Abstract
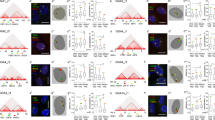
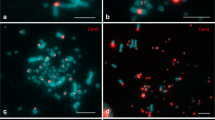
The chromosomal distribution of 41-bp repeats, known as CNM and PO41 repeats in the chicken genome and BglII repeats in the Japanese quail, was analyzed precisely using giant lampbrush chromosomes (LBC) from chicken, Japanese quail, and turkey growing oocytes. The PO41 repeat is conserved in all galliform species, whereas the other repeats are species specific. In chicken and quail, the centromere and subtelomere regions share homologous satellite sequences. RNA polymerase II transcribes the 41-bp repeats in both centromere and subtelomere regions. Ongoing transcription of these repeats was demonstrated by incorporation of BrUTP injected into oocytes at the lampbrush stage. RNA complementary to both strands of CNM and PO41 repeats is present on chicken LBC loops, whereas strand-specific G-rich transcripts are characteristic of BglII repeats in the Japanese quail. The RNA from 41-bp repeats does not undergo cotranscriptional U snRNP-dependent splicing. At the same time, the ribonucleoprotein matrix of transcription units with C-rich RNA of CNM and PO41 repeats was enriched with hnRNP protein K. Potential promoters for satellite transcription are discussed.






Similar content being viewed by others
References
Almedia R, Allshire RC (2005) RNA silencing and genome regulation. Trends Cell Biol 5:251–258
Baldwin L, Macgregor HC (1985) Centromeric satellite in the newt Triturus cristatus karelinii and related species: its distribution and transcription on lampbrush chromosomes. Chromosoma 92:100–107
Barsacchi-Pilone G, Batistoni R, Andronico F, Vitelli L, Nardi I (1986) Heterochromatic DNA in Triturus (Amphibia, Urodela). I. A satellite DNA component of the pericentric C-bands. Chromosoma 93:435–446
Bomsztyk K, Denisenko O, Ostrowski J (2004) hnRNP K: one protein multiple processes. Bioessays 26:629–638
Boyce-Jacino MT, Resnick R, Faras AJ (1989) Structural and functional characterization of the unusually short long terminal repeats and their adjacent regions of a novel endogenous avian retrovirus. Virology 173:157–166
Christy RJ, Huang RCC (1988) Functional analysis of the long terminal repeats of intracisternal A-particle genes: sequences within the U3 region determine both the efficiency and direction of promoter activity. Mol Cell Biol 8:1093–1102
Diaz MO, Gall JG (1985) Giant readthrough transcription units at the histone loci on lampbrush chromosomes of the newt Notophthalmus. Chromosoma 92:243–253
Diaz MO, Barsacchi-Pilone G, Mahon KA, Gall JG (1981) Transcripts from both strands of a satellite DNA occur on lampbrush chromosome loops of the newt Notophthalmus. Cell 24:649–659
Dreyfuss G, Kim VN, Kataoka N (2002) Messenger-RNA-binding proteins and the messages they carry. Nat Rev Mol Cell Biol 3:195–205
Dunn CA, Romanish MT, Gutierrez LE, van de Lagemaat LN, Mager DL (2006) Transcription of two human genes from a bi-directional endogenous retroviral promoter. Gene 366:335–342
Epstein LM, Mahon KA, Gall JG (1986) Transcription of a satellite DNA in the newt. J Cell Biol 103:1137–1144
Galkina S, Deryusheva S, Fillon V, Vignal A, Crooijmans R, Groenen M, Rodionov A, Gaginskaya E (2006) FISH on avian lampbrush chromosomes produces higher resolution gene mapping. Genetica 128:241–251
Hori T, Suzuki Y, Solovei I, Saitoh Y, Hatchison N, Ikeda J-E, Macgregor H, Mizuno S (1996) Characterization of DNA sequence constituting the terminal heterochromatin of the chicken Z chromosome. Chromosome Res 4:411–426
