Abstract
Essential tremor (ET) is a common, progressive neurological disease characterized by an 8–12-Hz kinetic tremor. Despite its high prevalence, the patho-mechanisms of tremor in ET are not fully known. Through comprehensive studies in postmortem brains, we identified major morphological changes in the ET cerebellum that reflect cellular damage in Purkinje cells (PCs), suggesting that PC damage is central to ET pathogenesis. We previously performed a transcriptome analysis in ET cerebellar cortex, identifying candidate genes and several dysregulated pathways. To directly target PCs, we purified RNA from PCs isolated by laser capture microdissection and performed the first ever PC-specific RNA-sequencing analysis in ET versus controls. Frozen postmortem cerebellar cortex from 24 ETs and 16 controls underwent laser capture microdissection, obtaining ≥2000 PCs per sample. RNA transcriptome was analyzed via differential gene expression, principal component analysis (PCA), and gene set enrichment analyses (GSEA). We identified 36 differentially expressed genes, encompassing multiple cellular processes. Some ET (13/24) had greater dysregulation of these genes and segregated from most controls and remaining ETs in PCA. Characterization of genes/pathways enriched in this PCA and GSEA identified multiple pathway dysregulations in ET, including RNA processing/splicing, synapse organization/ion transport, and oxidative stress/inflammation. Furthermore, a different set of pathways characterized marked heterogeneity among ET patients. Our data indicate a range of possible mechanisms for the pathogenesis of ET. Significant heterogeneity among ET combined with dysregulation of multiple cellular processes supports the notion that ET is a family of disorders rather than one disease entity.





Similar content being viewed by others
Data Availability
Raw sequencing data and DEseq2 analysis for whole cerebellar cortex (GSE134878) [33] and laser-captured PCs (GSE197345) can be found at the NIH Gene Expression Omnibus (GEO).
References
Louis ED, McCreary M. How common is essential tremor? Update on the worldwide prevalence of essential tremor. Tremor Other Hyperkinet Mov. 2021;11(28). https://doi.org/10.5334/tohm.632.
Clark LN, Louis ED. Essential tremor. Handb Clin Neurol. 2018;147:229–39. https://doi.org/10.1016/B978-0-444-63233-3.00015-4.
Odgerel Z, Hernandez N, Park J, et al. Whole genome sequencing and rare variant analysis in essential tremor families. PLoS One. 2019;14(8):e0220512. https://doi.org/10.1371/journal.pone.0220512.
Liu X, Hernandez N, Kisselev S, et al. Identification of candidate genes for familial early-onset essential tremor. Eur J Hum Genet. 2016;24(7):1009–15. https://doi.org/10.1038/ejhg.2015.228.
Shatunov A, Sambuughin N, Jankovic J, et al. Genomewide scans in North American families reveal genetic linkage of essential tremor to a region on chromosome 6p23. Brain. 2006;129:2318–3. https://doi.org/10.1093/brain/awl120.
Diez-Fairen M, Houle G, Ortega-Cubero S, et al. Exome-wide rare variant analysis in familial essential tremor. Parkinsonism Relat Disord. 2020;82:109–16. https://doi.org/10.1016/j.parkreldis.2020.11.021.
Muller SH, Girard SL, Hopfner F, et al. Genome-wide association study in essential tremor identifies three new loci. Brain. 2016;139(Pt 12):3163–9. https://doi.org/10.1093/brain/aww242.
Liao C, Castonguay C-E, Heilbron K, et al. Association of essential tremor with novel risk loci a genome-wide association study and meta-analysis. JAMA. Neurology. 2022. https://doi.org/10.1001/jamaneurol.2021.4781.
Handforth A. Linking essential tremor to the cerebellum-animal model evidence. Cerebellum. 2016;15(3):285–98. https://doi.org/10.1007/s12311-015-0750-0.
