Abstract
Genetic variation in brain size may provide the basis for the evolution of the brain and complex behaviours. The genetic substrate and the selective pressures acting on brain size are poorly understood. Here we use the Drosophila Genetic Reference Panel to map polymorphic variants affecting natural variation in mushroom body morphology. We identify 139 genes and 39 transcription factors and confirm effects on development and adult plasticity. We show correlations between morphology and aggression, sleep and lifespan. We propose that natural variation in adult brain size is controlled by interaction of the environment with gene networks controlling development and plasticity.
Similar content being viewed by others
Introduction
The brain plays a central role in controlling the social interactions of animals and their interactions with the environment. Visual, auditory, olfactory, tactile and gustatory sensory stimuli are integrated by the brain and result in a behavioural response appropriate for the context. Key higher order brain centres for integration and processing of sensory information are the mammalian cerebral cortex and the insect mushroom bodies (MBs). The size of the cerebral cortex and the MBs has been regarded as a proxy for cognitive ability and behavioural plasticity. For instance, the social brain hypothesis states that the evolution of the large brain size of primates is driven by the requirement to live in large groups1. By analogy, it was proposed that the large MBs in insects such as bees, wasps and ants are also driven by sociality2. However, the observation that solitary bee, wasp and ant species also have large MBs puts this notion in doubt and rather suggests that a common ancestor of these species had large MBs thereby providing the neural substrate on which sociality could evolve2,3. Other solitary insects with large MBs include cockroaches, herbivorous scarab beetles and some butterfly species4. Variation in brain size and MB size in particular is of great interest to understand the evolution of the brain and the mechanisms that govern this. The genetic substrate and the selective pressures acting on brain size as a quantitative trait are poorly defined.
Variation in brain size in the wild has been studied using two complementary approaches: interspecific comparative studies of the relationship between brain size and behavioural and environmental factors, and analyses of adaptive phenotypic plasticity5. Comparison of MBs in a broad spectrum of insects reveals big differences in size6,7. These size differences depend at least in part on significantly expanded neuroblast numbers. Drosophila MBs are derived from a total of eight MB neuroblasts, while Apis MBs are produced by four neuroblast clusters each consisting of 500 neuroblasts2,8. Thus, changes in developmental programmes are likely contributors to MB size differences. MB size in the adult insect also displays profound plasticity. Calyx volumes have been shown to be associated with age, sex and dominance behaviour in different paper wasp species9. In honeybees, experience modulates both dendritic spine morphology in the calyx and neuropil volume6,10. In ants, MB volumes vary between sexes and casts, and these volume changes are task dependent11,12. In Drosophila, MB fibre number has been related to age, sex and experience13,14.
The Drosophila MBs consist of ∼2,500 Kenyon cells projecting their axons rostroventrally through the peduncle, where at the MB heel they form distinct lobes8. All MB neurons are derived from eight MB neuroblasts and are subdivided in three groups: the α/β and α’/β’ neurons with neurites that project to a medial and a vertical lobe, and the γ neurons that only project medially. The γ neurons arise during early larval stages and initially project both medially and vertically. The γ neurons are remodelled during pupal stages to form their characteristic medial MB lobe. The α’/β’ and α/β neurons develop during late larval and pupal stages to form their respective lobes8.
In the current study, we use the inbred, sequenced lines of the Drosophila Genetic Reference Panel (DGRP)15,16 to demonstrate considerable natural genetic variation in length and width of the α and β MB lobes that is associated with variation in behavioural traits, and to perform genome wide association (GWA) analyses as a genetic screen to identify candidate genes affecting MB size. We identify the top candidate genes affecting MB size as those which have significantly associated genetic variants located within the gene and that are expressed in the MB neurons. In addition, we identify transcription factor-binding motifs that might be affected by variants in putative regulatory gene regions associated with MB size. We functionally validate the role of candidate genes and transcription factors in MB development and plasticity by knocking down their expression in the MBs during development and as adults using RNA interference (RNAi). We thus identify genes with known and unknown functions in MB development and plasticity; suggesting that the genes required to form these structures also cause naturally occurring quantitative genetic variation in size, and that GWA analyses of natural variation is an efficient gene discovery tool.
Results
Natural variation in mushroom body morphometry
We assessed morphology of the MB α and β lobes in 40 DGRP lines. These lobes are straightforward to visualize and we have previously optimized their morphometric analysis17,18,19. Animals of similar age and kept under constant environmental conditions were used throughout the study to minimize experience-dependent effects of mushroom body parameters (see Materials and Methods for more details).
Surprisingly, we found gross morphological defects in 31 of the lines (77.5%), including thin lobes, short lobes, missing lobes, abnormal lobes (such as defasciculation and general malformation or misguidance) and fusion of β lobes. Some of the phenotypes were highly penetrant in certain lines, while others were seen more sporadically (Fig. 1; Supplementary Data 1). We attribute these gross morphological defects to fixation of recessive mutations affecting MB architecture. We quantified subtle variations in MB morphology by measuring the length and width of the α and β lobes17,18,19. All four traits were significantly variable, with broad sense heritabilities of respectively 0.274 and 0.376 for the length and width of the α lobes; and 0.123 and 0.313 for the length and width of the β lobes (Fig. 2a; Supplementary Data 2). All four traits are positively correlated with each other (Supplementary Data 3).
(a–h) Anti-FasII staining visualizing the αβ lobes of the MBs in the adult brain of 3–7 day old males of the 40 DGRP lines (Scale bar, 50 μm). (a) Scheme representing the different MB lobes in the adult brain (b) DGRP–639 males show thin αβ lobes. (c) DGRP–705 males show missing α lobes (d) DGRP–517 show thin α lobes and missing β lobes. (e) DGRP–707 males show defasciculation of αβ lobes and abnormal guidance of β lobes. (f) DGRP–730 males show β lobe fusion. (g) DGRP–799 males show missing β lobes and abnormal guidance and outgrowth of α lobes (h) DGRP–303 males show thin and abnormally formed β lobes. (i,j) Quantification of variation in gross MB defects in the 40 DGRP lines (N=20 hemispheres). (i) α lobe defects. (j) β lobe defects.