Jamrich M, Warrion R, Steele R, Gall JG (1983) Transcription of repetitive sequences on Xenopus lampbrush chromosomes. Proc Natl Acad Sci USA 80:3364–3367
Jolly C, Lakhotia SC (2006) Human sat III and Drosophila hsrω transcripts: a common paradigm for regulation of nuclear RNA processing in stressed cells. Nucleic Acids Res 34:5508–5514
Kim VN (2005) Small RNAs: classification, biogenesis, and function. Mol Cells 19:1–15
Kim JH, Hahm B, Kim YK, Choi M, Jang SK (2000) Protein–protein interaction among hnRNPs shuttling between nucleus and cytoplasm. J Mol Biol 298:395–405
Klimek-Tomczak K, Mikula M, Dzwonek A, Paziewska A, Wyrwicz LS, Hennig EE, Ostrowski J (2006) Mitochondria-associated satellite I RNA binds to hnRNP K protein. Acta Biochim Pol 53:169–178
Krasikova A, Kulikova T, Saifitdinova A, Derjusheva S, Gaginskaya E (2004) Centromeric protein bodies on avian lampbrush chromosomes contain a protein detectable with an antibody against DNA topoisomerase II. Chromosoma 113:316–323
Krasikova A, Barbero JL, Gaginskaya E (2005) Cohesion proteins are present in centromere protein bodies associated with avian lampbrush chromosomes. Chromosome Res 13:675–685
Krasikova A, Deryusheva S, Galkina S, Kurganova A, Evteev A, Gaginskaya E (2006) On the positions of centromeres in chicken lampbrush chromosomes. Chromosome Res 14:777–789
Laurent A-M, Puechberty J, Roizes G (1999) Hypothesis: for the worst and for the best, L1Hs retrotransposons actively participate in the evolution of the centromeric alphoid sequences. Chromosome Res 7:305–317
Lerner EA, Lerner MR, Janeway CA, Steitz JA (1981) Monoclonal antibodies to nucleic acid-containing cellular constituents: probes for molecular biology and autoimmune disease. Proc Natl Acad Sci USA 78:2737–2741
Ma J, Jackson SA (2006) Retrotransposon accumulation and satellite amplification mediated by segmental duplication facilitate centromere expansion in rice. Genome Res 16:251–259
Masi T, Johnson AD (2003) Read-through histone transcripts containing 3′ adenylate tails are zygotically expressed in Xenopus embryos and undergo processing to mature transcripts when introduced into oocytes nuclei. Biochem Biophys Res Commun 304:612–618
Matunis MJ, Michael WM, Dreyfuss G (1992) Characterization and primary structure of the poly(C)-binding heterogeneous nuclear ribonucleoprotein complex K protein. Mol Cell Biol 12:164–171
Matzke MA, Birchler JA (2005) RNAi-mediated pathways in the nucleus. Nat Rev Genet 6:24–35
Matzke MA, Varga F, Berger H, Schernthaner J, Schweizer D, Mayr B, Matzke AJM (1990) A 41-42 bp tandemly repeated sequence isolated from nuclear envelopes of chicken erythrocytes is located predominantly on microchromosomes. Chromosoma 99:131–137
Matzke AJM, Varga F, Gruendler P, Unfried I, Berger H, Mayr B, Matzke MA (1992) Characterization of a new repetitive sequence that is enriched on microchromosomes of turkey. Chromosoma 102:9–14
Pinol-Roma S, Swanson MS, Gall JG, Dreyfuss G (1989) A novel heterogeneous nuclear RNP protein with unique distribution on nascent transcripts. J Cell Biol 109:2575–2587
Prades C, Laurent A-M, Puechberty J, Yurov Y, Roizes G (1996) SINE and LINE within human centromeres. J Mol Evol 42:37–43
Prasanth KV, Spector D (2007) Eukaryotic regulatory RNAs: an answer to the ‘genome complexity’ conundrum. Genes Dev 21:11–42
Prieto I, Tease C, Pezzi N, Buesa JM, Ortega S, Kremer L, Martinez A, Martinez-A C, Hulten MA, Barbero JL (2004) Cohesin component dynamics during meiotic prophase I in mammalian oocytes. Chromosome Res 12:197–213