Pan MK, Li YS, Wong SB, et al. Cerebellar oscillations driven by synaptic pruning deficits of cerebellar climbing fibers contribute to tremor pathophysiology. Sci Transl Med. 2020;12(526):eaay1769. https://doi.org/10.1126/scitranslmed.aay1769.
Pan M-K, Ni C-L, Wu Y-C, Li Y-S, Kuo S-H. Animal models of tremor: relevance to human tremor disorders. Tremor Other Hyperkinet Mov. 2018;8(587):1–13. https://doi.org/10.7916/D89S37MV.
Louis ED, Jurewicz EC, Parides MK. Case-control study of nutritional antioxidant intake in essential tremor. Neuroepidemiology. 2005;24(4):203–8. https://doi.org/10.1159/000084713.
Miura S, Kamada T, Fujioka R, Yamanishi Y. Plasma amino acids in patients with essential tremor. Clin Case Rep. 2021;9(8):e04580. https://doi.org/10.1002/ccr3.4580.
Wong S-B, Wang Y-M, Lin C-C, et al. Cerebellar oscillations in familial and sporadic essential tremor. Cerebellum. 2021. https://doi.org/10.1007/s12311-021-01309-9.
Kuo SH, Wang J, Tate WJ, et al. Cerebellar pathology in early onset and late onset essential tremor. Cerebellum. 2017;16(2):473–82. https://doi.org/10.1007/s12311-016-0826-5.
Louis ED, Kuo S-H, Wang J, et al. Cerebellar pathology in familial vs. sporadic essential tremor. Cerebellum. 2017;16(4):786–91. https://doi.org/10.1007/s12311-017-0853-x.
Louis ED, Kerridge CA, Chatterjee D, et al. Contextualizing the pathology in the essential tremor cerebellar cortex: a patholog-omics approach. Acta Neuropathol. 2019;138(5):859–76. https://doi.org/10.1007/s00401-019-02043-7.
Filip P, Lungu OV, Manto MU, Bares M. Linking essential tremor to the cerebellum: physiological evidence. Cerebellum. 2016;15(6):774–80. https://doi.org/10.1007/s12311-015-0740-2.
Trujillo Diaz D, Hernandez NC, Cortes EP, et al. Banking brains: a pre-mortem “how to” guide to successful donation. Cell Tissue Bank. 2018;19(4):473–88. https://doi.org/10.1007/s10561-018-9720-3.
Louis ED, Kuo SH, Vonsattel JP, Faust PL. Torpedo formation and Purkinje cell loss: modeling their relationship in cerebellar disease. Cerebellum. 2014;13(4):433–9. https://doi.org/10.1007/s12311-014-0556-5.
Louis ED, Faust PL, Vonsattel JP, et al. Torpedoes in Parkinson's disease, Alzheimer's disease, essential tremor, and control brains. Mov Disord. 2009;24(11):1600–5. https://doi.org/10.1002/mds.22567.
Louis ED, Yi H, Erickson-Davis C, Vonsattel JP, Faust PL. Structural study of Purkinje cell axonal torpedoes in essential tremor. Neurosci Lett. 2009;450(3):287–91. https://doi.org/10.1016/j.neulet.2008.11.043.
Babij R, Lee M, Cortes E, et al. Purkinje cell axonal anatomy: quantifying morphometric changes in essential tremor versus control brains. Brain. 2013;136(Pt 10):3051–61. https://doi.org/10.1093/brain/awt238.
Yu M, Ma K, Faust PL, et al. Increased number of Purkinje cell dendritic swellings in essential tremor. Eur J Neurol. 2012;19(4):625–30. https://doi.org/10.1111/j.1468-1331.2011.03598.x.
Louis ED, Lee M, Babij R, et al. Reduced Purkinje cell dendritic arborization and loss of dendritic spines in essential tremor. Brain. 2014;137(Pt 12):3142–8. https://doi.org/10.1093/brain/awu314.
