Adhesion Associated, Oncogene Regulated (BOC), Elongation of very long chain fatty acids protein 6 (ELOVL6), Vascular endothelial growth factor C (VEGFC), etc. The molecular aberrations in GBM include mutations in the p53 gene, Retinoblastoma, PI3K-Akt signaling, EGFR signaling, Neurogenic locus STAT homolog protein 1 (NOTCH 1), and NOTCH 2 signaling, etc. (Brennan et al. 2013). The essential gene that dictates a patient’s survival and responsiveness towards TMZ is MGMT. Methylation of MGMT promoter results in an increased survival rate when compared with the hypomethylated state. MGMT repairs the N7 and O6 positions of guanine, alkylated by TMZ. Kato et al. 2010 have reported that small interfering RNA (siRNA) targeting MGMT can enhance the ability of TMZ-induced cytotoxic effects.
Interestingly, O6-Benzylguanine (O6-BG) and O6-(4-Bromothenyl) guanine (O6-4-BTG) can access pseudo substrates and inhibit the catalytic activity of MGMT protein (10 q26.3). In addition, miRNAs such as miR-142, miR-181d, miR-221, miR-222, miR-603, and miR-767-3p bind to 3’ untranslated region (3’UTR) of MGMT and lead to degrade the mRNA. Studies by Matthew H. Kulke et al. 2009, have shown that the therapeutic efficiency of TMZ was enhanced during lower levels of MGMT (Yu et al. 2020). Identifying the molecular targets of GBM involved in disease pathogenesis and developing small molecules is essential to anticancer therapy. The nervous system cancers include astrocytoma, ependymoma, glioma, meningioma, medulloblastoma, and neuroblastoma. Chemotherapy has a role in treating almost all newly diagnosed diffuse gliomas (WHO I-IV). Emerging studies have identified potential benefits using TMZ for grade IV GBM, Procarbazole, and Vincristine for grade II and grade III cancers. Several alternatives for GBM care include systemic therapies and combined modality therapy. Furthermore, there are several factors, such as tumor size and Karnofsky's performance score (KPS). Interestingly, the tumor-treating fields (TTFields) with TMZ represent an effective therapeutic strategy for GBM therapy (Hottinger et al. 2016). In GBM cancers and GBM cancer stem cells (GSCs), several chromosomal alternations such as recurrent copy number changes, polysomy (chromosome 7), monosomy (chromosome 10), loss of chromosome 10 q, and deletions in 9p21, as well as cancer stemness markers such as CD15, CD31, CD34, CD45, CD133, CD90 lead to genomic instability and cancer stemness (Pesenti et al. 2019). The other genomic alterations include amplification of epidermal growth factor receptor (EGFR) and platelet-derived growth factor receptor (PDGFR), aberrations in RTK/Ras/PI3K signaling pathways, frequent mutations including alterations in Neurofibromatosis type 1 (NF1), Phosphatase and Tensin Homolog (PTEN), Mouse double minute 2 homolog (FGF), and telomerase reverse transcriptase (hTERT) (Reifenberger et al. 1993; Verhaak et al. 2010; Heidenreich et al. 2015). ncRNAs, a significant portion of the transcriptome that do not appear to have protein-coding functions, are crucial in various biological processes, including disease development (Sun and Chen 2020). ncRNAs are junk transcriptional products that regulate various cellular processes such as chromatin remodeling, transcription, post-transcription, and cancer signaling pathways. These pathways were broadly classified into oncogenic and tumor suppressive in nature. (Wang et al. 2019). Completing an understanding of ncRNAs will provide useful insights for better therapy against various diseases including cancer. LncRNAs, small interfering RNA (siRNA), enhancer RNA (eRNA), circular RNA (circRNA), Y RNA, miRNA, and piwi-interacting RNA (piRNA) are classified as regulatory ncRNAs. In contrast, ribosomal RNA (rRNA), transfer RNA (tRNA), small nuclear RNA (snRNA), telomerase RNA (TERC), tRNA-derived stress-induced RNAs (tiRNA), tRNA-Derived Fragments (tRF), and Small nucleolar RNA (snoRNA) classified as housekeeping ncRNAs (Zhang et al. 2019) rRNAs and tRNAs play a role in protein translation and that lncRNAs and miRNAs can control the expression of genes (Decoding noncoding RNAs). Targeting many transcripts that encode regulators of cell-cycle progression, migration, invasion, and metastasis (Ma et al. 2010; Kim et al. 2016; El Fatimy et al. 2017). miRNAs and lncRNAs are effective biomarkers for predicting treatment outcomes or monitoring therapeutic responses. Additionally, ncRNAs function as a nuclear receptor response element mimic for glucocorticoid receptor (GR), which reduces the expression of oncogenic miRNA and increases apoptosis while reducing proliferation, invasion, and migration (Zhang et al. 2013; Zhao et al. 2015). ncRNAs reduce the production of genes that improve lipid synthesis (SCAP, SREBP-1), proliferation (CDK6), and apoptosis (MCL-1), as well as B cell proliferation (Garzon et al. 2009; Santanam et al. 2010; Ru et al. 2016). The present review focuses on the emerging aspects of therapeutic, prognostic, and diagnostic aspects, as well as drug delivery approaches against GBM cancer. The regulation of various genes involved in cancer cell proliferation by miRNA and lnc RNA.
Drugs and importance
GBM patients have a survival period of 12–15 months. Most therapeutic drugs in clinical trials impair pathological processes of glioma formation and thus improve quality of life. Recent high-through-hit identification (hit-ID) strategies such as high throughput screening, DNA encoded library screening, and fragment-based drug identification led to drug discovery (Silvestri and Colbon 2021). Also, emerging trends have indicated that Drug repurposing is a novel concept for effectively treating cancers and many other health disorders. Also, drug repurposing saves time and is cost-effective (Tan et al. 2018). TMZ is the standard drug with 100% bioavailability and lipophilicity in GBM cancer therapy. In several GBM cases, the high expression of MGMT resulting unresponsiveness to chemotherapeutic drugs and enhancement in the GSCs. This indicates the need to focus on identifying effective therapeutic drugs against GBM cancers (William et al. 2018). To date, Food and Drug Administration (FDA) has approved several drugs such as Lomustine, Carmustine, Bevacizumab, carmustine wafer implants, Dabrafenib, Trametinib, Afinitor, Belzutifan, Danyelza, Welireg, and Tafinlar (Hadjipanayis and Stummer 2019; Odogwu et al. 2018; Novartis 2016; Fallah et al. 2022; Mullard 2021) (Fig. 1). Also, it is imperative to identify the specific, and effective drug molecules that specifically target the GBM cancer tissue and BBB, cancer stem cells, etc. The GBM anticancer drugs ideally possess less than 500 Da. The effective entry of drugs can be prevented by various proteins such as organic anion transporting polypeptide 1A2 (OATP1A2/SLCO1A2), organic anion transporter 3 (OAT3/SLC22AB), p-glycoprotein (P-gp), multi-drug resistance-associated protein 4 (MRP4/ABCC4), monocarboxylate transporter 1 (MCT1/SLC16A1) (Urquhart and Kim 2009).
Recent drug developments have suggested using high throughput (HTS), microwave-assisted organic synthesis (MAOS), combinatorial chemistry, and medicinal chemistry for drug discovery. The genomic-wide association studies (GWAS) about metabolic modeling, transcriptomic data, as well as system biology data have indicated that GBM patients with low survival periods have upregulated glycine (cytosol), methionine (cytosol), L-methionyl-tRNA (met) (cytosol), formate (mitochondria) and low levels of heparin sulfate precursor 14 (Golgi apparatus), taurine ( extracellular), trichloroacetate ( cytosol), etc. (Larsson et al. 2020).
Drugs in clinical trials and their importance
Drugs involved in clinical trial II against GBM are multi-kinase inhibitors, Poly ADP-Ribose Polymerase Inhibitor, proteasome inhibitors, Stromal Cell-Derived Factor Chemokine Receptor Type-4 (CXCR-4) Inhibitors, G-protein coupled receptor antagonists, BRAF inhibitors, etc. (Wang et al. 2021b). To date, only one drug that has reached clinical trial phase III is Bevacizumab, which is presently approved for GBM cancer therapy. Emerging studies have indicated that various drugs such as Tyrosine kinase inhibitors inhibit various tyrosine kinases such as Vascular endothelial growth factor receptor 1 (VEGFR1), Fibroblast growth factor receptor (FGFR), PDGFR, etc., and were found to inhibit tumor angiogenesis (Wilhelm et al. 2011). Several proteasome inhibitors disrupt the cell cycle and are also known to inhibit the proteasomal degradation of several proteins (Yin et al. 2005). Bortezomib inhibits angiogenesis and cytokine signaling and enhances chemotherapy by enhancing apoptosis (Yin et al. 2005). Iniparib (4-iodo-3-nitrobenzamide) (i.e., prodrug) and PARP inhibitor were used to treat breast cancer, pancreatic cancer, and GBM. Researchers have identified that Iniparib facilitates the release of the radical nitro ion that binds to selenium protein and thus enhances redox condition and cellular cytotoxicity (Fogelman et al. 2011; O'Shaughnessy et al. 2011). In addition, BRAF belongs to the serine-threonine kinase family and is an oncogene participating in the RAS-RAF-MEK-ERK pathway. Interestingly, mutation BRAFV600E was observed in 50% of anaplastic ganglions, xanthoastrocytomas, and high-grade gliomas (Chi et al. 2013; Brennan et al. 2013; Schindler et al. 2011). The most important drugs that are useful for treating GBM are presented below.
Lomustine (CCNU)
Chemotherapeutic drugs are also considered primary therapeutic agents to treat GBM. Lomustine was an approved drug for treating recurrent high-grade glioma (HGG). CCNU is an alkylating agent that crosslinks DNA as well as RNA in dividing cells, thereby inducing apoptosis in tumor cells. In GBM patients, CCNU is administered orally at 80–110 mg/m2 every 6 weeks (Wirsching et al. 2014; Wick et al. 2017). Wick et al. conducted a randomized clinical trial (RCT) with CCNU in combination with Bevacizumab, which improved overall survival (Weller and Rhun 2020). Common toxicities of chemotherapeutics were hematologic toxicity (49.7%) (Weller and Rhun 2020). CCNU was a critical factor in PCV (P: procarbazine, C: Lomustine, V: Vincristine) regimen against HGG (Lassen et al. 2014).
Carmustine: (BCNU; bis-chloroethyl nitrosourea)
The FDA approved Carmustine to treat HGGs (Hadjipanayis and Stummer 2019). Walker et al. conducted an RCT that reported a median overall survival (O.S.) of 11.75 months (Walker et al. 1978). Currently, BCNU is only FDA-approved to treat recurrent GBM. Surprisingly, Carmustine induces several nonspecific effects on normal lung and eye cells and pulmonary and ocular toxicity.
Carmustine wafer implants
The FDA approved carmustine wafer implants for recurrent HGGs 2003 (Hadjipanayis and Stummer 2019).
Bevacizumab
Bevacizumab (BVZ) is FDA-approved drug as a monotherapy and combined with Irinotecan (Cohen et al. 2013; Vredenburgh et al. 2007). In several clinical trials, combining Etoposide and Carboplatin with Bevacizumab has exhibited potential benefits against recurrent GBM (Carrillo et al. 2014; Mrugala et al. 2012) (Fig. 1; Table 1).
Temozolomide
TMZ, an oral alkylating agent, is the first-line treatment for GBM, resistance to TMZ is a significant hurdle in GBM patients. The combination of TMZ, difluoromethylornithine (DFMO), an inhibitor of ornithine decarboxylase, and radiation in GBM cell lines resulted in consistently higher suppression of proliferation, causing cell-cycle arrest in the G2/M phase and caspase-8 activity (Alexiou et al. 2019).
Nanoparticle-mediated drug delivery in GBM
Emerging studies have emphasized using nanoparticles to deliver chemotherapeutic drugs or agents specific to the tumor site. Thus, the application of nanoparticle-mediated drugs and siRNA delivery has enormous potential in drug discovery. These nanoparticles were classified as organic or inorganic (Bukhari et al. 2021). Throughout the body, the blood vessels supply nutrients and oxygen to various types of cells. It comprises endothelial cells, neurons, astrocytes, and pericytes (Keaney and Campbell 2015). BBB facilitates the movement of molecules with only molecular weight molecules of less than 500 Da. Nanoparticles carry small molecules such as siRNAs, miRNAs, and drugs into various cells and tissues. Thus, it aids in preventing cancer, cardiac diseases, and diabetes. Recently, several attempts were made to effectively deliver siRNA and miRNA to inhibit RNA-dependent RNA polymerase (RdRP) and viral replication against COVID-19. Organic nanoparticles, including dendrimers, chitosan, etc., and inorganic, iron, zinc oxide, etc., were found to have future therapeutic utility against various cancers. The most important property of an ideal nanoparticle is biodegradability, biostability, ease of synthesis, and multiple modifications (Kozielski et al. 2019). Multiple nanoparticle modifications will allow the incorporation of small molecules, siRNAs, and antibodies in multiple systems for targeting cancer (Pourgholi et al. 2016). Most of these nanoparticles are in clinical trials.
Further, exploring nanoparticles provides better results for effective stabilization in the bloodstream or circulating system. These nanoparticles have considerable advantages in the future therapy of GBM cancer. Nanoparticles that deliver siRNA have amino groups on the outer surface (Weber et al. 2000). siRNA easily binds with the amino group. For example, the PAMAM nanoparticle has 64 amino groups, but the potential application of the PAMAM nanoparticle was limited due to heavy cytotoxic effects on normal cells (Wu et al. 2013). The other types of PAMAM dendrimer modifications include hydroxy, carboxy, etc. In addition, Magnetic nanoparticles were also utilized for the miRNA delivery, which will be further tracked in vivo animal model systems using the magnetic resonance analytic technique.
Nano-formulations generally have a lesser particle size, bulky surface area, reactivity, and several active sites with sufficient adsorption ability. The key advantages of the nanoparticle are increased drug absorption, bioavailability, and prolonged circulating time (Yin et al. 2020). SGT-p53 comprises wild-type p53 plasmid DNA in a cationic liposome containing transferrin receptor and single-chain antibody. This nano-complex crosses the BBB and sensitizes the TMZ resistant GBM (Kim et al. 2015). It was found that SGT-p53 reverses TMZ resistance via abrogation of MGMT and enhances the TMZ-induced cytotoxic effect in TMZ-resistant GBM (Kim et al. 2016). The various nanoparticles employed for GBM cancer therapy are liposomes, polymers, magnetic nanoparticles, etc. Further, the inhibitors used in nanoparticle-mediated drug delivery in GBM are interferon β (IFN β), Signal transducer, and activator of Transcription 3 (STAT3) inhibitor, etc. (Yoshino et al. 2009; Koh Saka et al. 2012; Bobustuc et al. 2010; Hirose et al. 2001). A mutant form of p53 was observed in both primary and secondary GBM, and thus improving wild-type p53 has a beneficial effect (Steele and Lane 2005; ShChors et al. 2013). Transferrin receptor (TfR) is a receptor that was widely expressed in BBB and GBM cells (Ramalho et al. 2022; Voth et al. 2015). Interestingly, Transferrin receptor 1 was found to control the rate of iron uptake by fine-tuning the amount of iron delivered to the cells to meet metabolic needs (Cazolari et al. 2007). Recent studies have identified the critical hurdles in GBM therapy, such as the inability to cross BBB and ineffective penetration in the BBB.
Liposome (Yang et al. 2012)
The initiation, progression, of GBM cancer was dysregulation due to differential changes in miRNAs. Thus, targeting dysregulated miRNA with various oligonucleotides has clinical limitations, including poor RNA stability, off-target effects, and inefficiency in crossing the BBB. The BBB was found to prevent many small molecule drugs from entering the brain tumor environment. However, focused ultrasound (FUS) combined with intravenous microbubbles (MBs) have shown a promising result and enhanced the BBB permeability. Interestingly, modified liposomes (i.e., liposomal-doxorubicin) targeting Interleukin-4 (IL4) receptor has exhibited promising result in nonobese diabetic-severe combined immunodeficiency (NOD-scid) mice. Both drug delivery, and survival of NOD Mice of GBM cancer was enhanced.
Polymers
Several copolymers include polyethylene glycol (PEG), polypropylene glycol (PPG), and poly(L-lysine). These copolymers have properties such as self-assembly, siRNA binding, particle size, surface potential, the architecture of the complexes, and siRNA delivery. Silencing of green fluorescent protein (GFP) using copolymer to deliver GFP-specific siRNA to Neuro-2a cells expressing GFP was almost as effective as using Lipofectamine 2000, with minimal cytotoxicity. Thus, the copolymer platform for siRNA delivery exhibited improved siRNA delivery in vitro and in vivo (Dai et al. 2014). Recent studies have identified that specifically designed siRNA bind and induce post-transcriptional silencing of target genes (mRNA).
Magnetic hyperthermia therapy
Magnetic Hyperthermia Therapy (MHT) is a modern, advanced therapeutic option for treating GBM. Hyperthermia therapy (H.T.) involves exposure of a body region to elevated temperatures to achieve and potential anticancer effect. This includes radiofrequency, ultrasound, microwave, laser, and magnetic nanoparticles (MNPs) (Mahmoudi et al. 2018).