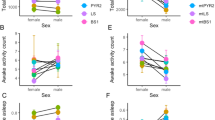
(a) Observed variation in MB morphology traits in the 40 lines of the DGRP (N=20 hemispheres, Mean±s.e.m.). (b) α-lobe length is negatively correlated with aggression (Pearson’s r=−0.410; P=0.009, N=20). (c) β-lobe width is negatively correlated with fitness (Pearson’s r=−0.322; P=0.042, N=20). (d) β-lobe length is negatively correlated with night time sleep (Pearson’s r=−0.323; P=0.044, N=20). (e) α-lobe width is negatively correlated with the number of sleep bouts during the day (Pearson’s r=−0.315; P=0.047, N=20). (f) β-lobe length is negatively correlated with the number of sleep bouts during the night (Pearson’s r=−0.361; P=0.023, N=20). (g) Genomic locations of variants associated with different MB morphology traits.
These DGRP lines have been extensively tested in various behavioural paradigms, enabling us to identify possible MB morphological correlates with behaviours. We assessed the correlations (r) for each MB parameter and aggressive behaviour, copulation latency, fitness, startle response, ethanol resistance, starvation resistance, lifespan, chill coma recovery and sleep traits20,21,22. All behavioural data were male specific except for mating and fitness traits, for which both sexes were pooled. We found significant negative correlations between aggression and α lobe length (r=−0.410, P=0.009), β lobe width and fitness (r=−0.322, P=0.042), β lobe length and night time sleep (r=−0.323, P=0.044), α lobe width with the number of sleep bouts during the day (r=−0.315, P=0.047) and β lobe length with the number of sleep bouts during the night (r=−0.361, P=0.023; Fig. 2b–f; Supplementary Data 4). Although MBs have been shown to be involved in the regulation of aggression and sleep, this is, to our knowledge, the first demonstration that natural variation in MB morphology is associated with these traits19,23,24,25,26.
GWA analyses of mushroom body morphological variation
We performed single marker GWA analyses for the four genetically variable traits (length and width of the α and β lobes). At a reporting P value threshold of P<10−5, we identified variants in annotated genes, as well as variants in intergenic regions (not located in an annotated gene) associated with MB morphology27. The former variants implicate candidate genes affecting MB size, while the latter could potentially contain regulatory motifs indicative of transcription factor-binding sites regulating these traits. We identified 104 variants associated with α lobe length, 109 variants associated with α lobe width, 90 variants associated with β lobe length and 138 variants associated with β lobe width (Supplementary Data 5). On average, 62% of the variants were associated with genes (variants in the coding sequence, introns and the 3′ and 5′ untranslated region (UTR)); the remainder was intergenic (variants outside the transcribed region of an annotated gene) (Fig. 2g).
We mapped 39 candidate genes associated with α lobe length, 37 with α lobe width, 35 with β lobe length and 54 with β lobe width. These comprised 139 unique genes, as 18 genes were associated with two MB traits and four were associated with three mushroom body traits (Supplementary Data 6). Six of the candidate genes are known to regulate MB development (starry night (stan), frizzled (fz), Protostome-specific GEF (PsGEF), Protein tyrosine phosphatase 10D (Ptp10D), misshapen (msn), vein (vn)), four act in pathways that modulate MB development but have themselves never been linked to MB development (APP-like protein interacting protein 1 (Aplip1), Ecdysone-induced protein 63E (Eip63E), Ecdysone-induced protein 75B (Eip75B), JIL-1) and two are involved in adult MB function (slowpoke (slo), pumilio (pum))28,29,30,31,32,33,34,35,36,37. No role in MB development or function has been reported for the remaining 127 genes. We performed gene ontology enrichment analyses to place the candidate genes into biological context38. Individual gene ontology categories and clusters were enriched for genes involved in development, morphogenesis, metamorphosis and behaviour (Supplementary Data 7 and 8).
To assess whether variants in non-coding regions that are associated with MB size could be in regulatory regions, we used the Multiple EM for Motif Elicitation tool to identify recurring motifs for these variants for each of the genetically variable traits39. We selected the three most significantly recurring motifs for each trait (Fig. 3). We then used the TOMTOM motif comparison tool to identify the transcription factor(s) most likely to bind to these motifs40, and categorized them based on their patterns of association with MB traits (all four traits, α and β lobe length, α and β lobe width, α lobe width and length, β width and length) (Supplementary Data 9). Of these 39 candidate transcription factors, longitudinals lacking (lola), tailless (tll) and ftz transcription factor 1 (ftz-f1) have been previously shown to be involved in MB development or function17,41,42. Dissatisfaction (dsf) and lethal of scute (l(1)sc) are expressed in the MBs43,44. Abdominal B (Abd-B) and hunchback (hb) interact with pathways with a previously described function in MB development or function45,46. Many of the other transcription factors are involved in (neuro-) developmental processes, but have no known role in the MBs.
(a) Motifs associated with α lobe length (motif 1: P=6,9 E-014; motif 2 P=5,4 E-010; motif 3: P=4,9 E-002). (b) Motifs associated with α lobe width (motif 1: P=2,6 E-015; motif 2 P=3,6 E-006; motif 3: P=2,5 E-012). (c) Motifs associated with β lobe length (motif 1: P=2,0 E-018; motif 2 P=3,2 E-010; motif 3: P=3,2 E-009). (d) Motifs associated with β lobe width (motif 1: P=4,1 E-014; motif 2 P=7,9 E-013; motif 3: P=1,4 E-005)
In summary, we identified 139 candidate genes and 39 candidate transcription factors associated with morphological variation in MB lobes. Although many of these genes are involved in development, morphogenesis and metamorphosis, only eight genes and four transcription factors have been previously shown to act in the MBs.