Choe M, Cortes E, Vonsattel JP, et al. Purkinje cell loss in essential tremor: random sampling quantification and nearest neighbor analysis. Mov Disord. 2016;31(3):393–401. https://doi.org/10.1002/mds.26490.
Louis ED, Kuo SH, Tate WJ, et al. Heterotopic Purkinje cells: a comparative postmortem study of essential tremor and spinocerebellar ataxias 1, 2, 3, and 6. Cerebellum. 2018;17(2):104–10. https://doi.org/10.1007/s12311-017-0876-3.
Lee PJ, Kerridge CA, Chatterjee D, et al. A quantitative study of empty baskets in essential tremor and other motor neurodegenerative diseases. J Neuropathol Exp Neurol. 2018. https://doi.org/10.1093/jnen/nly114.
Erickson-Davis CR, Faust PL, Vonsattel J-PG, et al. “Hairy baskets” associated with degenerative Purkinje cell changes in essential tremor. Neuropathol. Exp Neurol. 2010;69(3):262–71. https://doi.org/10.1097/NEN.0b013e3181d1ad04.
Lee D, Gan SR, Faust PL, Louis ED, Kuo SH. Climbing fiber-Purkinje cell synaptic pathology across essential tremor subtypes. Parkinsonism Relat Disord. 2018;51:24–9. https://doi.org/10.1016/j.parkreldis.2018.02.032.
Lin C-Y, Louis ED, Faust PL, et al. Abnormal climbing fibre-Purkinje cell synaptic connections in the essential tremor cerebellum. Brain. 2014;137:3149–59. https://doi.org/10.1093/brain/awu281.
Louis ED, Faust PL. Essential tremor pathology: neurodegeneration and reorganization of neuronal connections. Nat Rev Neurol. 2020;16(2):69–83. https://doi.org/10.1038/s41582-019-0302-1.
Martuscello RT, Kerridge CA, Chatterjee D, et al. Gene expression analysis of the cerebellar cortex in essential tremor. Neurosci Lett. 2020;721:134540. https://doi.org/10.1016/j.neulet.2019.134540.
Martuscello RT, Louis ED, Faust PL. A stainless protocol for high quality RNA isolation from laser capture microdissected Purkinje cells in the human post-mortem cerebellum. J Vis Exp. 2019;143. https://doi.org/10.3791/58953.
Gionco JT, Hartstone WG, Martuscello RT, et al. Essential tremor versus “ET-plus”: a detailed postmortem study of cerebellar pathology. Cerebellum. 2021;20(6):904–12. https://doi.org/10.1007/s12311-021-01263-6.
Dowd H, Zdrodowska MA, Radler KH, et al. Prospective longitudinal study of gait and balance in a cohort of elderly essential tremor patients. Front Neurol. 2020;11:1–14. https://doi.org/10.3389/fneur.2020.581703.
Consensus recommendations for the postmortem diagnosis of Alzheimer’s disease. The National Institute on Aging, and Reagan Institute Working Group on Diagnostic Criteria for the Neuropathological Assessment of Alzheimer’s Disease. Neurobiol Aging. 1997;18:S1–2. https://doi.org/10.1016/S0197-4580(97)00057-2.
Barton AJL, Pearson RCA, Najlerahim A, Harrison PJ. Pre- and postmortem influences on brain RNA. J Neurochem. 1993;61(1):1–11. https://doi.org/10.1111/j.1471-4159.1993.tb03532.x.
Popova T, Mennerich D, Weith A, Quast K. Effect of RNA quality on transcript intensity levels in microarray analysis of human post-mortem brain tissues. BMC Genomics. 2008;9(91). https://doi.org/10.1186/1471-2164-9-91.
Braak H, Alafuzoff I, Arzberger T, Kretzschmar H, Del Tredici K. Staging of Alzheimer disease-associated neurofibrillary pathology using paraffin sections and immunocytochemistry. Acta Neuropathol. 2006;112(4):389–404. https://doi.org/10.1007/s00401-006-0127-z.