Gold nanoparticle
Emerging studies have revealed that the surface reactivity of gold nanoparticles (AuNPs) has gained attention as a radiation therapy radiosensitizer for cancer cells and as a drug carrier to target cells. This calf thymus DNA with HAuCl4 solution as a radiosensitizer of human glioma cells with cancer stem cell (CSC)-like properties (e.g., U251MG-P1), to reduce their survival. The radiosensitivity of the AuNP-associated cells is significantly enhanced. Also, the generation of reactive oxygen species (ROS), apoptosis induction, or DNA damage was enhanced (Kunoh et al. 2019). In vivo, 1.9 nm nanoparticles were found to be toxic following intracerebral delivery in rats bearing glioma, while no toxicity was observed using 15 nm nanoparticles at the same concentration (50 mg/mL). Survival of rats that had received the combination of treatments (AuNPs:50 mg/mL, 15 Gy) was significantly increased compared with the survival of rats that had received irradiation alone (Bobyk et al. 2013). Gold-iron oxide nanoparticles (polyGIONs) surface loaded with therapeutic miRNAs (miR-100 and antimiR-21) inhibit GBM cancer cell proliferation and enhance apoptosis (Sukumar et al. 2019).
Dendrimers
2,2-bis(methylol)propionic acid (Bis-MPA) as "nonviral vectors" for transfection of siRNA in cell cultures. The study encompassed dendrimers of generation one to four (G1–G4), modified to bear 6–48 amino end-groups, where the G2–G4 proved capable of siRNA complexation and protection against RNase-mediated degradation. The G2 dendrimers were nontoxic to astrocytes, glioma (C6), and GBM cancer cells (U87), while G3 and G4 dendrimers exhibited concentration-dependent toxicity towards primary neurons (Stenström et al. 2018). Specific properties in cancer cells compared to normal cells, such as overexpression of various receptors and differences in biological conditions like pH, temperature, and redox of tumor microenvironment, cause an increase in site-specific targeting efficiency. Thus, modifications of dendrimers through the attachment of lipids, amino acids, proteins/peptides, aptamers, vitamins, antibody were effective against GBM (Ghaffari et al. 2018). Studies also proved that Poly (amidoamine) (PAMAM) dendrimers are well-defined, highly branched macromolecules with numerous active amine groups on the surface. These N.P.s carry drugs and genes (pDNA, siRNA) and deliver them to cancer cells.
Inorganic nanoparticles
Titanium dioxide nanoparticles (TiO2NPs) have attracted interest due to their use in various applications. TiO2NPs can enter the brain; toxicity was assessed at different levels: mitochondrial function (by MTT), membrane integrity, and cell morphology (by calcein AM/PI staining) after acute exposure at various doses ranging from 1.5 to 250 μg/ml for 7–10 days at sub-toxic concentrations (from 0.05 to 31 μg/ml). Prolonged exposure has revealed that the proliferative capacity (colony size) was compromised at the shallow TiO2NP doses, such as 1.5 μg/ml and 0.1 μg/ml, respectively, for D384 and SH-SY5Y (Coccini et al. 2015).
Polymicelles
Polymeric micelles are core–shell-type nanoparticles that act as promising nanoparticles due to their size, stability, and drug incorporation efficiency and release rate (Nishiyama et al. 2016).
Quantum dots
Quantum dots (Q.D.) nano-transporters are Carbon quantum dots (CQDs) that were successfully functionalized with Mal-PEG-NHS linked RGERPPR. They exhibit double functions of both tissue imaging and targeting train gliomas (Devi et al. 2022).
Nanogels
Nanogels are unique local tailorable drug delivery systems and consist of a three-dimensional polymeric network formed via physical or chemical assembly. Nanogel delivery systems (DPPC) with cell-penetrating peptides (CPP) are introduced into the astrocyte. DPPC is around 300 nm, the potential is about 0–5 Mv. The DPPC is verified as the `biocompatible carrier for further application by cell viability tests. The in vitro- constructed BBB model proves that Dipalmitoyl phosphatidylcholine (DPPC) can efficiently penetrate the BBB, attributed to both the temperature-sensitive passive targeting and the active cell-penetrating peptides (CPP) penetration. This indicates that the use of these nanoparticles acts effectively against glioma.
Graphene
Graphene, graphene oxide, and reduced graphene oxide were considered promising for industrial and biomedical applications due to their high mechanical stiffness and strength. Synthesized techniques, such as liquid phase exfoliation and wet chemical oxidation, often require toxic organic solvents, surfactants, strong acids, and oxidants for exfoliating graphite flakes. The residual contaminants cause of graphene-induced toxicity in biological cells. Pooresmaeil and co-workers developed the pH-responsive magnetic (Fe3O4 NPs)/G.O. hybrids to deliver doxorubicin. Approximately 65% drug release was observed at 40 °C and pH 5.0 in cancer cells, which was 22% in normal cells (37 °C and pH 7.4) (Borandeh et al. 2021). Liu et al. 2012 fabricated electrically responsive rGO / poly (vinyl alcohol) (PVA) membranes for the delivery of lidocaine hydrochloride (Liu et al. 2012). The G.O. nanocarrier system has more advantages such as anti–tumor drug delivery systems, like liposomes, ① good blood compatibility and optimal dispersibility in the liquid environment of the human body (Daniyal et al. 2020); ② sizeable specific surface area facilitating multi-functional modification by biomolecules and small molecules, such as proteins and single–stranded DNA bases (Shahmoradi et al. 2018). (Wang et al. 2022).
Nanopeptide-drug combination
Studies have indicated that platinum pro drug (Pt IV) was effectively transported into GBM tissue by M13 peptide, A cell-penetrating peptide transporter 10, that can deliver the drugs to the tumor site and inhibit the growth of the tumor. Emerging studies have identified the tumor homing peptide, also known as tumor-promoting peptides such as TT1 and its linear form Lin TT1 (AKRGARSTA), bind to cell surface receptors expressed on GBM cancer cells and follow the process of tumor accumulation and penetration. These peptides will process and generate the c-terminally expose a C-end rule motif RXXR/K-OH in TT1, which will further bind to another receptor neutrophilic-1 (NRP-1) which eventually results in endocytic/exocytic transcytosis, extravasation, and tumor penetration (Teesalu et al. 2013; Sugahara et al. 2009; Teesalu et al. 2009; Sharma et al. 2017). Recent studies have also indicated that Iron oxide nano worms (N.W.s) coated with LinTT1, a nanocarrier system optimized for peptide-mediated tumor targeting (Park et al. 2008, 2009, 2010; Agemy et al. 2011; Roth et al. 2012).
Signaling pathways in GBM cancer
GBM (grade IV cancer) has poor patient survival. GBM cancers are primarily primary and are up to 90% in incidence. The majority of patients are in the elderly category. Secondary-grade GBM cancers are low-grade astrocytomas. The critical genetic alterations and pathway alterations in GBM were found to be p53, EGFR, PDGFR, PTEN, MDM2, phosphatidyl inositol-3-kinase PI3K/ Akt/ mammalian target of rapamycin (mTOR), mitogen-activated protein kinase (MAPK), nuclear factor-kappa beta (NF-kB), Wnt, STAT-3, and NOTCH pathway. Understanding the complex disease biology and the signaling provides effective therapeutic strategies and improves the prognostic and diagnostic aspects of GBM pathogenesis and prevention.
P53pathway
The p53 gene is a regulator and a critical tumor suppressor that induces cell-cycle arrest and apoptosis (Huang et al. 2007; Huse and Holland 2010), many of which are involved in tumor development and invasion. GBMs are divided into primary and secondary subtypes. Primary GBMs develop quickly and robustly, while secondary GBMs develop progressively from low-grade astrocytoma. p53 mutations are the most common development of secondary GBMs, whereas mutations in the p53 pathway are also detected in primary gliomas at a lesser frequency (St Louis et al. 1999; Ohgaki and Kleihues 2013). PTEN mutations were found to be widely mutated in high-grade gliomas (Lespagnard et al. 1999; Fulci et al. 2000). The chromosomal regions (chro 9p, chro 10q23.3, and chro10q25.26) that encompass genes such as CDK2A cyclin-dependent kinase 2A), CDK2B (cyclin-dependent kinase 2B), ARF, MDM2, EGFR, and PTEN were also found to be mutated or deleted. p53 protein expression was associated with significantly longer survival rates, as observed in univariate analysis. Also, it was found that in multivariate analysis of overall survival (Cox regression), only postoperative Karnofsky performance status remained as an independent prognostic factor (Birner et al. 2002). Inactivation of p53 can result in resistance to apoptosis, a critical mechanism in treatment failure during DNA-damaging agents (Nieder et al. 2000). Mutations of the p53 gene on (exons 5 to 8) were found in many primary tumors and to a lesser extent in oligodendroglia, 1 oligoastrocytoma). (Reifenberger et al. 1996).
pRB-CDK2-CDK4 axis in GBM pathogenesis
Retinoblastoma (R.b) is located at the chromosome location 13q14.1-q14.2 and was found to be involved in the cancer progression of astrocytomas (Henson et al. 1994). Mutations in R.B. are detected in more than 20% of high-grade gliomas. Interestingly loss of 13q was associated with the transition from low- to intermediate-grade gliomas (Henson et al. 1994; Bahuau et al. 1998). CDKN2B, i.e., (p15), a CDK inhibitor commonly inactivated in GBM, forms a complex with CDK4 or CDK6, thus preventing the activation of CDKs leading to the inhibition of cell growth and the cell-cycle arrest at G1 phase.
PI3K-PTEN-Akt-mTOR pathway
The PI3K-PTEN-Akt-mTOR pathway regulates normal cellular functions and plays a critical regulatory role in cancer cell migration and metabolism. It was found that the PI3K pathway is altered in about 70% of GBMs, due to various biological aspects such as deletion of PTEN or amplification of EGFR and vascular endothelial growth factor receptor (VEGFR)/ platelet-derived growth factor receptor α (PDGFRα) (Zhi et al. 2009). Overexpression of EGFR, which is one of the most frequent signaling mutations in GBM, and its amplification leads to increased activation of the PI3K pathway (Peraud et al. 1997; Watanabe et al. 1996).
RAS/MAPK pathway
Human RAS genes (Rat Sarcoma) transform oncogenes, including H-Ras, N-Ras, and K-Ras. K-RAS belongs to the G protein family. The activation and deactivation of RAS are controlled by its binding to guanosine triphosphate (GTP) or guanosine diphosphate (GDP), respectively (Lu et al. 2005; Hurley et al. 1984). Activated RAS further activates RAF kinase through direct binding, regulating downstream signaling pathways such as the mitogen-activated protein kinase (MAPK) pathway (Moodie et al. 1993; Thomas et al. 1992). RAS also regulates the activities of other pathways, such as the PI3K pathway, and consequently, RAS regulates cancer cell proliferation, differentiation, signal transduction, apoptosis, and tumorigenesis.
STAT3 and zinc importer 4 (ZIP4) pathway
Signal transducer and activators of transcription (STAT) protein complexes are a family of cytoplasmic proteins with Scr Homology-2 (SH2) domains that functions as cytokines and transduce signals from the cytoplasm to the nucleus. Interestingly, these proteins acted as transcription factors and were found to regulate various biological processes related to cancer cells, such as proliferation, and migration (Abal et al. 2006; Rahaman et al. 2002). Interestingly, STAT3 was once thought to possess only oncogenic properties, but emerging studies have identified both the suppressive and oncogenic roles in GBM, depending on the genetic profile of the tumor (de la Iglesia et al. 2008).
WNT pathway
Wnt signaling is overexpressed GBM. Activation of the Wnt pathway in these tumors is associated with mutations in, Adenomatous polyposis coli (APC), β-catenin, AXIN, and transcription factor 4 (TCF4). Mutations in Wnt signaling genes were extensively characterized in colorectal cancers. Mutations in Wnt signaling components were observed in β-catenin, APC, and AXIN1 of colon cancer and medulloblastoma, hepatocellular carcinoma. In contrast, aberration of the key components of the Wnt pathway is not of common occurrence in GBM, and gastric cancers (Nageret al. 2012; Morris et al. 2013). Recent reports from a small cohort have reported the existence of APC mutations observed in 13% of GBM (Tang et al. 2015). Overexpression of leucine-rich protein 1 (PELP1) was observed in almost all GBM samples (Sareddy et al. 2019). Studies have identified the pivotal role of epigenetic modifications regulating the Wnt pathway. Further, Gene Expression Omnibus miRNARNA profiling of GBM versus the normal brain found that miR-138–2-3p and miR-770-5p were differentially expressed.
The mammalian homologs of the Drosophila melanogaster protein Van Gogh (VANGL). VANGL1, VANGL2, and frizzled protein 7 (FZD7) are transcriptionally upregulated in glioma and correlate with poor patient outcomes. Consequently, knocking down of VANGL1 suppresses the motility of GBM cell lines, restoration of Neuregulin receptor degradation protein-1 (NRDP1), a RING finger type E3 ubiquitin ligase whose decreased expression in GBM correlates with poor prognosis, reduces GBM cell migration, and invasiveness by suppressing Planar Cell Polarity (PCP) signaling. These findings revealed an essential mechanistic role for this pathway in GBM malignancy (Wald et al. 2017). In addition, receptor-like tyrosine kinase (RYK), a typical member of the receptor tyrosine kinase (RTK) family involved in the control of neuronal differentiation (Lyu et al. 2008), resulted in being essential for WNT5a-dependent invasiveness in glioma. (Hirano et al. 2014). MuTSigCV has indicated that in GBM cancer, the mutation of genes such as PTEN, EGFR, p53 (Lawrence et al. 2013). In addition, loss of chromosome 10q, alterations of p53, amplification of EGFR and PDGFR, and aberrant tyrosine kinase (RTK/Ras) signaling contributes to GBM cancer. Interestingly, studies have shown the differential expression of transforming growth factor beta 1 (TGFβ1) induced and SRY box 4 (SOX4) were also considered as therapeutic drug targets (Qiu et al. 2018). Furthermore, genes such as nucleolar and spindle-associated protein 1 (NUSAP1) and G-protein coupled receptor 65 (GPR65) were considered survival biomarkers.
Gene expression patterns in GBM cancer
In GBM cancer, three genes TP53, PTEN, and EGFR are the most significantly mutated genes (SMG) as observed by MuTSigCV (Lawrence et al. 2013). The loss of chromosome 10q, alteration of p53, R.B., amplification of EGFR, PDGFR, and aberrations in receptor tyrosine kinase (RTK/ Ras) signaling. Other frequent alterations include NF1 and MDM2. Differentially expressed genes such as transforming growth factor beta-induced (TGFβ1) and SRY box 4 (SOX4) were also considered therapeutic target genes. Further, in survival analysis, nucleolar and spindle-associated protein 1 (NUSAP1) and G-protein coupled receptor 65 (GPR65) were significant genes.
Dynein, cytoplasmic 1, intermediate chain 1 (DYNC1I1) was down-regulated in glioma. The lower expression of DYNC1I1 was correlated with poor patient survival. The epigenetic mark H3K27me3 on lysine 4 was found in the promoter region, revealing the active transcription region. The SEMA3C is another essential gene with multiple mutations observed in GBM cancer. Telomerase reverse transcriptase (TERT) gene -124 bp (hg19chr5:1; 295, 228 C > T; -146 bp (hg 19 chr5, 1, 295, 250 C > T (Heidenreich et al. 2015). The homeobox gene 5 DLX5, distal-less homeobox 5, affects glioma cell motility via PAX6/DLX5-WNT5A axis (Hu et al. 2016; Cell; 167:1281–1295). Among GBM and ISL1 are. 90% of GBM cases belong to the IDH-WT type ( high grade), and 10% of cases IDH mutant type ( lower grade). The other vital mutations are protein tyrosine phosphatase receptor type Z1 (PTPRZ1 and promoter methylation in MGMT. Thus, it is imperative to investigate the mutations, methylation, and other epigenetic changes to identify the prognostic and diagnostic markers and therapeutic response.
Prognostic and diagnostic markers
Glioma cells can invade the neighboring tissues beyond detection leading to tumor relapse. This leads to an inevitably critical recurrence even after the surgical removal of GBM tissue (Jacobs et al. 2011; Stylli et al. 2005).
IDH
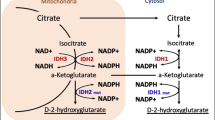
IDH mutation was prognostic of more prolonged survival in low-grade Glioma (Leu et al. 2013; Sabha et al. 2014; Metellus et al. 2010; Gorovets et al. 2012). IDH mutations were most common in cases of oligodendroglioma (94%) and little lesser extent in cases of astrocytoma or mixed tumors (Leu et al. 2013). Metabolic mapping and image analysis on GBM cancer samples revealed the occurrence of IDH1 (R132) mutation. NADP + -dependent IDH activity and other NADPH-producing dehydrogenases, glucose-6-phosphate dehydrogenase, 6-phosphogluconate dehydrogenase, malate dehydrogenase, and hexose-6-phosphate dehydrogenase. Mutation of Isocitrate dehydrogenase (IDH) gains oncogenic function. It converts alpha-ketoglutarate to the oncometabolite 2-hydroxyglutarate (2-HG), ultimately leading to genome-wide methylation change in GBM patients, leading to altered gene expression (Najafi et al. 2022). Heterozygous mutations of isocitrate dehydrogenase-1 (IDH1) dominantly inhibit wild-type IDH in GBM cancer. Studies have revealed that IDH1 activity results in the inactivation of enzymes. Interestingly, forced expression of a mutant form of IDH1 in cultured cells decreased alpha-ketoglutarate (alpha-KG) and thus enhances hypoxia-inducible factor subunit HIF-1 alpha (HIF-α) that facilitates tumor growth during low oxygen availability. Thus, HIF-1α levels were higher in tumors with mutant IDH than in wild-type tumors (Zhao et al. 2009). Interestingly, IDH mutation was observed to be associated with enhanced DNA methylation, called Glioma CpG island methylation phenotype (G-CIMP) (Noushmehr et al. 2010). It was observed that in secondary GBMs, IDH mutation and G-CIMP phenotype exist exclusively (Cohen et al. 2013). The key IDH mutations include those associated with R132 (CGT): 12R132H (C•G → T•A) and 1 R132G (C•G → G•C) that functions as a homodimer. The mutant IDH1 protein acts as a dominant negative by combining with the wild-type (W.T.) allele product to afford a dysfunctional heterodimer of the wild-type and mutant (Zhang et al. 2019).
