Abstract
Durum and bread wheat are two related species with different ploidy levels but a high similarity between the common A and B genomes. This feature, which allows a continuous gene flow between the two species, can be exploited in breeding programs to improve key traits in both crops. Therefore, durum wheat, despite covering only 5% of cultivated wheat worldwide, also represents an asset for the genetic improvement of bread wheat. Tetraploid wheat, with a very large availability of wild and domesticated accessions, durum landraces, and cultivars, offers a large gene reservoir to increase the genetic diversity of A and B genomes in bread wheat. Moreover, thanks to the possibility of crossing durum wheat with Aegilops tauschii, synthetic hexaploid lines are generated which show a much larger genetic diversity also in the D genome compared to common wheat. The genome sequences of wild emmer, durum, and bread wheat provide power tools for gene cloning and comparative genomics that will also facilitate the shuttling of genes between tetraploid and hexaploid wheats.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
8.1 Introduction
Durum wheat (tetraploid) and bread wheat (hexaploid) are two closely related species with potentially different adaptation capacities and only a few distinct technological properties that make durum semolina and wheat flour more suitable for pasta or bread and bakery products, respectively (Mastrangelo and Cattivelli 2021).
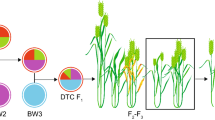
The history of wheat began with the domestication of wild emmer wheat (WEW, Triticum turgidum ssp. dicoccoides) in the mountains of the Fertile Crescent around 12–10 thousand years ago, which gave rise to the first domesticated form (domesticated emmer wheat, DEW, T. turgidum ssp. dicoccum) and a first domestication sweep related to Brittle rachis (Btr) trait. Then, human selection of natural mutations at a few loci associated with the domestication syndrome (i.e., Tg tenacious glume and Q compact spike) allowed for the selection of wheat forms with square-shaped spikes, soft glumes, and non-hulled grains improving with an improved threshing efficiency, grain size and uniformity, productivity, and suitable for a more widespread cultivation. This phenotypic evolution together with hybridization between different forms (Matsuoka 2011) led to free-threshing subspecies (T. turgidum ssp. turgidum, ssp. turanicum, ssp. polonicum, ssp. carthlicum, and ssp. durum, Fig. 8.1), all inter-fertile and sharing the same AABB genomic configuration. Hulled and free-threshing forms played a crucial role in the development of Mediterranean civilizations. They were at the base of early agricultural movements leading to agriculture systems based on tetraploid wheat. Among the different subspecies, durum wheat (T. turgidum ssp. durum) became the major cultivated form of tetraploid wheat during the last 3000 years. Nowadays, élite durum wheat cultivars (DWCs) and durum wheat landraces (DWLs) grow in different environments around the Mediterranean Basin and are of major importance for grain production and for staple food, respectively (Fig. 8.2).
The expansion of emmer cultivation toward the Transcaucasian corridor promoted an additional natural hybridization of tetraploid forms with Aegilops tauschii (genome DD) and the emergence of the hexaploid bread wheat (T. aestivum L. ssp. aestivum, genome AABBDD) (Dubcoksky and Dvorak 2007). As a result, durum and bread wheat share the A and B genomes and a long evolutionary history.
8.2 Tetraploid Genetic Resources
Most of wheat genetic diversity is contributed by tetraploid wheat genetic resources, particularly primitive tetraploids and wild and domesticated emmer. Indeed, the bottleneck effect, caused by the evolutionary recent hybridization events from which hexaploid wheat has evolved, has strongly limited its genetic diversity compared to tetraploid and diploid wheats (Cox 1997). Therefore, tetraploid wheat germplasm represents a strategic reservoir of alleles for both durum and bread wheat improvement (Marone et al. 2021). The International Maize and Wheat Improvement Center (CIMMYT) and the International Center for Agricultural Research in the Dry Areas (ICARDA) have the largest collections of tetraploid wheat, with approximately 27,500 and 22,500 accessions, respectively, including 22,000 and 20,000 durum wheat accessions. The ICARDA gene bank stores more than 15,700 accessions of DWL and traditional cultivars, while CIMMYT retains a larger collection of domesticated emmer wheat (around 3000 accessions). A wide tetraploid wheat diversity is conserved by the National Small Grains Germplasm Research Facility at USDA-ARS where approximately 12,500 accessions are conserved, with a large representation of primitive tetraploid subspecies (T. turgidum ssp. polonicum, ssp. carthlicum, ssp. turanicum, and ssp. turgidum) and more than 900 WEW accessions. Many national gene banks also retain important local germplasm resources which include historical materials and the predominant remaining landraces (Robbana et al. 2019).
Many studies have reported on the assembly and characterization of panels of tetraploid genotypes, from tens of genotypes to many hundreds of entries of wider origin. These germplasm collections are representative of (i) subspecies, (ii) specific geographic regions including local DWLs, historical cultivars, and modern DWCs, and (iii) breeding programs. They have been characterized for population structure, genome-wide molecular diversity, and linkage disequilibrium (LD)-decay rate as estimated with either multi-allelic (SSRs) and/or bi-allelic (DArT™, AFLPs) markers in earlier studies (Maccaferri et al. 2005; Mantovani et al. 2008; Laidò et al. 2013; Roncallo et al. 2019) or more recently with the Illumina iSelect 90K SNP array (Maccaferri et al. 2016; Saccomanno et al. 2018; N’Diaye et al. 2018) and the Axiom 35K array (Kabbaj et al. 2017). All these studies have generated an in-depth description of genetic diversity and differentiation within/among subspecies and subgroups with a focus on both temporal and spatial trends, particularly targeting the cultivated and DWL germplasm.
Following the publication of the first high-density SNP-based consensus map of tetraploid wheat (Maccaferri et al. 2015) and the first release of the durum wheat reference genome of cultivar SVEVO, many SSRs and iSelect 90K wheat SNPs have been anchored on the durum genome sequence, thus providing opportunities for genetic insights on relevant genomic regions (Maccaferri et al. 2019). Two recent collaborative studies provided germplasm panels and advanced in-depth analysis supporting a detailed knowledge at the molecular level of the historical loss of diversity events. The identification of favorable allelic combinations progressively accumulated over repeated breeding cycles is instrumental for a more effective management of breeding. In the first study, the International Durum Wheat Sequencing Consortium supported a comprehensive analysis of genetic diversity in tetraploids which entailed the organization of the single seed descent Tetraploid Germplasm Collection (TGC) and its genetic diversity analysis using the Illumina iSelect 90K SNP array, projected onto the Svevo genome (Maccaferri et al. 2019; Fig. 8.3).
Population structure of the Tetraploid Germplasm Collection (TGC) composed of 1856 accessions of tetraploid wheat. a Neighbor joining tree from Nei’s genetic distances on the TGC. b Principal component analysis plot of the TGC calculated based on genome-wide linkage disequilibrium-pruned SNPs. c Admixture analyses of the TGC with k (number of populations assumed for the analysis) from 2 to 20. Correspondence between branches and main tetraploid wheat taxa/populations based on Nei’s genetic distances, PCA and Admixture are indicated by color code (modified from Maccaferri et al. 2019). The analyses of population structure concord to highlight five major subpopulations: wild emmer wheat, domesticated emmer wheat, durum wheat landraces from Ethiopian, durum wheat landraces from Asian and Mediterranean regions, and durum wheat cultivars
At the same time, the Wheat Initiative through the durum wheat-expert working group supported the development of the Global Durum Panel (GDP), a collection targeting mainly the cultivated durum germplasm, and fully genotyped with the Illumina iSelect 90K wheat SNP array. The GDP genetic diversity is described in Mazzucotelli et al. (2020). The two collections have been assembled, seed increased and made freely available for research with the aim to facilitate the inventory, molecular and phenotypic characterization, and use of tetraploid genetic resources for durum and bread wheat improvement. The collections are maintained at ICARDA (Morocco), University of Bologna (Italy), and CREA-Research Centre for Genomics and Bioinformatics (Italy). Related information and genotypic data are accessible at the GrainGenes database (https://wheat.pw.usda.gov/GG3/global_durum_genomic_resources).
8.2.1 Wild Emmer Shows the Widest Range of Adaptation to Environment and Retains the Highest Level of Genetic Diversity Genome-Wide
WEW is an annual, predominantly self-pollinating allotetraploid species with large, elongated grains and brittle ears disarticulating at maturity into spikelets. Molecular data indicate that WEW is about 500,000 years old, resulting from a hybridization event between two wild diploid grasses that took place in the Fertile Crescent, probably in the vicinity of Mt. Hermon and the catchment area of the Jordan River where a center of WEW diversity has been reported (Dvorak and Akhunov 2005; Feldman and Kislev 2007). WEW is naturally distributed in the Near East Fertile Crescent with two major races which are geographically, morphologically, and genetically distinct: (1) the north-eastern part of the Fertile Crescent, with main populations found in north- and central-eastern Turkey, western Iran, and northern Iraq; (2) the western race found in the southern Levant, including Syria, Lebanon, Jordan, and Israel. Apart from dense and frequent natural populations found in the upper Jordan valley catchment area in Israel, and massive stands on the basalt slopes of the Karacadağ (Şanlıurfa and Diyarbakır provinces) in Turkey, WEW currently displays a patchy distribution in the region, with populations being semi-isolated or isolated. Its habitats range in altitude from 100 m below sea level up to 1800 m above sea level, with very different climatic regions from cool and humid Karacadağ Mountains to hot and dry valleys in Israel (Nevo et al. 2002).
The domestication dynamics of WEW are still unclear, though several pieces of the puzzle have been identified. In present days, the south-eastern Turkish subpopulations are more closely related to DEWs than any other wild emmer populations, but the monophyletic origin of DEW is still debated. Whole genome analysis based on multi-locus assays pointed out that the wild emmer populations from the Karacadağ region west of Diyarbakir and from the Sulaimanyia region along the Iraq/Iran border appeared the most closely related to DEW (Ozkan et al. 2011) with a further molecular indication in favor of the Diyarbakir WEW (Luo et al. 2007). Later analyses supported the reticulated origin including sharing phylogenetic signals with wild populations from all parts of the wild range (Civan et al. 2013). Recently, a study from Nave et al. (2021) focused on the Brittle Rachis gene (BTR1), a fundamental gene for wheat domestication present in the two homeologous copies BTR1-A and BTR1-B. Haplotype sequences showed that for the BTR1-A locus, the domestic BTR1-A-hap11 is highly related to the WEW founder haplotype BTR1-A-hap10 which is ubiquitous in both northern and southern Fertile Crescent, while for the BTR1-B copy, the domesticated haplotype BTR1-B-hap8 was derived from the wild haplotype BTR1-B-hap7 found only in the southern Levant. This indicated that at least part of the domestication process of WEW occurred outside of the “core area” of the northern part of the Fertile Crescent (Nave et al. 2021).
Each WEW race is genetically further subdivided in subpopulations with a pattern that mirrors the geographic origin (Luo et al. 2007; Ozkan et al. 2011; Badaeva et al. 2015; Maccaferri et al. 2019). Up to 12 well-distinct populations and subpopulations were identified by Admixture analysis in the TGC (Maccaferri et al. 2019; Fig. 8.4a). Moreover, it was shown that populations belonging to the eastern race were less diverse than those collected in the Levant. Notably, the genetic structure of the western population also correlates with differences in morphologic features. Indeed, most of the western populations belong to the horanum botanical variety and include accessions with a slender habit, while northern Israel is specifically inhabited by the subpopulation judaicum which includes tall accessions with upright habitus, wide spikes with large grains, and more fertile than the rest of WEWs in the western area (Poyarkova et al. 1991). Ecological variables also play an important role in shaping the genetic structure of WEW. Indeed, loci under positive selection significantly correlated with eco-geographical factors (e.g., geographic location, temperature, water availability, singly or in combination) for allele frequency suggesting that natural selection could have created regional divergence in WEW (Ren et al. 2013). An example of natural selection shaping WEW genetic diversity is provided by Yr15, a broad spectrum disease resistance gene cloned in WEW belonging to the family of tandem kinase-pseudokinase proteins (Klymiuk et al. 2018). Northern regions of Israel show climatic conditions more favorable for stripe rust pathogen development with respect to the southern regions. A large screening of wild emmer natural populations confirmed that Yr15 gene is present only in northern Israeli populations and distributed along a narrow mountain ridge of about 100 km from Mt. Carmel to Mt. Hermon regions, mainly at an elevation higher than 500 m above sea level (Klymiuk et al. 2019a; He et al. 2020). Thus, it seems that selection pressure exerted by the pathogen is affecting the host-parasite interactions and co-evolution and shapes the distribution of resistance genes among wild emmer populations.
Admixture analysis of wild emmer, emmer, and landraces included in the TGC, represented as bar plots of Q membership coefficients (modified from Maccaferri et al. 2019). More in detail results of: a wild emmer wheat accessions with K from 2 to 12; b domesticated emmer wheat accessions with K from 2 to 20; c durum wheat landrace accessions with K from 2 to 12
8.2.2 Emmer Wheat, the First Domesticated Wheat
DEW was a widely cultivated staple crop in the Near East, ancient Mesopotamia, and Egypt for over 7000 years during the Neolithic period. The decline started in Turkey during the Bronze Age about 5000 years ago when it was replaced by naked wheats (T. durum and/or T. aestivum), while in Europe its cultivation continued until about 2000 years ago with a long and slow decline. Today, DEW can be found only in marginal areas and some isolated traditional farming communities in the Balkans and Mediterranean countries, Iran, Armenia, Ethiopia, Yemen, Oman, and India. Recently, it has been re-discovered by the organic food industry for bread and cookie production.