Functional validation analyses
We hypothesized that the most promising candidates for functional validation would be candidate genes that are expressed in the MBs. Therefore, we used the FlyLight database to restrict the number of candidate genes for functional validation tests to those with MB expression47. A total of 264 lines, containing enhancer sequences of 26 genes of the total 139 candidate genes, were present in the FlyLight database; of these, 44 drove expression in the MBs. Note that this may deviate from the actual number as the FlyLight lines may not always be a complete representation of the real expression pattern (own unpublished observations for the eyeless gene). In total, we found a total of 20 unique genes which showed expression in the MBs. We included pum as a candidate gene for functional validation because it is known to be expressed in the MB and play a role in synaptic plasticity in the adult brain, although a FlyLight line was not available37. We also used the FlyLight database to determine MB expression for the candidate transcription factors, of which 44 lines, containing enhancer sequences of 21 genes of the total 39 candidate transcription factors, were present in the FlyLight database (Supplementary Data 10). Thirty-two of these lines showed MB expression. In total these represented 17 unique transcription factors. We verified the MB expression of all lines by driving green fluorescent protein (GFP)–CD8 expression with the different GAL4 lines followed by anti-GFP immunostaining (Supplementary Data 10), and confirmed all but one (acj6). In addition, we determined that all enhancer sequences in the FlyLight lines corresponded to the reported gene by means of PCR (Supplementary Data 10). In summary, we confirmed MB expression of 20 candidate genes and 16 candidate transcription factors associated with one or more MB traits, the large majority of which are novel with respect to a role in MB function.
We used targeted RNAi knockdown in the MBs using the OK107-Gal4 driver to validate the role of the candidate genes and transcription factors in MB morphology (Supplementary Data 10). This driver is expressed in all MB neuroblasts, ganglion mother cells and neurons from embryonic stages onwards48. For the candidate genes we focused on the genes that are expressed in the MBs themselves, for the transcription factors we analysed all 39 genes. We tested one RNAi line for each of the 60 genes. OK107-Gal4 driven knockdown of Eip75B, Mrtf and ftz-f1 resulted in lethality, therefore these genes could not be evaluated for MB phenotypes. For the remaining genes, we evaluated gross as well as subtle MB phenotypes of the RNAi knockdown genotypes compared to the control. No control animals had gross MB abnormalities, but 24 of the 57 remaining genes tested showed gross defects in MB morphology. These defects included thin and missing α lobes and thin, short, fused and missing β lobes (Fig. 4; Supplementary Data 11). A total of 10 genes had more subtle, quantitative effects on MB size. misshapen (msn) affected all four traits; lola affected α lobe length and width and β lobe length, and slamdance (sda) affected α lobe width and β lobe length. crooked legs (crol), pou domain motif 3 (pdm3) and bric a brac 1 (bab1) specifically affected α lobe length and tailup (tup), jim, lethal (3) neo8 (l(3)neo8) and trachealess (trh) specifically affected α lobe width (Fig. 5).
(a–d) Anti-FasII staining visualizing the αβ lobes of the MBs in the adult brain of 3–7 day old males (scale bar, 50 μm) (a) UAS-RNAi-AbdBTRIP35647, OK107-Gal4 (b) UAS-RNAi-CG33557VDRC23517, OK107-Gal4 (c) UAS-RNAi-lolaTRIP35721, OK107-Gal4 (d) UAS-RNAi-sdaTRIP 37494, OK107-Gal4. (e,f) Quantification of variation in gross MB defects on RNAi-mediated knockdown (N=20 hemispheres). (e) α lobe defects. (f) β lobe defects.
In many insect species, the MB is a dynamic structure in the adult brain, able to change in an experience-dependent manner6,9,13,14. Thus, it is possible that the candidate genes and transcription factors do not influence MB development per se, but rather adult plasticity. Support for this hypothesis comes from our observation that we identified pum, which is known for its role in plasticity in the adult MBs, as a candidate gene37. We could not include pum in the expression analysis since no FlyLight lines were available, but we did include it in our plasticity analysis as a positive control. To determine whether the candidate genes act in adult MB plasticity, we crossed RNAi lines targeting these genes with OK107-Gal4 in combination with a TubP-Gal80TS transgene to restrict the RNAi-mediated knockdown to adult stages by elevating the temperature to 29 °C. We only performed these analyses for the candidate genes. We identified eight genes that modulate adult MB plasticity (Fig. 6). pum and hedgehog (hh) affect plasticity in α lobe length and width and β lobe width and jim lovell (lov) affects plasticity of α and β lobe width. The other genes have specific effects on plasticity: slamdance (sda) and doublesex-Mab related 99B (dmrt99B) for α lobe length; bubblegum (bgm) for β lobe length; and spire (spir) and schlank for β lobe width.
(a) α lobe length. (b) α lobe width. (c) β lobe length. (d) β lobe width. (Kruskall–Wallis tests followed by Mann–Whitney tests with a Bonferroni correction; *P<0.05; **P<0.01; ***P<0.001; ****P<0.0001, N=20, Mean±s.e.m.) Red bars represent flies switched to 29 °C after eclosion and 4 days before dissection, blue bars represent control flies kept at 18 °C.