MGMT
MGMT was known to play an essential role in chemoresistance to TMZ. Thus, MGMT is considered a promising target in GBM treatment (Yu et al. 2020). GBM patients with MGMT methylation were associated with more prolonged overall survival but not progression-free survival (PFS) (Binabaj et al. 2018). It was also established that methylation of MGMT promoter is a crucial predictor of alkylating agents’ ability to kill glioma cells. The methylation sites and rich CpG islands vary in MGMT-deficient GBM cancer cells. However, the change in the methylation status of the MGMT promoter after chemotherapy, radiotherapy, or both still need to be fully explored. Several studies have demonstrated that chemotherapy can induce MGMT expression in gliomas. Thus, researchers have employed several strategies that have been pursued to improve the anti-tumor effects of TMZ which include the synthesis of analogs of O6-methylguanine (O6-meG) such as O6-benzylguanine (O6-BG) and O6-(4-bromothenyl) guanine (O6-BTG), RNA interference (RNAi), and viral proteins (Yu et al. 2020). The expression level of MGMT in glioma has no relation with gender, age, tumor size, surgical approach, and Karnofsky Performance status (KPS) score. MGMT has shown a profound influence on cancer cell survival and proliferation during the treatment with O6-alkylating agents such as TMZ, Carmustine, Lomustine, etc. In addition, to these drugs, MGMT deficiency causes O6-methylguanine lesions to mispair with thymine, which leads to mismatch repair (MMR) mediated apoptosis in gliomas. Thus, MGMT, MMR, and DNA replication were found to be crucial factors in cell resistance (Li et al. 2017a, b). In general, as a part of diagnostics as well as prognostics, the methylation of MGMT promoter was detected by methylation-specific PCR, and bisulfite sequencing (BiSEQ). EGFR amplification was detected by fluorescence in situ hybridization (FISH) or next-generation sequencing was co-related (Goldstein et al. 2019). Studies have indicated that optimal assessment of MGMT status function as a prognostic biomarker for patients with newly diagnosed GBM treated with chemo-radiation requires determination of both promoter methylation and immunohistochemistry (IHC) protein expression (Lalezari et al. 2013). Interestingly concordant MGMT methylation and lack of protein expression result in a response in TMZ therapy-treated patient subgroups with HR of 2.02 and 0.76 (p < 0.05) (Pandith et al. 2018; Uno et al. 2011).
3.P53
Expression of the neoplastic phenotype in GBM cells was inhibited when rat cells were transfected with the murine wild-type p53 gene, mutant p53 gene, and other oncogenes (Finlay et al. 1989). Moreover, the short arm of chromosome 17 is often deleted in human tumors. In colorectal cancers, deletion of 17pl3.1 (Baker et al. 1989); harbors the p53 gene. In general, p53 gene mutations are clustered in four 'hot spots' that coincide with highly conserved regions. DNA sequence analysis has revealed that p53 mutations were rare in primary GBMs (11%), whereas secondary GBMs were characterized by a high number of p53 mutations (67%). The incidence of p53 protein accumulation (nuclear immunoreactivity to PAb 1801) was also lower in primary (37%) than in secondary GBMs (97%) (Watanabe et al. 1996). Progression of low-grade astrocytomas to anaplastic astrocytoma or GBM occurred at an equal frequency (Watanabe et al. 1997).
PTEN
PTEN deleted on chromosome 10 is a tumor suppressor gene that regulates various biological processes such as proliferation, survival, genomic stability, and cell motility (Bazzichetto et al. 2019). Regulation of PTEN function involves genetic, transcriptional, post-transcriptional, and post-translational events (Yang et al. 2017a, b). Recent meta-analysis studies indicated that PTEN mutation is associated with poor prognosis and shorter survival time (Han et al. 2016a, b; Sasaki et al. 2001). (Han et al. 2016a, b). Mutation of PTEN is the second common oncogenic mutation in GBM, occurring in 30% of the tumors (Gao et al. 2013; Cerami et al. 2012). PTEN mutation and mutations in EGFR are critical prognostic factors in anaplastic astrocytoma with GBM (Smith et al. 2001). PTEN on chromosome 10q23.3 regulates the Akt signaling pathway and thus modulates cell growth and apoptosis. It was found that the PTEN gene is mutated in 20–40% of GBM. Very few GBMs showed loss of PTEN. PTEN methylation frequently occurs in GBMs leading to loss of PTEN expression. Loss of Heterozygosity (LOH) at the PTEN locus and loss of PTEN protein expression was inconsistent. (Baeza et al. 2003). The novel chromatin-associated function of PTEN in complex with the histone chaperone death domain associated protein (DAXX) and the histone variant H3.3. Interestingly, PTEN interacts with DAXX and, regulates oncogene expression by modulating DAXX-H3.3 association on the chromatin, independently of PTEN enzymatic activity. DAXX inhibition inhibits tumor growth explicitly and improves the survival of orthotopically engrafted mice implanted with human PTEN-deficient glioma samples, associated with global H3.3 genomic distribution changes leading to upregulation of tumor suppressor genes and downregulation of oncogenes.
EGFR & CDKN2A
EGFR gene amplification and overexpression are striking features of GBM in up to 40% of tumors (Hatanpaa et al. 2010). It was found that the Median survival was longer in the high-amplifier group (Hobbs et al. 2012). Overexpression of EGFR was an indicator of poor prognosis in overall survival in glioma patients (Li et al. 2018a, b). The serum levels of EGFR are enhanced many folds in patients with malignant Glioma, suggesting poor survival (Quaranta et al. 2007; Li et al. 2018a, b). The highly oncogenic mutant is produced by the deletion of exons 2 to 7 of the EGFR causing a loss of 267 amino acids from the receptor's external domain. Since EGFRvIII cannot attach a ligand, it signals automatically. EGFRvIII shares the same signaling domain as the wild-type EGFR, but it appears to produce a unique set of downstream signals that may boost tumorigenicity. EGFR and Cyclin-Dependent Kinase Inhibitor 2A (CDKN2A) alterations and determine the prognostic significance in lower-grade glioma (LGG).
Neurofibromin 1 (NF-1)
NF1 is a tumor suppressor gene and a RAS-GTPase. Dysregulated NF1 expression activates cancer cell proliferation, migration, and invasion. Loss of NF1 expression in GBM was associated with increased tumor aggressiveness. The neurofibromin protein contains at least four significant domains. The leucine-rich domain (LRD) of neurofibromin inhibits the invasion of human GBM cells without affecting their proliferation. NF1-LRD fails to hydrolyze Ras-GTP to Ras-GDP. Thus, NF1-LRD inhibits glioma invasion (Fadhlullah et al. 2019).
MDM2
Whole transcriptome analysis has identified MDM2 as associated with sensitivity and resistance to the chemotherapeutic drugs against GBM. MDM2 amplification occurred in 2 primary (7%) GBMs but none of the secondary GBMs. Only one out of 15 primary GBMs overexpressing MDM2 contained a p53 mutation (Biernat et al. 1997).
MALAT1
MALAT1 is a prognostic factor in GBMs that induces chemoresistance to TMZ via suppression of miR-203, thereby promoting thymidylate synthase (TS) expression. MALAT1 knockdown reversed TMZ resistance in GBM cells. In contrast, MALAT1 overexpression induced chemoresistance by suppressing miR-203, promoting TS expression. (Chen et al. 2017).
HOTAIR
HOTAIR is an adverse prognostic factor overexpressed in multiple human cancers.
HOTAIR has overexpressed in GBM. HOTAIR was frequently co-expressed with HOXA9 in high-grade gliomas. Integrated into silico analyses, chromatin immunoprecipitation (chIP) and quantitative RT-PCR (qPCR) data showed. GBM patients with high HOTAIR expression have a significantly reduced overall survival, (Xavier-Magalhães et al. 2018).
Maternally Expressed Gene 3 (MEG 3)
MEG3 expression, when observed in studies, was significantly downregulated in GBM cancer, and negatively correlated with WHO grade in glioma patients. Low MEG3 expression was associated with the advanced WHO grade. This indicates the role of MEG3 in glioma cell proliferation, apoptosis, and autophagy (Zhao et al. 2018).
Plasmacytoma Variant Translocation 1 (PVT1)
The expression of lncRNA PVT1 oncogene (PVT1) in Glioma and its clinical samples of gliomas have shown that its level is positively related to WHO glioma grade and prognosis of gliomas.
Urothelial Carcinoembryonic Antigen 1 (UCA 1)
Urothelial carcinoembryonic antigen 1 (UCA1), overexpressing glioma cell lines. LncRNA UCA1 can promote glioma cells' proliferation by upregulating cyclin D1 transcription. Thus, UCA1 may serve as a prognostic indicator (Zhao et al. 2017).
The epigenetic mechanism, genetics, and epigenetics in GBM cancer
LncRNA is an epigenetic player of the size of more than 200 nucleotides in length. Studies have identified that a delicate balance between genetic and epigenetic alterations drives tumor recurrence and chemoresistance. The Cancer Genome Atlas (TCGA) has identified the involvement of a considerable number of lncRNAs which can specifically be expressed in low-grade glioma (LGG) subtypes (IDH1/2 wt, IDH1/2 mut, and IDH mut 1p19q co-deletion), as well as classical, mesenchymal, neural, and pro-neural types of GBM (Reon et al. 2016). Microarray analysis has shown few lncRNA, such as ACo16745.3, XL0C_001711, and RP11-128A17.1 were involved in glioma recurrence. The critical target of ACo16745.3 is fork-head box protein D4- like1 (FOXD L1). Higher expression of FOXDL1 correlates with higher expression of lncRNA. LncRNA (up to 50,000) and miRNA (up to 3,000) broadly and profoundly regulate gene expression. Like in other cancers, the expression status of various lncRNAs in GBM cancer can be used as diagnostic and prognostic markers. Frequent deregulation of lncRNAs was observed in cancer cells which regulate several aspects of malignancy, including tumor cell proliferation, survival, invasion, and migration, as well as cancer stemness, angiogenesis, tumor immune responses, therapy resistance, and microenvironment (Fig. 2A-D). Lnc RNA participates in the differentiation and maintenance of pluripotency of stem cells. Chromatin RNA In situ reverse transcription sequencing (CRIST-seq) has emphasized that the lncRNA interacts with promoters of Oct4, and Sox-2 dictates the fate of stem cells (Chen et al. 2020).
Nuclear RNAs interact with various RNA types, such as transcription factors, chromatin modifying factors, and several RNA binding factors to regulate gene expression. The lncRNA Gm15055 was found to be induced by Oct4 and regulated the HOX gene expression by interacting with PRC2, which was involved in the maintenance of H3K27me3 (Liu et al. 2016). Mechanistically, lncRNA, which is p53-regulated and ESC-associated 1 (lncPRESS1), interacts with SIRT6. Inhibiting SIRT6 decreases the histone H3K56 and H3K9 acetylation levels to safeguard human embryonic stem cell (hESC) pluripotency (Jain et al. 2016). P53 inhibits the lncPRESS1, and the knockdown of lncPRESS1 result in the differentiation of hESC by increased expression of HOXA2, HOXB1, and FOXA2 and decreased expression of c-Myc, oct-4, and Nanog, etc. (Jain et al. 2016).
LncRNAs were involved in various cancers, including hepatocellular carcinoma and ovarian cancers. RNA sequencing analysis has identified vital lncRNA SOX2OT, and its nearby Sex determining region Y-box 2 (SOX2) was upregulated in TMZ -resistant cancer cells (Shahryari et al. 2015). SOX2 activates the Wnt / β-catenin pathway and induces cisplatin-resistance of lung adenocarcinoma (He et al. 2017). SOX2OT can regulate SOX2 to promote cancer cell growth and proliferation via regulating the miRNAs such as miR-195-5p and miR-122 in glioma cells (Su et al. 2017). SOX2OT is positively regulated with tumor grade, and the level of SOX2OT is higher in relapsed GBM patients than in primary GBM patients. Cancer stem cell-associated distal enhancer of SOX2 (CASCADES) functions as an epigenetic regulator, and the knockdown of CASCADE in GSCs results in the differentiation of neurons (Shahzad et al. 2020). Another essential lncRNA MATN-AS1 and its regulation of RELA genes, such as p65, p50, p52, c-Rel, and RelB, is involved in GBM cancer stem cell proliferation (Han et al. 2019) (Fig. 3; Table 2).ncRNAs such as TALC, MALAT1, OIPS-AS1, HOXD-AS1, H19, UCA1, NEAT1, and HOTAIR regulate several miRNAs that can regulate several genes involved in carcinogenesis.
RNA-guided RNA modification in GBM diagnosis
RNA–protein complexes involved in the RNA-dependent modifications. In human RNAs, approximately 200 types of 2′-O-methylations and pseudouridylations are introduced by two RNA-guided RNA modification systems, such as box C/D and H/ACA RNA–protein complexes (RNPs) (Table 3). A distinct guide RNA belongs to each complex for determining the target RNAs and binding to their complementary regions. Similarly, a group of proteins with the modifying enzyme (2′-O-methylase or pseudouridylase) belongs to each complex (Ye et al. 2009). For example, snoRNPs, a small nucleolar ribonucleoprotein complex, are involved in RNA-guided RNA modification. Box C/ An RNA-guided RNA modification system carries out alteration of the primary sequence and modulation of the function of target RNAs, including rRNAs, snRNAs, tRNAs, and perhaps mRNAs. From various eukaryotic RNAs, uridines are converted to pseudouridines with the help of H/ACA RNPs. The functional H/ACA RNP complex consists of a guide RNA and four proteins such as Cbf5, Gar1, L7Ae, and Nop10. L7Ae and Cbf5 interact with guide RNA. Cbf5 catalyzes the modification via explicitly identifying and binding to H/ACA guide RNAs. Guide RNAs help modify specific ribonucleotides by base pair with target RNAs (Baker et al. 2005).
D guide RNAs involved in the 2′-O-methylation of specific nucleotides. C/D RNAs are the methylation guide RNAs with a C (RUGAUAG, R is purine) and D (CUGA) box near the 5′ end and 3′ end, respectively. A bipartite C/D RNA functional enzyme complex consists of a guide RNA and three different proteins: methyltransferase fibrillarin, Nop5 (Nop56/58), and L7Ae. In the eukaryotic complex, a single Nop5 protein is replaced by paralogs Nop56 and Nop58 and a 15.5-kDa protein instead of L7Ae. Fibrillarin catalyzes the transfer of a methyl group to the ribose 2′-OH group from the unbound SAM. A scaffold protein, Nop5 consists of three domains, (i) a coiled-coil domain for self-dimerization; (ii) an N-terminal domain (NTD) for binds to fibrillarin; (iii) a C-terminal domain (CTD) for binding to the L7Ae–RNA complex. L7Ae–C/D RNA complex formation is the initial step, followed by Nop5 association with the preassembled L7Ae–C/D RNP in the C/D RNP assembly process. Finally, fibrillarin is recruited into the assembly by its interaction with Nop5. The activity of this complex is dependent mainly upon the integrity of the symmetric structure (Ye et al. 2009) (Table 3).
Conclusion
GBM is the most malignant and aggressive type of Glioma. Understanding GBM progression and epigenetic regulation by ncRNA will help in future diagnostic tools and therapeutic strategies. A further role of lncRNAs in gliomas may lead to the discovery of novel molecular mechanisms behind glioma biological features. It also enables the development of new solutions to overcome the most significant obstacles in treating glioma patients. Epigenetic alterations can cause the mis-regulation of ncRNAs. Primarily, lncRNAs will act as a scaffold for various epigenetic proteins, such as EZH2 and LSD1, and influence the epigenetic chromatin state at various genomic loci in cancer cells. Both miRNAs and lncRNAs can interact with numerous epigenetic modifiers and transcription factors to influence gene expression. Studies found that most abnormally expressed ncRNAs impact cellular proliferation and apoptotic pathways, and such changes are cancer-dependent. Further, the nature of miRNA binding to multiple mRNAs and the precise molecular and biological mechanisms targeting a miRNA should be carefully studied. These studies should be conducted not only in the tumor cells but also in the tumor microenvironment. LncRNAs hold great promise in the treatment of cancer. Up to 102,000 lncRNAs were known to regulate various processes by various interactions with DNA, mRNA, and protein in cancer cells. Recent studies have revealed urine, and blood-based lncRNAs as key diagnostic markers. For example, PCA3 was approved for the detection of prostate cancer. Similarly, lncRNAs such as CASC2, and CRNDE were found to be effective biomarkers against GBM. HOTAIR, and MALAT1 in case of breast and gastric cancers, etc. Several miRNAs and lncRNAs act as oncogenes as well as tumor suppressors. The important miRNAs include miR-10b in GBM and breast cancers, miR-21 in the case of B-chronic lymphocytic leukemia, miR-155 in the case of lymphoma, and miR-221 in liver cancer. Similarly, the key lncRNAs include GAS5 in the case of GBM, MEG3 in lung cancer, MALAT1 in lung cancer, and breast cancer, etc. Targeting some miRNAs and lncRNAs with RNA-interfering molecules in GBM cell lines and GBM mouse models has resulted in beneficial effects. However, delivering RNAi molecules to the brain is challenging as BBB precludes most substances' passage into the brain. Developing a complete network of all ncRNAs involved in glioma formation, and progression could supplement other therapeutic approaches such as immunotherapy and gene therapy.