Geographical expansion of DEW was intimately associated with historical human migrations and spread from the Fertile Crescent with a typical star-like dispersal mode. Indeed, four major diffusion routes out of the Fertile Crescent have been postulated (Badaeva et al. 2015). The expansion of DEW was a long and complex process in which emmer genotypes became adapted to new habitats and climates. The genetic structure of DEW populations was affected, among other factors, by exchange of seed stock during migration and by gene flow between wild and domesticated wheats or between different locally adapted DEW populations.
Few studies have focused on the genetic structure of the DEW germplasm. A cluster analysis, based on karyotypic information on a comprehensive collection of 446 DEW lines from 47 countries, identified four groups (Balkan, Asian, European, and Ethiopian) that allowed the authors to postulate four major diffusion routes of the crop out of the Fertile Crescent (Badaeva et al. 2015). Notably, although specifically evolved in certain geographic regions, populations of DEW usually included representatives of more than one karyotypic group at different frequencies. This mixture of karyotypic groups probably originated from multiple crop introductions/exchanges by successive waves of colonizing civilizations, which swept across Europe, the Mediterranean, and Asia. This clustering partially agrees with the population structure highlighted by Liu et al. (2017a, 2017b) on a collection of 176 spring accessions representing a large portion of the worldwide genetic diversity in the gene pool of cultivated spring emmer wheat. Three major groups were recognized: an “African subpopulation” with mostly accessions from Ethiopia, Kenya, and Morocco, the “European subpopulation” with accessions from southern and western Europe, and an “Asian subpopulation” grouping accessions from eastern and western Asia. About DEW, the TGC collection included up to 335 unique, non-admixed DEWs comprehensively sampled from gene banks worldwide. Their analysis clearly evidenced the presence of at least six well-distinct main populations evolved based on the human-driven dispersal along the main already described migration routes (Maccaferri et al. 2019; Fig. 8.4b). The diversity analysis evidenced the already described high stratification level consisting of six main populations and up to 18 subpopulations corresponding to: (1–2) two distinct and ancestral populations from the southern Levant, (3) a population close relative of the southern Levant populations but distinct and evolved in southern Europe, (4) a population evolved along the dispersal route Turkey-to-Balkans, (5) a distinct population evolved along the Turkey-to-Transcaucasia/Iran, and (6) an early-separated population evolved and spread from Oman-to-India/Ethiopia.
All these data pointed out the presence of a considerable level of diversity naturally evolved post-domestication in adaptation to environments and well differentiated from the native Fertile Crescent (southern Levant and Turkey). The unique application of genome-wide association studies (GWASs) reported so far on DEW also indicated high genetic diversity. Indeed, the same collection showed to be a rich source of stripe rust resistance loci very useful for wheat improvement (Liu et al. 2017a, 2017b). Among the 51 loci for resistance including genes effective in multiple field environments or against multiple races, a large proportion mapped distantly from previously reported stripe rust resistance genes or QTLs and provide novel resistance loci. Notably, African germplasm showed a higher frequency of resistant genotypes to stripe rust than the other two subpopulations.
8.2.3 Variable Human and Environmental Pressures Have Affected Divergence of Durum Wheat Landraces
Similar to DEW, the southern Levant is the center of origin of T. turgidum ssp. durum (Vavilov 1951; Feldman 2001). The first evidence of durum wheat dates ~7500–6500 years ago (Faris 2014). Then, it spread throughout the same migration routes already described for DEW through substantially independent pathways with limited evidence of gene flow and/or admixture between DEW and DWL (Maccaferri et al. 2019).
The dispersal routes moved durum wheat west throughout the Mediterranean Basin up to the Iberian Peninsula, probably via trading by Phoenician merchants and along the caravan’ routes along the Sahara desert or the North African coasts (Bozzini 1988), and east through the Silk Road to Asia (Waugh 2010). Following another early dispersal route to Ethiopia, an independent origin of durum wheat by a separate domestication of naked emmer has been suggested to have occurred in Ethiopia and have originated T. durum ssp. abyssinicum which is morphologically different from other durum wheat accessions, with uncompact spikes and small purple seeds (Mengistu et al. 2015, 2016). In addition, natural and anthropogenic selection in DEW during human migration resulted in the establishment of local DWLs specifically adapted to a diversity of agro-ecological zones (Nazco et al. 2012).
Local landraces were progressively abandoned starting from the early 1970s due to their replacement with the improved, more productive, and genetically uniform semi-dwarf cultivars derived from the Green Revolution. This notwithstanding, empiric breeding aimed to exploit the phenotypic variability of DWLs resulted in traditional varieties still preferred by smallholder farmers in traditional farming systems of rural/marginal areas where modern intensive ones cannot be adopted and/or where this germplasm provides the required higher stress tolerance. These traditional DWLs are usually tall plants and are often cultivated for both grain and straw, where in case yield is too low or even fail due to high temperature and drought the straw can still be harvested. Thus, crop diversity managed by smallholder farmers in traditional agro-systems is the outcome of historical and current processes interacting at various spatial scales and influenced by local factors such as farming practices and environmental constraints. Due to their evolutionary dynamics, landraces strongly represent the diversity of semiarid and marginal conditions. Indeed, evidence supports the hypothesis that DWLs harbor the largest source of biodiversity within the cultivated durum germplasm, including documented resilience to abiotic stresses and resistance to pests and diseases which could be used to enrich the modern wheat genetic repertoire for the improvement of commercially valuable traits (Lopes et al. 2015).
Many studies have focused on panels of landrace accessions from a restricted country/area, as those from southern Italy (Marzario et al. 2018; Mangini et al. 2018), Iran (Seyedimoradi et al. 2016), Spain (Giraldo et al. 2016; Ruiz et al. 2012), Tunisia (Robbana et al. 2019; Slim et al. 2019; Ouaja et al. 2021), Turkey and Syria (Baloch et al. 2017), Palestine, Jordan and Israel (Abu-Zaitoun et al. 2018), Morocco (Kehel et al. 2013; Sahri et al. 2014), and Ethiopia (Mengistu et al. 2016; Alemu et al. 2020).
Different drivers of population restructuring have emerged from the analysis of these collections. A collection of 91 DWLs originating from a wide range of ecological conditions of soil, temperature, and water availability in Turkey and Syria showed a grouping pattern not associated with the geographical distribution of durum wheat, suggesting a high mixing of Turkish and Syrian landraces due to large exchange of genetic material among farmers, as an alternative to the lack of commercial varieties (Baloch et al. 2017). Higher admixture among landraces was also observed in Ethiopia although it is a country characterized by a wide range of agro-ecological conditions coupled with diverse farmers’ culture. Indeed, both the clustering of 167 DWLs by Alemu et al. (2020) and 287 Ethiopian DWLs by Mengistu et al. (2016) collected from major wheat-growing areas of the country did not reflect their geographical origin, suggesting admixture arose from the existence of historical seed exchanges involving regional and countrywide farming communities in Ethiopia. Moreover, Seyedimoradi et al. (2016) reported low correlation between genetic distances and geographical origin in a small panel of DWLs from different zones of Iran. Notably, the genetic analysis showed that the country of origin did not have any genetic footprint within the core collection of DWLs from Jordan, Palestine, and Israel, despite historical sociopolitical barriers present in this area during the last decades (Abu-Zaitoun et al. 2018). However, in the latter case adaptations to similar semiarid conditions might reflect no separation between neighboring countries. Conversely, the fingerprinting of a collection of the National Gene Bank of Tunisia (NGBT) for traditional varieties from different Tunisian agro-ecological zones has found a strong genetic stratification from north to south in Tunisia (Slim et al. 2019). Indeed, five subpopulations were identified, two of which appeared more strongly represented in germplasm collected in central and southern Tunisia, where environmental conditions at critical development phases of the plant are harsher. Notably, these subpopulations were underrepresented in modern varieties which were instead prevalent in the north, suggesting that traits for breeding more resilient varieties might be present in central and southern Tunisian traditional varieties. In Morocco, a stratification of the genetic diversity according to agro-ecological conditions (geography, but also water and temperature regimes) was recorded, related to the two distant regions Pre-Rif and Atlas Mountains which display very different environmental, cultural, and agronomic conditions (Kehel et al. 2013; Sahri et al. 2014). However, within each region, only a few patterns emerged from the genetic and morphologic characterization, as if distance does not represent a consistent barrier to genetic exchange (Sahri et al. 2014). Different hypotheses can explain these results, ranging from unconscious mixing by farmers in threshing areas, to unreliability of seed exchange networks (e.g., seed lots that do not correspond to the declared names) as well as limited farmers’ interest for durum wheat cultivation and seed production. These mixtures create strong opportunities to generate diversity through cross-fertilization and recombination but also would homogenize the pool of traditional varieties in the absence of human or environmental divergent pressures that maintain some differentiation between them.
In the TGC collection (Maccaferri et al. 2019), up to 947 accessions of durum and durum-related unique and relatively low in admixture were analyzed for population structure (Fig. 8.4c). The results showed six main populations corresponding to: (1) a local Turkey-to-Levant (in particular Syria) population, (2) the main Southern Levant-to-North Africa and Southern Europe (Italy, Spain, and Portugal) migration route, (3) a highly differentiated Ethiopian population subdivided into two subpopulations, (4) a Turkey-to-Transcaucasia/Russia route, (5) a well-distinct T. turgidum ssp. turanicum population developed in Iran/Iraq up to Afghanistan, and (6) a localized Greece-to-Balkans population including representatives of the T. turgidum ssp. turgidum.
Other comprehensive studies considering both molecular and phenotypic data considered wide panels of landrace accessions from a larger geographic area as the collection of 172 DWLs from 21 Mediterranean countries characterized by Royo et al. (2014) and Soriano et al. (2016, 2018). Germplasm from the Mediterranean area is of interest for traits relevant for adaptation to the climate changes since in this region wheat is mainly grown under rain-fed conditions and yield is often constrained by water and heat stress that are common during the grain-filling period due to the low and unpredictable seasonal rainfall. Thus, the above collection highlighted an evident relationship between the genetic stratification and the eco-geographic patterning, which was suggested to be the result of different physiological and genetic strategies to sustain yield according to prevalent climate conditions. Indeed, the 172 DWLs showed a genetic structure related to an eastern–western geographical pattern formed by four clearly defined groups: eastern Mediterranean, eastern Balkans and Turkey, western Balkans and Egypt, and western Mediterranean, in agreement with the dispersal pattern of wheat from east to west in the Mediterranean Basin (Soriano et al. 2016). Interestingly, this study also showed a reliable relationship between genetic and phenotypic population structures, the latter being based on yield, yield components, and crop phenology-related traits. A high number of spikes and harvest index were recorded in DWLs from the eastern Mediterranean Basin, in agreement with the findings of previous studies (Moragues et al. 2006; Royo et al. 2014) which demonstrated that durum wheat yield under warm and dry environments is determined mostly by the number of spikes per unit area, whereas kernel weight predominantly influences grain production in colder and wetter environments. Interestingly, using a subset of this collection, Soriano et al. (2018) identified 23 marker alleles with a differential frequency in DWLs from east and west regions of the Mediterranean Basin, which affected the mentioned agronomic traits. Eastern DWLs had higher frequencies than the western ones of alleles for increasing the number of spikes (chr. 1B), grains per m2 (chr. 7B), grain-filling duration (several marker-trait associations), reduced cycle length, and lighter grains (chr. 4A, 5B, and 6B).
8.2.4 Main Breeding Gene Pools Within the DWC Germplasm
Durum wheat genetic makeup became more complex at the beginning of the twentieth century when conscious breeding started by applying artificial hybridization and selection pressure for commercial purposes (Autrique et al. 1996; Pecetti and Annicchiarico 1998; De Vita et al. 2007). The first durum wheat breeding program, setup in southern Italy by Nazareno Strampelli, was initially based on the selection of pure lines from local landraces (Scarascia Mugnozza 2005), which in 1915 led to the release of the cultivar CAPPELLI, a pioneer cultivar which had a major global impact in the following years and to which many modern varieties can be traced back to. A second major impact was provided by the deployment of lines carrying dwarfing genes to increase harvest index. This was first carried out by Nazareno Strampelli in Italy using Rht8 from the variety AKAKOMUGI, from Japan, and several years later by Norman Borlaug in Mexico using Rht-B1b also from a Japanese variety called NORIN10. The dwarfing gene Rht-B1b was successfully transferred to the durum wheat CANDO in the 1960s and widely used in durum wheat breeding (Quick et al. 1976). The last decades have been characterized by several hybridizations occurring between different breeding programs or with relatives aiming at increasing productivity while ensuring genetic diversity and mega-cultivars that have crossed the boundaries of their country of origin (Ren et al. 2013). Thus, although Autrique et al. (1996) observed that a limited number of ancestral lines have contributed largely to the development of the modern durum wheat materials and that the molecular fingerprints of a few ancestors accounted for most of the molecular diversity detected in the cultivated gene pool, a more complex network has emerged from studies on the genetic diversity pattern of the most recent durum wheat germplasm.