Braak H, Braak E. Diagnostic criteria for neuropathologic assessment of Alzheimer’s disease. Neurobiol Aging. 1997;18:S85–8. https://doi.org/10.1016/s0197-4580(97)00062-6.
Mirra S. The CERAD neuropathology protocol and consensus recommendations for the postmortem diagnosis of Alzheimer's disease: a commentary. Neurobiol Aging. 1997;18:S91–4. https://doi.org/10.1016/s0197-4580(97)00058-4.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12(4):357–60. https://doi.org/10.1038/nmeth.3317.
Anders S, Pyl PT, Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31(2):166–9. https://doi.org/10.1093/bioinformatics/btu638.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550. https://doi.org/10.1186/s13059-014-0550-8.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–50. https://doi.org/10.1073/pnas.0506580102.
Mootha VK, Lindgren CM, Eriksson K-F, et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34(3):267–73. https://doi.org/10.1038/ng1180.
Szklarczyk D, Morris JH, Cook H, et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017;45(D1):D362–D8. https://doi.org/10.1093/nar/gkw937.
Tio M, Tan EK. Genetics of essential tremor. Parkinsonism Relat Disord. 2016;22(Suppl 1):S176–8. https://doi.org/10.1016/j.parkreldis.2015.09.022.
Louis ED. ‘Essential tremor’ or ‘the essential tremors’: is this one disease or a family of diseases? Neuroepidemiology. 2014;42(2):81–9. https://doi.org/10.1159/000356351.
Jimenez-Jimenez FJ, Alonso-Navarro H, Garcia-Martin E, et al. Genomic markers for essential tremor. Pharmaceuticals (Basel). 2021;14(6). https://doi.org/10.3390/ph14060516.
Louis ED, Bares M, Benito-Leon J, et al. Essential tremor-plus: a controversial new concept. Lancet Neurol. 2019;19(3):266–70. https://doi.org/10.1016/S1474-4422(19)30398-9.
Bhatia KP, Bain P, Bajaj N, et al. Consensus statement on the classification of tremors. from the task force on tremor of the International Parkinson and Movement Disorder Society. Mov Disord. 2018;33(1):75–87. https://doi.org/10.1002/mds.27121.
Stefansson H, Steinberg S, Petursson H, et al. Variant in the sequence of the LINGO1 gene confers risk of essential tremor. Nat Genet. 2009;41:277–9. https://doi.org/10.1038/ng.299.
Clark LN, Park N, Kisselev S, et al. Replication of the LINGO1 gene association with essential tremor in a North American population. Eur J Hum Genet. 2010;18:838–43. https://doi.org/10.1038/ejhg.2010.27.
Thier S, Lorenz D, Nothnagel M, et al. Polymorphisms in the glial glutamate transporter SLC1A2 are associated with essential tremor. Neurology. 2012;79(3). https://doi.org/10.1212/WNL.0b013e31825fdeed.
Lee M, Cheng MM, Lin C-Y, et al. Decreased EAAT2 protein expression in the essential tremor cerebellar cortex. Acta Neuropathol Commun. 2014;2(157):1–11. https://doi.org/10.1186/s40478-014-0157-z.
Xiao B, Deng X, Ng EY, et al. GWAS-linked PPARGC1A variant in Asian patients with essential tremor. Brain. 2017;140(4):e24. https://doi.org/10.1093/brain/awx027.
Merner ND, Girard SL, Catoire H, et al. Exome sequencing identifies FUS mutations as a cause of essential tremor. Am J Hum Genet. 2012;91:313–9. https://doi.org/10.1016/j.ajhg.2012.07.002.
Parmalee N, Mirzozoda K, Kisselev S, et al. Genetic analysis of the FUS/TLS gene in essential tremor. Eur J Neurol. 2013;20(3):534–9. https://doi.org/10.1111/ene.12023.