Data availability
Data sharing is not applicable to this article as no datasets were generated or analysed during the current study.
References
Abal M, Planaguma J, Gil-Moreno A, Monge M, Gonzalez M, Baro T, Garcia A, Castellvi J, Ramon Y Cajal S, Xercavins J, Alameda F, Reventos J (2006) Molecular pathology of endometrial carcinoma: transcriptional signature in endometrioid tumors. Histol Histopathol 21(2):197–204 https://doi.org/10.14670/HH-21.197
Agemy L, Friedmann-Morvinski D, Kotamraju VR, Roth L, Sugahara KN, Girard OM et al (2011) Targeted nanoparticle-enhanced proapoptotic peptide as potential therapy for glioblastoma. Proc Natl Acad Sci 108(42):17450–17455. https://doi.org/10.1073/pnas.1114518108
Alexiou GA, Vartholomatos E, Tsamis IK, Peponi E, Markopoulos G, Papathanasopoulou AV, Tasiou I, Ragos V, Tsekeris P, Kyritsis AP, Galani V (2019) Combination treatment for glioblastoma with temozolomide, DFMO and radiation. J BUON 24(1):397–404
Baeza N, Weller M, Yonekawa Y, Kleihues P, Ohgaki H (2003) PTEN methylation and expression in glioblastomas. Acta Neuropathol 106(5):479–485. https://doi.org/10.1007/s00401-003-0748-4
Bahuau M, Vidaud D, Jenkins RB, Bièche I, Kimmel DW, Assouline B, Smith JS, Alderete B, Cayuela JM, Harpey JP, Caille B, Vidaud M (1998) Germ-line deletion involving the INK4 locus in familial proneness to melanoma and nervous system tumors. Can Res 58(11):2298–2303
Baker SJ, Fearon ER, Nigro JM, Hamilton SR, Preisinger AC, Jessup JM, van Tuinen P, Ledbetter DH, Barker DF, Nakamura Y, White R, Vogelstein B (1989) Chromosome 17 deletions and p53 gene mutations in colorectal carcinomas. Science (New York, N.Y.) 244(4901):217–221 https://doi.org/10.1126/science.2649981
Baker DL, Youssef OA, Chastkofsky MI, Dy DA, Terns RM, Terns MP (2005) RNA-Guided RNA modification: functional organization of the archaeal H/ACA RNP. Genes Dev 19(10):1238–1248. https://doi.org/10.1101/gad.1309605
Batchelor TT, Mulholland P, Neyns B, Nabors LB, Campone M, Wick A et al (2013) Phase III randomized trial comparing the efficacy of cediranib as monotherapy, and in combination with lomustine, versus lomustine alone in patients with recurrent glioblastoma. J Clin Oncol 31:3212–3218. https://doi.org/10.1200/JCO.2012.47.2464
Bazzichetto C, Conciatori F, Pallocca M, Falcone I, Fanciulli M, Cognetti F, Milella M, Ciuffreda L (2019) PTEN as a Prognostic/Predictive Biomarker in Cancer: An Unfulfilled Promise? Cancers 11(4):435. https://doi.org/10.3390/cancers11040435
Biernat W, Kleihues P, Yonekawa Y, Ohgaki H (1997) Amplification and overexpression of MDM2 in primary (de novo) glioblastomas. J Neuropathol Exp Neurol 56(2):180–185. https://doi.org/10.1097/00005072-199702000-00009
Binabaj MM, Bahrami A, ShahidSales S, Joodi M, Joudi Mashhad M, Hassanian SM, Anvari K, Avan A (2018) The prognostic value of MGMT promoter methylation in glioblastoma: A meta-analysis of clinical trials. J Cell Physiol 233(1):378–386. https://doi.org/10.1002/jcp.25896
Birner P, Piribauer M, Fischer I, Gatterbauer B, Marosi C, Ungersböck K, Rössler K, Budka H, Hainfellner JA (2002) Prognostic relevance of p53 protein expression in glioblastoma. Oncol Rep 9(4):703–707
Bobustuc GC, Baker CH, Limaye A, Jenkins WD, Pearl G, Avgeropoulos NG, Konduri SD (2010) Levetiracetam enhances p53-mediated MGMT inhibition and sensitizes glioblastoma cells to temozolomide. Neuro-Oncology 12(9):917–927. https://doi.org/10.1093/neuonc/noq044
Bobyk L, Edouard M, Deman P, Vautrin M, Pernet-Gallay K, Delaroche J, Adam JF, Estève F, Ravanat JL, Elleaume H (2013) Photoactivation of gold nanoparticles for glioma treatment. Nanomed Nanotechnol Biol Med 9(7):1089–1097. https://doi.org/10.1016/j.nano.2013.04.007
Boccaletto P, Machnicka MA, Purta E, Piatkowski P, Baginski B, Wirecki TK, de Crécy-Lagard V, Ross R, Limbach PA, Kotter A, Helm M, Bujnicki J M (2018) MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res 46(D1):D303–D307 https://doi.org/10.1093/nar/gkx1030
Borandeh S, Hosseinbeigi H, Abolmaali SS, Monajati M, Tamaddon AM (2021) Steric stabilization of β-cyclodextrin functionalized graphene oxide by host-guest chemistry: A versatile supramolecule for dual-stimuli responsive cellular delivery of doxorubicin. J Drug Deliv Sci Technol 63:102536
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR et al (2013) The somatic genomic landscape of glioblastoma. Cell 155(2):462–477. https://doi.org/10.1016/j.cell.2013.09.034
Brock CS, Newlands ES, Wedge SR, Bower M, Evans H, Colquhoun I, Roddie M, Glaser M, Brampton MH, Rustin GJ (1998) Phase I trial of temozolomide using an extended continuous oral schedule. Can Res 58(19):4363–4367
Bukhari B, Naveed M, Makhdoom SI, Jabeen K, Asif MF, Batool H et al (2021) A comparison between organic and inorganic nanoparticles: Prime nanoparticles for tumor curation. NANO 16(13):2130011. https://doi.org/10.1142/S1793292021300115
Cao S, Wang Y, Li J, Lv M, Niu H, Tian Y (2016) Tumor-suppressive function of long non-coding RNA MALAT1 in glioma cells by suppressing miR-155 expression and activating FBXW7 function. Am J Cancer Res 6(11):2561–2574
Carella A, Tejedor JR, García MG, Urdinguio RG, Bayón GF, Sierra M, López V, García-Toraño E, Santamarina-Ojeda P, Pérez RF, Bigot T, Mangas C, Corte-Torres MD, Sáenz-de-Santa-María I, Mollejo M, Meléndez B, Astudillo A, Chiara MD, Fernández AF, Fraga MF (2020) Epigenetic downregulation of TET3 reduces genome-wide 5hmC levels and promotes glioblastoma tumorigenesis. Int J Cancer 146(2):373–387. https://doi.org/10.1002/ijc.32520
Carrillo JA, Hsu FP, Delashaw J, Bota D (2014) Efficacy and safety of bevacizumab and etoposide combination in patients with recurrent malignant gliomas who have failed bevacizumab. Rev Health Care 5(1):23–32
Cavenee WK, Hastie ND, Stanbridge EJ (eds) (1989) Current Communications in Molecular Biology: Recessive Oncogenes and Tumor Suppression. Cold Spring Harbor Press, New York
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, Sander C, Schultz N (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2(5):401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Chen W, Xu XK, Li JL, Kong KK, Li H, Chen C, He J, Wang F, Li P, Ge XS, Li FC (2017) MALAT1 is a prognostic factor in glioblastoma multiforme and induces chemoresistance to temozolomide through suppressing miR-203 and promoting thymidylate synthase expression. Oncotarget 8(14):22783–22799 https://doi.org/10.18632/oncotarget.15199
Chen X, Li Y, Zuo C, Zhang K, Lei X, Wang J, Yang Y, Zhang J, Ma K, Wang S, Mu N, Yang C, Xian J, Feng H, Tang R, Chen T (2021) Long Non-Coding RNA H19 Regulates Glioma Cell Growth and Metastasis via miR-200a-Mediated CDK6 and ZEB1 Expression. Front Oncol 11:757650 https://doi.org/10.3389/fonc.2021.757650.
Chen Y, Zhao F, Cui D, Jiang R, Chen J, Huang Q, Shi J (2018) HOXD-AS1/miR-130a sponge regulates glioma development by targeting E2F8. Int J Cancer 142(11):2313–2322. https://doi.org/10.1002/ijc.31262
Chen J, Wang Y, Wang C, Hu JF, Li W (2020) LncRNA functions as a new emerging epigenetic factor in determining the fate of stem cells. Front Genet 11:277. https://doi.org/10.3389/fgene.2020.00277
Cheng H, Zhao H, Xiao X, Huang Q, Zeng W, Tian B, Ma T, Lu D, Jin Y, Li Y (2021) Long Non-coding RNA MALAT1 Upregulates ZEB2 Expression to Promote Malignant Progression of Glioma by Attenuating miR-124. Mol Neurobiol 58(3):1006–1016. https://doi.org/10.1007/s12035-020-02165-0
Chi AS, Batchelor TT, Yang D, Dias-Santagata D, Borger DR, Ellisen LW et al (2013) BRAF V600E Mutation Identifies a Subset of Low-Grade Diffusely Infiltrating Gliomas in Adults. J Clin Oncol off J Am Soc Clin Oncol 31(14):e233–e236. https://doi.org/10.1200/jco.2012.46.0220
Chinot OL, Wick W, Mason W, Henriksson R, Saran F, Nishikawa R, Carpentier AF, Hoang-Xuan K, Kavan P, Cernea D, Brandes AA, Hilton M, Abrey L, Cloughesy T (2014) Bevacizumab plus radiotherapy-temozolomide for newly diagnosed glioblastoma. N Engl J Med 370(8):709–722. https://doi.org/10.1056/NEJMoa1308345
Coccini T, Grandi S, Lonati D, Locatelli C, De Simone U (2015) Comparative cellular toxicity of titanium dioxide nanoparticles on human astrocyte and neuronal cells after acute and prolonged exposure. Neurotoxicology 48:77–89. https://doi.org/10.1016/j.neuro.2015.03.006
Cohen AL, Holmen SL, Colman H (2013) IDH1 and IDH2 mutations in gliomas. Curr Neurol Neurosci Rep 13(5):345. https://doi.org/10.1007/s11910-013-0345-4
Cui X, Liang Z, Shen L, Zhang Q, Bao S, Geng Y, Zhang B, Leo V, Vardy LA, Lu T, Gu X, Yu H (2017) 5-Methylcytosine RNA Methylation in Arabidopsis Thaliana. Mol Plant 10(11):1387–1399. https://doi.org/10.1016/j.molp.2017.09.013
Dai Z, Arévalo MT, Li J, Zeng M (2014) Addition of poly (propylene glycol) to multiblock copolymer to optimize siRNA delivery. Bioengineered 5(1):30–37. https://doi.org/10.4161/bioe.27339
Daniyal M, Liu B, Wang W (2020) Comprehensive Review on Graphene Oxide for Use in Drug Delivery System. Curr Med Chem 27(22):3665–3685. https://doi.org/10.2174/13816128256661902011296290
David R, Burgess A, Parker B, Li J, Pulsford K, Sibbritt T, Preiss T, Searle IR (2017) Transcriptome-Wide Mapping of RNA 5-Methylcytosine in Arabidopsis mRNAs and Non-coding RNAs. Plant Cell 29(3):445–460. https://doi.org/10.1105/tpc.16.00751
de la Iglesia N, Konopka G, Puram SV, Chan JA, Bachoo RM, You MJ, Levy DE, Depinho RA, Bonni A (2008) Identification of a PTEN-regulated STAT3 brain tumor suppressor pathway. Genes Dev 22(4):449–462. https://doi.org/10.1101/gad.1606508
Devi S, Kumar M, Tiwari A, Tiwari V, Kaushik D, Verma R et al. (2022) Quantum Dots: An Emerging Approach for Cancer Therapy. Front Mater 8:798440 https://doi.org/10.3389/fmats
Dong Z, Cui H (2020) The Emerging Roles of RNA Modifications in Glioblastoma. Cancers 12(3):736. https://doi.org/10.3390/cancers12030736
Du P, Zhao H, Peng R, Liu Q, Yuan J, Peng G, Liao Y (2017) LncRNA-XIST interacts with miR-29c to modulate the chemoresistance of glioma cells to TMZ through DNA mismatch repair pathway. Biosci Rep 37(5):BSR20170696 10.1042/BSR20170696
El Fatimy R, Subramanian S, Uhlmann EJ, Krichevsky AM (2017) Genome editing reveals glioblastoma addiction to microRNA-10b. Mol Ther 25(2):368–378. https://doi.org/10.1016/j.ymthe.2016.11.004
Fadhlullah S, Halim N, Yeo J, Ho R, Um P, Ang BT, Tang C, Ng WH, Virshup DM, Ho I (2019) Pathogenic mutations in neurofibromin identifies a leucine-rich domain regulating glioma cell invasiveness. Oncogene 38(27):5367–5380. https://doi.org/10.1038/s41388-019-0809-3
Fallah J, Brave MH, Weinstock C, Mehta GU, Bradford D, Gittleman H et al (2022) FDA Approval Summary: Belzutifan for von Hippel-Lindau Disease-Associated Tumors. Clin Cancer Res 28(22):4843–4848. https://doi.org/10.1158/1078-0432.CCR-22-1054
Farwell JR, Dohrmann GJ, Flannery JT (1977) Central nervous system tumors in children cns tumors in children. Cancer 40(6):3123–3132. https://doi.org/10.1002/1097-0142(197712)40:6%3C3123::AID-CNCR2820400656%3E3.0.CO;2-6
Finlay CA, Hinds PW, Levine AJ (1989) The p53 proto-oncogene can act as a suppressor of transformation. Cell 57(7):1083–1093. https://doi.org/10.1016/0092-8674(89)90045-7
Fogelman DR, Wolff RA, Kopetz S, Javle M, Bradley C, Mok I et al (2011) Evidence for the Efficacy of Iniparib, a PARP-1 Inhibitor, in BRCA2-Associated Pancreatic Cancer. Anticancer Res 31(4):1417–1420
Fu L, Guerrero CR, Zhong N, Amato NJ, Liu Y, Liu S, Cai Q, Ji D, Jin SG, Niedernhofer LJ, Pfeifer GP, Xu GL, Wang Y (2014) Tet-mediated formation of 5-hydroxymethylcytosine in RNA. J Am Chem Soc 136(33):11582–11585. https://doi.org/10.1021/ja505305z
Fu Z, Luo W, Wang J, Peng T, Sun G, Shi J, Li Z, Zhang B (2017) Malat1 activates autophagy and promotes cell proliferation by sponging miR-101 and upregulating STMN1, RAB5A and ATG4D expression in Glioma. Biochem Biophys Res Commun 492(3):480–486. https://doi.org/10.1016/j.bbrc.2017.08.070
Fulci G, Labuhn M, Maier D, Lachat Y, Hausmann O, Hegi ME, Janzer RC, Merlo A, Van Meir EG (2000) p53 gene mutation and ink4a-arf deletion appear to be two mutually exclusive events in human glioblastoma. Oncogene 19(33):3816–3822. https://doi.org/10.1038/sj.onc.1203700
Furuichi Y, LaFiandra A, Shatkin AJ (1977) 5’-Terminal structure and mRNA stability. Nature 266(5599):235–239. https://doi.org/10.1038/266235a0
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6(269):l1 https://doi.org/10.1126/scisignal.2004088
García MG, Carella A, Urdinguio RG, Bayón GF, Lopez V, Tejedor JR, Sierra MI, García-Toraño E, Santamarina P, Perez RF, Mangas C, Astudillo A, Corte-Torres MD, Sáenz-de-Santa-María I, Chiara MD, Fernández AF, Fraga MF (2018) Epigenetic dysregulation of TET2 in human glioblastoma. Oncotarget 9(40):25922–25934 https://doi.org/10.18632/oncotarget.25406
Garzon R, Heaphy CE, Havelange V, Fabbri M, Volinia S, Tsao T et al (2009) MicroRNA 29b functions in acute myeloid leukemia. Blood J Am Soc Hematol 114(26):5331–5341. https://doi.org/10.1182/blood-2009-03-211938
Ge J, Liu H, Yu YT (2010) Regulation of pre-mRNA splicing in Xenopus oocytes by targeted 2'-O-methylation. RNA (New York, N.Y.) 16(5):1078–1085 https://doi.org/10.1261/rna.2060210
Geula S, Moshitch-Moshkovitz S, Dominissini D, Mansour AA, Kol N, Salmon-Divon M, Hershkovitz V, Peer E, Mor N, Manor YS, Ben-Haim MS, Eyal E, Yunger S, Pinto Y, Jaitin DA, Viukov S, Rais Y, Krupalnik V, Chomsky E, Zerbib M, et al (2015) Stem cells. m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science (New York, N.Y.) 347(6225):1002–1006 https://doi.org/10.1126/science.1261417
Ghaffari M, Dehghan G, Abedi-Gaballu F, Kashanian S, Baradaran B, Ezzati Nazhad Dolatabadi J, Losic D (2018) Surface functionalized dendrimers as controlled-release delivery nanosystems for tumor targeting. Eur J Pharm Sci 122:311–330https://doi.org/10.1016/j.ejps.2018.07.020