Numerous studies reported on characterization of diverse panels made of DWCs (Maccaferri et al. 2005, 2006, 2011; Reimer et al. 2008; Condorelli et al. 2018) or related to a breeding program (N’Diaye et al. 2018). These works have identified a few main gene pools reflecting the genetic basis and breeding strategies involved in their development. Maccaferri et al. (2005) subdivided the DWCs into six major gene pools: (1) Italian group which includes varieties selected and released in Italy, (2) CIMMYT-ICARDA group with hallmark accessions derived from the CIMMYT-ICARDA breeding program and released in Mexico, Spain, Italy, and in several West Asia and North Africa countries, (3) French group encompassing lines released by French breeders and well adapted to a range of environments throughout central and south Europe, (4) Austrian–Australian group derived from Austrian or Australian breeding programs, (5) North American group with accessions selected in the Great Plains of the USA and Canada, and (6) southwestern US group constituted by representative of the germplasm cultivated in the southwestern region of the US under irrigation and commonly referred to as desert durum.
More recently, the analysis of wider collections, as reported by Kabbaj et al. (2017) and by Mazzucotelli et al. (2020), provided the basis for separating the two CGIAR breeding programs (CIMMYT and ICARDA), as well as defining a subgroup of highly admixed varieties derived by exchange of materials among different breeding pools. This high exchange of materials was confirmed by the analysis of genetic diversity on the Global Durum Panel (GDP) clusters based on geography and breeding program of origin. Indeed, it was shown that most diversity remained among individuals within clusters, and only 13% of the total genetic variance could be captured by groups (Mazzucotelli et al. 2020). These insights also indicated that a good level of genetic diversity remains available within the breeding groups for direct exploitation, and there is even greater potential when considering exchanges between breeding groups. The 473 modern cultivars/breeding lines of the GDP were grouped into nine distinct groups organized as follows: old Italian elite, ICARDA, CIMMYT, Spanish/Argentinian elite germplasm, US desert durum, Australian elite, and at the opposite of diversity, the North American, Canadian, and French germplasm, the latter including the Evolutive Population (EPO, David et al. 2014) (Fig. 8.5). Founders of these modern cultivars were identified in western Asian durum landraces, North African Mediterranean landraces and central Asian, Turkey to Transcaucasia/Russian landraces, indicating that some landraces and durum primitive groups did not contribute to the genetic makeup of modern cultivars.
Neighbor joining tree of the Global Durum Panel (GDP) collection (modified from Mazzucotelli et al. 2020)
Interestingly, the level of LD decay rate, an important feature to be assessed when implementing GWAS, is quite differentiated comparing modern durum cultivars to landraces (Fig. 8.6). On average, LD decays to r2 = 0.3 (a generally accepted reference threshold in GWAS) in a range of physical distances of 4.21 Mb in modern durum, while the range decreases to 0.94 Mb in landraces. Thus, the landrace germplasm potentially offers higher genetic resolution in QTL mapping than modern varieties. However, these metrics are highly differentiated when referring to pericentromeric versus distal chromosome regions, with the physical-to-genetic ratio showing differences of 102–103 magnitude (Maccaferri et al. 2019).
Genome-wide linkage disequilibrium (LD) decay in respect to physical distance in the GDP collection for the two main groups of: a modern durum wheat germplasm and b durum landraces; critical distances are provided to three different r2 threshold values (0.5, 0.3, and 0.09 as for unliked markers) (Figure from Mazzucotelli et al. 2020)
Through GWAS on mentioned germplasm collections, specific phenotypic traits and/or the frequency of alleles at known loci for critical adaptation traits (vernalization requirement, response to photoperiod, heading date, plant height), for disease resistance, and for root morphology have been interestingly related to the population structure. For instance, Maccaferri et al. (2011) identified different patterns of allele frequency at the three major genes for wheat phenology and plant architecture (Vrn-A1, Ppd-A1, and Rht-B1) across the five subgroups present in a collection of elite durum mostly composed of Mediterranean germplasm. Most of the accessions were vernalization-insensitive and semi-dwarf, as expected for élite durum wheat materials. The vernalization-sensitive allele vrn-A1 was present in only six accessions, all but one (CLAUDIO) from ICARDA germplasm. Interestingly, in a subgroup made of cultivars bred for the semiarid areas, most genotypes carried the wild-type Rht-B1a allele and an almost fixed Ppd-A1 wild-type allele. In addition to conferring a tall phenotype, the wild Rht allele also increases the coleoptile length hence allowing for deeper sowing and better exploitation of soil moisture, thus making genotypes better suited for drought prone areas (Rebetzke et al. 2007). As to the Ppd locus, the wild allele confers lateness through photoperiod sensitivity in the Mediterranean environments, while the photoperiod-insensitive alleles dominate in most of the modern germplasm accessions. The same collection was evaluated for root system architecture, with a focus on root growth angle which is considered a fundamental trait to enhance the genetic capacity of the plant to acquire soil resources (Sanguineti et al. 2007; Maccaferri et al. 2016). Indeed, a narrow and deep root in contrast to a shallow ideotype can contribute to drought resistance and was found correlated with grain yield under harsh rain-fed conditions. Alleles contributing a narrow root growth angle were found to be present at relatively high frequencies in the modern high-yielding germplasm including the most recent cultivars from CIMMYT/ICARDA programs, the Italian, and the desert durum cultivars. The major root growth angle QTL detected on chromosome 6AL (Maccaferri et al. 2016) was also reported in a study on Ethiopian germplasm (Alemu et al. 2021).
Resistance to diseases is a relevant trait for durum varieties. Notably, the analysis of panels made of cultivars has frequently identified the leaf rust-resistant gene Lr14, a locus originally transferred from DEW YAROSLAV to common wheat (McFadden 1930), then identified in the Chilean DWC LLARETA-INIA and in diverse loosely related genetic materials, such as the CIMMYT line Somateria (Herrera-Foessel et al. 2008a), the Italian cultivars COLOSSEO (Maccaferri et al. 2008), and CRESO (Marone et al. 2009). The resistant haplotype at the Lr14 locus was found in many cultivars from Italian, CIMMYT and ICARDA breeding programs suggesting it was the most important source of resistance to leaf rust exploited by durum breeders. Another interesting example has been provided by the breeding for resistance to Hessian fly (Bassi et al. 2019). A major locus was identified on chromosome 6B in a group of Moroccan DWCs related to the cultivar NASSIRA. Pedigree analysis demonstrated kinship of these lines and traced back the origin of the locus to a resistant T. araraticum accession which had been used to introgress the resistance in locally adapted elite lines.
8.2.5 Selection Signatures
An exhaustive genome-wide analysis of changes in genetic diversity imposed by thousands of years of empirical selection and breeding was enabled by the Global Tetraploid Wheat Collection consisting of 1856 accessions representing the four main germplasm groups involved in tetraploid wheat domestication history and breeding (T. dicoccoides, T. dicoccum, DWLs, and DWCs) (Maccaferri et al. 2019). For each germplasm group, the pattern of diversity was assessed through a SNP-based gene diversity index (Fig. 8.7), then different metrics were used to detect selection signatures between evolutionary transitions, including both diversity reduction, and divergence/differentiation of allele frequency. WEWs showed the highest average diversity with only two pericentromeric regions (chr. 2A and 4A) with a lower-than-average diversity, thus the authors referred to WEW as the reference for assessing the reduction of diversity associated with domestication and breeding in tetraploid wheat. In total, 104 pericentromeric (average size 107.7 Mb) and 350 non-pericentromeric (average size 11.4 Mb) genomic regions reported co-occurrence of signals of selections in one or more evolutionary transitions.
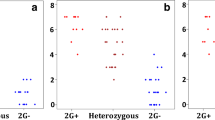
SNP-based diversity index (DI) for the main germplasm groups identified in the TGC (WEW, DEW, DWL, and DWC). DI is reported as a centered 25 SNP-based average sliding window (single SNP step). Top and bottom 2.5% DI quantile distributions are highlighted as red- and blue-filled dots, respectively (modified from Maccaferri et al. 2019)
Compared to WEW, each of the subsequently domesticated/improved germplasm group showed several strong diversity depletions that arose independently and were progressively consolidated through domestication and breeding. Consequently, the genome of DWCs revealed numerous regions showing near fixation of allelic diversity. Exceptions were observed for chromosomes 2A and 3A in the pericentromeric region where the DWCs showed an increased diversity as compared to DWL and DEW groups. The pericentromeric regions showed extensive signals of divergence/selection in the WEW-DEW and WEW-DWL transitions which highlights that most of the loss of diversity and divergence signatures occurred during domestication. The combination of this analysis with availability of the durum wheat reference genome allowed higher resolution analysis for the non-pericentromeric regions based on a comparative alignment between selection signals and wheat genes and QTLs relevant for domestication/improvement. For instance, the transition from DEW to durum wheat showed different depletion of diversity related to technological quality improvements. Indeed, the locus Glu-A1, coding for glutenin subunits and located at 500.8 Mb on chromosome 1A, which was reported to be nearly fixed in modern germplasm for null allele, was associated to a local strong signal of diversity reduction. Analogously, other extreme reductions in diversity were found colocated with grain yellow pigment content loci, including Psy-B1. Interestingly, among a set of 41 previously cloned loci, that have been most probably the target of selection, many colocated with regions marked by strong selection metrics. Intriguing examples were detected at critical loci for grain weight, a trait that has been strongly modified across the domestication and selection history for its relationship with grain yield (Fig. 8.8, Desiderio et al. 2019). Co-location was found for TaGW2 on chromosome 6A in the WEW-to-DEW transition and TaGW2 on 2B for both WEW-to-DEW and DEW-to-DWL transitions. Additionally, TaSus2-A1, TaSdr-A1, and TaCWI-A1 on chromosome 2A and their homeologs on 2B were associated to multiple extended signals in WEW-to-DEW and in DEW-to-DWL transitions, while the durum germplasm showed extended regions of low diversity.
Variation for kernel size and shape in tetraploid wheat (modified from Desiderio et al. 2019)
8.3 Tetraploid Wheat Genomes
The reference genome of the DWC (SVEVO v.1, Maccaferri et al. 2019) and of the WEW accession ZAVITAN (WEWseq v.1.0, Avni et al. 2017; assembly reviewed in Zhu et al. 2019 based on optical mapping) have been sequenced in recent years, hence allowing for a better exploitation of the genetic diversity of the tetraploid gene pool. The SVEVO genome sequence was assembled in 10.46 Gb including 0.5 Gb of unassigned scaffolds. Very similar numbers were produced for the genome sequence of ZAVITAN with an assembly size of 10.5 Gb including 0.4 Gb of unassigned scaffolds. The alignment of the durum wheat genome with high-density SNP genetic maps showed the typical pattern of recombination with highly recombinogenic distal chromosome regions and large pericentromeric regions nearly devoid of recombination.
A comparison of the two assemblies revealed strong overall synteny with high similarity in total gene number (66,559 high confidence genes in SVEVO vs. 65,012 in ZAVITAN) and repetitive element content (82.2% of the total assembly) and composition (Maccaferri et al. 2019). Nevertheless, a comparison of the orthologous gene pairs has highlighted several examples of presence–absence variations and of copy number variations as expected in a context of pangenome analysis where deletions and gene family expansions are frequently found (Walkowiak et al. 2020). For many years, genomic studies in tetraploid wheats were carried out based on the genomic resources developed in bread wheat thanks to the extensive sequence similarity and gene collinearity between the A and B genomes of the two species. As an example, the SNPs carried on the wheat 90K iSelect Infinium SNP assay (Wang et al. 2014), most of which originated from bread wheat, have been extensively used in genetic diversity and mapping studies in tetraploid wheat.
The availability of genome sequences for wild emmer and durum wheat is expected to facilitate genomic studies in tetraploid wheats. The projection of the 90K iSelect Infinium SNPs to the SVEVO and ZAVITAN genomes allowed to map genes and QTLs with higher precision and resolution. Moreover, once identified, the QTL confidence interval can now be used to search for candidate genes directly in the tetraploid genome. Such approaches have been used in mapping studies based on biparental segregating populations, as in the case of the identification of the SrKN gene for stem rust resistance from the tetraploid DWC KRONOS (Li et al. 2021); GWAS for different traits (Saccomanno et al. 2018; Aoun et al. 2021), and ultimately to better refine QTL regions through meta-QTL analysis for quality, abiotic and biotic stresses in durum wheat (Soriano et al. 2021). The molecular characterization of gene families can greatly help in the search of candidate genes for a particular trait, in studies on gene mapping, functional analysis, and comparative genomics, as in the case of the analysis of Hsp70 and glutathione S-transferases (GSTs) genes in different Triticum subspecies, with implications on evolution of these gene families and molecular mechanisms of their involvement in response to stress (Lai et al. 2021; Hao et al. 2021). The approach can also be focused on the characterization of a chromosomal locus, as seen for Gli-2 locus regions containing α-gliadin genes on A and B genomes of WEW (Huo et al. 2019). The availability of genome sequences is also of particular interest in transcriptomic studies such as those based on RNA-seq, in which projecting the reads onto a high-quality genome can greatly improve the accuracy and completeness of the analysis (Arenas et al. 2022). In a recent study, both the analysis of the translatome, the collection of all open reading frames that are actively translated, and in vivo RNA structure profiling were carried out to investigate the complex wheat RNA structure landscape in durum wheat. The translatome revealed subgenome asymmetry at the translational level, due to the strong impact of mRNA structure on translation, independent of GC content (Yang et al. 2021).