Discussion
Analysis of the genomic architecture of the DGRP panel highlighted extensive natural variation in the genomes of these flies16. These natural variations have been associated with multiple behaviours20,49,50. Furthermore, natural variation in volumes of brain regions, as well as in neuron numbers has been observed in humans and mice51,52. However, this is the first (to our knowledge) systematic analysis of natural variation in brain structures. We show that the DGRP exhibits variation in MB morphology with broad sense heritabilities ranging from 12 to 38%, which correlates with behavioural variation. In many lines, this variation is characterized by profound morphological differences. In humans and primates, overall brain size has been reported to be highly heritable, however, the reported heritability of the size of different substructures ranges from <5% to >80% (refs 53, 54).
Our GWA analyses identified many variants that are associated with variation in MB morphology, allowing us to identify genes and putative transcription factors involved in MB development. Of the 139 unique candidate genes, only eight have been previously reported to be involved in MB morphology or function28,29,30,32,33,34,35,37. Furthermore, more than one-third of the polymorphisms associated with variation in MB morphology were located in intergenic regions. We hypothesized that these intergenic regions, as well as other non-coding regions, would contain binding sites for transcription factors regulating MB morphology. Analysis of recurring motifs allowed us to select 39 putative binding transcription factors. Four of these had a known role in the MBs, including lola which was linked to all four MB traits, providing a proof of concept for our approach17; the other transcription factors were not known to affect MB morphology. We showed that the large majority of identified genes and transcription factors for which FlyLight lines are available indeed show expression in the MBs. Furthermore, we validated a functional role for these genes in MB development. Knockdown of many of the identified genes resulted in variation in MB morphology, ranging from sporadic gross defects to more subtle variations. The observed phenotypes may, however, be an underestimate as some RNAi constructs may not be sufficiently efficient to produce prominent phenotypes. At the same time, some false positives due to off-target effects cannot be excluded. Overall, the prominent MB expression of the identified genes and transcription factors and their causal effects reveal the biological relevance of our analysis and highlight the strength of GWA analyses of natural variation as an efficient gene discovery tool.
In addition to our analysis of the developmental roles of the candidate genes in the MBs, we performed the first systematic study of the genetic basis of variation in adult brain morphometry. We showed that our approach is valid to identify genes involved in structural plasticity in the MB. We investigated 22 genes, of which eight showed significant effects on adult MB morphology. Interestingly, pum, which is known for its role in behavioural and synaptic plasticity and is expressed in the MBs, had large effects37. We conclude that pum is also required for adult MB structural plasticity.
In mice, different brain regions are not constrained by developmental programmes and can thus evolve independently of other regions or overall brain size55. We focus on branches within one neuropil that derive from one neuron. The different MB traits are correlated, suggesting that these branches do not develop completely independently of each other. However, different variants or knockdown of different genes can have α or β lobe-specific effects, arguing for at least partially independent development within one neuropil. Branch-specific effects have been previously reported for RhoGAPp190 (ref. 56). This protein is important for dorsal branch stability both during development and in the adult brain. Interestingly, this protein and two of its interactors, Integrin and Src, have both been implicated in learning and memory56. Hence, it was hypothesized that RhoGap signalling could play a role in adult brain plasticity underlying behavioural changes56. Of note, two of the genes we identified, psGEF and CG30440, are involved in Rho signalling.
Increases in brain volumes, including cortex and hippocampus, have been associated with enriched environments and learning in many species6. MBs are known to play a prominent role in multiple behaviours. We show that morphometric variations in this neuropil correlate with changes in aggression, fitness and sleep, implicating natural variation in MB morphology in behavioural differences. It remains to be determined whether the morphological differences are causally linked to the variation in behaviour. Complex behaviours and MB morphology are known to be influenced by pleiotropic genes affecting distinct processes, and thus the two sets of observations could be independent consequences of the action of pleiotropic genes18,19,21. However, it is equally possible that the observed changes in MBs represent a direct physical correlate that forms the basis of behavioural alterations. Lobe-specific morphological differences have been shown to underlie different aspects of memory formation57. Our current work confirms a previously reported correlation between the length of the MB α lobe and aggressive behaviour18, thereby demonstrating that this relation is robust and independent of genetic background. The relationship between MB structure and aggression is further supported by the fact that 19 of the identified genes and transcription factors are also candidate genes for aggressive behaviour18,26. MB size has also been shown to be correlated with aggressive behaviour in wasps, suggesting a conservation of the role of MB in aggression across species9. The MBs have previously been implicated in the regulation of sleep and many other behaviours important for survival, such as olfaction, interpretation of visual input and learning and memory23,58,59. We find correlations of MB structure with sleep and fitness. Interestingly, many of the variants that we identified have been reported to be associated with both sleep and fitness traits in the DGRP (22 with sleep, 12 with fitness traits)20,60. We propose that the changes in structure reflect changes in MB function affecting these traits. Interestingly, sleep and fitness traits have also been associated with alterations in brain size in humans61.
Changes in adult MBs have been observed in many insects. The mechanisms underlying these alterations are unknown. In certain insects, including crickets, neurogenesis of kenyon cells in the adult brain has been shown. However, in Drosophila and honeybees, this process is absent and can thus not explain the observed plasticity in MB morphometry6,62. Alterations in MB lobe morphometry are reminiscent of metamorphosis in both honeybees and Drosophila. During metamorphosis, MBs are remodelled through MB fibre outgrowth and regression independent of cell body proliferation or death13. We propose that a similar process of fibre shedding and regrowth could form the basis of MB volume changes during adulthood. Previously, it has been hypothesized that such processes form the mechanistic basis of plasticity in odour templates in the MB63,64. However, many other processes could be involved in morphometric or volumetric changes in adult MB. These include changes in the size and number of terminal branches, swelling of the involved axons or axonal branches, as well as spine formation and retraction13,65. In addition, non-neuronal alterations, such as swelling of glia, could be contributing factors. Our data provide a first insight into the genetic mechanisms underlying structural plasticity in the adult MB. We identified genes with a variety of different functions, suggesting a complex interplay between different processes influencing MB morphology. Identification of spir, an actin nucleation protein involved in cytoskeleton organization, and hh, shown to modulate adult brain regeneration, can argue for remodelling due to fibre outgrowth66. On the other hand, identification of genes involved in synaptic plasticity, such as pum, can argue that volumetric change can be due to local remodelling of synapses on present axons37.