Hor H, Francescatto L, Bartesaghi L, et al. Missense mutations in TENM4, a regulator of axon guidance and central myelination, cause essential tremor. Hum Mol Genet. 2015;24(20):5677–86. https://doi.org/10.1093/hmg/ddv281.
Sánchez E, Bergareche A, Krebs CE, et al. SORT1 mutation resulting in sortilin deficiency and p75NTR upregulation in a family with essential tremor. ASN Neuro. 2015;7(4):1–13. https://doi.org/10.1177/1759091415598290.
Gulsuner HU, Gulsuner S, Mercan FN, et al. Mitochondrial serine protease HTRA2 p.G399S in a kindred with essential tremor and Parkinson disease. PNAS. 2014;111(51):18285–90. https://doi.org/10.1073/pnas.1419581111.
Sun Q-Y, Xu Q, Tian Y, et al. Expansion of GGC repeat in the human-specific NOTCH2NLC gene is associated with essential tremor. Brain. 2020;143(1):222–33. https://doi.org/10.1093/brain/awz372.
Leng X-R, Qi X-H, Zhou Y-T, Wang Y-P. Gain-of-function mutation p.Arg225Cys in SCN11A causes familial episodic pain and contributes to essential tremor. J Hum Genet. 2017;62:641–6. https://doi.org/10.1038/jhg.2017.21.
Liao C, Sarayloo F, Rochefort D, et al. Multiomics analyses identify genes and pathways relevant to essential tremor. Mov Disord. 2020;35(7):1153–62. https://doi.org/10.1002/mds.28031.
Neueder A. RNA-mediated disease mechanisms in neurodegenerative disorders. J Mol Biol. 2019;431(9):1780–91. https://doi.org/10.1016/j.jmb.2018.12.012.
Montes M, Sanford BL, Comiskey DF, Chandler DS. RNA splicing and disease: animal models to therapies. Trends Genet. 2019;35(1):68–87. https://doi.org/10.1016/j.tig.2018.10.002.
Hopfner F, Stevanin G, Müller SH, et al. The impact of rare variants in FUS in essential tremor. Mov Disord. 2015;30(5):721–4. https://doi.org/10.1002/mds.26145.
Ortega-Cubero S, Lorenzo-Betancor O, Lorenzo E, et al. Fused in sarcoma (FUS) gene mutations are not a frequent cause of essential tremor in Europeans. Neurobiol Aging. 2013;34:2441.e9–e11. https://doi.org/10.1016/j.neurobiolaging.2013.04.024.
Kino Y, Washizu C, Kurosawa M, et al. FUS/TLS deficiency causes behavioral and pathological abnormalities distinct from amyotrophic lateral sclerosis. Acta Neuropathol Commun. 2015;3(24). https://doi.org/10.1186/s40478-015-0202-6.
Uhlén M, Fagerberg L, Hallström BM, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;347(6220). https://doi.org/10.1126/science.1260419.
Santulli G, Marks A. Essential roles of intracellular calcium release channels in muscle, brain, metabolism, and aging. Curr Mol Pharmacol. 2015;8:1–17. https://doi.org/10.2174/1874467208666150507105105.
Khodakhah K, Armstrong CM. Inositol trisphosphate and ryanodine receptors share a common functional Ca2+ pool in cerebellar Purkinje neurons. Biophys J. 1997;73:3349–57. https://doi.org/10.1016/S0006-3495(97)78359-0.
Gomez LC, Kawaguchi S-Y, Collin T, et al. Influence of spatially segregated IP 3-producing pathways on spike generation and transmitter release in Purkinje cell axons. Proc Natl Acad Sci U S A. 2020;117(20):11097–108. https://doi.org/10.1073/pnas.2000148117.
Martinez Leo E, Secura Campos M. Systemic oxidative stress: a key point in neurodegeneration - a review. J Nutr Health Aging. 2019;23(8):694–9. https://doi.org/10.1007/s12603-019-1240-8.