Matching mapping results with information regarding the gene content and annotation of genomic regions provides a huge advantage for fine mapping and gene/QTL cloning in tetraploid wheat. With the fully assembled ZAVITAN genome, the causal mutations in Brittle Rachis 1 (TtBtr1) genes controlling shattering, a key domestication trait, were identified (Avni et al. 2017). More recently, TdHMA3-B1, a gene encoding a metal transporter with a non-functional variant causing high accumulation of cadmium in grain, was rapidly cloned in SVEVO. Moreover, a wild functional allele, characterized by a very low frequency among DWCs, was rescued with great advantage for durum wheat breeding for cadmium accumulation in grain (Maccaferri et al. 2019). The utility of tetraploid wheat genome has also been shown for improvement of resistance to fungal diseases. The WEW derived Yr15, a gene for broad-spectrum resistance to stripe rust, was identified and cloned in a large mapping population developed by crossing the susceptible durum wheat line “D447” with introgression lines carrying Yr15 in the genetic background of “D447” (Klymiuk et al. 2018). Both ZAVITAN and SVEVO genomes were used to clone and functionally characterize Pm41, a powdery mildew resistance gene derived from WEW, which encodes a coiled-coil, nucleotide-binding site, and leucine-rich repeat protein (CNL) (Li et al. 2020).
Interestingly, all available wheat genomes are important in comparative genomic approaches to precisely characterize a chromosomal region and its gene content in gene cloning studies. This way, tetraploid genomes are instrumental for fine mapping and cloning of genes not only in emmer or durum wheat, but also in bread wheat as in the case of Ne2, a typical CNL gene responsible for hybrid necrosis in wheat (Si et al. 2021).
All these data indicate that wheat genomes for bread (IWGSC 2018), durum (Maccaferri et al. 2019), and wild emmer (Avni et al. 2017) wheat once merged in a unique wheat pangenome will provide an excellent asset for genetic studies in both tetraploid and hexaploid wheat. At the same time, the tetraploid and hexaploid wheat germplasms could be considered a unique gene pool from which to recruit genes and alleles useful for breeding in both species (Mastrangelo and Cattivelli 2021).
8.4 Tetraploid Germplasm for Bread Wheat Improvements (and Vice Versa)
The increasing demand for food and more sustainable crops and the increasingly evident climatic changes require the selection of new wheat DWCs with improved grain yield, protein content, and resistance to biotic and abiotic stresses (Foley et al. 2011; Soares et al. 2019). Achievement of this goal is hindered by the limited genetic variability present in wheat modern cultivars resulting from multiple domestication bottlenecks and breeding involving a few selected progenitors. This situation prompted the attention toward the use of DWLs, non-cultivated wheat subspecies, and wild relatives to contribute genes conferring traits of interest and, more in general, to increase the genetic diversity of the cultivated gene pool (Reif et al. 2005). Nowadays, the identification and characterization of these genes and/or related regions (QTLs) are assisted and accelerated by the recent technological advances in wheat genomics (Tuberosa and Pozniak 2014; Vendramin et al. 2019; Rasheed and Xia 2019), and their introgression into elite cultivars can be performed both with marker-assisted selection (MAS) and/or with transgenic strategies (Cobb et al. 2019; Gadaleta et al. 2008; Mores et al. 2021).
The close phylogenetic proximity between tetraploid and hexaploid wheat allows for the transfer of specific genes between the two species, as already occurred during wheat evolution (Dubcoksky and Dvorak 2007). Crosses between the two species are feasible, overcoming the problems due to necrosis and low fertility of hybrids (Klymiuk et al. 2019b; Othmeni et al. 2019). Although the superior adaptability of the hexaploid genome has made bread wheat the most cultivated wheat worldwide, tetraploid wheat exhibits greater genetic diversity, a highly desirable feature for present and future wheat breeding programs (Mastrangelo and Cattivelli 2021). WEW is probably the most relevant reservoirs of genetic diversity for durum and bread wheat with durum acting as a bridge between the wild relative and the bread wheat to facilitate the introgression of WEW traits in modern bread wheat lines (Maccaferri et al. 2015; Klymiuk et al. 2019b).
8.4.1 Transfer of Disease Resistance Genes
A number of important genes for biotic stress resistance have been transferred into common wheat from the primary gene pool of tetraploid wheats (T. turgidum ssp. dicoccoides, ssp. dicoccum, and ssp. durum), such as those related to the most dreadful and economically important diseases of wheat: rust, namely yellow rust (Yr—Puccinia striiformis f. sp. tritici); leaf rust (Lr—Puccinia triticina Eriks); stem rust (Sr—P. graminis f. sp. tritici), powdery mildew (Pm—Blumeria graminis f. sp. tritici), and Fusarium head blight (FHB; Fusarium graminearum). The list of rust resistance genes identified and transferred from durum to hexaploid wheat includes: the yellow rust resistance genes Yr53 (chr. 2BL, Xu et al. 2013), Yr64, and Yr65 (chr. 1BS, Cheng et al. 2014); the leaf rust resistance genes Lr23 (chr. 2BS, McIntosh et al. 1995; Sibikeev et al. 2020), Lr61 (chr. 6BS, Herrera-Foessel et al. 2008b), and Lr79 (chr. 3BL, Qureshi et al. 2018), and the stem rust resistance genes Sr12 (chr. 3BL, Sheen and Snyder 1964), Sr13 (chr. 6AL, Simons et al. 2011; Zhang et al. 2017), Sr8155B1 (chr. 6AS, Nirmala et al. 2017), and SrKN (chr. 2BL, Li et al. 2021).
Other stem rust-resistant genes (Sr2, Sr13, and Sr14) have been transferred into common wheat from cultivated emmer. Sr2 (3BS, McIntosh et al. 1995), a recessive and race non-specific adult plant resistance gene, was transferred from the emmer variety YAROSLAV into common wheat Hope and represents a major success in resistant wheat breeding which has been deployed in many cultivars in the last 80 years and still confers an effective rust resistance. Sr14 (chr. 1BL) was identified in the cultivated emmer Khapli and introgressed into hexploid cultivar Steinwedel. Both Sr2 and Sr14 are currently important sources of resistance to Ug99 lineage races of stem rust (Singh et al. 2011). Sr13 (chr. 6AL), present in both in durum and cultivated emmer, was transferred to the common wheat variety KHAPSTEIN from the DEW KHAPLI. Sr13 confers resistance to all races in the Ug99 group (Jin et al. 2007), but the resistant responses are influenced by temperature and genetic background (McIntosh et al. 1995; Roelfs and Mcvey 1979). Lr53 and Lr64 mapped on chromosome 6BS and 6AL, respectively, were transferred from WEW to common wheat (Kolmer 2008; Dadkhodaie et al. 2011; Huang et al. 2016). Several stripe rust resistance genes derived from WEW (Yr15 on chr. 1BS, Yr35-6BS, Yr36-6BS, YrH52-1BS, and YrSM139-1BS) were mapped using T. durum × T. dicoccoides segregating populations and transferred into bread wheat using durum wheat as a “bridge” (Peng et al. 2000; Dadkhodaie et al. 2011; Hale et al. 2012; Yaniv et al. 2015; Zhang et al. 2016).
Durum wheat has been used as source of powdery mildew resistance genes (Mld, Pm3h and PmDR147) for bread wheat improvement (Miedaner et al. 2019). Mld (chr. 4B, recessive) was employed in wheat breeding in combination with other Pm resistance genes, such as Pm2 (chr. 5DS, Bennett 1984) and Pm3h (chr. 1AS, dominant, Yahiaoui et al. 2006), and probably originated from an Ethiopian durum wheat accession (Srichumpa et al. 2005). PmDR147 (chr. 2AL, dominant) was transferred into bread wheat cv. LAIZHOU 953 from the durum wheat accession DR147 (Zhu et al. 2004). Two powdery mildew resistance genes, formally named Pm5a and Pm4a, identified in cultivated emmer, were used for bread wheat improvement. Pm5a (chr. 7BL, recessive) (McIntosh et al. 1967) appeared in the varieties Hope and H-44 along with Sr2, while the dominant gene Pm4a (2AL, dominant; The et al. 1979) was transferred to bread wheat variety chancellor from the Indian emmer landrace Khapli (Briggle 1966). WEW is a main source of Pm resistance genes—twenty-one—for hexaploid wheat (Huang et al. 2016). A direct transfer from WEW into bread wheat was done for 13 of these, while for the others an identification/mapping after a crossing with durum wheat or a validation/mapping in durum background, followed by transfer into hexaploid wheat, was undertaken (Klymiuk et al. 2019b).
Even if several FHB resistance regions were identified in Triticum turgidum ssp., none provides a level of resistance comparable to that of Fhb1 (3BS) in bread wheat. Until now, only two hard red spring wheat cultivars resistant to FHB, STEELE, and REEDER, were developed from crosses in which two cultivated emmer accessions resistant to FHB were involved (Mergoum et al. 2005; Stack et al. 2003). Hessian fly [Hf—Mayetiola destructor (Say) (Diptera: Cecidomyiidae)] is an important pest of durum and bread wheat (Stuart et al. 2012). To date, 37 Hf resistance genes have been identified (Bassi et al. 2019; Li et al. 2013; Zhao et al. 2020). Among them, 15 (H6, H9-H11, H14-H19, H28, and H29 (all on chr. 1AS), H31 on chr. 5BS and H33 on chr. 3A) were identified in durum wheat, and one, Hdic (1AS), was derived from an accession of cultivated emmer wheat. Most of them, as for example H9-H11 and Hdic, have been introgressed into common wheat (Patterson et al. 1994; Carlson et al. 1978; Stebbins et al. 1982; Liu et al. 2005), but only few have been deployed in commercial cultivars.
While all the genes described above were identified in tetraploid wheat and then introgressed into bread wheat, several disease-resistant genes moved in the opposite direction. Noteworthy is the case of the introgression and validation of the bread wheat locus Fhb1 in three European durum wheat genotypes for resistance to Fusarium head blight (FHB) (Prat et al. 2017). Indeed, durum wheat is particularly susceptible to FHB, and limited genetic variation has been found so far within durum modern germplasm. On the contrary, Fhb1 is a major determinant of FHB resistance found in the hexaploid wheat SUMAI-3 (Anderson et al. 2001). Another successful example is provided by the common wheat broad resistance gene Lr34/Yr18/Sr57/Pm38/Ltn1 that was transferred into a Canadian cultivar by transgenesis (Rinaldo et al. 2017).
8.4.2 Transfer of Quality-Related Loci
Gpc-B1 (chr. 6BS, also known as NAM-B1), a gene responsible for high grain protein and mineral content, was first identified in WEW and cloned by a map-based approach (Uauy et al. 2006), then successfully introgressed into many durum and bread wheat cultivars (Tabbita et al. 2017). While many of the genes described above were identified in tetraploid wheat and then introgressed into bread wheat, several genes moved in the opposite direction. For instance, some glutenin loci responsible for gluten elasticity and extensibility were introgressed into durum to improve the quality of bread made with semolina. Two approaches have been undertaken to introgress bread wheat loci into durum wheat. The Glu-D1 (chr. 1DL) alleles associated with good baking quality were introgressed in durum wheat either using lines carrying a mutation in Pairing homolog-1 (ph1b) gene, thus promoting homoeologous recombination (Gennaro et al. 2012) or by standard crosses with triticale as intermediate (Lukaszewsky 2003). Similarly, the introgression of Glu-D1-1d and Glu-D1-2b from bread to durum wheat resulted in dough with stronger mixing features (Gadaleta et al. 2008). Ph1b-mediated chromosomal translocations (5DS–5BS) were also employed to produce a tetraploid wheat with soft grains by transferring the Hardness locus controlling kernel texture from bread wheat chromosome 5D (Boehm et al. 2017).
8.4.3 Transfer of Abiotic Stress-Related Loci
The introgression of specific alleles at Vernalization loci from winter bread wheat has led to durum wheat more adapted to cold environments (Longin et al. 2013). The tolerance to Al3+ in acidic soils of durum wheats was also improved by the transfer of TaMATE1B and TaALMT1 (chr. 4B and 4D, respectively) which confer a large tolerance in Al3+ tolerance among bread wheats (Delhaize et al. 2012; Han et al. 2016). The TaMATE1B gene, responsible for constitutive citrate efflux from root tips (Tovkach et al. 2013), showed a positive effect also on grain yield, probably due to an increased root growth and proliferation. Indeed, the transgenic durum line JANDAROI–TaMATE1B compared to JANDAROI per se showed a significantly higher total root biomass and produced from 25.3 to 49.0% higher grain yield under both well-watered and terminal drought conditions (Pooniya et al. 2020).
The other way round, examples of gene transfer between durum and bread wheat for abiotic stress tolerance are more limited due to the complex genetic bases of this trait. The great genetic diversity present in collections of tetraploid wheat accessions, from WEWs to DWCs, represents an asset for future efforts aimed at the identification of loci explaining a good fraction of the phenotypic variation for resistance abiotic stresses, to be introgressed into hexaploid wheat.