We report the first systematic analysis of the genetic basis of natural variation in brain structures in Drosophila. Our approach offers an unbiased identification of candidate genes involved in both development and adult plasticity. We provide insights in the genetic mechanisms underlying these variations, as well as in the biological relevance of these morphological changes in the light of behavioural alterations. The overall genetic architecture of variation in brain structure and other complex traits in Drosophila is very similar to high resolution GWAS in humans and involves a large number of loci with relatively small effect size. Furthermore, epistasis and pleiotropy have also been demonstrated to be conserved features of the genes involved in complex traits67. However, our results are different from the situation in humans where ‘missing heritability’ is frequently seen in studies of complex phenotypic traits68. We suspect that the main differences between our study and GWAS in human populations are that (1) we have used full sequence data; (2) we have stringently controlled the environment; (3) we have measured multiple individuals per genotype, thus improving the accuracy in estimating the true genetic value of each line; and (4) under a strictly additive model, the genetic variance of a population of inbred lines is twice that of the outbred population from which they were derived.
Methods
Fly stocks and husbandry
Flies were reared on standard cornmeal-agar-molasses Drosophila medium with a 12:12 light-dark cycle. For each cross, we used four to five virgin females and two males to obtain comparable levels of offspring density. Flies without tubP-Gal80TS were maintained at 25 °C. Flies with the tubP-Gal80TS allele were kept at 18 °C during development and switched to 29 °C 1–3 days after eclosion and 4 days before dissecting. All flies were between 3 and 7 days old at the time of dissection. We used the first 40 DGRP lines for which full sequencing data were available. Stocks from the Janelia Farm FlyLight Project, the TRIP collection, OK107-Gal4 and tubP-Gal80TS were obtained from the Bloomington stock center (Bloomington, IN, USA)69. KK and GD RNAi lines and their respective co-isogenic controls were ordered from the Vienna Drosophila Resource Center (Vienna, Austria).
Immunohistochemistry
Adult brains from male flies were dissected and processed for immunohistochemistry. Mouse monoclonal anti-fasciclin 2 antibody (1:200; Developmental Studies Hybridoma Bank, University of Iowa, IO, USA) was used to visualize MB α and β lobes17,18,19,26. Anti-GFP (1:500) was obtained from Abcam, Cambridge, USA. Immunostainings were documented with an Olympus BX61 epifluorescence microscope equipped with a DP70 digital camera. Confocal imaging was performed using an Olympus FV1000 confocal microscope. Since the possibility existed that fasII expression levels themselves differ between DGRP lines (for example, due to effects of SNPs on fasII regulation) we adjusted fluorescence intensities whenever needed so that unambiguous measurements could be made. Overall, we never observed lobe-specific changes in FasII expression levels or any other difference that could impair accurately measuring mushroom body lobe parameters.
Morphometric measurements
Length and width of the α and β lobes of the MBs were measured by using the analySIS FIVE software and expressed as values relative to the distance between the α lobe heels17,18,19. This internal calibration controls for differences in brain size when assessing variation in morphometric parameters among genotypes. Values were obtained for 10 brains for all genotypes, thus allowing analysis of 20 hemispheres.
Quantitative genetic analyses
We partitioned the variation in the length and width of the α and β MB lobes using random effects factorial analysis of variance models of form Y=L+H+L × H+ɛ, where L denotes DGRP line, H is hemisphere and ɛ the within line variance. We computed the variance components (σ2) for each of these terms using restricted maximum likelihood and estimated broad sense heritabilities (H2) as H2=(σL2+σL × H2)/(σL2+σL × H2+σɛ2).
GWA analyses of mushroom body size
We associated the line mean of each of the four MB traits with all segregating sites in the DGRP. We used the analysis of variance model Y=μ+SNP+ ɛ to evaluate each segregating site15, where Y is the phenotype, μ is the overall mean, SNP is the genotype and ɛ is the variance among line means within each genotype class. We used a nominal P value<10−5 as the reporting threshold for nominating candidate genes for functional validation. We annotated the site class of each significant variant as genic (variants in coding sequences, introns and UTR) or non-coding (variants outside the transcribed region of an annotated gene, UTR and introns)27.
Identification of putative transcription factors
We inferred possible regulatory functions of variants associated with MB size located in non-coding regions. We analysed 40 basepairs up- and down-stream of each of these variants as this size is likely to span possible transcription factor-binding sites70. We entered these sequences in the Multiple EM for Motif Elicitation tool (version 4.9.0), separately for each MB trait, to identify shared motifs associated with each trait39. We selected the three most significant motifs (P<0.05) with an occurrence of zero or one per sequence. We then used the TOMTOM Motif Comparison Tool to compare the significant motifs with a database of known Drosophila motifs and to predict putative transcription factors (P<0.05) binding to these motifs40. We then classified the transcription factors based on their association with MB traits: all four traits, β and α lobe length, β and α lobe width, α lobe width and length, β lobe width and length. Finally, we selected the most significant predicted transcription factors from each group for further analysis.