Gilbert MR, Dignam JJ, Armstrong TS, Wefel JS, Blumenthal DT, Vogelbaum MA, Colman H, Chakravarti A, Pugh S, Won M, Jeraj R, Brown PD, Jaeckle KA, Schiff D, Stieber VW, Brachman DG, Werner-Wasik M, Tremont-Lukats IW, Sulman EP, Aldape KD et al (2014) A randomized trial of bevacizumab for newly diagnosed glioblastoma. N Engl J Med 370(8):699–708. https://doi.org/10.1056/NEJMoa1308573
Goldstein M, Rudra S, Dahiya S, Tsien C, Huang J (2019) Prognostic value of EGFR amplification in glioblastoma patients treated with radiation therapy and concurrent temozolomide. Int J Radiat Oncol Biol Phys 105(1):E98–E99
Goll MG, Kirpekar F, Maggert KA, Yoder JA, Hsieh CL, Zhang X, Golic KG, Jacobsen SE, Bestor TH (2006) Methylation of tRNA-Asp by the DNA methyltransferase homolog Dnmt2. Science (New York, N.Y.) 311(5759):395–398 https://doi.org/10.1126/science.1120976
Gorovets D, Kannan K, Shen R, Kastenhuber ER, Islamdoust N, Campos C, Pentsova E, Heguy A, Jhanwar SC, Mellinghoff IK, Chan TA, Huse JT (2012) IDH mutation and neuroglial developmental features define clinically distinct subclasses of lower grade diffuse astrocytic glioma. Clin Cancer Res 18(9):2490–2501. https://doi.org/10.1158/1078-0432.CCR-11-2977
Guy MP, Phizicky EM (2014) Two-subunit enzymes involved in eukaryotic post-transcriptional tRNA modification. RNA Biol 11(12):1608–1618. https://doi.org/10.1080/15476286.2015.1008360
Hadjipanayis CG, Stummer W (2019) 5-ALA and FDA approval for glioma surgery. J Neurooncol 141(3):479–486. https://doi.org/10.1007/s11060-019-03098-y
Hambarde S, Sharpe M, Baskin D, Helekar S (2020) CBIO-07. Cell Death Induced by an Oscillating Magnetic Field in Patient Derived Glioblastoma Cells is Mediated by Reactive Oxygen Species. Neuro-Oncol 22(Suppl 2):ii17
Han F, Hu R, Yang H, Liu J, Sui J, Xiang X, Wang F, Chu L, Song S (2016a) PTEN gene mutations correlate to poor prognosis in glioma patients: a meta-analysis. OncoTargets Ther 9:3485–3492https://doi.org/10.2147/OTT.S99942
Han K, Peyret T, Marchand M, Quartino A, Gosselin NH, Girish S et al (2016b) Population pharmacokinetics of bevacizumab in cancer patients with external validation. Cancer Chemother Pharmacol 78:341–351
Han B, Cai J, Gao W, Meng X, Gao F, Wu P et al (2018) Loss of ATRX Suppresses ATM Dependent DNA Damage Repair by Modulating H3K9me3 to Enhance Temozolomide Sensitivity in Glioma. Cancer Lett 419:280–290. https://doi.org/10.1016/j.canlet.2018.01.056
Han N, Yang L, Zhang X, Zhou Y, Chen R, Yu Y et al. (2019) LncRNA MATN1-AS1 prevents glioblastoma cell from proliferation and invasion via RELA regulation and MAPK signaling pathway. Ann Transl Med 7(23) https://doi.org/10.21037/2Fatm.2019.11.36
Hatanpaa KJ, Burma S, Zhao D, Habib AA (2010) Epidermal growth factor receptor in Glioma: signal transduction, neuropathology, imaging, and radioresistance. Neoplasia (New York, N.Y.) 12(9):675–684 https://doi.org/10.1593/neo.10688
He J, Shi J, Zhang K, Xue J, Li J, Yang J et al (2017) Sox2 inhibits Wnt-β-catenin signaling and metastatic potency of cisplatin-resistant lung adenocarcinoma cells. Mol Med Rep 15(4):1693–1701. https://doi.org/10.3892/mmr.2017.6170
He Z, You C, Zhao D (2018) Long non-coding RNA UCA1/miR-182/PFKFB2 axis modulates glioblastoma-associated stromal cells-mediated glycolysis and invasion of glioma cells. Biochem Biophys Res Commun 500(3):569–576. https://doi.org/10.1016/j.bbrc.2018.04.091
Heidenreich B, Rachakonda PS, Hosen I, Volz F, Hemminki K, Weyerbrock A, Kumar R (2015) TERT promoter mutations and telomere length in adult malignant gliomas and recurrences. Oncotarget 6:10617–33 https://doi.org/10.18632/2Foncotarget.3329
Helekar SA, Voss HU (2016) Transcranial brain stimulation with rapidly spinning high-field permanent magnets. IEEE Access 4:2520–2528
Helekar S, Hambarde S, Baskin D, Sharpe M (2020) EXTH-13. Potent Anticancer Effects of a New Wearable Noninvasive Oncomagnetic Device: Cellular Mechanisms of Action. Neuro-Oncol 22(Suppl 2):ii89
Helekar SA, Convento S, Nguyen L, John BS, Patel A, Yau JM, Voss HU (2018) The strength and spread of the electric field induced by transcranial rotating permanent magnet stimulation in comparison with conventional transcranial magnetic stimulation. J Neurosci Methods 309:153–160. https://doi.org/10.1016/j.jneumeth.2018.09.002
Henson JW, Schnitker BL, Correa KM, von Deimling A, Fassbender F, Xu HJ, Benedict WF, Yandell DW, Louis DN (1994) The retinoblastoma gene is involved in the malignant progression of astrocytomas. Ann Neurol 36(5):714–721. https://doi.org/10.1002/ana.410360505
Hirano H, Yonezawa H, Yunoue S, Habu M, Uchida H, Yoshioka T, Kishida S, Kishida M, Oyoshi T, Fujio S, Sugata S, Yamahata H, Hanaya R, Arita K (2014) Immunoreactivity of Wnt5a, Fzd2, Fzd6, and Ryk in glioblastoma: evaluative methodology for DAB chromogenic immunostaining. Brain Tumor Pathol 31(2):85–93. https://doi.org/10.1007/s10014-013-0153-1
Hirose Y, Berger MS, Pieper RO (2001) p53 effects both the duration of G2/M arrest and the fate of temozolomide-treated human glioblastoma cells. Cancer Res 61(5):1957–1963
Hobbs J, Nikiforova MN, Fardo DW, Bortoluzzi S, Cieply K, Hamilton RL, Horbinski C (2012) Paradoxical relationship between the degree of EGFR amplification and outcome in glioblastomas. Am J Surg Pathol 36(8):1186–1193. https://doi.org/10.1097/PAS.0b013e3182518e12
Hottinger AF, Pacheco P, Stupp R (2016) Tumor treating fields: A novel treatment modality and its use in brain tumors. Neuro Oncol 18(10):1338–1349. https://doi.org/10.1093/neuonc/now182
Hu B, Wang Q, Wang YA, Hua S, Sauvé CEG, Ong D et al (2016) Epigenetic activation of WNT5A drives glioblastoma stem cell differentiation and invasive growth. Cell 167(5):1281–1295. https://doi.org/10.1016/j.cell.2016.10.039
Hu Q, Yin J, Zeng A, Jin X, Zhang Z, Yan W, You Y (2018) H19 Functions as a Competing Endogenous RNA to Regulate EMT by Sponging miR-130a-3p in Glioma. Cell Physiol Biochem 50(1):233–245. https://doi.org/10.1159/000494002
Huang W, Lan MD, Qi CB, Zheng SJ, Wei SZ, Yuan BF, Feng YQ (2016) Formation and determination of the oxidation products of 5-methylcytosine in RNA. Chem Sci 7(8):5495–5502. https://doi.org/10.1039/c6sc01589a
Huang PH, Cavenee WK, Furnari FB, White FM (2007) Uncovering therapeutic targets for glioblastoma: a systems biology approach. Cell Cycle (Georgetown, Tex.) 6(22):2750–2754 https://doi.org/10.4161/cc.6.22.4922
Huang Z, Zhao X, Wu X, Xiang L, Yuan Y, Zhou S, Yu W (2019) LncRNA UCA1 facilitated cell growth and invasion through the miR-206/CLOCK axis in Glioma. Cancer Cell Int 19:316. https://doi.org/10.1186/s12935-019-1023-7
Hurley JB, Simon MI, Teplow DB, Robishaw JD, Gilman AG (1984) Homologies between signal transducing G proteins and ras gene products. Science (New York, N.Y.) 226(4676):860–862 https://doi.org/10.1126/science.6436980
Huse JT, Holland EC (2010) Targeting brain cancer: advances in the molecular pathology of malignant Glioma and medulloblastoma. Nat Rev Cancer 10(5):319–331. https://doi.org/10.1038/nrc2818
Jacobs VL, Valdes PA, Hickey WF, De Leo JA (2011) Current review of in vivo GBM rodent models: emphasis on the CNS-1 tumour model. ASN Neuro 3(3):AN20110014 https://doi.org/10.1042/AN20110014
Jain AK, Xi Y, McCarthy R, Allton K, Akdemir KC, Patel LR et al (2016) LncPRESS1 is a p53-regulated LncRNA that safeguards pluripotency by disrupting SIRT6-mediated de-acetylation of histone H3K56. Mol Cell 64(5):967–981. https://doi.org/10.1016/j.molcel.2016.10.039
Jeuken JW, Deimling AV, Wesseling P (2004) Molecular pathogenesis of oligodendroglial tumors. J Neurooncol 70:161–181. https://doi.org/10.1007/s11060-004-2748-1
Ji L, Chen X (2012) Regulation of small RNA stability: methylation and beyond. Cell Res 22(4):624–636. https://doi.org/10.1038/cr.2012.36
Ji W, Wang Q, Yang J (2020) LncRNA HOXD-AS1 promotes the metastasis of human hepatocellular carcinoma via modulating miR-326/SLC27A4. Cancer Cell Int 20:161. https://doi.org/10.1186/s12935-020-01217-8
Kato T, Natsume A, Toda H, Iwamizu H, Sugita T, Hachisu R et al (2010) Efficient delivery of liposome-mediated MGMT-siRNA reinforces the cytotoxity of temozolomide in GBM-initiating cells. Gene Ther 17(11):1363–1371. https://doi.org/10.1038/gt.2010.88
Keaney J, Campbell M (2015) The dynamic blood–brain barrier. FEBS J 282(21):4067–4079. https://doi.org/10.1111/febs.13412
Ke J, Yao YL, Zheng J, Wang P, Liu YH, Ma J, Li Z, Liu XB, Li ZQ, Wang ZH, Xue YX (2015) Knockdown of long non-coding RNA HOTAIR inhibits malignant biological behaviors of human glioma cells via modulation of miR-326. Oncotarget 6(26):21934–21949 https://doi.org/10.18632/oncotarget.4290
Kim SS, Rait A, Kim E, Pirollo KF, Chang EH (2015) A tumor-targeting p53 nanodelivery system limits chemoresistance to temozolomide prolonging survival in a mouse model of glioblastoma multiforme. Nanomedicine 11(2):301–311. https://doi.org/10.1016/j.nano.2014.09.005
Kim J, Siverly AN, Chen D, Wang M, Yuan Y, Wang Y et al (2016) Ablation of miR-10b Suppresses Oncogene-Induced Mammary Tumorigenesis and Metastasis and Reactivates Tumor-Suppressive PathwaysGenetic Dissection of miR-10b and Breast Cancer. Can Res 76(21):6424–6435. https://doi.org/10.1158/0008-5472.CAN-16-1571
Kozielski KL, Ruiz-Valls A, Tzeng SY, Guerrero-Cázares H, Rui Y, Li Y et al (2019) Cancer-selective nanoparticles for combinatorial siRNA delivery to primary human GBM in vitro and in vivo. Biomaterials 209:79–87. https://doi.org/10.1016/j.biomaterials.2019.04.020
Kulke MH, Hornick JL, Frauenhoffer C, Hooshmand S, Ryan DP, Enzinger PC et al (2009) O 6-methylguanine DNA methyltransferase deficiency and response to temozolomide-based therapy in patients with neuroendocrine tumors. Clin Cancer Res 15(1):338–345. https://doi.org/10.1158/1078-0432.CCR-08-1476
Kunoh T, Shimura T, Kasai T, Matsumoto S, Mahmud H, Khayrani AC, Seno M, Kunoh H, Takada J (2019) Use of DNA-generated gold nanoparticles to radiosensitize and eradicate radioresistant glioma stem cells. Nanotechnology 30(5):055101 https://doi.org/10.1088/1361-6528/aaedd5
Lalezari S, Chou AP, Tran A, Solis OE, Khanlou N, Chen W, Li S, Carrillo JA, Chowdhury R, Selfridge J, Sanchez DE, Wilson RW, Zurayk M, Lalezari J, Lou JJ, Ormiston L, Ancheta K, Hanna R, Miller P, Piccioni D et al (2013) Combined analysis of O6-methylguanine-DNA methyltransferase protein expression and promoter methylation provides optimized prognostication of glioblastoma outcome. Neuro Oncol 15(3):370–381. https://doi.org/10.1093/neuonc/nos308
Larsson I, Uhlén M, Zhang C, Mardinoglu A (2020) Genome-scale metabolic modeling of glioblastoma reveals promising targets for drug development. Front Genet 11:381. https://doi.org/10.3389/fgene.2020.00381
Lassen U, Mau-Sørensen M, Poulsen HS (2014) Orphan drugs in glioblastoma multiforme: a review. Orphan Drugs Res Rev 4:83–91
Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A et al (2013) Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499(7457):214–218. https://doi.org/10.1038/nature12213
Lee M, Kim B, Kim VN (2014) Emerging roles of RNA modification: m(6)A and U-tail. Cell 158(5):980–987. https://doi.org/10.1016/j.cell.2014.08.005
Lespagnard L, Gancberg D, Rouas G, Leclercq G, de Saint-Aubain Somerhausen N, Di Leo A, Piccart M, Verhest A, Larsimont D (1999) Tumor-infiltrating dendritic cells in adenocarcinomas of the breast: a study of 143 neoplasms with a correlation to usual prognostic factors and to clinical outcome. Int J Cancer 84(3):309–314. https://doi.org/10.1002/(sici)1097-0215(19990621)84:3%3c309::aid-ijc19%3e3.0.co;2-3
Leu S, von Felten S, Frank S, Vassella E, Vajtai I, Taylor E, Schulz M, Hutter G, Hench J, Schucht P, Boulay JL, Mariani L (2013) IDH/MGMT-driven molecular classification of low-grade Glioma is a strong predictor for long-term survival. Neuro Oncol 15(4):469–479. https://doi.org/10.1093/neuonc/nos317
Li H, Guan C (2020a) HOTAIR inhibits the proliferation of glioblastoma cells by targeting miR-219. Cancer Biomark Sect A Dis Mark 28(1):41–47. https://doi.org/10.3233/CBM-190467
Li H, Guan C (2020b) HOTAIR inhibits the proliferation of glioblastoma cells by targeting miR-219. Cancer Biomark 28(1):41–47. https://doi.org/10.3233/CBM-190467
Li Q, Guo J, Wang W, Wang D (2017a) Relationship between MGMT gene expression and treatment effectiveness and prognosis in Glioma. Oncol Lett 14(1):229–233. https://doi.org/10.3892/ol.2017.6123
Li Z, Xu C, Ding B, Gao M, Wei X, Ji N (2017b) Long non-coding RNA MALAT1 promotes proliferation and suppresses apoptosis of glioma cells through derepressing Rap1B by sponging miR-101. J Neurooncol 134(1):19–28. https://doi.org/10.1007/s11060-017-2498-5
Li Y, Wang X, Zhao Z, Shang J, Li G, Zhang R (2021) LncRNA NEAT1 promotes glioma cancer progression via regulation of miR-98-5p/BZW1. Biosci Rep 41(7):BSR20200767 10.1042/BSR20200767
Li J, Liang H, Liu J, Wang Z (2018a) Poly (amidoamine) (PAMAM) dendrimer mediated delivery of drug and pDNA/siRNA for cancer therapy. Int J Pharm 546(1–2):215–225. https://doi.org/10.1016/j.ijpharm.2018.05.045
Li J, Liang R, Song C, Xiang Y, Liu Y (2018b) Prognostic significance of epidermal growth factor receptor expression in glioma patients. Onco Targets Ther 11:731–742. https://doi.org/10.2147/OTT.S155160
Liao Y, Shen L, Zhao H, Liu Q, Fu J, Guo Y, Peng R, Cheng L (2017) LncRNA CASC2 Interacts With miR-181a to Modulate Glioma Growth and Resistance to TMZ Through PTEN Pathway. J Cell Biochem 118(7):1889–1899. https://doi.org/10.1002/jcb.25910
