Abstract
Long terminal repeat (LTR) retrotransposons are major components of plant genomes influencing genome size and evolution. Using two separate approaches, we identified the Ty1-copia retrotransposon families Cotzilla and SALIRE in the Beta vulgaris (sugar beet) genome. While SALIRE elements are similar to typical Ty1-copia retrotransposons, Cotzilla elements belong to a lineage called Sireviruses. Hallmarks of Cotzilla retrotransposons are the existence of an additional putative env-like open reading frame upstream of the 3′LTR, an extended gag region, and a frameshift separating the gag and pol genes. Detected in a c 0 t-1 DNA library, Cotzilla elements belong to the most abundant retrotransposon families in B. vulgaris and are relatively homogenous and evolutionarily young. In contrast, the SALIRE family has relatively few copies, is diverged, and most likely ancient. As revealed by fluorescent in situ hybridization, SALIRE elements target predominantly gene-rich euchromatic regions, while Cotzilla retrotransposons are abundant in the intercalary and pericentromeric heterochromatin. The analysis of two retrotransposons from the same subclass contrasting in abundance, age, sequence diversity, and localization gives insight in the heterogeneity of LTR retrotransposons populating a plant genome.






Similar content being viewed by others
Abbreviations
- BAC:
-
Bacterial artificial chromosome
- DAPI:
-
4′,6′-diamidino-2-phenylindole
- env :
-
Envelope
- FISH:
-
Fluorescent in situ hybridization
- FITC:
-
Fluorescein isothiocyanate
- LTR:
-
Long terminal repeat
- ORF:
-
Open reading frame
- PBS:
-
Primer-binding site
- PPT:
-
Polypurine tract
- RT:
-
Reverse transcriptase
- SDS:
-
Sodium dodecyl sulfate
- SSC:
-
Standard saline citrate (1× SSC = 0.15 M NaCl, 0.015 M Na3-citrate)
- TE:
-
Transposable element
References
Arumuganathan K, Earle E (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Report 9:208–218
Bennetzen JL (2002) Mechanisms and rates of genome expansion and contraction in flowering plants. Genetica 115:29–36
Birney E, Clamp M, Durbin R (2004) GeneWise and Genomewise. Genome Res 14:988–995
Boeke JD, Corces VG (1989) Transcription and reverse transcription of retrotransposons. Annu Rev Microbiol 43:403–434
Brandes A, Heslop-Harrison JS, Kamm A, Kubis S, Doudrick RL, Schmidt T (1997) Comparative analysis of the chromosomal and genomic organization of Ty1-copia-like retrotransposons in pteridophytes, gymnosperms and angiosperms. Plant Mol Biol 33:11–21
Casacuberta J, Vernhettes S, Audeon C, Grandbastien MA (1997) Quasispecies in retrotransposons: a role for sequence variability in Tnt1 evolution. Genetica 100:109–117
Dechyeva D, Gindullis F, Schmidt T (2003) Divergence of satellite DNA and interspersion of dispersed repeats in the genome of the wild beet Beta procumbens. Chromosome Res 11:3–21
de Felice B, Wilson R, Argenziano C, Kafantaris I, Conicella C (2008) A transcriptionally active copia-like retroelement in Citrus limon. Cell Mol Biol Lett. doi:10.2478/s11658-008-0050-5
de Jong HJ, Fransz P, Zabel P (1999) High resolution FISH in plants—techniques and applications. Trends Plant Sci 4:258–263
Desel C, Jung C, Cai D, Kleine M, Schmidt T (2001) High-resolution mapping of YACs and the single-copy gene Hs1pro1 on Beta vulgaris chromosomes by multi-colour fluorescence in situ hybridization. Plant Mol Biol 45:113–122
Devos KM, Brown JKM, Bennetzen JL (2002) Genome size reduction through illegitimate recombination counteracts genome expansion in Arabidopsis. Genome Res 12:1075–1079
Edgar R (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5:113
Francki MG (2001) Identification of Bilby, a diverged centromeric Ty1-copia retrotransposon family from cereal rye (Secale cereale L.). Genome 44:266–274
Gallo SA, Finnegan CM, Viard M et al (2003) The HIV Env-mediated fusion reaction. BBA - Biomembranes 1614:36–50