8.5 Synthetic Wheats
Hexaploid wheat (Triticum aestivum L.), subgenomes AABBDD, is a natural amphiploid derived from interspecific cross between the tetraploid wheat species T. turgidum L. (AABB) and the diploid grass Aegilops tauschii Coss. (DD). The origin of common wheat is non-monophyletic, indeed, at the time of wheat's origin 10,000 years ago, the formation of more than one interspecific amphiploid contributed to the creation of common wheat (Caldwell et al. 2004). The resulting bottleneck effect has limited its genetic diversity compared with tetraploid and diploid wheats (Cox 1997). For this reason, breeders are looking at tools to increase the genetic diversity to be exploited in mainstream breeding on a worldwide scale. Based on those efforts, two distinct approaches were deployed: production of amphiploids, known as synthetic hexaploids, between T. turgidum and Ae. tauschii, and direct hybridization between T. aestivum and Ae. tauschii. Both approaches involve backcrossing to T. aestivum (Cox et al. 2017). The direct hybridization approach aims at increasing the genetic diversity for the D genome, and this is an important issue in wheat breeding, as very low diversity values characterize this genome compared to A and B genomes (Poland et al. 2012; Maccaferri et al. 2015). Studies reporting the improvement of bread wheat for traits related to both agronomic performance (grain yield and quality) and resistance to fungal diseases and pests have been carried out via direct hybridization (Cox et al. 2017). Nevertheless, the reduced genetic diversity following the bottleneck at the origin of bread wheat also involves A and B genomes.
Accordingly, the development of synthetic hexaploid wheats (SHWs) allows for enhancing the diversity for all the three wheat genomes and for the direct transfer of loci for traits of interest from tetraploid to hexaploid wheat. The analysis of the population structure of a panel of 121 SHW lines genotypically characterized with 35,939 high-quality SNPs derived from genotyping-by-sequencing revealed that the percentage of SNPs on the D genome was nearly the same as the other two genomes (nearly 30%), demonstrating the effectiveness of this approach to enhance genetic diversity of the D genome (Bhatta et al. 2018a). When SHW and bread wheat groups were compared at level of the entire genome, the gene diversity of SHWs was from 33.2 to 50% higher compared with a sample of elite bread wheat cultivars in two distinct SHW panels (Bhatta et al. 2018a, 2019a). When these panels were used in GWAS, QTLs for yield and quality-related traits, as well as disease resistance, were identified on all the three genomes, underlying the importance of both durum wheat and Aegilops parents in increasing genetic diversity and providing alleles of interest for breeding of bread wheat (Bhatta et al. 2018b, c, 2019b). A relevant contribution of the durum parents has been revealed also for traits usually associated to the D genome. As an example, although tolerance to Al3+ toxicity has been mainly linked to the TaALMT1 gene carried by the D genome (Han et al. 2016), a GWAS with 300 SHW lines besides the effect of TaALMT1 has identified many other QTLs, mostly located on A and B genomes (Emebiri et al. 2020). Similar results were found in a GWAS with 173 SHWs for leaf, stem and yellow rusts, yellow leaf spot, Septoria nodorum, and crown rot (Jighly et al. 2016).
SHW lines have been also used as parents of segregating populations to identify QTLs for specific traits. In these studies, the SHW parent usually incorporates a variable number of loci from the Ae. tauschii parent, but some important QTLs are also contributed by the tetraploid parent of the SHW line. Some examples are available for root traits under drought stress conditions. Liu et al. (2020) analyzed a RIL population of 111 individuals derived from a bread wheat cultivar crossed to a SHW line, which incorporated mainly QTLs on the D genome, but also some QTLs, as those on chromosome 2B, which were probably derived from the tetraploid parent. A RIL mapping population derived from a cross between W7984 (synthetic) and OPATA 85 was evaluated for root length and root dry weight under water stress and control conditions. QTLs common to both water conditions and stress specific were identified on A, B, and D genomes (Ayalew et al. 2017). The same population together with a doubled haploid population derived from the same parents (W7984 and OPATA) were used to identify QTLs for number of crossover (Gutierrez-Gonzalez et al. 2019). Similar results, in terms of genetic contribution from A and B genomes, were found for traits related to resistance to stem rust (Dunckel et al. 2015; Sharma et al. 2021) and root rot (Mahoney et al. 2017).
Pshenichnikova et al. (2020) developed a SHW line from a cross between accessions of T. dicoccoides and Ae. tauschii. The resulting line (SYN6) was crossed with CHINESE SPRING to obtain a set of 21 substitution lines, each containing 20 chromosomes from CHINESE SPRING and one from SYN6. The 1A substitution resulted in a substantial reduction of root length and weight, while a chromosome 5D substitution led to a significant increase compared to the recipient and the donor lines under two contrasting irrigation regimes. Developing synthetic lines through interspecific crosses can have consequences on the genomic asset of the resulting lines. Sequence elimination can happen after allopolyploidization, increasing the divergence among homoeologous chromosomes. This phenomenon has practical consequences on bread wheat breeding, as it can cause the loss of important genes. A recent study in which the differences between synthetic and natural hexaploid wheat lines were investigated by utilizing a large germplasm set of primary synthetics and synthetic derivatives revealed that reproducible segment elimination occurrence was highly dependent on the choice of diploid and tetraploid parental lines and, that the almost complete short arm of chromosome 1B carrying loci important for grain quality, was eliminated in one line (Jighly et al. 2019). In a different study in which 1862 mapped loci were compared between synthetic wheat SHW-L1 and its parental lines Ae. tauschii AS60 (DD) and T. turgidum AS2255 (AABB), the D genome of SHW-L1 showed a higher number of eliminated loci following the allopolyploidization compared to the A and B genomes (Yu et al. 2017). At a phenotypic level, hybrid chlorosis can be observed in SHW lines, and genetic loci involved in this phenomenon have been identified on D genome (Nakano et al. 2015; Nishijima et al. 2018).
Despite the possibility of losing some loci of interest, SHW lines have been extensively used in bread wheat breeding. An example is given by the importance of SHW lines in breeding programs at CIMMYT, where more than 1500 SHWs have been developed since 1980s and thousands of crosses have been generated with bread wheat to obtain synthetic lines. With this approach, advanced lines with excellent performance for yield and other traits have been obtained, and more than 80 have also been released as cultivars and are widely grown (Rosyara et al. 2019). Some very promising lines were obtained with adaptation to specific environments. As an example, a large breeding program was aimed at developing and evaluating SHW lines derived from winter durum wheat germplasm from Ukraine and Romania crossed with Ae. tauschii accessions from the Caspian Sea region at CIMMYT. These populations, subjected to rigorous pedigree selection under dry, cold, disease-affected environments of the Central Anatolian Plateau, provided superior lines characterized by resistance to leaf, stripe and stem rust, common bunt, and soil-borne pathogens, with the contribution of both durum and Aegilops parents (Morgounov et al. 2018).
The breeding programs involving SHWs are also deploying genomic selection. Ninety-seven populations were developed using first back-cross, biparental, and three-way crosses between 33 primary SHW genotypes and 20 spring bread wheat cultivars at CIMMYT. Genomic estimated breeding values (GEBVs) of parents and synthetic derived lines were estimated using a genomic best linear unbiased prediction (GBLUP) model, and higher GEBVs of progenies were related to introgression and retention of positive alleles from SHW parents (Jafarzadeh et al. 2016). Different results were shown by Dunckel et al. (2017), who analyzed selected lines from double haploid and RIL populations between six different primary synthetics and the elite cultivar OPATA M85 chosen for grain yield and other important agronomic traits. Overall, the prediction models had only moderate predictive ability, slightly lower than expected based on traits’ heritability. Nevertheless, a more recent study, based on SHW populations and SHW derivatives coming from crosses between the primary SHWs and bread wheat cultivars, suggested that models with heterogeneous additive genetic variances may be suitable to predict breeding values in wheat crosses with variable ploidy levels (Puhl et al. 2021).
In conclusion, the loss of genetic diversity in bread wheat due to bottlenecks from polyploidy, domestication, and modern plant breeding can be compensated by introducing diversity in all the three wheat genomes from Ae. tauschii and durum wheat. The increasing use of SHWs and SHW derivatives worldwide and at CIMMYT indicates the success of these approaches in improving bread wheat for many traits of interest, from yield and yield-related traits to resistance to biotic and/or abiotic stresses. Considering the studies based on SHWs, a major limiting factor could be the low number of durum wheat genotypes used as tetraploid parents of the crosses for the development of primary SHWs, ALTAR 84 and LANGDON being among the most frequently employed. This is in contrast with the large number of Ae. tauschii accessions used to broaden the genetic diversity of the D genome. As durum wheat genomics comes of age (Tuberosa and Pozniak 2014) and haplotype variation linked with target phenotypes of key traits becomes increasingly available (Maccaferri et al. 2019; Mazzucotelli et al. 2020), genomics-assisted breeding for durum wheat undergoes a notable evolution, one that will be accelerated by widening the basis of the tetraploid germplasm harnessed for SHW development by choosing the most suitable parents among the most recent and diverse products of the durum wheat breeding, hence increasing the contribution of the tetraploid gene pool to hexaploid wheat improvement.
8.6 Conclusions and Perspectives
Recent studies and breeding approaches clearly indicate that the tetraploid and hexaploid wheats are merging in a unique gene pool, where it will become increasingly easy to recruit genes and alleles for tetraploid and hexaploid wheat breeding regardless of the genome configuration of the donor genotype (Mastrangelo and Cattivelli 2021). Wild and domesticated emmer wheat and T. turgidum subspecies and landraces are resources of outstanding importance for breeding of both durum and bread wheat, and there is increasing evidence of élite cultivars carrying genes recruited from emmer and wheat landraces. The different wheat genomes will soon merge in a unique wheat pangenome that will change forever the current breeding strategies. Genes will “shuttle” easily from wild accessions to cultivars and between tetraploid and hexaploid wheats making possible the fast introgression of new traits (e.g., diseases resistance and traits for coping with the effect of climate change) and the selection of varieties increasingly based on genome knowledge which will provide a accurate and holistic view of the wheat genome, while safeguarding and ensuring an adequate level of food security for mankind.