Singh A, Kukreti R, Saso L, Kukreti S. Oxidative stress: a Key modulator in neurodegenerative diseases. Molecules. 2019;24:1–20. https://doi.org/10.3390/molecules24081583.
Dogu O, Louis ED, Tamer LT, et al. Elevated blood lead concentrations in essential tremor: a case–control study in Mersin, Turkey. Environ Health Perspect. 2007;115(11):1564–8. https://doi.org/10.1289/ehp.10352.
Louis ED, Jurewicz EC, Applegate L, et al. Association between essential tremor and blood lead concentration. Environ Health Perspect. 2003;111(14):1707–11. https://doi.org/10.1289/ehp.6404.
Louis ED, Factor-Litvak P, Gerbin M, et al. Blood harmane, blood lead, and severity of hand tremor: evidence of additive effects. Neurotoxicology. 2011;32:227–32. https://doi.org/10.1016/j.neuro.2010.12.002.
Almeida Lopes ACB, Peixe TS, Mesas AE, Paoliello MMB. Lead exposure and oxidative stress: a systematic review. Rev Environ Contam Toxicol. 2016;236:193-238. Springer.
Sammi SR, Agim ZS, Cannon JR. Harmane-induced selective dopaminergic neurotoxicity in Caenorhabditis elegans. Toxicol Sci. 2018;161(2):335–48. https://doi.org/10.1093/toxsci/kfx223.
Wilkins HM, Kirchhof D, Manning E, Joseph JW, Linseman DA. Mitochondrial glutathione transport is a key determinant of neuronal susceptibility to oxidative and nitrosative stress. J Biol Chem. 2013;288(7):5091–101. https://doi.org/10.1074/jbc.M112.405738.
Gu F, Chauhan V, Chauhan A. Glutathione redox imbalance in brain disorders. Curr Opin Clin Nutr Metab Care. 2015;19(1):89–95. https://doi.org/10.1097/MCO.0000000000000134.
Chen WW, Zhang X, Huang WJ. Role of neuroinflammation in neurodegenerative diseases (Review). Mol Med Rep. 2016;13(4):3391–6. https://doi.org/10.3892/mmr.2016.4948.
Louis ED, Faust PL. Essential tremor: the most common form of cerebellar degeneration? Cereb Ataxias. 2020;7(12):1–10. https://doi.org/10.1186/s40673-020-00121-1.
Muruzheva ZM, Ivleva IS, Traktirov DS, Zubov AS, Karpenko MN. The relationship between serum interleukin-1beta, interleukin-6, interleukin-8, interleukin-10, tumor necrosis factor-alpha levels and clinical features in essential tremor. Int J Neurosci. 2021;1-10. https://doi.org/10.1080/00207454.2020.1865952.
Louis ED. The essential tremors: evolving concepts of a family of diseases. Front Neurol. 2021;12. https://doi.org/10.3389/fneur.2021.650601.
Acknowledgements
The authors would like to acknowledge their funding for this project from the NIH (R01 NS088257-01A1). There are no other financial disclosures or conflicts of interest to report for any author as it pertains to this research. Full list of financial support from the last 12-months for each author is provided. We would like to thank all patients that donated their brains for banking and the staff at the ETCBR at the New York Brain Bank, the NIH NeuroBioBank, and the Rush Alzheimer’s Disease Center Brain Bank. We thank Columbia University Genome Center and Dr. Erin Bush for their assistance and expertise on this project. The first author personally thanks William Zeizer for his instrumental knowledge of −80°C freezers.
Funding
NINDS RO1# NS088257-01A1
Author information
Authors and Affiliations
Contributions
Regina T. Martuscello: Designed and executed laser capture collection of PCs. Extracted RNA, quality control, prepared libraries, and sequenced samples. Analyzed data, prepared statistics, prepared figures, and wrote first draft manuscript.