Gao X, Havecker ER, Baranov PV, Atkins JF, Voytas DF (2003) Translational recoding signals between gag and pol in diverse LTR retrotransposons. RNA 9:1422–1430
Gindullis F, Dechyeva D, Schmidt T (2001a) Construction and characterization of a BAC library for the molecular dissection of a single wild beet centromere and sugar beet (Beta vulgaris) genome analysis. Genome 44:846–855
Gindullis F, Desel C, Galasso I, Schmidt T (2001b) The large-scale organization of the centromeric region in Beta species. Genome Res 11:253–265
Grandbastien MA, Audeon C, Casacuberta JM et al (1994) Functional analysis of the tobacco Tnt1 retrotransposon. Genetica 93:181–189
Havecker ER, Gao X, Voytas DF (2005) The Sireviruses, a plant-specific lineage of the Ty1/copia retrotransposons, interact with a family of proteins related to dynein light chain 8. Plant Physiol 139:857–868
Heslop-Harrison JS, Schwarzacher T, Anamthawat-Jónsson K, Leitch AR, Shi M, Leitch IJ (1991) In-situ hybridization with automated chromosome denaturation. Technique 3:109–116
Heslop-Harrison JS, Brandes A, Taketa S et al (1997) The chromosomal distributions of Ty1-copia group retrotransposable elements in higher plants and their implications for genome evolution. Genetica 100:197–204
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300
Hirochika H, Fukuchi A, Kikuchi F (1992) Retrotransposon families in rice. Mol Gen Genet 233:209–216
Hirochika H, Otsuki H, Yoshikawa M, Otsuki Y, Sugimoto K, Takeda S (1996) Autonomous transposition of the tobacco retrotransposon Tto1 in rice. Plant Cell 8:725–734
Holligan D, Zhang X, Jiang N, Pritham EJ, Wessler SR (2006) The transposable element landscape of the model legume Lotus japonicus. Genetics 174:2215–2228
Hull R (2001) Classifying reverse transcribing elements: a proposal and a challenge to the ICTV. Arch Virol 146:2255–2261
Jia J, Yang Z, Li G, Liu C, Lei M, Zhang T, Zhou J, Ren Z (2009) Isolation and chromosomal distribution of a novel Ty1-copia-like sequence from Secale, which enables identification of wheat—Secale africanum introgression lines. J Appl Genet 50:25–28
Kapitonov V, Jurka J (1999) Molecular paleontology of transposable elements from Arabidopsis thaliana. Genetica 107:27–37
Kimura Y, Tosa Y, Shimada S, Sogo R, Kusaba M, Sunaga T, Betsuyaku S, Eto Y, Nakayashiki H, Mayama S (2001) OARE-1, a Ty1-copia retrotransposon in oat activated by abiotic and biotic stresses. Plant Cell Physiol 42:1345–1354
Kloc A, Martienssen R (2008) RNAi, heterochromatin and the cell cycle. Trends Genet 24:511–517
Koch MA, Haubold B, Mitchell-Olds T (2000) Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis, Arabis, and related genera (Brassicaceae). Mol Biol Evol 17:1483–1498
Kubis S, Heslop-Harrison JS, Desel C, Schmidt T (1998) The genomic organization of non-LTR retrotransposons (LINEs) from three Beta species and five other angiosperms. Plant Mol Biol 36:821–831
Kumar A (1998) The evolution of plant retroviruses: moving to green pastures. Trends Plant Sci 3:371–374
Kumar A, Bennetzen JL (1999) Plant retrotransposons. Annu Rev Genet 33:479–532
Kuykendall D, Shao J, Trimmer K (2009) A nest of LTR retrotransposons adjacent the disease resistance-priming gene NPR1 in Beta vulgaris L. U.S. Hybrid H20. Int J Plant Genomics. doi:10.1155/2009/576742
Lange C, Holtgräwe D, Schulz B, Weisshaar B, Himmelbauer H (2008) Construction and characterization of a sugar beet (Beta vulgaris) fosmid library. Genome 51:948–951
Laten HM, Majumdar A, Gaucher EA (1998) SIRE-1, a copia/Ty1-like retroelement from soybean, encodes a retroviral envelope-like protein. Proc Natl Acad Sci USA 95:6897–6902
Le Q, Melayah D, Bonnivard E, Petit M, Grandbastien MA (2007) Distribution dynamics of the Tnt1 retrotransposon in tobacco. Mol Genet Genomics 278:639–651
Lerat E, Capy P (1999) Retrotransposons and retroviruses: analysis of the envelope gene. Mol Biol Evol 16:1198–1207