References
Abu-Zaitoun SY, Chandrasekhar K, Assili S et al (2018) Unlocking the genetic diversity within a middle-east panel of durum wheat landraces for adaptation to semi-arid climate. Agronomy 8:233. https://doi.org/10.3390/agronomy8100233
Alemu A, Feyissa T, Letta T et al (2020) Genetic diversity and population structure analysis based on the high density SNP markers in Ethiopian durum wheat (Triticum turgidum ssp. durum). BMC Genet 21:18. https://doi.org/10.1186/s12863-020-0825-x
Alemu A, Feyissa T, Maccaferri M et al (2021) Genome-wide association analysis unveils novel QTLs for seminal root system architecture traits in Ethiopian durum wheat. BMC Genomics 22:1–16. https://doi.org/10.1186/s12863-020- 0825-x
Anderson JA, Stack RW, Liu S et al (2001) DNA markers for Fusarium head blight resistance QTLs in two wheat populations. Theor Appl Genet 102:1164–1168. https://doi.org/10.1007/s001220000509
Aoun M, Rouse MN, Kolmer JA et al (2021) Genome-wide association studies reveal all-stage rust resistance loci in elite durum wheat genotypes. Front Plant Sci 12:640739. https://doi.org/10.3389/fpls.2021.640739
Arenas-M A, Castillo FM, Godoy D et al (2022) Transcriptomic and physiological response of durum wheat grain to short-term heat stress during early grain filling. Plants 11:59. https://doi.org/10.3390/plants11010059
Autrique E, Nachit MM, Monneveux P et al (1996) Genetic diversity in durum wheat based on RFLPs, morphophysiological traits, and coefficient of parentage. Crop Sci 36:735–742. https://doi.org/10.2135/cropsci1996.0011183X003600030036x
Avni R, Nave M, Barad O et al (2017) Wild emmer genome architecture and diversity elucidate wheat evolution and domestication. Science 357:93–97. https://doi.org/10.1126/science.aan0032
Ayalew H, Liu H, Yan G (2017) Identification and validation of root length QTLs for water stress resistance in hexaploid wheat (Triticum aestivum L.). Euphytica 213:126. https://doi.org/10.1007/s10681-017-1914-4
Badaeva ED, Keilwagen J, Knüpffer H et al (2015) Chromosomal passports provide new insights into diffusion of emmer wheat. PLoS ONE 10(5):e0128556. https://doi.org/10.1371/journal.pone.0128556
Baloch FS, Alsaleh A, Shahid MQA et al (2017) Whole genome DArTseq and SNP analysis for genetic diversity assessment in durum wheat from central fertile crescent. PLoS ONE 12:e0167821. https://doi.org/10.1371/journal.pone.0167821
Bassi F, Brahmi H, Sabraoui A et al (2019) Genetic identification of loci for Hessian fly resistance in durum wheat. Mol Breed 39:24. https://doi.org/10.1007/s11032-019-0927-1
Bennett FGA (1984) Resistance to powdery mildew in wheat: a review of its use in agriculture and breeding programmes. Plant Pathol 33:279–300. https://doi.org/10.1111/j.1365-3059.1984.tb01324.x
Bhatta M, Morgounov A, Belamkar V et al (2018a) Unlocking the novel genetic diversity and population structure of synthetic hexaploid wheat. BMC Genomics 19:591. https://doi.org/10.1186/s12864-018-4969-2
Bhatta M, Morgounov A, Belamkar V et al (2018b) Genome-wide association study reveals novel genomic regions for grain yield and yield-related traits in drought-stressed synthetic hexaploid wheat. Int J Mol Sci 19:3011. https://doi.org/10.3390/ijms19103011
Bhatta M, Morgounov A, Belamkar V et al (2018c) Genome-wide association study reveals favorable alleles associated with common bunt resistance in synthetic hexaploid wheat. Euphytica 214:200. https://doi.org/10.1007/s10681-018-2282-4
Bhatta M, Shamanin V, Shepelev S et al (2019a) Genetic diversity and population structure analysis of synthetic and bread wheat accessions in Western Siberia. J App Genet 60:283–289. https://doi.org/10.1007/s13353-019-00514-x
Bhatta M, Morgounov A, Belamkar V et al (2019b) Genome-wide association study for multiple biotic stress resistance in synthetic hexaploid wheat. Int J Mol Sci 20:3667. https://doi.org/10.3390/ijms20153667
Boehm JD Jr, Zhang M, Cai X, Morris CF (2017) Molecular and cytogenetic characterization of the 5DS-5BS chromosome translocation conditioning soft kernel texture in durum wheat. Plant Genome 10:3. https://doi.org/10.3835/plantgenome2017.04.0031
Bozzini A (1988) Origin, distribution, and production of durum wheat in the world. In: Fabriani G, Lintas C (eds) Durum: chemistry and technology. American Association of Cereal Chemists, St. Paul, MN, pp 1–16
Briggle LW (1966) Transfer of resistance to Erysiphe graminis f. sp. tritici from Khapli emmer and Yuma durum to hexaploid wheat. Crop Sci 6:459–461. https://doi.org/10.2135/cropsci1966.0011183X000600050020x
Caldwell KS, Dvorak J, Lagudah ES et al (2004) Sequence polymorphism in polyploid wheat and their D-genome diploid ancestor. Genetics 167:941–947. https://doi.org/10.1534/genetics.103.016303
Carlson SK, Patterson FL, Gallun RL (1978) Inheritance of resistance to Hessian fly derived from Triticum turgidum L. Crop Sci 18:1011–1014. https://doi.org/10.2135/cropsci1978.0011183X001800060027x
Cheng P, Xu LS, Wang MN et al (2014) Molecular mapping of genes Yr64 and Yr65 for stripe rust resistance in hexaploid derivatives of durum wheat accessions PI 331260 and PI 480016. Theor Appl Genet 127:2267–2277. https://doi.org/10.1007/s00122-014-2378-8
Civáň P, Ivaničová Z, Brown TA (2013) Reticulated origin of domesticated emmer wheat supports a dynamic model for the emergence of agriculture in the fertile crescent. PLoS ONE 8(11):e81955. https://doi.org/10.1371/journal.pone.0081955
Cobb JN, Juma RU, Biswas PS et al (2019) Enhancing the rate of genetic gain in public-sector plant breeding programs: lessons from the Breeder’s equation. Theor Appl Genet 132:627–645. https://doi.org/10.1007/s00122-019-03317-0
Condorelli GE, Maccaferri M, Newcomb M et al (2018) Comparative aerial and ground based high throughput phenotyping for the genetic dissection of NDVI as a proxy for drought adaptive traits in durum wheat. Front Plant Sci 9:893. https://doi.org/10.3389/fpls.2018.00893
Cox TS (1997) Deepening the wheat gene pool. J Crop Prod 1:145–168. https://doi.org/10.1300/J144v01n01_01
Cox TS, Wu J, Wang S et al (2017) Comparing two approaches for introgression of germplasm from Aegilops tauschii into common wheat. Crop J 5:355–362. https://doi.org/10.1016/j.cj.2017.05.006
Dadkhodaie NA, Karaoglou H, Wellings CR et al (2011) Mapping genes Lr53 and Yr35 on the short arm of chromosome 6B of common wheat with microsatellite markers and studies of their association with Lr36. Theor Appl Genet 122:479–487. https://doi.org/10.1007/s00122-010-1462-y
David J, Holtz Y, Ranwez V et al (2014) Genotyping by sequencing transcriptomes in an evolutionary pre-breeding durum wheat population. Mol Breed 34:1531–1548. https://doi.org/10.1007/s11032-014-0179-z
De Vita P, Li Destri Nicosia O, Nigro F et al (2007) Breeding progress in morpho-physiological, agronomical and qualitative traits of durum wheat cultivars released in Italy during the 20th century. Eur J Agr 26:39–53. https://doi.org/10.1016/j.eja.2006.08.009
Delhaize E, Ma JF, Ryan PR (2012) Transcriptional regulation of aluminium tolerance genes. Trends Plant Sci 17:341–348. https://doi.org/10.1016/j.tplants.2012.02.008
Desiderio F, Zarei L, Licciardello S et al (2019) Genomic regions from an Iranian landrace increase kernel size in durum wheat. Front Plant Sci 10:448. https://doi.org/10.3389/fpls.2019.00448
Dubcoksky J, Dvorak J (2007) Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316:1862–1866. https://doi.org/10.1126/science.1143986
Dunckel S, Crossa J, Wu S et al (2017) genomic selection for increased yield in synthetic-derived wheat. Crop Sci 57:713–725. https://doi.org/10.2135/cropsci2016.04.0209
Dunckel SM, Olson EL, Rouse MN et al (2015) Genetic mapping of race-specific stem rust resistance in the synthetic hexaploid W7984 x Opata M85 mapping population. Crop Sci 55:2580–2588. https://doi.org/10.2135/cropsci2014.11.0755
Dvorak J, Akhunov E (2005) Tempos of gene locus deletions and duplications and their relationship to recombination rate during diploid and polyploid evolution in the Aegilops-Triticum alliance. Genetics 171:323–332. https://doi.org/10.1534/genetics.105.041632
Emebiri LC, Raman H, Ogbonnaya FC (2020) Synthetic hexaploid wheat as a source of novel genetic loci for aluminium tolerance. Euphytica 216:135. https://doi.org/10.1007/s10681-020-02669-9
Faris J (2014) Wheat domestication: key to agricultural revolutions past and future. In: Tuberosa R, Graner A, Frison E (eds) Genomics of plant genetic resources. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-7572-5_18
Feldman M (2001) Origin of cultivated wheat. In Bonjean AP, Angus WJ (eds) The world wheat book: a history of wheat breeding. Intercept Limited, Andover, England, pp 3–58
Feldman M, Kislev EM (2007) Domestication of emmer wheat and evolution of free-threshing tetraploid wheat. Isr J Plant Sci 55:207–221. https://doi.org/10.1560/IJPS.55.3-4.207
Foley JA, Ramankutty N, Brauman KA et al (2011) Solutions for a cultivated planet. Nature 478:337–342. https://doi.org/10.1038/nature10452
Gadaleta A, Giancaspro A, Blechl AE et al (2008) A transgenic durum wheat line that is free of marker genes and expresses 1Dy10. J Cereal Sci 48:439–445. https://doi.org/10.1016/j.jcs.2007.11.005
Gennaro A, Forte P, Panichi D et al (2012) Stacking small segments of the 1D chromosome of bread wheat containing major gluten quality genes into durum wheat: transfer strategy and breeding prospects. Mol Breed 30:149–167. https://doi.org/10.1007/s11032-011-9606-6
Giraldo P, Royo C, González M et al (2016) Genetic diversity and association mapping for agro-morphological and grain quality traits of a structured collection of durum wheat landraces including subsp. durum, turgidum and diccocon. PLoS One 11:e0166577. https://doi.org/10.1371/journal.pone.0166577
Gutierrez-Gonzalez JJ, Mascher M, Poland J et al (2019) Dense genotyping-by-sequencing linkage maps of two Synthetic W7984×Opata reference populations provide insights into wheat structural diversity. Sci Rep 9:1793. https://doi.org/10.1038/s41598-018-38111-3
Hale I, Zhang X, Fu D et al (2012) Registration of wheat lines carrying the partial stripe rust resistance gene Yr36 without the Gpc-B1 high grain protein content allele. J Plant Regist 7:108–112. https://doi.org/10.3198/jpr2012.03.0150crg
Han C, Zhang P, Ryan PR et al (2016) Introgression of genes from bread wheat enhances the aluminium tolerance of durum wheat. Theor Appl Genet 129:729–739. https://doi.org/10.1007/s00122-015-2661-3
Hao Y, Xu S, Lyu Z et al (2021) Comparative analysis of the glutathione S-transferase gene family of four Triticeae species and transcriptome analysis of GST genes in common wheat responding to salt stress. Int J Genomics 6289174. https://doi.org/10.1155/2021/6289174
He Y, Feng L, Jiang Y et al (2020) Distribution and nucleotide diversity of Yr15 in wild emmer populations and chinese wheat germplasm. Pathogens 9:212. https://doi.org/10.3390/pathogens9030212
Herrera-Foessel SA, Singh RP, Huerta-Espino J et al (2008a) Identification and molecular characterization of leaf rust resistance gene Lr14a in durum wheat. Plant Dis 92:469–473. https://doi.org/10.1094/PDIS-92-3-0469
Herrera-Foessel S, Singh R, Huerta-Espino et al (2008b) Molecular mapping of a leaf rust resistance gene on the short arm of chromosome 6B of durum wheat. Plant Dis 92:1650–1654. https://doi.org/10.1094/PDIS-92-12-1650
Huang L, Raats D, Sela H et al (2016) Evolution and adaptation of wild emmer wheat populations to biotic and abiotic stresses. Annu Rev Phytopathol 54:279–301. https://doi.org/10.1146/annurev-phyto-080614-120254
Huo N, Zhu T, Zhang S et al (2019) Rapid evolution of α-gliadin gene family revealed by analyzing Gli-2 locus regions of wild emmer wheat. Func Int Genomics 19:993–1005. https://doi.org/10.1007/s10142-019-00686-z
IWGSC (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:7191. https://doi.org/10.1126/science.aar7191
Jafarzadeh J, Bonnett D, Jannink J-L et al (2016) Breeding value of primary synthetic wheat genotypes for grain yield. PLoS ONE 11:e0162860. https://doi.org/10.1371/journal.pone.0162860
Jighly A, Joukhadar R, Sehgal D et al (2019) Population-dependent reproducible deviation from natural bread wheat genome in synthetic hexaploid wheat. Plant J 100:801–812. https://doi.org/10.1111/tpj.14480
Jighly A, Alagu M, Makdis F et al (2016) Genomic regions conferring resistance to multiple fungal pathogens in synthetic hexaploid wheat. Mol Breed 36:127. https://doi.org/10.1007/s11032-016-0541-4
Jin Y, Singh RP, Ward RW et al (2007) Characterization of seedling infection types and adult plant infection responses of monogenic Sr gene lines to race TTKS of Puccinia graminis f. sp tritici. Plant Dis 91:1096–1099. https://doi.org/10.1094/PDIS-91-9-1096
Kabbaj H, Sall AT, Al-Abdallat A et al (2017) Genetic diversity within a global panel of durum wheat (Triticum durum) landraces and modern germplasm reveals the history of alleles exchange. Front Plant Sci 8:1277. https://doi.org/10.3389/fpls.2017.01277
Kehel Z, Garcia-Ferrer A, Nachit MM (2013) Using bayesian and eigen approaches to study spatial genetic structure of Moroccan and Syrian durum wheat landraces. Am J Mol Biol 3:17–31. https://doi.org/10.4236/ajmb.2013.31003
Klymiuk V, Yaniv E, Huang L et al (2018) Cloning of the wheat Yr15 resistance gene sheds light on the plant tandem kinase-pseudokinase family. Nat Commun 9:3735. https://doi.org/10.1038/s41467-018-06138-9
Klymiuk V, Fatiukha A, Fahima T (2019a) Wheat tandem kinases provide insights on disease-resistance gene flow and host-parasite co-evolution. Plant J 98:667–679. https://doi.org/10.1111/tpj.14264
Klymiuk V, Fatiukha A, Huang L et al (2019b) Durum wheat as a bridge between wild emmer wheat genetic resources and bread wheat. In: Miedaner T, Korzun V (eds) Applications of genetic and genomic research in cereals. Woodhead Publishing series in food science, technology and nutrition. Elsevier Ltd., Duxford, U.K, pp 201–230. https://doi.org/10.1016/B978-0-08-102163-7.00010-7
Kolmer J (2008) Lr63, Lr64. In: McIntosh RA, Dubcovsky J, Rogers WJ et al (eds) Catalogue of gene symbols for wheat: 2009 supplement, p 271 (Reference10550, p 273). Ann Wheat Newsl 55:256–278.