Expression and functional analyses
We used the FlyLight database69 to analyze the expression patterns of the candidate genes and transcription factors. We checked the availability of reporter lines in the FlyLight database. We included all transcription factors in this analysis. For genic variants we selected genes expressed in the MB for further analyses. We verified all FlyLight line inserts using PCR. The forward primer was vector specific (5′-AAATAGGGGTTCCGCGCACAT-3′) and reverse primers were made against the relevant fragments for each line (Supplementary Data 12). We verified MB expression of all FlyLight candidate genes described as showing MB expression, as well as for all available candidate transcription factor lines using immunohistochemistry (anti-GFP).
We performed MB specific, RNAi-mediated knockdown of the selected genes and transcription factors using OK107-Gal4 (Supplementary Data 10). For experiments without tubP-Gal80TS,, dissected males were between 4 and 7 days old and kept at 25 °C. OK107-Gal4 was crossed to the RNAi empty vector progenitor strain to obtain heterozygous control males. For experiments with tubP-Gal80TS, males were kept at 18 °C during development and switched to 29 °C after 1–3 days after eclosion and 4 days before dissection. Control males were kept at 18 °C throughout. For each condition, we analysed 20 hemispheres. The effect of RNAi-mediated knockdown on MB parameters during development was statistically analysed using non-parametric Kruskall–Wallis tests followed by Dunn’s post hoc tests. All lines were compared with the heterozygous y1,v1;OK107-Gal4 line. The effects of RNAi-mediated knockdown on MB parameters in the adult brain were statistically analysed using non-parametric Kruskall–Wallis tests followed by Mann–Whitney tests with a Bonferroni correction. Each genotype at 29 °C was compared with the same genotype at 18 °C. To control for possible effects due to the temperature shift, we used heterozygous y1,v1;OK107-Gal4; tubP-Gal80TS flies at 18 °C and shifted to 29 °C as a control.
Additional information
How to cite this article: Zwarts, L. et al. The genetic basis of natural variation in mushroom body size in Drosophila melanogaster. Nat. Commun. 6:10115 doi: 10.1038/ncomms10115 (2015).
References
Dunbar, R. I. The social brain hypothesis. Evol. Anthropol. 6, 178–190 (1998).
Farris, S. M. Evolution of complex higher brain centers and behaviors: behavioral correlates of mushroom body elaboration in insects. Brain Behav. Evol. 82, 9–18 (2013).
Withers, G. S., Day, N. F., Talbot, E. F., Dobson, H. E. & Wallace, C. S. Experience-dependent plasticity in the mushroom bodies of the solitary bee Osmia lignaria (Megachilidae). Dev. Neurobiol. 68, 73–82 (2008).
Lihoreau, M., Latty, T. & Chittka, L. An exploration of the social brain hypothesis in insects. Front. Physiol. 3, 442 (2012).
Gonda, A., Herczeg, G. & Merila, J. Evolutionary ecology of intraspecific brain size variation: a review. Ecol. Evol. 3, 2751–2764 (2013).
Fahrbach, S. E. Structure of the mushroom bodies of the insect brain. Annu. Rev. Entomol. 51, 209–232 (2006).
Strausfeld, N. J., Sinakevitch, I., Brown, S. M. & Farris, S. M. Ground plan of the insect mushroom body: functional and evolutionary implications. J. Comp. Neurol. 513, 265–291 (2009).
Lee, T., Lee, A. & Luo, L. Development of the Drosophila mushroom bodies: sequential generation of three distinct types of neurons from a neuroblast. Development 126, 4065–4076 (1999).
Molina, Y. & O'Donnell, S. Age, sex, and dominance-related mushroom body plasticity in the paperwasp Mischocyttarus mastigophorus. Dev. Neurobiol. 68, 950–959 (2008).
Coss, R. G., Brandon, J. G. & Globus, A. Changes in morphology of dendritic spines on honeybee calycal interneurons associated with cumulative nursing and foraging experiences. Brain Res. 192, 49–59 (1980).
Ehmer, B. & Gronenberg, W. Mushroom body volumes and visual interneurons in ants: comparison between sexes and castes. J. Comp. Neurol. 469, 198–213 (2004).
Gronenberg, W., Heeren, S., Ouml, & Lldobler, B. Age-dependent and task-related morphological changes in the brain and the mushroom bodies of the ant Camponotus floridanus. J. Exp. Biol. 199, 2011–2019 (1996).
Heisenberg, M., Heusipp, M. & Wanke, C. Structural plasticity in the Drosophila brain. J. Neurosci. 15, 1951–1960 (1995).
Technau, G. M. Fiber number in the mushroom bodies of adult Drosophila melanogaster depends on age, sex and experience. J. Neurogenet. 1, 113–126 (1984).
Mackay, T. F. et al. The Drosophila melanogaster Genetic Reference Panel. Nature 482, 173–178 (2012).
Huang, W. et al. Natural variation in genome architecture among 205 Drosophila melanogaster Genetic Reference Panel lines. Genome Res. 24, 1193–1208 (2014).
Yamamoto, A. et al. Neurogenetic networks for startle-induced locomotion in Drosophila melanogaster. Proc. Natl Acad. Sci. USA 105, 12393–12398 (2008).
Zwarts, L. et al. Complex genetic architecture of Drosophila aggressive behavior. Proc. Natl Acad. Sci. USA 108, 17070–17075 (2011).
Rollmann, S. M. et al. Pleiotropic effects of Drosophila neuralized on complex behaviors and brain structure. Genetics 179, 1327–1336 (2008).
Harbison, S. T., McCoy, L. J. & Mackay, T. F. Genome-wide association study of sleep in Drosophila melanogaster. Bmc Genomics 14, 281 (2013).
Ayroles, J. F. et al. Systems genetics of complex traits in Drosophila melanogaster. Nat. Genet. 41, 299–307 (2009).
Morozova, T. V. et al. Alcohol sensitivity in Drosophila: translational potential of systems genetics. Genetics 183, 733–745 (2009).