Lescot M, Dehais P, Thijs G et al (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Lupas A (1996) Prediction and analysis of coiled-coil structures. Methods Enzymol 266:513–525
Ma J, Bennetzen JL (2004) Rapid recent growth and divergence of rice nuclear genomes. Proc Natl Acad Sci USA 101:12404–12410
Ma J, Devos KM, Bennetzen JL (2004) Analyses of LTR-retrotransposon structures reveal recent and rapid genomic DNA loss in rice. Genome Res 14:860–869
Malik HS, Henikoff S, Eickbush TH (2000) Poised for contagion: evolutionary origins of the infectious abilities of invertebrate retroviruses. Genome Res 10:1307–1318
McGrath JM, Shaw RS, de los Reyes BG, Weiland JJ (2004) Construction of a sugar beet BAC library from a hybrid with diverse traits. Plant Mol Biol Report 22:23–28
Menzel G, Dechyeva D, Wenke T, Holtgräwe D, Weisshaar B, Schmidt T (2008) Diversity of a complex centromeric satellite and molecular characterization of dispersed sequence families in sugar beet (Beta vulgaris). Ann Bot 102:521–530
Miguel C, Simões M, Oliveira M, Rocheta M (2008) Envelope-like retrotransposons in the plant kingdom: evidence of their presence in gymnosperms (Pinus pinaster). J Mol Evol 67:517–525
Moisy C, Garrison K, Meredith C, Pelsy F (2008) Characterization of ten novel Ty1/copia-like retrotransposon families of the grapevine genome. BMC Genomics 9:469
Mroczek RJ, Dawe RK (2003) Distribution of retroelements in centromeres and neocentromeres of maize. Genetics 165:809–819
Nagaki K, Cheng Z, Ouyang S et al (2004) Sequencing of a rice centromere uncovers active genes. Nat Genet 36:138–145
Pagni M, Ioannidis V, Cerutti L, Zahn-Zabal M, Jongeneel CV, Falquet L (2004) MyHits: a new interactive resource for protein annotation and domain identification. Nucleic Acids Res 32:W332
Pearce SR (2007) SIRE-1, a putative plant retrovirus is closely related to a legume Ty1-copia retrotransposon family. Cell Mol Biol Lett 12:120–126
Pearce SR, Harrison G, Li D, Heslop-Harrison JS, Kumar A, Flavell AJ (1996) The Ty1-copia group retrotransposons in Vicia species: copy number, sequence heterogeneity and chromosomal localisation. Mol Gen Genet 250:305–315
Peterson-Burch BD, Voytas DF (2002) Genes of the Pseudoviridae (Ty1/copia retrotransposons). Mol Biol Evol 19:1832–1845
Peterson-Burch BD, Wright DA, Laten HM, Voytas DF (2000) Retroviruses in plants? Trends Genet 16:151–152
Ramallo E, Kalendar R, Schulman A, Martínez-Izquierdo J (2008) Reme1, a copia retrotransposon in melon, is transcriptionally induced by UV light. Plant Mol Biol 66:137–150
Rico-Cabanas L, Martínez-Izquierdo J (2007) CIRE1, a novel transcriptionally active Ty1-copia retrotransposon from Citrus sinensis. Mol Genet Genomics 277:365–377
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
SanMiguel P, Tikhonov A, Jin YK (1996) Nested retrotransposons in the intergenic regions of the maize genome. Science 274:765–768
SanMiguel P, Gaut BS, Tikhonov A, Nakajima Y, Bennetzen JL (1998) The paleontology of intergene retrotransposons of maize. Nat Genet 20:43–45
Schmidt T, Heslop-Harrison JS (1998) Genomes, genes and junk: the large-scale organization of plant chromosomes. Trends Plant Sci 3:195–199
Schmidt T, Jung C, Metzlaff M (1991) Distribution and evolution of two satellite DNAs in the genus Beta. Theor Appl Genet 82:793–799
Schmidt T, Schwarzacher T, Heslop-Harrison JS (1994) Physical mapping of rRNA genes by fluorescent in-situ hybridization and structural analysis of 5S rRNA genes and intergenic spacer sequences in sugar beet (Beta vulgaris). Theor Appl Genet 88:629–636
Schmidt T, Kubis S, Heslop-Harrison JS (1995) Analysis and chromosomal localization of retrotransposons in sugar beet (Beta vulgaris L.): LINEs and Ty1-copia-like elements as major components of the genome. Chromosome Res 3:335–345
Shen Y, Ford-Lloyd BV, Newbury HJ (1998) Genetic relationships within the genus Beta determined using both PCR-based marker and DNA sequencing techniques. Heredity 80:624–632