Lai D‑l, Yan J, Fan Y et al (2021) Genome‑wide identification and phylogenetic relationships of the Hsp70 gene family of Aegilops tauschii, wild emmer wheat (Triticum dicoccoides) and bread wheat (Triticum aestivum). 3 Biotech 11:301. https://doi.org/10.1007/s13205-021-02639-5
Laidò G, Mangini G, Taranto F et al (2013) Genetic diversity and population structure of tetraploid wheats (Triticum turgidum L.) estimated by SSR, DArT and pedigree data. PLoS One 8:e67280. https://doi.org/10.1371/journal.pone.0067280
Li CL, Chen MS, Chao SM et al (2013) Identification of a novel gene, H34, in wheat using recombinant inbred lines and single nucleotide polymorphism markers. Theor Appl Genet 126:2065–2071. https://doi.org/10.1007/s00122-013-2118-5
Li M, Dong L, Li B et al (2020) A CNL protein in wild emmer wheat confers powdery mildew resistance. New Phytol 228:1027–1037. https://doi.org/10.1111/nph.16761
Li H, Hua L, Rouse MN et al (2021) Mapping and characterization of a wheat stem rust resistance gene in durum wheat “Kronos.” Front Plant Sci 12:751398. https://doi.org/10.3389/fpls.2021.751398
Liu XM, Brown-Guedira GL, Hatchett JH et al (2005) Genetic characterization and molecular mapping of a Hessian fly-resistance gene transferred from T. turgidum ssp. dicoccum to common wheat. Theor Appl Genet 111:1308–1315. https://doi.org/10.1007/s00122-005-0059-3
Liu W, Maccaferri M, Chen X et al (2017a) Genome-wide association mapping reveals a rich genetic architecture of stripe rust resistance loci in emmer wheat (Triticum turgidum ssp. dicoccum). Theor Appl Genet 130:2249–2270. https://doi.org/10.1007/s00122-017-2957-6
Liu W, Maccaferri M, Rynearson S et al (2017b) Novel sources of stripe rust resistance identified by genome-wide association mapping in Ethiopian durum wheat (Triticum turgidum ssp. durum). Front Plant Sci 8 774. https://doi.org/10.1023/B:GRES.0000024164.80444.f0
Liu R-x, Wu F-k, Yi X et al (2020) Quantitative trait loci analysis for root traits in synthetic hexaploid wheat under drought stress conditions. J Int Agric 19:1947–1960. https://doi.org/10.1016/S2095-3119(19)62825-X
Longin CFH, Sieber A-N, Reif JC et al (2013) Combining frost tolerance, high grain yield and good pasta quality in durum wheat. Plant Breed 132:353–358. https://doi.org/10.1111/pbr.12064
Lopes MS, El-Basyoni I, Baenziger PS et al (2015) Exploiting genetic diversity from landraces in wheat breeding for adaptation to climate change. J Exp Bot 66:3477–3486. https://doi.org/10.1093/jxb/erv122
Lukaszewsky AJ (2003) Registration of six germplasms of bread wheat having variations of cytogenetically engineered wheat–rye translocation 1RS.1BL. Crop Sci 43:1137–1138
Luo MC, Yang ZL, You FM et al (2007) The structure of wild and domesticated emmer wheat populations, gene flow between them, and the site of emmer domestication. Theor Appl Genet 114:947–959. https://doi.org/10.1007/s00122-006-0474-0
Maccaferri M, Sanguineti MC, Noli E et al (2005) Population structure and long-range linkage disequilibrium in a durum wheat elite collection. Mol Breed 15:271–290. https://doi.org/10.1007/s11032-004-7012-z
Maccaferri M, Sanguineti MC, Natoli V et al (2006) A panel of elite accessions of durum wheat (Triticum durum Desf.) suitable for association mapping studies. Plant Genet Res 4:79–85. https://doi.org/10.1079/PGR2006117
Maccaferri M, Mantovani P, Tuberosa R et al (2008) A major QTL for durable leaf rust resistance widely exploited in durum wheat breeding programs maps on the distal region of chromosome arm 7BL. Theor Appl Genet 117:1225–1240. https://doi.org/10.1007/s00122-008-0857-5
Maccaferri M, Sanguineti MC, Demontis A et al (2011) Association mapping in durum wheat grown across a broad range of water regimes. J Exp Bot 62:409–438. https://doi.org/10.1093/jxb/erq287
Maccaferri M, Ricci A, Salvi S et al (2015) A high-density, SNP-based consensus map of tetraploid wheat as a bridge to integrate durum and bread wheat genomics and breeding. Plant Biotechnol J 13:648–663. https://doi.org/10.1111/pbi.12288
Maccaferri M, El-Feki W, Nazemi G et al (2016) Prioritizing quantitative trait loci for root system architecture in tetraploid wheat. J Exp Bot 67:1161–1178. https://doi.org/10.1093/jxb/erw039
Maccaferri M, Harris NS, Twardziok SO et al (2019) Durum wheat genome highlights past domestication signatures and future improvement targets. Nat Genet 51:885–895. https://doi.org/10.1038/s41588-019-0381-3
Mahoney AK, Babiker EM, See DR et al (2017) Analysis and mapping of Rhizoctonia root rot resistance traits from the synthetic wheat (Triticum aestivum L.) line SYN-172. Mol Breed 37:130. https://doi.org/10.1007/s11032-017-0730-9
Mangini G, Nigro D, Margiotta B et al (2018) Exploring SNP diversity in wheat landraces germplasm and setting of a molecular barcode for fingerprinting. Cereal Res Commun 46:377–387. https://doi.org/10.1556/0806.46.2018.033
Mantovani P, Maccaferri M, Sanguineti MC et al (2008) An integrated DArT-SSR linkage map of durum wheat. Mol Breed 4:629–648
Marone D, Del Olmo AI, Laidò G et al (2009) Genetic analysis of durable resistance against leaf rust in du-rum wheat. Mol Breed 24:25–39. https://doi.org/10.1007/s11032-009-9268-9
Marone D, Russo MA, Mores A et al (2021) Importance of landraces in cereal breeding for stress tolerance. Plants 10:1267. https://doi.org/10.3390/plants10071267
Marzario S, Logozzo G, David JL et al (2018) Molecular genotyping (SSR) and agronomic phenotyping for utilization of durum wheat (Triticum durum Desf.) ex situ collection from Southern Italy: a combined approach including pedigreed varieties. Genes 9:1–20. https://doi.org/10.3390/genes9100465
Mastrangelo AM, Cattivelli L (2021) What makes bread and durum wheat different? Trends Plant Sci 26:7. https://doi.org/10.1016/j.tplants.2021.01.004
Matsuoka Y (2011) Evolution of polyploid Triticum wheats under cultivation: The role of domestication natural hybridization and allopolyploid speciation in their diversification. Plant Cell Physiol 52:750–764. https://doi.org/10.1093/pcp/pcr018
Mazzucotelli E, Sciara G, Mastrangelo AM et al (2020) The Global Durum Wheat Panel (GDP): an international platform to identify and exchange beneficial alleles. Front Plant Sci 11:569905. https://doi.org/10.3389/fpls.2020.569905
McFadden ES (1930) A successful transfer of emmer characters to vulgare wheat. J Am Soc Agron 22:1020–1034
McIntosh RA, Luig NH, Baker EP (1967) Genetic and cytogenetic studies of stem rust, leaf rust, and powdery mildew resistances in Hope and related wheat cultivars. Aust J Biol Sci 20:1181–1192. https://doi.org/10.1071/BI9671181
McIntosh RA, Wellings CR, Park RF (1995) Wheat Rusts: An atlas of resistance genes (Jean K ed). CSIRO, Melbourne, VIC
Mengistu DK, Kiros AY, Pè ME (2015) Phenotypic diversity in Ethiopian durum wheat (Triticum turgidum var. durum) landraces. Crop J 3:190–199. https://doi.org/10.1016/j.cj.2015.04.003
Mengistu DK, Kidane YG, Catellani M et al (2016) High-density molecular characterization and association mapping in Ethiopian durum wheat landraces reveals high diversity and potential for wheat breeding. Plant Biotechnol J 14:1800–1812. https://doi.org/10.1111/pbi.12538
Mergoum M, Frohberg RC, Miller JD et al (2005) Registration of ‘Steele-ND’ wheat. Crop Sci 45:1163–1164
Miedaner T, Rapp M, Flath K et al (2019) Genetic architecture of yellow and stem rust resistance in a durum wheat diversity panel. Euphytica 215:71. https://doi.org/10.1007/s10681-019-2394-5
Moragues M, García del Moral LF, Moralejo M et al (2006) Yield formation strategies of durum wheat landraces with distinct pattern of dispersal within the Mediterranean Basin: I. Yield components. Field Crops Res 95:194–205. https://doi.org/10.1016/j.fcr.2005.02.009
Mores A, Borrelli GM, Laidò G et al (2021) Genomic approaches to identify molecular bases of crop resistance to diseases and to develop future breeding strategies. Int J Mol Sci 22(11):5423. https://doi.org/10.3390/ijms22115423
Morgounov A, Abugalieva A, Akan K et al (2018) High-yielding winter synthetic hexaploid wheats resistant to multiple diseases and pests. Plant Genet Res: Charact Utilization 16:273–278. https://doi.org/10.1017/S147926211700017X
N’Diaye A, Haile JK, Nilsen KT et al (2018) Haplotype loci under selection in Canadian durum wheat germplasm over 60 years of breeding: association with grain yield, quality traits, protein loss, and plant height. Front Plant Sci 9:1589. https://doi.org/10.3389/fpls.2018.01589
Nakano H, Mizuno N, Tosa Y et al (2015) Accelerated senescence and enhanced disease resistance in hybrid chlorosis lines derived from interspecific crosses between tetraploid wheat and Aegilops tauschii. PLoS ONE 10:e0121583. https://doi.org/10.1371/journal.pone.0121583
Nave M, Taş M, Raupp J et al (2021) The independent domestication of Timopheev’s wheat: Insights from haplotype analysis of the Brittle rachis 1 (BTR1-A) gene. Genes 12:338. https://doi.org/10.3390/genes12030338
Nazco R, Villegas D, Ammar K et al (2012) Can Mediterranean durum wheat landraces contribute to improved grain quality attributes in modern cultivars? Euphytica 185:1–17. https://doi.org/10.1007/s10681-011-0588-6
Nevo E, Korol AB, Beiles A et al (2002) Evolution of wild emmer and wheat improvement. Springer, Berlin
Nirmala J, Saini J, Newcomb M et al (2017) Discovery of a novel stem rust resistance allele in durum wheat that exhibits differential reactions to Ug99 isolates. G3-Genes Genom Genet 7:3481–3490. https://doi.org/10.1534/g3.117.300209
Nishijima R, Yoshida K, Sakaguchi K et al (2018) RNA sequencing-based bulked segregant analysis facilitates efficient D-genome marker development for a specific chromosomal region of synthetic hexaploid wheat. Int J Mol Sci 19:3749. https://doi.org/10.3390/ijms19123749
Othmeni L, Grewal S, Hubbart-Edwards S et al (2019) The use of pentaploid crosses for the introgression of Amblyopyrum muticum and D-genome chromosome segments into durum wheat. Front Plant Sci 10:1110. https://doi.org/10.3389/fpls.2019.01110
Ouaja M, Bahri BA, Aouini L et al (2021) Morphological characterization and genetic diversity analysis of Tunisian durum wheat (Triticum turgidum var. durum) accessions. BMC Genom Data 22:3. https://doi.org/10.1186/s12863-021-00958-3
Ozkan H, Willcox G, Graner A et al (2011) Geographic distribution and domestication of wild emmer wheat (Triticum dicoccoides). Genet Resour Crop Evol 58:11–15. https://doi.org/10.1007/s10722-010-9581-5
Patterson FL, Mass FB III, Foster JE et al (1994) Registration of eight Hessian fly resistant common winter wheat germplasm lines (Carol, Erin, Flynn, Iris, Joy, Karen, Lola, and Molly). Crop Sci 34:315–316. https://doi.org/10.2135/cropsci1994.0011183X003400010084x
Pecetti L, Annicchiarico P (1998) Agronomic value and plant type of Italian durum wheat cultivars from different eras of breeding. Euphytica 99:9–15. https://doi.org/10.1023/A:1018346901579
Peng JH, Fahima T, Roder MS et al (2000) High-density molecular map of chromosome region harboring stripe rust resistance genes YrH52 and Yr15 derived from wild emmer wheat, Triticum dicoccoides. Genetica 109:199–210. https://doi.org/10.1023/A:1017573726512
Poland JA, Brown PJ, Sorrells ME et al (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping by-sequencing approach. PLoS ONE 7:e32253. https://doi.org/10.1371/journal.pone.0032253
Pooniya V, Palta JA, Chen Y et al (2020) Impact of the TaMATE1B gene on above and below-ground growth of durum wheat grown on an acid and Al3+-toxic soil. Plant Soil 447:73–84. https://doi.org/10.1007/s11104-019-04231-6
Poyarkova H, Gerechteramitai ZK, Genizi A (1991) 2 Variants of wild emmer (Triticum dicoccoides) native to Israel: morphology and distribution. Can J Bot 69:2772–2789. https://doi.org/10.1139/b91-348
Prat N, Guilbert C, Prah U et al (2017) QTL mapping of Fusarium head blight resistance in three related durum wheat populations. Theor Appl Genet 130:13–27. https://doi.org/10.1007/s00122-016-2785-0
Pshenichnikova TA, Smirnova OG, Simonov AV et al (2020) The relationship between root system development and vernalization under contrasting irrigation in bread wheat lines with the introgressions from a synthetic hexaploid. Plant Growth Regul 92:583–595. https://doi.org/10.1007/s10725-020-00666-5