Liao K, Lin Y, Gao W, Xiao Z, Medina R, Dmitriev P, Cui J, Zhuang Z, Zhao X, Qiu Y, Zhang X, Ge J, Guo L (2019) Blocking lncRNA MALAT1/miR-199a/ZHX1 Axis Inhibits Glioblastoma Proliferation and Progression. Mol Ther Nucleic Acids 18:388–399. https://doi.org/10.1016/j.omtn.2019.09.005
Lin S, Liu Q, Lelyveld VS, Choe J, Szostak JW, Gregory RI (2018) Mettl1/Wdr4-Mediated m7G tRNA Methylome Is Required for Normal mRNA Translation and Embryonic Stem Cell Self-Renewal and Differentiation. Mol Cell 71(2):244-255.e5. https://doi.org/10.1016/j.molcel.2018.06.001
Liu HW, Hu SH, Chen YW, Chen SY (2012) Characterization and drug release behavior of highly responsive chip-like electrically modulated reduced graphene oxide–poly (vinyl alcohol) membranes. J Mater Chem 22(33):17311–17320
Liu GY, Zhao GN, Chen XF, Hao DL, Zhao X, Lv X, Liu DP (2016) The long non-coding RNA Gm15055 represses Hoxa gene expression by recruiting PRC2 to the gene cluster. Nucleic Acids Res 44(6):2613–2627. https://doi.org/10.1093/nar/gkv1315
Liu YJ, Ma YC, Zhang WJ, Yang ZZ, Liang DS, Wu ZF, Qi XR (2017) Combination therapy with micellarized cyclopamine and temozolomide attenuate glioblastoma growth through Gli1 down-regulation. Oncotarget 8(26):42495–42509 https://doi.org/10.18632/oncotarget.17205
Liu X, Zheng J, Xue Y, Yu H, Gong W, Wang P, Li Z, Liu Y (2018) PIWIL3/OIP5-AS1/miR-367-3p/CEBPA feedback loop regulates the biological behavior of glioma cells. Theranostics 8(4):1084–1105. https://doi.org/10.7150/thno.21740
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A et al (2007) The 2007 WHO Classification of Tumours of the Central Nervous System. Acta Neuropathol 114(2):97–109. https://doi.org/10.1007/s00401-007-0243-4
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK et al (2016) The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A Summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR (2005) MicroRNA expression profiles classify human cancers. Nature 435(7043):834–838. https://doi.org/10.1038/nature03702
Luo C, Quan Z, Zhong B, Zhang M, Zhou B, Wang S, Luo X, Tang C (2020) lncRNA XIST promotes glioma proliferation and metastasis through miR-133a/SOX4. Exp Ther Med 19(3):1641–1648. https://doi.org/10.3892/etm.2020.8426
Lutterbach J, Fauchon F, Schild SE, Chang SM, Pagenstecher A, Volk B et al (2002) Malignant pineal parenchymal tumors in adult patients: patterns of care and prognostic factors. Neurosurgery 51(1):44–56. https://doi.org/10.1097/00006123-200207000-00006
Lyu J, Yamamoto V, Lu W (2008) Cleavage of the Wnt receptor Ryk regulates neuronal differentiation during cortical neurogenesis. Dev Cell 15(5):773–780. https://doi.org/10.1016/j.devcel.2008.10.004
Ma L, Reinhardt F, Pan E, Soutschek J, Bhat B, Marcusson EG et al (2010) Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat Biotechnol 28(4):341–347. https://doi.org/10.1038/nbt.1618
Ma J, Wang P, Yao Y, Liu Y, Li Z, Liu X, Li Z, Zhao X, Xi Z, Teng H, Liu J, Xue Y (2016) Knockdown of long non-coding RNA MALAT1 increases the blood-tumor barrier permeability by upregulating miR-140. Biochem Biophys Acta 1859(2):324–338. https://doi.org/10.1016/j.bbagrm.2015.11.008
Ma R, Zhang BW, Zhang ZB, Deng QJ (2020) LncRNA MALAT1 knockdown inhibits cell migration and invasion by suppressing autophagy through miR-384/GOLM1 axis in Glioma. Eur Rev Med Pharmacol Sci 24(5):2601–2615. https://doi.org/10.26355/eurrev_202003_20529
Mahmoudi K, Bouras A, Bozec D, Ivkov R, Hadjipanayis C (2018) Magnetic hyperthermia therapy for the treatment of glioblastoma: a review of the therapy’s history, efficacy and application in humans. Int J Hyperth 34(8):1316–1328. https://doi.org/10.1080/02656736.2018.1430867
Metellus P, Coulibaly B, Colin C, de Paula AM, Vasiljevic A, Taieb D, Barlier A, Boisselier B, Mokhtari K, Wang XW, Loundou A, Chapon F, Pineau S, Ouafik L, Chinot O, Figarella-Branger D (2010) Absence of IDH mutation identifies a novel radiologic and molecular subtype of WHO grade II gliomas with dismal prognosis. Acta Neuropathol 120(6):719–729. https://doi.org/10.1007/s00401-010-0777-8
Meyer KD, Jaffrey SR (2017) Rethinking m6A Readers, Writers, and Erasers. Annu Rev Cell Dev Biol 33:319–342. https://doi.org/10.1146/annurev-cellbio-100616-060758
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 149(7):1635–1646. https://doi.org/10.1016/j.cell.2012.05.003
Miao FA, Chu K, Chen HR, Zhang M, Shi PC, Bai J, You YP (2019) Increased DKC1 expression in Glioma and its significance in tumor cell proliferation, migration and invasion. Invest New Drugs 37(6):1177–1186. https://doi.org/10.1007/s10637-019-00748-w
Moodie SA, Willumsen BM, Weber MJ, Wolfman A (1993) Complexes of Ras.GTP with Raf-1 and mitogen-activated protein kinase kinase. Science (New York, N.Y.) 260(5114):1658–1661 https://doi.org/10.1126/science.8503013
Moon HJ, Redman KL (2014) Trm4 and Nsun2 RNA:m5C methyltransferases form metabolite-dependent, covalent adducts with previously methylated RNA. Biochemistry 53(45):7132–7144. https://doi.org/10.1021/bi500882b
Morris LG, Kaufman AM, Gong Y, Ramaswami D, Walsh LA, Turcan Ş, Eng S, Kannan K, Zou Y, Peng L, Banuchi VE, Paty P, Zeng Z, Vakiani E, Solit D, Singh B, Ganly I, Liau L, Cloughesy TC, Mischel PS et al (2013) Recurrent somatic mutation of FAT1 in multiple human cancers leads to aberrant Wnt activation. Nat Genet 45(3):253–261. https://doi.org/10.1038/ng.2538
Mrugala MM, Crew LK, Fink JR, Spence AM (2012) Carboplatin and bevacizumab for recurrent malignant Glioma. Oncol Lett 4(5):1082–1086. https://doi.org/10.3892/ol.2012.839
Mullard A (2021) 2020 FDA drug approvals. Nat Rev Drug Discov 20(2):85–91
Nager M, Bhardwaj D, Cantí C, Medina L, Nogués P, Herreros J (2012) β-Catenin Signalling in Glioblastoma Multiforme and Glioma-Initiating Cells. Chemother Res Pract 2012:192362 https://doi.org/10.1155/2012/192362
Najafi S, Esmaeili S, Zhaleh H, Rahmati Y (2022) The role of IDH1 mutation on gene expression in glioblastoma. Inf Med Unlocked 28:100812
Newlands ES, Stevens MF, Wedge SR, Wheelhouse RT, Brock C (1997) Temozolomide: a review of its discovery, chemical properties, pre-clinical development and clinical trials. Cancer Treat Rev 23(1):35–61. https://doi.org/10.1016/s0305-7372(97)90019-0
Nieder C, Petersen S, Petersen C, Thames HD (2000) The challenge of p53 as prognostic and predictive factor in gliomas. Cancer Treat Rev 26(1):67–73. https://doi.org/10.1053/ctrv.1999.0145
Nishiyama N, Matsumura Y, Kataoka K (2016) Development of polymeric micelles for targeting intractable cancers. Cancer Sci 107(7):867–874. https://doi.org/10.1111/cas.12960
Nitta Y, Shimizu S, Shishido-Hara Y, Suzuki K, Shiokawa Y, Nagane M (2016) Nimotuzumab enhances temozolomide-induced growth suppression of glioma cells expressing mutant EGFR in vivo. Cancer Med 5(3):486–499. https://doi.org/10.1002/cam4.614
Noushmehr H, Weisenberger DJ, Diefes K, Phillips HS, Pujara K, Berman BP, Pan F, Pelloski CE, Sulman EP, Bhat KP, Verhaak RG, Hoadley KA, Hayes DN, Perou CM, Schmidt HK, Ding L, Wilson RK, Van Den Berg D, Shen H, Bengtsson H et al (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of Glioma. Cancer Cell 17(5):510–522. https://doi.org/10.1016/j.ccr.2010.03.017
Novartis A (2016) FDA approves afinitor (everolimus) for gi & lung nets. Oncol Times. https://doi.org/10.1097/01.COT.0000482575.83197.4b
Odogwu L, Mathieu L, Blumenthal G, Larkins E, Goldberg KB, Griffin N et al (2018) FDA approval summary: dabrafenib and trametinib for the treatment of metastatic non-small cell lung cancers harboring BRAF V600E mutations. Oncologist 23(6):740–745. https://doi.org/10.1634/theoncologist.2017-0642
Ohgaki H, Kleihues P (2013) The definition of primary and secondary glioblastoma. Clin Cancer Res 19:764–772. https://doi.org/10.1158/1078-0432.CCR-12-3002
O’Shaughnessy J, Osborne C, Pippen JE, Yoffe M, Patt D, Rocha C et al (2011) Iniparib Plus Chemotherapy in Metastatic Triple-Negative Breast Cancer. N Engl J Med 364(3):205–214. https://doi.org/10.1056/NEJMoa1011418
Ozpolat B, Sood AK, Lopez-Berestein G (2014) Liposomal siRNA nanocarriers for cancer therapy. Adv Drug Deliv Rev 66:110–116. https://doi.org/10.1016/j.addr.2013.12.008
Packer RJ, MacDonald T, Vezina G (2008) Central nervous system tumors. Pediatr Clin North Am 55(1):121–145. https://doi.org/10.1016/j.pcl.2007.10.010
Pan T, Xue M (2021) LncRNA-NNT-AS1 contributes to the progression of Glioma by miR-582-5p/EZH2 axis. Cytotechnology 73(3):473–482. https://doi.org/10.1007/s10616-021-00471-6
Pandith AA, Qasim I, Zahoor W, Shah P, Bhat AR, Sanadhya D, Shah ZA, Naikoo NA (2018) Concordant association validates MGMT methylation and protein expression as favorable prognostic factors in glioma patients on alkylating chemotherapy (Temozolomide). Sci Rep 8(1):6704. https://doi.org/10.1038/s41598-018-25169-2
Pandolfini L, Barbieri I, Bannister AJ, Hendrick A, Andrews B, Webster N, Murat P, Mach P, Brandi R, Robson SC, Migliori V, Alendar A, d’Onofrio M, Balasubramanian S, Kouzarides T (2019) METTL1 Promotes let-7 MicroRNA Processing via m7G Methylation. Mol Cell 74(6):1278-1290.e9. https://doi.org/10.1016/j.molcel.2019.03.040
Park JH, von Maltzahn G, Zhang L, Schwartz MP, Ruoslahti E, Bhatia SN, Sailor MJ (2008) Magnetic iron oxide nanoworms for tumor targeting and imaging. Adv Mater 20(9):1630–1635. https://doi.org/10.1002/adma.200800004
Park JH, von Maltzahn G, Zhang L, Derfus AM, Simberg D, Harris TJ et al. (2009) Systematic surface engineering of magnetic nanoworms for in vivo tumor targeting. Small 5(6):694–700 https://doi.org/10.1002/smll.200801789
Park JH, von Maltzahn G, Xu MJ, Fogal V, Kotamraju VR, Ruoslahti E et al (2010) Cooperative nanomaterial system to sensitize, target, and treat tumors. Proc Natl Acad Sci 107(3):981–986. https://doi.org/10.1073/pnas.0909565107
Patil DP, Chen CK, Pickering BF, Chow A, Jackson C, Guttman M, Jaffrey SR (2016) m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537(7620):369–373. https://doi.org/10.1038/nature19342
Peraud A, Watanabe K, Plate KH, Yonekawa Y, Kleihues P, Ohgaki H (1997) p53 mutations versus EGF receptor expression in giant cell glioblastomas. J Neuropathol Exp Neurol 56(11):1236–1241. https://doi.org/10.1097/00005072-199711000-00008
Pesenti C, Navone SE, Guarnaccia L, Terrasi A, Costanza J, Silipigni R et al. (2019) The genetic landscape of human glioblastoma and matched primary cancer stem cells reveals intratumour similarity and intertumour heterogeneity. Stem Cells Int 2019 https://doi.org/10.1155/2019/2617030
Pourgholi F, Farhad JN, Kafil HS, Yousefi M (2016) Nanoparticles: Novel vehicles in treatment of glioblastoma. Biomed Pharmacother 77:98–107. https://doi.org/10.1016/j.biopha.2015.12.014
Qiu J, Kong L, Cao X, Li A, Wei P, Wang L, Mignani S, Caminade AM, Majoral JP, Shi X (2018) Enhanced Delivery of Therapeutic siRNA into Glioblastoma Cells Using Dendrimer-Entrapped Gold Nanoparticles Conjugated with β-Cyclodextrin. Nanomaterials (Basel, Switzerland) 8(3):131. https://doi.org/10.3390/nano8030131
Quaranta M, Divella R, Daniele A, Di Tardo S, Venneri MT, Lolli I, Troccoli G (2007) Epidermal growth factor receptor serum levels and prognostic value in malignant gliomas. Tumori 93(3):275–280
Quinn JA, Jiang SX, Carter J, Reardon DA, Desjardins A, Vredenburgh JJ, Rich JN, Gururangan S, Friedman AH, Bigner DD, Sampson JH, McLendon RE, Herndon JE 2nd, Threatt S, Friedman HS (2009) Phase II trial of Gliadel plus O6-benzylguanine in adults with recurrent glioblastoma multiforme. Clin Cancer Res 15(3):1064–1068. https://doi.org/10.1158/1078-0432.CCR-08-2130
Rahaman SO, Harbor PC, Chernova O, Barnett GH, Vogelbaum MA, Haque SJ (2002) Inhibition of constitutively active Stat3 suppresses proliferation and induces apoptosis in glioblastoma multiforme cells. Oncogene 21(55):8404–8413. https://doi.org/10.1038/sj.onc.1206047
Ramalho MJ, Loureiro JA, Coelho MA, Pereira MC (2022) Transferrin receptor-targeted nanocarriers: overcoming barriers to treat glioblastoma. Pharmaceutics 14(2):279. https://doi.org/10.3390/pharmaceutics14020279
Reardon DA, Brandes AA, Omuro A, Mulholland P, Lim M, Wick A, Baehring J, Ahluwalia MS, Roth P, Bähr O, Phuphanich S, Sepulveda JM, De Souza P, Sahebjam S, Carleton M, Tatsuoka K, Taitt C, Zwirtes R, Sampson J, Weller M (2020) Effect of Nivolumab vs Bevacizumab in Patients With Recurrent Glioblastoma: The CheckMate 143 Phase 3 Randomized Clinical Trial. JAMA Oncol 6(7):1003–1010. https://doi.org/10.1001/jamaoncol.2020.1024
Reifenberger G, Liu L, Ichimura K, Schmidt EE, Collins VP (1993) Amplification and overexpression of the MDM2 gene in a subset of human malignant gliomas without p53 mutations. Cancer Res 53:2736–2739
Reifenberger J, Ring GU, Gies U, Cobbers L, Oberstrass J, An HX, Niederacher D, Wechsler W, Reifenberger G (1996) Analysis of p53 mutation and epidermal growth factor receptor amplification in recurrent gliomas with malignant progression. J Neuropathol Exp Neurol 55(7):822–831. https://doi.org/10.1097/00005072-199607000-00007
Ren X, Ai D, Li T, Xia L, Sun L (2021) Effectiveness of Lomustine Combined With Bevacizumab in Glioblastoma: A Meta-Analysis. Front Neurol 11:603947 https://doi.org/10.3389/fneur.2020.603947
Reon BJ, Anaya J, Zhang Y, Mandell J, Purow B, Abounader R, Dutta A (2016) Expression of lncRNAs in low-grade gliomas and glioblastoma multiforme: an in silico analysis. PLoS Med 13(12):e1002192 https://doi.org/10.1371/journal.pmed.1002192
Roth L, Agemy L, Kotamraju VR, Braun G, Teesalu T, Sugahara KN et al (2012) Transtumoral targeting enabled by a novel neuropilin-binding peptide. Oncogene 31(33):3754–3763. https://doi.org/10.1038/onc.2011.537
Ru P, Hu P, Geng F, Mo X, Cheng C, Yoo JY et al (2016) Feedback loop regulation of SCAP/SREBP-1 by miR-29 modulates EGFR signaling-driven glioblastoma growth. Cell Rep 16(6):1527–1535. https://doi.org/10.1016/j.celrep.2016.07.017