Slotkin RK, Martienssen R (2007) Transposable elements and the epigenetic regulation of the genome. Nat Rev Genet 8:272–285
Song SU, Gerasimova T, Kurkulos M, Boeke JD, Corces VG (1994) An env-like protein encoded by a Drosophila retroelement: evidence that gypsy is an infectious retrovirus. Genes Dev 8:2046–2057
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Vershinin AV, Ellis THN (1999) Heterogeneity of the internal structure of PDR1, a family of Ty1/copia-like retrotransposons in pea. Mol Gen Genet 262:703–713
Wang H, Liu JS (2008) LTR retrotransposon landscape in Medicago truncatula: more rapid removal than in rice. BMC Genomics 9:382
Weber B, Schmidt T (2009) Nested Ty3-gypsy retrotransposons of a single Beta procumbens centromere contain a putative chromodomain. Chromosome Res 17:379–396
Wicker T, Keller B (2007) Genome-wide comparative analysis of copia retrotransposons in Triticeae, rice, and Arabidopsis reveals conserved ancient evolutionary lineages and distinct dynamics of individual copia families. Genome Res 17:1072–1081
Wright DA, Voytas DF (2002) Athila4 of Arabidopsis and Calypso of soybean define a lineage of endogenous plant retroviruses. Genome Res 12:122–131
Wu J, Yamagata H, Hayashi-Tsugane M et al (2004) Composition and structure of the centromeric region of rice chromosome 8. Plant Cell 16:967–976
Xiong Y, Eickbush TH (1990) Origin and evolution of retroelements based upon their reverse transcriptase sequences. EMBO J 9:3353–3362
Zakrzewski F, Wenke T, Holtgräwe D, Weisshaar B, Schmidt, T (2009) Analysis of a c0t-1 library enables the targeted identification of minisatellite and satellite families in Beta vulgaris. BMC Plant Biol (in press)
Zdobnov EM, Apweiler R (2001) InterProScan—an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17:847–848
Acknowledgements
We thank Ines Walter for excellent technical assistance. Tony Heitkam acknowledges a fellowship and financial support of the FAZIT foundation. Torsten Wenke is funded by the BMBF grant “KMU-innovativ-2: Entwicklung von Retrotransposon-basierten molekularen Werkzeugen für die Züchtung, Sortenidentifizierung und Genbankerhaltung von Kartoffeln” (0315425B).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: JS (Pat) Heslop-Harrison.
Beatrice Weber and Torsten Wenke contributed equally to the manuscript.
Electronic Supplementary Materials
Below is the link to the electronic supplementary material.
S1
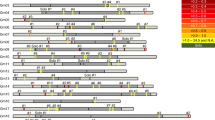
Schematic representation of an alignment containing the Cotzilla1 LTRs and 20 c 0 t-1 sequences similar to the Cotzilla1 LTR. Black bars represent sequences. The percentage identity relative to the Cotzilla1 5′LTR is indicated. These c 0 t-1 sequences were crucial for the identification of Cotzilla1. A BLAST search in the EMBL database using their consensus as query showed homology to BAC EF101866 containing the full-length retrotransposon Cotzilla1. (PDF 69 kb)
S2
Schematic representation of an alignment containing the complete Cotzilla1 retrotransposon and 347 homologous BAC sequences. Black bars represent sequences. Compared with Cotzilla1, the minimum sequence identity is 59%, the maximum sequence identity 99%. In average, the identity is 94% indicating that Cotzilla1 is a typical member of the Cotzilla family. Available BAC sequences have been generated by end-sequencing of HindIII cloned Beta vulgaris fragments (McGrath et al. 2004). Several regions of Cotzilla1 are underrepresented by BAC sequences since they do not contain HindIII restriction sites. (PDF 251 kb)
Rights and permissions
About this article
Cite this article
Weber, B., Wenke, T., Frömmel, U. et al. The Ty1-copia families SALIRE and Cotzilla populating the Beta vulgaris genome show remarkable differences in abundance, chromosomal distribution, and age. Chromosome Res 18, 247–263 (2010). https://doi.org/10.1007/s10577-009-9104-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-009-9104-4