Karthigayini Sivaprakasam: Analyzed raw RNA-sequencing data. Critiqued data and reviewed manuscript.
Whitney Hartstone: Assisted with laser capturing of PCs.
Sheng-Han Kuo: Critiqued data and reviewed manuscript.
Genevieve Konopka: Project design for RNA-sequencing analysis. Critiqued data and reviewed manuscript.
Elan D. Louis: Conceptualized project, critiqued data, and reviewed manuscript.
Phyllis L. Faust: Conceptualized project, project design, critiqued data, and manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
Dr. Martuscello, Ms. Hartstone and Dr. Sivaprakasam have no individual funding to report. Dr. Kuo has received research support from the National Institutes of Health: NINDS #R01 NS104423 (principal investigator), NINDS #R01 NS118179 (principal investigator), NINDS #R01 NS124854 (principal investigator). Dr. Konopka has received research support from the National Institutes of Health, NIMH #R01MH126481 (principal Investigator), NHGRI #R01HG011641 (principal Investigator), NINDS #UF1NS115821 (principal investigator), NICHD #R01HD099162 (co-investigator), NIMH #R01MH103517 (co-investigator); Simons Foundation for Autism Research #573689 (principal investigator); and The Welch Foundation #I-1997-20190330 (principal investigator). Dr. Louis has received research support from the National Institutes of Health: NINDS #R01 NS094607 (principal investigator), NINDS #R01 NS088257 (principal investigator), NINDS #R01 NS117745 (principal investigator), NINDS #R01 NS086736 (principal investigator), and NINDS #R01 NS124854 (co-investigator). Dr. Faust has received research support from the National Institutes of Health: NINDS #R01 NS086736 (co-investigator), NINDS #R01 NS088257 (principal investigator), NINDS #R01 NS117745 (principal investigator), NINDS #R01 NS124854 (principal investigator). No conflicts of interest to report from any author.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information

Supplemental Table 1
Demographic, clinical and neuropathological information for all samples sequenced (PNG 7496 kb)

Supplemental Table 2
Top expressed genes in PCs isolated by laser capture microdissection. The top 20 genes expressed across all samples sequenced are shown. Gene localization in cerebellar cortex showing PC specific = only expressed in PCs; PC enriched = strongly expressed in PCs along with limited expression in other cerebellar cell types; PC expressed (high cerebellum expression) = expressed in PCs and high expression throughout the cerebellum. Localization in the cerebellum obtained from The Human Protein Atlas and The Allen Brain Atlas(PNG 3868 kb)

Supplemental Table 3
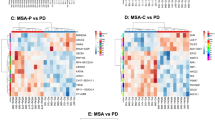
Differentially expressed pathways from ET versus Control analyzed by ssGSEA and compared via hierarchical cluster analysis. Pathways from heatmap (Fig. 4A) grouped with kindred gene ontology (GO) pathways shows various levels of dysregulation in control and ET cases. False discovery rate (FDR) is shown (PNG 5808 kb)

Supplemental Table 4
Differentially expressed pathways among ET samples analyzed by ssGSEA and compared via hierarchical cluster analysis. Pathways from heatmap (Fig. 5A) grouped with kindred pathways shows various levels of dysregulation and heterogeneity in ET cases. False discovery rate (FDR) is shown (PNG 6517 kb)

Supplemental Table 5
Cross validation analysis comparing PC enriched sequencing (LCM-seq) results to prior whole cerebellar cortex sequencing (whole lysate-seq) results (PNG 249 kb)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Martuscello, R.T., Sivaprakasam, K., Hartstone, W. et al. Gene Expression Analysis of Laser-Captured Purkinje Cells in the Essential Tremor Cerebellum. Cerebellum 22, 1166–1181 (2023). https://doi.org/10.1007/s12311-022-01483-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12311-022-01483-4