Rudert F, Bronner S, Garnier J-M, Dolle P (1995) Transcripts from opposite strands of γ satellite DNA are differentially expressed during mouse development. Mamm Genome 6:76–83
Schmid M, Nanda I, Hoehn H, Schartl M, Haaf T, Buerstedde J-M, Arakawa H, Caldwell RB, Weigend S, Burt DW, Smith J, Griffin DK, Masabanda JS, Groenen MAM, Crooijmans RPMA, Vignal A, Fillon V, Morisson M, Pitel F, Vignoles M, Garrigues A, Gellin J, Rodionov AV, Galkina SA, Lukina NA, Ben-Ari G, Blum S, Hillel J, Twito T, Lavi U, David L, Feldman MW, Delany ME, Conley CA, Fowler VM, Hedges SB, Godbout R, Katyal S, Smith C, Hudson Q, Sinclair A, Mizuno S (2005) Second report on chicken genes and chromosomes. Cytogenet Genome Res 109:415–479
Solovei I, Gaginskaya ER, Macgregor HC (1994) The arrangement and transcription of telomere DNA sequences at the ends of lampbrush chromosomes of birds. Chromosome Res 2:460–470
Solovei I, Macgregor H, Gaginskaya E (1995) Single stranded nucleic acid binding structures on chicken lampbrush chromosomes. J Cell Sci 108:1391–1396
Solovei I, Joffe BI, Gaginskaya ER, Macgregor HC (1996) Transcription on lampbrush chromosomes of a centromerically localized highly repeated DNA in pigeon (Columba) relates to sequence arrangement. Chromosome Res 4:588–603
Sun X, Le HD, Wahlstrom JM, Karpen GH (2003) Sequence analysis of a functional Drosophila centromere. Genome Res 13:182–194
Tanaka K, Suzuki T, Nojiri T, Yamagata T, Namikawa T, Matsuda Y (2000) Characterization and chromosomal distribution of a novel satellite DNA sequence of Japanese quail (Coturnix coturnix japonica). J Hered 91:412–415
Topp CN, Zhong CX, Dawe RK (2004) Centromere-encoded RNAs are integral components of the maize kinetochore. Proc Natl Acad Sci USA 101:15986–15991
Varley JM, Macgregor HC, Erba HP (1980) Satellite DNA is transcribed on lampbrush chromosomes. Nature 283:686–688
Wang X, Li J, Leung FC (2002) Partially inverted tandem repeat isolated from pericentric region of chicken chromosome 8. Chromosome Res 10:73–82
Wicker T, Robertson JS, Schulze SR, Feltus FA, Magrini V, Morrison JA, Mardis ER, Wilson RK, Peterson DG, Paterson AH, Ivarie R (2005) The repetitive landscape of the chicken genome. Genome Res 15:126–136
Yamada K, Shibusawa M, Tsudzuki M, Matsuda Y (2002) Molecular cloning and characterization of novel centromeric repetitive DNA sequences in the blue-breasted quail (Coturnix chinensis, Galliformes). Cytogenet Genome Res 98:255–261
Acknowledgments
We gratefully acknowledge the following people for providing antibodies: J. L. Barbero (Centro Nacional de Biotecnologia, Madrid, Spain), M. V. Filatov (Saint-Petersburg Institute of Nuclear Physics, Russia), J. G. Gall (Carnegie Institution of Washington, Baltimore, MD, USA), and U. Scheer and R. Hock (University of Wuerzburg, Germany). We are grateful to J. G. Gall for the critical reading and careful editing of the manuscript. We would like to thank our former graduate student A. Kurganova for her assistance in some experiments. This work was supported by the Russian Foundation for Basic Research (grant 05-04-48252). We used the equipment of the Core Facility “CHROMAS” (Biological Institute, Saint-Petersburg State University).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Matzke
Rights and permissions
About this article
Cite this article
Deryusheva, S., Krasikova, A., Kulikova, T. et al. Tandem 41-bp repeats in chicken and Japanese quail genomes: FISH mapping and transcription analysis on lampbrush chromosomes. Chromosoma 116, 519–530 (2007). https://doi.org/10.1007/s00412-007-0117-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-007-0117-5