Joiner, W. J., Crocker, A., White, B. H. & Sehgal, A. Sleep in Drosophila is regulated by adult mushroom bodies. Nature 441, 757–760 (2006).
Baier, A., Wittek, B. & Brembs, B. Drosophila as a new model organism for the neurobiology of aggression? J. Exp. Biol. 205, 1233–1240 (2002).
Zwarts, L., Versteven, M. & Callaerts, P. Genetics and neurobiology of aggression in Drosophila. Fly (Austin) 6, 35–48 (2012).
Edwards, A. C., Zwarts, L., Yamamoto, A., Callaerts, P. & Mackay, T. F. Mutations in many genes affect aggressive behavior in Drosophila melanogaster. BMC Biol. 7, 29 (2009).
dos Santos, G. et al. FlyBase: introduction of the Drosophila melanogaster Release 6 reference genome assembly and large-scale migration of genome annotations. Nucl. Acids Res. 43, D690–D697 (2015).
Shimizu, K., Sato, M. & Tabata, T. The Wnt5/planar cell polarity pathway regulates axonal development of the Drosophila mushroom body neuron. J. Neurosci. 31, 4944–4954 (2011).
Zhang, W. et al. A developmentally regulated splice variant from the complex lola locus encoding multiple different zinc finger domain proteins interacts with the chromosomal kinase JIL-1. J. Biol. Chem. 278, 11696–11704 (2003).
Higuchi, N., Kohno, K. & Kadowaki, T. Specific retention of the protostome-specific PsGEF may parallel with the evolution of mushroom bodies in insect and lophotrochozoan brains. BMC Biol. 7, 21 (2009).
Leyssen, M. et al. Amyloid precursor protein promotes post-developmental neurite arborization in the Drosophila brain. EMBO J. 24, 2944–2955 (2005).
Lee, T., Marticke, S., Sung, C., Robinow, S. & Luo, L. Cell-autonomous requirement of the USP/EcR-B ecdysone receptor for mushroom body neuronal remodeling in Drosophila. Neuron 28, 807–818 (2000).
Becker, M. N., Brenner, R. & Atkinson, N. S. Tissue-specific expression of a Drosophila calcium-activated potassium channel. J. Neurosci. 15, 6250–6259 (1995).
Qian, M. et al. Receptor-like tyrosine phosphatase PTP10D is required for long-term memory in Drosophila. J. Neurosci. 27, 4396–4402 (2007).
Bates, K. E., Sung, C., Hilson, L. & Robinow, S. Unfulfilled interacting genes display branch-specific roles in the development of mushroom body axons in Drosophila melanogaster. G3 (Bethesda) 4, 693–706 (2014).
King, I. F., Eddison, M., Kaun, K. R. & Heberlein, U. EGFR and FGFR pathways have distinct roles in Drosophila mushroom body development and ethanol-induced behavior. PLoS ONE 9, e87714 (2014).
Dubnau, J. et al. The staufen/pumilio pathway is involved in Drosophila long-term memory. Curr. Biol. 13, 286–296 (2003).
Huang,, da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2008).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Gupta, S., Stamatoyannopolous, J., Bailey, T. & Stafford Noble, W. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Kurusu, M. et al. A conserved nuclear receptor, Tailless, is required for efficient proliferation and prolonged maintenance of mushroom body progenitors in the Drosophila brain. Dev. Biol. 326, 224–236 (2009).
Lin, S., Huang, Y. & Lee, T. Nuclear receptor unfulfilled regulates axonal guidance and cell identity of Drosophila mushroom body neurons. PLoS ONE 4, e8392 (2009).
Kunz, T., Kraft, K. F., Technau, G. M. & Urbach, R. Origin of Drosophila mushroom body neuroblasts and generation of divergent embryonic lineages. Development 139, 2510–2522 (2012).
Finley, K. D. et al. Dissatisfaction encodes a tailless-like nuclear receptor expressed in a subset of CNS neurons controlling Drosophila sexual behavior. Neuron 21, 1363–1374 (1998).
Sonoda, J. & Wharton, R. P. Recruitment of Nanos to hunchback mRNA by Pumilio. Genes Dev. 13, 2704–2712 (1999).
Kim, K. H. & Yoo, S. Sequence-specific interaction between ABD-B homeodomain and castor gene in Drosophila. BMB Rep. 47, 92–97 (2014).
Pfeiffer, B. D. et al. Tools for neuroanatomy and neurogenetics in Drosophila. Proc. Natl Acad. Sci. USA 105, 9715–9720 (2008).
Adachi, Y. et al. Conserved cis-regulatory modules mediate complex neural expression patterns of the eyeless gene in the Drosophila brain. Mech. Dev. 120, 1113–1126 (2003).
Swarup, S., Huang, W., Mackay, T. F. & Anholt, R. R. Analysis of natural variation reveals neurogenetic networks for Drosophila olfactory behavior. Proc. Natl Acad. Sci. USA 110, 1017–1022 (2013).
Jordan, K. W. et al. Genome-wide association for sensitivity to chronic oxidative stress in Drosophila melanogaster. PLoS ONE 7, e38722 (2012).
Beatty, J. & Laughlin, R. E. Genomic regulation of natural variation in cortical and noncortical brain volume. BMC Neurosci. 7, 16 (2006).
Williams, R. W., Strom, R. C. & Goldowitz, D. Natural variation in neuron number in mice is linked to a major quantitative trait locus on Chr 11. J. Neurosci. 18, 138–146 (1998).
Miller, G. F. & Penke, L. The evolution of human intelligence and the coefficient of additive genetic variance in human brain size. Intelligence 35, 97–114 (2007).
Cheverud, J. M. et al. Heritability of brain size and surface features in rhesus macaques (Macaca mulatta). J. Hered. 81, 51–57 (1990).