Sabha N, Knobbe CB, Maganti M, Al Omar S, Bernstein M, Cairns R, Çako B, von Deimling A, Capper D, Mak TW, Kiehl TR, Carvalho P, Garrett E, Perry A, Zadeh G, Guha A, Croul S (2014) Analysis of IDH mutation, 1p/19q deletion, and PTEN loss delineates prognosis in clinical low-grade diffuse gliomas. Neuro Oncol 16(7):914–923. https://doi.org/10.1093/neuonc/not299
Santanam U, Zanesi N, Efanov A, Costinean S, Palamarchuk A, Hagan JP et al (2010) Chronic lymphocytic leukemia modeled in mouse by targeted miR-29 expression. Proc Natl Acad Sci 107(27):12210–12215. https://doi.org/10.1073/pnas.1007186107
Sareddy GR, Pratap UP, Viswanadhapalli S, Venkata PP, Nair BC, Krishnan SR, Zheng S, Gilbert AR, Brenner AJ, Brann DW, Vadlamudi RK (2019) PELP1 promotes glioblastoma progression by enhancing Wnt/β-catenin signaling. Neuro-Oncol Adv 1(1):vdz042 https://doi.org/10.1093/noajnl/vdz042
Sasaki H, Zlatescu MC, Betensky RA, Ino Y, Cairncross JG, Louis DN (2001) PTEN is a target of chromosome 10q loss in anaplastic oligodendrogliomas and PTEN alterations are associated with poor prognosis. Am J Pathol 159(1):359–367. https://doi.org/10.1016/S0002-9440(10)61702-6
Schindler G, Capper D, Meyer J, Janzarik W, Omran H, Herold-Mende C et al (2011) Analysis of BRAF V600E Mutation in 1,320 Nervous System Tumors Reveals High Mutation Frequencies in Pleomorphic Xanthoastrocytoma, Ganglioglioma and Extra-Cerebellar Pilocytic Astrocytoma. Acta Neuropathol 121(3):397–405. https://doi.org/10.1007/s00401-011-0802-6
Shahmoradi S, Golzar H, Hashemi M, Mansouri V, Omidi M, Yazdian F, Yadegari A, Tayebi L (2018) Optimizing the nanostructure of graphene oxide/silver/arginine for effective wound healing. Nanotechnology 29(47):475101 https://doi.org/10.1088/1361-6528/aadedc
Shahryari A, Jazi MS, Samaei NM, Mowla SJ (2015) Long non-coding RNA SOX2OT: expression signature, splicing patterns, and emerging roles in pluripotency and tumorigenesis. Front Genet 6:196. https://doi.org/10.3389/fgene.2015.00196
Shahzad U, Li C, Johnston M, Wang JJ, Sabha N, Varn FS et al. (2020) CASCADES, a novel SOX2 super-enhancer associated long non-coding RNA, regulates cancer stem cell specification and differentiation in glioblastoma multiforme. bioRxiv 2020–09 https://doi.org/10.1101/2020.09.05.284349
Shanmugam R, Fierer J, Kaiser S, Helm M, Jurkowski TP, Jeltsch A (2015) Cytosine methylation of tRNA-Asp by DNMT2 has a role in translation of proteins containing poly-Asp sequences. Cell Discovery 1:15010. https://doi.org/10.1038/celldisc.2015.10
Sharma S, Kotamraju VR, Molder T, Tobi A, Teesalu T, Ruoslahti E (2017) Tumor-penetrating nanosystem strongly suppresses breast tumor growth. Nano Lett 17(3):1356–1364. https://doi.org/10.1021/acs.nanolett.6b03815
Shchors K, Persson AI, Rostker F, Tihan T, Lyubynska N, Li N et al (2013) Using a preclinical mouse model of high-grade astrocytoma to optimize p53 restoration therapy. Proc Natl Acad Sci 110(16):E1480–E1489. https://doi.org/10.1073/pnas.1219142110
Shimotohno K, Kodama Y, Hashimoto J, Miura KI (1977) Importance of 5’-terminal blocking structure to stabilize mRNA in eukaryotic protein synthesis. Proc Natl Acad Sci USA 74(7):2734–2738. https://doi.org/10.1073/pnas.74.7.2734
Silvestri IP, Colbon PJ (2021) The Growing Importance of Chirality in 3D Chemical Space Exploration and Modern Drug Discovery Approaches for Hit-ID: Topical Innovations. ACS Med Chem Lett 12(8):1220–1229. https://doi.org/10.1021/acsmedchemlett.1c00251
Sloan KE, Warda AS, Sharma S, Entian KD, Lafontaine D, Bohnsack MT (2017) Tuning the ribosome: The influence of rRNA modification on eukaryotic ribosome biogenesis and function. RNA Biol 14(9):1138–1152. https://doi.org/10.1080/15476286.2016.1259781
Smith JS, Tachibana I, Passe SM, Huntley BK, Borell TJ, Iturria N et al (2001) PTEN mutation, EGFR amplification, and outcome in patients with anaplastic astrocytoma and glioblastoma multiforme. J Natl Cancer Inst 93(16):1246–1256. https://doi.org/10.1093/jnci/93.16.1246
Steele RJC, Lane DP (2005) P53 in cancer: a paradigm for modern management of cancer. Surgeon 3(3):197–205. https://doi.org/10.1016/S1479-666X(05)80041-1
Stenström P, Manzanares D, Zhang Y, Ceña V, Malkoch M (2018) Evaluation of Amino-Functional Polyester Dendrimers Based on Bis-MPA as Nonviral Vectors for siRNA Delivery. Molecules (basel, Switzerland) 23(8):2028. https://doi.org/10.3390/molecules23082028
St Louis DC, Woodcock JB, Franzoso G, Blair PJ, Carlson LM, Murillo M, Wells MR, Williams AJ, Smoot DS, Kaushal S, Grimes JL, Harlan DM, Chute JP, June CH, Siebenlist U, Lee KP (1999) Evidence for distinct intracellular signaling pathways in CD34+ progenitor to dendritic cell differentiation from a human cell line model. J Immunol (Baltimore, Md. : 1950) 162(6):3237–3248
Stupp R, Gander M, Leyvraz S, Newlands E (2001) Current and future developments in the use of temozolomide for the treatment of brain tumours. Lancet Oncol 2(9):552–560. https://doi.org/10.1016/S1470-2045(01)00489-2
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn U, Curschmann J, Janzer RC, Ludwin SK, Gorlia T, Allgeier A, Lacombe D, Cairncross JG, Eisenhauer E, Mirimanoff RO, European Organisation for Research and Treatment of Cancer Brain Tumor and Radiotherapy Groups et al. (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352(10):987–996 https://doi.org/10.1056/NEJMoa043330
Stupp R, Wong ET, Kanner AA, Steinberg D, Engelhard H, Heidecke V, Kirson ED, Taillibert S, Liebermann F, Dbalý V, Ram Z, Villano JL, Rainov N, Weinberg U, Schiff D, Kunschner L, Raizer J, Honnorat J, Sloan A, Malkin M et al. (2012) NovoTTF-100A versus physician's choice chemotherapy in recurrent glioblastoma: a randomised phase III trial of a novel treatment modality. Eur J Cancer (Oxford, England : 1990) 48(14):2192–2202 https://doi.org/10.1016/j.ejca.2012.04.011
Stupp R, Taillibert S, Kanner AA, Kesari S, Steinberg DM, Toms SA, Taylor LP, Lieberman F, Silvani A, Fink KL, Barnett GH, Zhu JJ, Henson JW, Engelhard HH, Chen TC, Tran DD, Sroubek J, Tran ND, Hottinger AF, Landolfi J et al (2015) Maintenance Therapy With Tumor-Treating Fields Plus Temozolomide vs Temozolomide Alone for Glioblastoma: A Randomized Clinical Trial. JAMA 314(23):2535–2543. https://doi.org/10.1001/jama.2015.16669
Stupp R, Taillibert S, Kanner A, Read W, Steinberg D, Lhermitte B, Toms S, Idbaih A, Ahluwalia MS, Fink K, Di Meco F, Lieberman F, Zhu JJ, Stragliotto G, Tran D, Brem S, Hottinger A, Kirson ED, Lavy-Shahaf G, Weinberg U et al (2017) Effect of Tumor-Treating Fields Plus Maintenance Temozolomide vs Maintenance Temozolomide Alone on Survival in Patients With Glioblastoma: A Randomized Clinical Trial. JAMA 318(23):2306–2316. https://doi.org/10.1001/jama.2017.18718
Stylli SS, Kaye AH, MacGregor L, Howes M, Rajendra P (2005) Photodynamic therapy of high grade glioma–long term survival. J Clin Neurosci 12(4):389–398. https://doi.org/10.1016/j.jocn.2005.01.006
Su R, Cao S, Ma J, Liu Y, Liu X, Zheng J et al (2017) Knockdown of SOX2OT inhibits the malignant biological behaviors of glioblastoma stem cells via upregulating the expression of miR-194-5p and miR-122. Mol Cancer 16:1–22. https://doi.org/10.1186/s12943-017-0737-1
Sugahara KN, Teesalu T, Karmali PP, Kotamraju VR, Agemy L, Girard OM et al (2009) Tissue-penetrating delivery of compounds and nanoparticles into tumors. Cancer Cell 16(6):510–520. https://doi.org/10.1016/j.ccr.2009.10.013
Sukumar UK, Bose R, Malhotra M, Babikir HA, Afjei R, Robinson E, Zeng Y, Chang E, Habte F, Sinclair R, Gambhir SS, Massoud TF, Paulmurugan R (2019) Intranasal delivery of targeted polyfunctional gold-iron oxide nanoparticles loaded with therapeutic microRNAs for combined theranostic multimodality imaging and presensitization of glioblastoma to temozolomide. Biomaterials 218:119342 https://doi.org/10.1016/j.biomaterials.2019.119342
Sun YM, Chen YQ (2020) Principles and innovative technologies for decrypting noncoding RNAs: from discovery and functional prediction to clinical application. J Hematol Oncol 13:1–27. https://doi.org/10.1186/s13045-020-00945-8
Sun WL, Kang T, Wang YY, Sun JP, Li C, Liu HJ, Yang Y, Jiao BH (2019) Long non-coding RNA OIP5-AS1 targets Wnt-7b to affect glioma progression via modulation of miR-410. Biosci Rep 39(1):BSR20180395 10.1042/BSR20180395
Takai H, Masuda K, Sato T, Sakaguchi Y, Suzuki T, Suzuki T, Koyama-Nasu R, Nasu-Nishimura Y, Katou Y, Ogawa H, Morishita Y, Kozuka-Hata H, Oyama M, Todo T, Ino Y, Mukasa A, Saito N, Toyoshima C, Shirahige K, Akiyama T (2014) 5-Hydroxymethylcytosine plays a critical role in glioblastomagenesis by recruiting the CHTOP-methylosome complex. Cell Rep 9(1):48–60. https://doi.org/10.1016/j.celrep.2014.08.071
Tan AC, Ashley DM, López GY, Malinzak M, Friedman HS, Khasraw M (2020) Management of glioblastoma: State of the art and future directions. CA Cancer J Clin 70(4):299–312 https://doi.org/10.3322/caac.21613
Tan SK, Jermakowicz A, Mookhtiar AK, Nemeroff CB, Schürer SC, Ayad NG (2018) Drug Repositioning in Glioblastoma: A Pathway Perspective. Front Pharmacol 9:218. https://doi.org/10.3389/fphar.2018.00218
Tang C, Guo J, Chen H, Yao CJ, Zhuang DX, Wang Y, Tang WJ, Ren G, Yao Y, Wu JS, Mao Y, Zhou LF (2015) Gene mutation profiling of primary glioblastoma through multiple tumor biopsy guided by 1H-magnetic resonance spectroscopy. Int J Clin Exp Pathol 8(5):5327–5335
Teesalu T, Sugahara KN, Kotamraju VR, Ruoslahti E (2009) C-end rule peptides mediate neuropilin-1-dependent cell, vascular, and tissue penetration. Proc Natl Acad Sci 106(38):16157–16162. https://doi.org/10.1073/pnas.0908201106
Teesalu T, Sugahara KN, Ruoslahti E (2013) Tumor-penetrating peptides. Front. Oncol 3:216. https://doi.org/10.3389/fonc.2013.00216
Thomas SM, DeMarco M, D’Arcangelo G, Halegoua S, Brugge JS (1992) Ras is essential for nerve growth factor- and phorbol ester-induced tyrosine phosphorylation of MAP kinases. Cell 68(6):1031–1040. https://doi.org/10.1016/0092-8674(92)90075-n
Tolcher AW, Gerson SL, Denis L, Geyer C, Hammond LA, Patnaik A, Goetz AD, Schwartz G, Edwards T, Reyderman L, Statkevich P, Cutler DL, Rowinsky EK (2003) Marked inactivation of O6-alkylguanine-DNA alkyltransferase activity with protracted temozolomide schedules. Br J Cancer 88(7):1004–1011. https://doi.org/10.1038/sj.bjc.6600827
Tuorto F, Liebers R, Musch T, Schaefer M, Hofmann S, Kellner S, Frye M, Helm M, Stoecklin G, Lyko F (2012) RNA cytosine methylation by Dnmt2 and NSun2 promotes tRNA stability and protein synthesis. Nat Struct Mol Biol 19(9):900–905. https://doi.org/10.1038/nsmb.2357
Tuszynski JA, Wenger C, Friesen DE, Preto J (2016) An Overview of Sub-Cellular Mechanisms Involved in the Action of TTFields. Int J Environ Res Public Health 13(11):1128. https://doi.org/10.3390/ijerph13111128
Uno M, Oba-Shinjo SM, Camargo AA, Moura RP, Aguiar PH, Cabrera HN, Begnami M, Rosemberg S, Teixeira MJ, Marie SK (2011) Correlation of MGMT promoter methylation status with gene and protein expression levels in glioblastoma. Clinics (Sao Paulo, Brazil) 66(10):1747–1755. https://doi.org/10.1590/s1807-59322011001000013
Urquhart BL, Kim RB (2009) Blood− brain barrier transporters and response to CNS-active drugs. Eur J Clin Pharmacol 65(11):1063–1070. https://doi.org/10.1007/s00228-009-0714-8
Verhaak RGW, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP et al (2010) An integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17:98. https://doi.org/10.1016/j.ccr.2009.12.020
Voth B, Nagasawa DT, Pelargos PE, Chung LK, Ung N, Gopen Q et al (2015) Transferrin receptors and glioblastoma multiforme: Current findings and potential for treatment. J Clin Neurosci 22(7):1071–1076. https://doi.org/10.1016/j.jocn.2015.02.002
Vredenburgh JJ, Desjardins A, Herndon JE 2nd, Marcello J, Reardon DA, Quinn JA, Rich JN, Sathornsumetee S, Gururangan S, Sampson J, Wagner M, Bailey L, Bigner DD, Friedman AH, Friedman HS (2007) Bevacizumab plus irinotecan in recurrent glioblastoma multiforme. Journal of Clinical Oncology : Official Journal of the American Society of Clinical Oncology 25(30):4722–4729. https://doi.org/10.1200/JCO.2007.12.2440
Wald JH, Hatakeyama J, Printsev I, Cuevas A, Fry W, Saldana MJ, VanderVorst K, Rowson-Hodel A, Angelastro JM, Sweeney C, Carraway KL (2017) Suppression of planar cell polarity signaling and migration in glioblastoma by Nrdp1-mediated Dvl polyubiquitination. Oncogene 36(36):5158–5167https://doi.org/10.1038/onc.2017.126
Walker MD, Alexander E Jr, Hunt WE, MacCarty CS, Mahaley MS Jr, Mealey J Jr, Norrell HA, Owens G, Ransohoff J, Wilson CB, Gehan EA, Strike TA (1978). Evaluation of BCNU and/or radiotherapy in the treatment of anaplastic gliomas. A cooperative clinical trial. J Neurosurg 49(3):333–343 https://doi.org/10.3171/jns.1978.49.3.0333
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, Fu Y, Parisien M, Dai Q, Jia G, Ren B, Pan T, He C (2014) N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505(7481):117–120. https://doi.org/10.1038/nature12730
Wang P, Liu YH, Yao YL, Li Z, Li ZQ, Ma J, Xue YX (2015a) Long non-coding RNA CASC2 suppresses malignancy in human gliomas by miR-21. Cell Signal 27(2):275–282. https://doi.org/10.1016/j.cellsig.2014.11.011
Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, Weng X, Chen K, Shi H, He C (2015b) N(6)-methyladenosine Modulates Messenger RNA Translation Efficiency. Cell 161(6):1388–1399. https://doi.org/10.1016/j.cell.2015.05.014
Wang WT, Han C, Sun YM, Chen TQ, Chen YQ (2019) Noncoding RNAs in cancer therapy resistance and targeted drug development. J Hematol Oncol 12:1–15. https://doi.org/10.1186/s13045-019-0748-z