Puhl LE, Crossa J, Munilla S et al (2021) Additive genetic variance and covariance between relatives in synthetic wheat crosses with variable parental ploidy levels. Genetics 217:iyaa048. https://doi.org/10.1093/genetics/iyaa048
Quick JS, Miller JD, Donnelly BJ (1976) Cando North Dakota’s first semidwarf durum. North Dakota Farm Res 33:15–18
Qureshi N, Bariana H, Kumran VV et al (2018) A new leaf rust resistance gene Lr79 mapped in chromosome 3BL from the durum wheat landrace Aus26582. Theor Appl Genet 131:1091–1098. https://doi.org/10.1007/s00122-018-3060-3
Rasheed A, Xia X (2019) From markers to genome-based breeding in wheat. Theor Appl Genet 132:767–784. https://doi.org/10.1007/s00122-019-03286-4
Rebetzke GJ, Richards RA, Fettell NA et al (2007) Genotypic increases in coleoptile length improves stand establishment, vigour and grain yield of deep-sown wheat. Field Crop Res 100:10–23. https://doi.org/10.1016/j.fcr.2006.05.001
Reif JC, Zhang P, Dreisigacker S et al (2005) Wheat genetic diversity trends during domestication and breeding. Theor Appl Genet 110:859–864. https://doi.org/10.1007/s00122-004-1881-8
Reimer S, Pozniak CJ, Clarke FR et al (2008) Association mapping of yellow pigment in an elite collection of durum wheat cultivars and breeding lines. Genome 51:1016–1025. https://doi.org/10.1139/g08-083
Ren J, Sun D, Chen L et al (2013) Genetic diversity revealed by single nucleotide polymorphism markers in a worldwide germplasm collection of durum wheat. Int J Mol Sci 14:7061–7088. https://doi.org/10.3390/ijms14047061
Rinaldo A, Gilbert B, Boni R et al (2017) The Lr34 adult plant rust resistance gene provides seedling resistance in durum wheat without senescence. Plant Biotechnol J 15:894–905. https://doi.org/10.1111/pbi.12684
Robbana C, Kehel Z, Ben Naceur M et al (2019) Genome-Wide genetic diversity and population structure of Tunisian durum wheat landraces based on DArTseq technology. Int J Mol Sci 20:1352. https://doi.org/10.3390/ijms20061352
Roelfs AP, McVey DV (1979) Low infection types produced by Puccinia graminis f. sp. tritici and wheat lines with designated genes for resistance. Phytopathol 69:722–730. https://doi.org/10.1094/phyto-69-722
Roncallo PF, Beaufort V, Larsen AO et al (2019) Genetic diversity and linkage disequilibrium using SNP (KASP) and AFLP markers in a worldwide durum wheat (Triticum turgidum L. var durum) collection. PLoS One 14:e0218562. https://doi.org/10.1371/journal.pone.0218562
Rosyara U, Kishii M, Payne T et al (2019) Genetic contribution of synthetic hexaploid wheat to CIMMYT’s spring bread wheat breeding germplasm. Sci Rep 9:12355. https://doi.org/10.1038/s41598-019-47936-5
Royo C, Nazco R, Villegas D (2014) The climate of the zone of origin of Mediterranean durum wheat (Triticum durum Desf.) landraces affects their agronomic performance. Genet Resour Crop Evol 61:1345–1358. https://doi.org/10.1007/s10722-014-0116-3
Ruiz M, Giraldo P, Royo C et al (2012) Diversity and genetic structure of a collection of Spanish durum wheat landraces. Crop Sci 52:2262–2275. https://doi.org/10.1371/journal.pone.0166577
Saccomanno A, Matny O, Marone D et al (2018) Genetic mapping of loci for resistance to stem rust in a tetraploid wheat collection. Int J Mol Sci 19:3907. https://doi.org/10.3390/ijms19123907
Sahri A, Chentoufi L, Arbaoui M et al (2014) Towards a comprehensive characterization of durum wheat landraces in Moroccan traditional agrosystems: analysing genetic diversity in the light of geography, farmers’ taxonomy and tetraploid wheat domestication history. BMC Evol Biol 14:264. https://doi.org/10.1186/s12862-014-0264-2
Sanguineti MC, Li S, Maccaferri M et al (2007) Genetic dissection of seminal root architecture in elite durum wheat germplasm. Ann Appl Biol 151:291–305 https://doi.org/10.1111/j.1744-7348.2007.00198.x
Scarascia Mugnozza GT (2005) The contribution of Italian wheat geneticists: from Nazareno Strampelli to Francesco D’Amato. In: Proceedings of International Congress on the wake of the double helix: from the green devolution to the gene revolution, Bologna, Italy, pp 52–75
Seyedimoradi H, Talebi R, Fayaz F (2016) Geographical diversity pattern in Iranian landrace durum wheat (Triticum turgidum) accessions using start codon targeted poly morphism and conserved DNA-derived polymorphism markers. Env Expl Bio 14:63–68. https://doi.org/10.22364/eeb.14.09
Sharma JS, Overlander M, Faris JD et al (2021) Characterization of synthetic wheat line Largo for resistance to stem rust. G3 11:jkab193. https://doi.org/10.1093/g3journal/jkab193
Sheen SJ, Snyder LA (1964) Studies on the inheritance of resistance to six stem rust cultures using chromosome substitution lines of a Marquis wheat selection. Can J Genet Cytol 6:74–82. https://doi.org/10.1139/g64-010
Si Y, Zheng S, Niu J et al (2021) Ne2, a typical CC–NBS–LRR-type gene, is responsible for hybrid necrosis in wheat. New Phytol 232:279–289. https://doi.org/10.1111/nph.17575
Sibikeev S, Druzhin A, Gultyaeva E et al (2020) Use of the durum wheat gene pool in breeding of spring bread wheat. Russ Agric Sci 46:432–436. https://doi.org/10.3103/S1068367420050201
Simons K, Abate Z, Chao S et al (2011) Genetic mapping of stem rust resistance gene Sr13 in tetraploid wheat (Triticum turgidum ssp. durum L.). Theor Appl Genet 122:649–658. https://doi.org/10.1007/s00122-010-1444-0
Singh RP, Hodson DP, Huerta-Espino J et al (2011) The emergence of Ug99 races of the stem rust fungus is a threat to world wheat pro-duction. Annu Rev Phytopathol 49:465–481. https://doi.org/10.1146/annurev-phyto-072910-095423. (PMID: 21568701)
Slim A, Piarulli L, Chennaoui Kourda H et al (2019) Genetic structure analysis of a collection of Tunisian durum wheat germplasm. Int J Mol Sci 20:3362. https://doi.org/10.3390/ijms20133362
Soares JC, Santos CS, Carvalho SMP et al (2019) Preserving the nutritional quality of crop plants under a changing climate: importance and strategies. Plant Soil 443:1–26. https://doi.org/10.1007/s11104-019-04229-0
Soriano JM, Villegas D, Aranzana MJ et al (2016) Genetic structure of modern durum wheat cultivars and Mediterranean landraces matches with their agronomic performance. PLoS ONE 11:e0160983. https://doi.org/10.1371/journal.pone.0160983
Soriano JM, Villegas D, Sorrells ME et al (2018) Durum wheat landraces from east and west regions of the Mediterranean basin are genetically distinct for yield components and phenology. Front Plant Sci 9:80. https://doi.org/10.3389/fpls.2018.00080
Soriano JM, Colasuonno P, Marcotuli I et al (2021) Meta-QTL analysis and identification of candidate genes for quality, abiotic and biotic stress in durum wheat. Sci Rep 11:11877. https://doi.org/10.1038/s41598-021-91446-2
Srichumpa P, Brunner S, Keller B et al (2005) Allelic series of four powdery mildew resistance genes at the Pm3 locus in hexaploid bread wheat. Plant Physiol 139:885–895. https://doi.org/10.1104/pp.105.062406
Stack RW, Frohberg RC, Hansen JM et al (2003) Transfer and expression of resistance to Fusarium head blight from wild emmer chromosome 3A to bread wheat. In: Lewis J, Ward RW (eds) Proceedings of national fusarium head blight forum, Bloomington, MN. Wheat and Bar-ley Scab Initiative, Washington, DC, p 232
Stebbins NB, Patterson FL, Gallun RL (1982) Interrelationships among wheat genes H3, H6, H9, and H10 for Hessian fly resistance. Crop Sci 22:1029–1032. https://doi.org/10.2135/cropsci1982.0011183X002200050032x
Stuart JJ, Chen MS, Shukle R et al (2012) Gall midges (Hessian flies) as plant pathogens. Annu Rev Phytopathol 50:339–357. https://doi.org/10.1146/annurev-phyto-072910-095255
Tabbita F, Pearcec S, Barneix AJ (2017) Breeding for increased grain protein and micronutrient content in wheat: Ten years of the GPC-B1 gene. J Cereal Sci 73:183–191. https://doi.org/10.1016/j.jcs.2017.01.003
The TT, McIntosh RA, Bennett FGA (1979) Cytogenetical studies in wheat. IX. Monosomic analyses, telocentric mapping and linkage relationships of genes Sr21, Pm4 and Mle. Aust J Biol Sci 32:115–125. https://doi.org/10.1071/BI9790115
Tovkach A, Ryan PR, Richardson AE et al (2013) Transposon-mediated alteration of TaMATE1B expression in wheat confers constitutive citrate efflux from root apices. Plant Physiol 161:880–888. https://doi.org/10.1104/pp.112.207142
Tuberosa R, Pozniak C (2014) Durum wheat genomics comes of age. Mol Breed 34:1527–1530. https://doi.org/10.1007/s11032-014-0188-y
Uauy C, Distelfeld A, Fahima T et al (2006) A NAC gene regulating senescence improves grain protein, zinc, and iron content in wheat. Science 314:1298–1301. https://doi.org/10.1126/science.1133649
Vavilov N (1951) The origin, variation, immunity and breeding of cultivated crops. Chronicles Bot 13:1–36
Vendramin V, Ormanbekova D, Scalabrin S et al (2019) Genomic tools for durum wheat breeding: de novo assembly of Svevo transcriptome and SNP discovery in elite germplasm. BMC Genomics 20:1–16. https://doi.org/10.1186/s12864-019-5645-x
Walkowiak S, Gao L, Monat C et al (2020) Multiple wheat genomes reveal global variation in modern breeding. Nature 588:277–283. https://doi.org/10.1038/s41586-020-2961-x
Wang S, Wong D, Forrest K et al (2014) Characterization of polyploid wheat genomic diversity using a high-density 90,000 SNP array. Plant Biotechnol J 12:787–796. https://doi.org/10.1111/pbi.12183
Waugh DC (2010) The silk roads in history. Expedition 52:9–22
Xu LS, Wang MN, Cheng P et al (2013) Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI 480148 and its transfer to common wheat. Theor Appl Genet 126:523–533. https://doi.org/10.1007/s00122-012-1998-0
Yahiaoui N, Brunner S, Keller B (2006) Rapid generation of new powdery mildew resistance genes after wheat domestication. Plant J 47:85–98. https://doi.org/10.1111/j.1365-313X.2006.02772.x
Yang X, Yu H, Sun W et al (2021) Wheat in vivo RNA structure landscape reveals a prevalent role of RNA structure in modulating translational subgenome expression asymmetry. Genome Biol 22:326. https://doi.org/10.1186/s13059-021-02549-y
Yaniv E, Raats D, Ronin Y et al (2015) Evaluation of marker-assisted selection for the stripe rust resistance gene Yr15, introgressed from wild emmer wheat. Mol Breed 35:43. https://doi.org/10.1007/s11032-015-0238-0
Yu M, Guan L-L, Chen G-Y et al (2017) Allopolyploidy-induced rapid genomic changes in newly generated synthetic hexaploid wheat. Biotechnol Biotechnol Equip 31:236–242. https://doi.org/10.1080/13102818.2016.1273797
Zhang H, Zhang L, Wang C et al (2016) Molecular mapping and marker development for the Triticum dicoccoides-derived stripe rust resistance gene YrSM139-1B in bread wheat cv. Shaanmai 139. Theor Appl Genet 129:369–376. https://doi.org/10.1007/s00122-015-2633-7
Zhang W, Chen S, Abate Z et al (2017) Identification and characterization of Sr13, a tetraploid wheat gene that confers resistance to the Ug99 stem rust race group. Proc Natl Acad Sci USA 114:E9483–E9492. https://doi.org/10.1073/pnas.1706277114
Zhao L, Abdelsalam NR, Xu Y et al (2020) Identification of two novel Hessian fly resistance genes H35 and H36 in a hard winter wheat line SD06165. Theor Appl Genet 133:2343–2353. https://doi.org/10.1007/s00122-020-03602-3
Zhu Z, Kong X, Zhou R et al (2004) Identification and microsatellite markers of a resistance gene to powdery mildew in common wheat introgressed from Triticum durum. Acta Botan Sin 46:867–872
Zhu T, Wang L, Rodriguez JC et al (2019) Improved genome sequence of wild emmer wheat Zavitan with the aid of optical maps. G3 (Bethesda) 9:619–624. https://doi.org/10.1534/g3.118.200902
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2024 The Author(s)
About this chapter
Cite this chapter
Mazzucotelli, E. et al. (2024). Gene Flow Between Tetraploid and Hexaploid Wheat for Breeding Innovation. In: Appels, R., Eversole, K., Feuillet, C., Gallagher, D. (eds) The Wheat Genome. Compendium of Plant Genomes. Springer, Cham. https://doi.org/10.1007/978-3-031-38294-9_8
Download citation
DOI: https://doi.org/10.1007/978-3-031-38294-9_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-38292-5
Online ISBN: 978-3-031-38294-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)