Hager, R., Lu, L., Rosen, G. D. & Williams, R. W. Genetic architecture supports mosaic brain evolution and independent brain-body size regulation. Nat. Commun. 3, 1079 (2012).
Billuart, P., Winter, C. G., Maresh, A., Zhao, X. & Luo, L. Regulating axon branch stability: the role of p190 RhoGAP in repressing a retraction signaling pathway. Cell 107, 195–207 (2001).
Pascual, A. & Preat, T. Localization of long-term memory within the Drosophila mushroom body. Science 294, 1115–1117 (2001).
Liu, L., Wolf, R., Ernst, R. & Heisenberg, M. Context generalization in Drosophila visual learning requires the mushroom bodies. Nature 400, 753–756 (1999).
Busto, G. U., Cervantes-Sandoval, I. & Davis, R. L. Olfactory learning in Drosophila. Physiology (Bethesda) 25, 338–346 (2010).
Durham, M. F., Magwire, M. M., Stone, E. A. & Leips, J. Genome-wide analysis in Drosophila reveals age-specific effects of SNPs on fitness traits. Nat. Commun. 5, 4338 (2014).
Zhu, N. et al. Cardiorespiratory fitness and brain volume and white matter integrity: The CARDIA Study. Neurology 84, 2347–2353 (2015).
Balling, A., Technau, G. M. & Heisenberg, M. Are the structural changes in adult Drosophila mushroom bodies memory traces? Studies on biochemical learning mutants. J. Neurogenet. 4, 65–73 (1987).
Heisenberg, M. Central brain function in insects: genetic studies on the mushroom bodies and central complex in Drosophila. Fort. Zool. 39, 30–39 (1994).
Heisenberg, M. Genetic approach to learning and memory (mnemogenetics) in Drosophila melanogaster. Forts. Zool. 37, 3–45 (1989).
Fields, R. D. Signaling by neuronal swelling. Sci. Signal. 4, tr1 (2011).
Ferreira, T., Ou, Y., Li, S., Giniger, E. & van Meyel, D. J. Dendrite architecture organized by transcriptional control of the F-actin nucleator Spire. Development 141, 650–660 (2014).
Flint, J. & Mackay, T. F. Genetic architecture of quantitative traits in mice, flies, and humans. Genome Res. 19, 723–733 (2009).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Jenett, A. et al. A GAL4-driver line resource for Drosophila neurobiology. Cell Rep. 2, 991–1001 (2012).
Stewart, A. J., Hannenhalli, S. & Plotkin, J. B. Why transcription factor binding sites are ten nucleotides long. Genetics 192, 973–985 (2012).
Acknowledgements
We thank the TRiP at Harvard Medical School (NIH/NIGMS R01-GM084947) for providing transgenic RNAi fly stocks and/or plasmid vectors used in this study. This work was supported by VIB funding and FWO grants (G065408.N10 and G078914N) to PC and National Institute of Health grants R01 GM45146 and R01 GM076083 to TFCM. L.V.B. is the recipient of a postdoctoral fellowship of the FWO.
Author information
Authors and Affiliations
Contributions
L.Z., L.V.B. designed the experiments, measured the data, analysed the results and wrote the manuscript. J.F.A., M.M.M. performed the GWAS. E.C., V.V. and J.C. assisted in performing the experiments. T.F.C.M. assisted in designing the project, interpreting the GWAS data and writing the manuscript. P.C. designed the project, assisted in data interpretation and writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data 1
Percentage of gross MB phenotypes observed in the 40 DGRP lines (XLSX 10 kb)
Supplementary Data 2
ANOVAs of mushroom body traits. (XLSX 9 kb)
Supplementary Data 3
Correlation between the analyzed MB traits in the 40 DGRP lines (Pearson's correlation coefficient) (XLSX 8 kb)
Supplementary Data 4
Correlation between the analyzed MB traits and different behaviors in the 40 DGRP lines (Pearson's correlation coefficient) (XLSX 10 kb)
Supplementary Data 5
GWA analyses of mushroom body morphological variation (XLSX 50 kb)
Supplementary Data 6
Genes with variants associated with multiple MB traits (XLSX 11 kb)
Supplementary Data 7
Enriched gene ontology categories among genes with variants associated with MB morphology traits (XLSX 12 kb)
Supplementary Data 8
Cluster analyses of gene ontology terms (XLSX 20 kb)
Supplementary Data 9
Selected transcription factors (XLSX 10 kb)
Supplementary Data 10
used FlyLight, TRIP and VDRC lines (XLSX 17 kb)
Supplementary Data 11
Percentage of gross MB phenotypes observed upon RNAi mediated knock down of the investigated genes (XLSX 10 kb)
Supplementary Data 12
Primers used for the verification of used FlyLight lines (XLSX 10 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zwarts, L., Vanden Broeck, L., Cappuyns, E. et al. The genetic basis of natural variation in mushroom body size in Drosophila melanogaster. Nat Commun 6, 10115 (2015). https://doi.org/10.1038/ncomms10115
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms10115
- Springer Nature Limited
This article is cited by
-
Genome-wide association in Drosophila identifies a role for Piezo and Proc-R in sleep latency
Scientific Reports (2024)
-
Modelling TDP-43 proteinopathy in Drosophila uncovers shared and neuron-specific targets across ALS and FTD relevant circuits
Acta Neuropathologica Communications (2023)
-
Lipophorin receptors regulate mushroom body development and complex behaviors in Drosophila
BMC Biology (2022)
-
Identification and Characterization of Gene SpDMRT99B and Its Sex-Biased Expression Profile in the Mud Crab, Scylla paramamosain
Journal of Ocean University of China (2021)
-
Genome-Wide Association Study of Circadian Behavior in Drosophila melanogaster
Behavior Genetics (2019)