Abstract
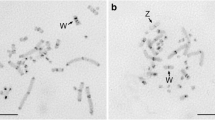
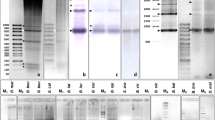
Two abundant satellite DNA sequences have been identified in and cloned from the DNA of Triturus cristatus karelinii. The smaller of these with a repeat unit of 33 base pairs (bp) is designated TkS1, the larger with 68 bp is designated TkS2. These satellites are also present in DNA from T.c. cristatus, T.c. carnifex and T. marmoratus but in substantially lower copy number. In situ hybridisations to lampbrush chromosomes of T.c. karelinii and T.c. cristatus have shown that the satellites are concentrated in the heterochromatic centromere bars of T.c. karelinii and in a region around the centromere granule in T.c. cristatus. The satellites also bind specifically to the centromere regions of mitotic metaphase chromosomes. They do not bind to the heteromorphic arms of chromosome 1, which have previously been shown to be rich in highly repeated DNA. DNA/RNA-transcript in situ hybrids to lampbrush chromosomes with TkS1 suggest that this sequence is occasionally transcribed on lampbrush loops near the centromeres.
Similar content being viewed by others
References
Arnheim N (1983) Concerted evolution of multigene families. In: Nei M, Koehn RK (eds) Evolution of genes and proteins. Sinauer Associates, Sunderland Mass., pp 38–61
Barsacchi G, Gall JG (1972) Chromosomal localisation of repetitive DNA in the newt Triturus. J Cell Biol 54:580–591
Britten RJ, Kohne DE (1968) Repeated sequences in DNA. Science 161:529–540
Callan HG (1982) Lampbrush chromosomes. Proc R Soc Lond B214:417–448
Callan HG, Lloyd L (1960) Lampbrush chromosomes of crested newts Triturus cristatus (Laurenti). Phil Trans R Soc Lond B243:135–219
Diaz MO, Barsacchi-Pilone G, Mahon KA, Gall JG (1981) Transcripts of both strands of a satellite DNA occur on lampbrush chromosome loops of the newt Notophthalmus. Cell 24:649–659
Dover GA, Flavell RB (eds) (1982) Genome evolution. Academic Press, New York, London
Gall JG, Atherton D (1974) Satellite DNA sequences in Drosophila virilis. J Mol Biol 85:633–664
Gall JG, Cohen EH, Polan ML (1971) Repetitive DNA sequences in Drosophila. Chromosoma 33:319–344
Hennig W, Hennig I, Stein H (1970) Repeated sequences in the DNA of Drosophila and their localisation in giant chromosomes. Chromosoma 32:31–63
Jamrich M, Warrior R, Steele R, Gall JG (1983) Transcription of repetitive sequences on Xenopus lampbrush chromosomes. Proc Natl Acad Sci USA 80:3364–3367
Jeffreys AJ, Flavell RA (1977) A physical map of the DNA regions flanking the rabbit β-globin gene. Cell 12:429–439
Kalezic ML, Hedgecock D (1980) Genetic variation and differentiation of three common European newts (Triturus) in Yugoslavia. Br J Herpetol 6:49–57
La Marca M, Allison D, Skinner DM (1981) Irreversible denaturation mapping of a pyrimidine-rich segment of a complex satellite DNA. J Biol Chem 256:6475–6479
Lantz LA (1947) Hybrids between Triturus cristatus Laur. and Triturus marmoratus Latr. Proc Zool Soc Lond 117:247–258
Macgregor HC (1979) In situ hybridisation of highly repetitive DNA to chromosomes of Triturus cristatus. Chromosoma 71:57–64
Macgregor HC, Kezer J (1971) The chromosomal localisation of a heavy satellite DNA in the testis of Plethodon c. cinereus. Chromosoma 33:167–182
Macgregor HC, Varley J (1984) Working with animal chromosomes. John Wiley and Sons, Chichester New York
Macgregor HC, Varley JM, Morgan GT (1981) The transcription of satellite and ribosomal DNA sequences on lampbrush chromosomes of crested newts. In: Schweiger HG (ed) International cell biology, Springer-Verlag, Berlin Heidelberg, pp 33–46
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: A laboratory manual. Cold Spring Harbor Laboratory
Miklos GL, Gill AC (1982) Nucleotide sequences of highly repeated DNAs: Compilation and comments. Genet Res Camb 39:1–30
Mizuno S, Macgregor HC (1974) Chromosomes, DNA sequences, and evolution in salamanders of the genus Plethodon. Chromosoma 48:239–296
Morgan GT, Macgregor HC, Colman A (1980) Multiple ribosomal gene sites revealed by in situ hybridisation of Xenopus rDNA to Triturus lampbrush chromosomes. Chromosoma 80:309–330
Pardue ML, Gall JG (1970) Chromosomal localisation of mouse satellite DNA. Science 168:1356–1358
Roizes GP, Pages M (1982) Conserved and divergent sequences of bovine satellite DNA. In: Dover GA, Flavell RB (eds) Genome evolution. Academic Press, New York London, pp 95–111
Sims SH (1984) Some aspects of the cytology of European newts (genus Triturus). University of Leicester Ph D Thesis
Sims SH, Macgregor HC, Pellatt PA, Horner HA (1984) Chromosome 1 in crested and marbled newts (Triturus). An extraordinary case of heteromorphism and independent chromosome evolution. Chromosoma 89:169–185
Skinner DM (1977) Satellite DNAs. BioScience 27:790–795
Smith GP (1973) Unequal crossing over and the evolution of multigene families. Cold Spring Harbor Symp Quant Biol 38:507–513
Smith GP (1976) Evolution of repeated DNA sequences by unequal crossover. Science 191:528–535
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98:503–517
Varley JM, Macgregor HC, Nardi I, Andrews C, Erba HP (1980) Cytological evidence of transcription of highy repeated DNA sequences during the lampbrush stage in Triturus cristatus carnifex. Chromosoma 80:289–307
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Baldwin, L., Macgregor, H.C. Centromeric satellite DNA in the newt Triturus cristatus karelinii and related species: Its distribution and transcription on lampbrush chromosomes. Chromosoma 92, 100–107 (1985). https://doi.org/10.1007/BF00328461
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00328461