Wang J, Qin C, Zhong C, Wen Y, Ke S, Liao BO (2020) Long non-coding RNA CASC2 targeting miR-18a suppresses glioblastoma cell growth, metastasis and EMT in vitro and in vivo. J Biosci 45:107. https://doi.org/10.1007/s12038-020-00077-8
Wang B, Guo H, Xu H, Chen Y, Zhao G, Yu H (2022) The Role of Graphene Oxide Nanocarriers in Treating Gliomas. Front Oncol 12:736177 https://doi.org/10.3389/fonc.2022.736177
Wang X, Li X, Zhou Y, Huang X, Jiang X (2021a) Long non-coding RNA OIP5-AS1 inhibition upregulates microRNA-129-5p to repress resistance to temozolomide in glioblastoma cells via downregulating IGF2BP2. Cell Biol Toxicol. (Advanceonlinepublication) https://doi.org/10.1007/s10565-021-09614-z
Wang Y, Chen W, Shi Y, Yan C, Kong Z, Wang Y et al. (2021b) Imposing Phase II and Phase III Clinical Trials of Targeted Drugs for Glioblastoma: Current Status and Progress. Front Oncol 3611 https://doi.org/10.3389/fonc.2021.719623
Warren KE (2012) Diffuse intrinsic pontine glioma: poised for progress. Front Oncol 2:205. https://doi.org/10.3389/fonc.2012.00205
Watanabe K, Tachibana O, Sata K, Yonekawa Y, Kleihues P, Ohgaki VH (1996) Overexpression of the EGF receptor and p53 mutations are mutually exclusive in the evolution of primary and secondary glioblastomas. Brain Pathol (Zurich, Switzerland) 6(3):217–24. https://doi.org/10.1111/j.1750-3639.1996.tb00848.x
Watanabe K, Sato K, Biernat W, Tachibana O, von Ammon K, Ogata N, Yonekawa Y, Kleihues P, Ohgaki H (1997) Incidence and timing of p53 mutations during astrocytoma progression in patients with multiple biopsies. Clin Cancer Res 3(4):523–530
Weber C, Coester C, Kreuter J, Langer K (2000) Desolvation process and surface characterisation of protein nanoparticles. Int J Pharm 194(1):91–102. https://doi.org/10.1016/S0378-5173(99)00370-1
Weller M, Le Rhun E (2020) How did lomustine become standard of care in recurrent glioblastoma? Cancer Treat Rev 87:102029 https://doi.org/10.1016/j.ctrv.2020.102029
Wick W, Puduvalli VK, Chamberlain MC, van den Bent MJ, Carpentier AF, Cher LM, Mason W, Weller M, Hong S, Musib L, Liepa AM, Thornton DE, Fine HA (2010) Phase III study of enzastaurin compared with lomustine in the treatment of recurrent intracranial glioblastoma. J Clin Oncol 28(7):1168–1174. https://doi.org/10.1200/JCO.2009.23.2595
Wick W, Gorlia T, Bendszus M, Taphoorn M, Sahm F, Harting I, Brandes AA, Taal W, Domont J, Idbaih A, Campone M, Clement PM, Stupp R, Fabbro M, Le Rhun E, Dubois F, Weller M, von Deimling A, Golfinopoulos V, Bromberg JC et al (2017) Lomustine and Bevacizumab in Progressive Glioblastoma. N Engl J Med 377(20):1954–1963. https://doi.org/10.1056/NEJMoa1707358
Wilhelm SM, Dumas J, Adnane L, Lynch M, Carter CA, Schutz G et al (2011) Regorafenib (BAY 73–4506): A New Oral Multikinase Inhibitor of Angiogenic, Stromal and Oncogenic Receptor Tyrosine Kinases With Potent Preclinical Antitumor Activity. Int J Cancer 129(1):245–255. https://doi.org/10.1002/ijc.25864
William D, Walther M, Schneider B, Linnebacher M, Classen CF (2018) Temozolomide-induced increase of tumorigenicity can be diminished by targeting of mitochondria in in vitro models of patient individual glioblastoma. PLoS One 13:e0191511 https://doi.org/10.1371/journal.pone.0191511
Wirsching HG, Tritschler I, Palla A, Renner C, Weller M, Tabatabai G (2014) The management of lomustine overdose in malignant glioma patients. Neuro-Oncol Pract 1(4):178–183. https://doi.org/10.1093/nop/npu023
Wu J, Huang W, He Z (2013) Dendrimers as carriers for siRNA delivery and gene silencing: a review. Sci World J 2013 https://doi.org/10.1155/2013/630654
Wu P, Cai J, Chen Q, Han B, Meng X, Li Y, Li Z, Wang R, Lin L, Duan C, Kang C, Jiang C (2019) Lnc-TALC promotes O6-methylguanine-DNA methyltransferase expression via regulating the c-Met pathway by competitively binding with miR-20b-3p. Nat Commun 10(1):2045. https://doi.org/10.1038/s41467-019-10025-2
Xavier-Magalhães A, Gonçalves CS, Fogli A, Lourenço T, Pojo M, Pereira B, Rocha M, Lopes MC, Crespo I, Rebelo O, Tão H, Lima J, Moreira R, Pinto AA, Jones C, Reis RM, Costello JF, Arnaud P, Sousa N, Costa BM (2018) The long non-coding RNA HOTAIR is transcriptionally activated by HOXA9 and is an independent prognostic marker in patients with malignant Glioma. Oncotarget 9(21):15740–15756 https://doi.org/10.18632/oncotarget.24597
Xiao W, Adhikari S, Dahal U, Chen YS, Hao YJ, Sun BF, Sun HY, Li A, Ping XL, Lai WY, Wang X, Ma HL, Huang CM, Yang Y, Huang N, Jiang GB, Wang HL, Zhou Q, Wang XJ, Zhao YL et al (2016) Nuclear m(6)A Reader YTHDC1 Regulates mRNA Splicing. Mol Cell 61(4):507–519. https://doi.org/10.1016/j.molcel.2016.01.012
Xing J, Yi J, Cai X, Tang H, Liu Z, Zhang X, Martindale JL, Yang X, Jiang B, Gorospe M, Wang W (2015) NSun2 Promotes Cell Growth via Elevating Cyclin-Dependent Kinase 1 Translation. Mol Cell Biol 35(23):4043–4052. https://doi.org/10.1128/MCB.00742-15
Xue W, Chen J, Liu X, Gong W, Zheng J, Guo X, Liu Y, Liu L, Ma J, Wang P, Li Z, Xue Y (2018) PVT1 regulates the malignant behaviors of human glioma cells by targeting miR-190a-5p and miR-488–3p. Biochimica et biophysica acta. Mol Basis Dis 1864(5 Pt A):1783–1794 https://doi.org/10.1016/j.bbadis.2018.02.022
Yang SH, Hong YK, Yoon SC, Kim BS, Lee YS, Lee TK, Lee KS, Jeun SS, Kim MC, Park CK (2007) Radiotherapy plus concurrent and adjuvant procarbazine, lomustine, and vincristine chemotherapy for patients with malignant Glioma. Oncol Rep 17(6):1359–1364
Yang FY, Wong TT, Teng MC, Liu RS, Lu M, Liang HF, Wei MC (2012) Focused ultrasound and interleukin-4 receptor-targeted liposomal doxorubicin for enhanced targeted drug delivery and antitumor effect in glioblastoma multiforme. J Control Release 160(3):652–658. https://doi.org/10.1016/j.jconrel.2012.02.023
Yang P, Cai J, Yan W, Zhang W, Wang Y, Chen B et al (2016) Classification Based on Mutations of TERT Promoter and IDH Characterizes Subtypes in Grade II/III Gliomas. Neuro Oncol 18(8):1099–1108. https://doi.org/10.1093/neuonc/now021
Yang JM, Schiapparelli P, Nguyen HN, Igarashi A, Zhang Q, Abbadi S, Amzel LM, Sesaki H, Quiñones-Hinojosa A, Iijima M (2017a) Characterization of PTEN mutations in brain cancer reveals that pten mono-ubiquitination promotes protein stability and nuclear localization. Oncogene 36(26):3673–3685. https://doi.org/10.1038/onc.2016.493
Yang X, Yang Y, Sun BF, Chen YS, Xu JW, Lai WY, Li A, Wang X, Bhattarai DP, Xiao W, Sun HY, Zhu Q, Ma HL, Adhikari S, Sun M, Hao YJ, Zhang B, Huang CM, Huang N, Jiang GB et al (2017b) 5-methylcytosine promotes mRNA export - NSUN2 as the methyltransferase and ALYREF as an m5C reader. Cell Res 27(5):606–625. https://doi.org/10.1038/cr.2017.55
Yao Y, Ma J, Xue Y, Wang P, Li Z, Liu J, Chen L, Xi Z, Teng H, Wang Z, Li Z, Liu Y (2015) Knockdown of long non-coding RNA XIST exerts tumor-suppressive functions in human glioblastoma stem cells by upregulating miR-152. Cancer Lett 359(1):75–86. https://doi.org/10.1016/j.canlet.2014.12.051
Ye K, Jia R, Lin J, Ju M, Peng J, Xu A, Zhang L (2009) Structural organization of box C/D RNA-guided RNA methyltransferase. Proc Natl Acad Sci 106(33):13808–13813. https://doi.org/10.1073/pnas.0905128106
Yin D, Zhou H, Kumagai T, Liu G, Ong JM, Black KL et al (2005) Proteasome Inhibitor PS-341 Causes Cell Growth Arrest and Apoptosis in Human Glioblastoma Multiforme (GBM). Oncogene 24(3):344–354. https://doi.org/10.1038/sj.onc.1208225
Yin Y, Yuan X, Gao H, Yang Q (2020) Nanoformulations of small molecule protein tyrosine kinases inhibitors potentiate targeted cancer therapy. Int J Pharm 573:118785. https://doi.org/10.1016/j.ijpharm.2019.118785
Yoshino A, Ogino A, Yachi K, Ohta T, Fukushima T, Watanabe T et al (2009) Effect of IFN-β on human glioma cell lines with temozolomide resistance. Int J Oncol 35(1):139–148. https://doi.org/10.3892/ijo_00000322
Yu H, Xue Y, Wang P, Liu X, Ma J, Zheng J, Li Z, Li Z, Cai H, Liu Y (2017) Knockdown of long non-coding RNA XIST increases blood-tumor barrier permeability and inhibits glioma angiogenesis by targeting miR-137. Oncogenesis 6(3):e303 https://doi.org/10.1038/oncsis.2017.7
Yu J, Chen M, Huang H, Zhu J, Song H, Zhu J, Park J, Ji SJ (2018) Dynamic m6A modification regulates local translation of mRNA in axons. Nucleic Acids Res 46(3):1412–1423. https://doi.org/10.1093/nar/gkx1182
Yu W, Zhang L, Wei Q, Shao A (2020) O6-Methylguanine-DNA Methyltransferase (MGMT): Challenges and New Opportunities in Glioma Chemotherapy. Front Oncol 9:1547. https://doi.org/10.3389/fonc.2019.01547
Yung WK, Albright RE, Olson J, Fredericks R, Fink K, Prados MD, Brada M, Spence A, Hohl RJ, Shapiro W, Glantz M, Greenberg H, Selker RG, Vick NA, Rampling R, Friedman H, Phillips P, Bruner J, Yue N, Osoba D et al. (2000) A phase II study of temozolomide vs. procarbazine in patients with glioblastoma multiforme at first relapse. Br J Cancer 83(5):588–593 https://doi.org/10.1054/bjoc.2000.1316
Zhang S, Guo W (2019) Long non-coding RNA MEG3 suppresses the growth of glioma cells by regulating the miR 96 5p/MTSS1 signaling pathway. Mol Med Rep 20(5):4215–4225. https://doi.org/10.3892/mmr.2019.10659
Zhang Z, Zhu Z, Watabe K, Zhang X, Bai C, Xu M et al (2013) Negative regulation of lncRNA GAS5 by miR-21. Cell Death Differ 20(11):1558–1568. https://doi.org/10.1038/cdd.2013.110
Zhang L, Liang X, Li Y (2017a) Long non-coding RNA MEG3 inhibits cell growth of gliomas by targeting miR-93 and inactivating PI3K/AKT pathway. Oncol Rep 38(4):2408–2416. https://doi.org/10.3892/or.2017.5871. (RetractionpublishedOncolRep.2021Jan;45(1):407)
Zhang J, Chen G, Gao Y, Liang H (2020) HOTAIR/miR-125 axis-mediated Hexokinase 2 expression promotes chemoresistance in human glioblastoma. J Cell Mol Med 24(10):5707–5717. https://doi.org/10.1111/jcmm.15233
Zhang J, Li Y, Liu Y, Xu G, Hei Y, Lu X, Liu W (2021) Long non-coding RNA NEAT1 regulates glioma cell proliferation and apoptosis by competitively binding to microRNA 324 5p and upregulating KCTD20 expression. Oncol Rep 46:125. https://doi.org/10.3892/or.2021.8076
Zhao X, Wang P, Liu J, Zheng J, Liu Y, Chen J, Xue Y (2015) Gas5 exerts tumor-suppressive functions in human glioma cells by targeting miR-222. Mol Ther 23(12):1899–1911. https://doi.org/10.1038/mt.2015.170
Zhao W, Sun C, Cui Z (2017) A long non-coding RNA UCA1 promotes proliferation and predicts poor prognosis in Glioma. Clin Transl Oncol 19(6):735–741. https://doi.org/10.1007/s12094-016-1597-7
Zhao H, Wang X, Feng X, Li X, Pan L, Liu J, Wang F, Yuan Z, Yang L, Yu J, Su R, Zhang Y, Ma L (2018) Long non-coding RNA MEG3 regulates proliferation, apoptosis, and autophagy and is associated with prognosis in Glioma. J Neurooncol 140(2):281–288. https://doi.org/10.1007/s11060-018-2874-9
Zhen L, Yun-Hui L, Hong-Yu D, Jun M, Yi-Long Y (2016) Long non-coding RNA NEAT1 promotes glioma pathogenesis by regulating miR-449b-5p/c-Met axis. Tumour Biol 37(1):673–683. https://doi.org/10.1007/s13277-015-3843-y
Zhi YH, Song MM, Wang PL, Zhang T, Yin ZY (2009) Suppression of matrix metalloproteinase-2 via RNA interference inhibits pancreatic carcinoma cell invasiveness and adhesion. World J Gastroenterol 15(9):1072–1078. https://doi.org/10.3748/wjg.15.1072
Zhang L, Liang X, Li Y (2021) Long non-coding RNA MEG3 inhibits cell growth of gliomas by targeting miR-93 and inactivating PI3K/AKT pathway. Oncol Rep 2017b Oct 38(4):2408-2416 https://doi.org/10.3892/or.2017.5871. Retraction in: Oncol Rep. 45(1):407
Zhang P, Wu W, Chen Q, Chen M (2019) Noncoding RNAs and their integrated networks. J Integr bioinformatics 16(3) https://doi.org/10.1515/jib-2019-0027
Zhao S, Lin Y, Xu W, Jiang W, Zha Z, Wang P, Yu W, Li Z, Gong L, Peng Y, Ding J, Lei Q, Guan KL, Xiong Y (2009) Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1alpha. Science (New York, N.Y.) 324(5924):261–265 https://doi.org/10.1126/science.1170944
Zhou K, Zhang C, Yao H, Zhang X, Zhou Y, Che Y, Huang Y (2018) Knockdown of long non-coding RNA NEAT1 inhibits glioma cell migration and invasion via modulation of SOX2 targeted by miR-132. Mol Cancer 17(1):105. https://doi.org/10.1186/s12943-018-0849-2
Zhou H, Ma Y, Zhong D, Yang L (2019) Knockdown of lncRNA HOXD-AS1 suppresses proliferation, migration and invasion and enhances cisplatin sensitivity of glioma cells by sponging miR-204. Biomed Pharmacother 112:108633 https://doi.org/10.1016/j.biopha.2019.108633.
Funding
SERB supported this work, Government of INDIA, with grant number EMR/2017/001201, and DBT, Govt. of INDIA, for funding with grant number BT/PR20836/MED/30/1727/2016. The grants were received by corresponding Dr. M. Janaki Ramaiah. All the authors thank KLEF management for its infrastructural support and encouragement.
Author information
Authors and Affiliations
Contributions
Janaki Ramaiah Mekala conceived the idea, written, planned, edited, and guided it. Kowsalya and Sahiti Chamarthy has written part of manuscript and A. Harisiaram Angirekula has made the figures.
Corresponding author
Ethics declarations
Competing interests
Authors don’t have any financial or non-financial interests directly or indirectly related to the work submitted for publication.
Ethical approval
This article contains no studies with human participants or animals performed by authors.
Consent to participate
N/A
Consent to publish
N/A
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Janaki Ramaiah Mekala, Kowsalya Adusumilli, Sahiti Chamarthy Equal 2nd authors.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mekala, J.R., Adusumilli, K., Chamarthy, S. et al. Novel sights on therapeutic, prognostic, and diagnostics aspects of non-coding RNAs in glioblastoma multiforme. Metab Brain Dis 38, 1801–1829 (2023). https://doi.org/10.1007/s11011-023-01234-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11011-023-01234-2