Abstract
Background
The complexity of the upper gastrointestinal (UGI) multidisciplinary team (MDT) is continually growing, leading to rising clinician workload, time pressures, and demands. This increases heterogeneity or ‘noise’ within decision-making for patients with oesophageal cancer (OC) and may lead to inconsistent treatment decisions. In recent decades, the application of artificial intelligence (AI) and more specifically the branch of machine learning (ML) has led to a paradigm shift in the perceived utility of statistical modelling within healthcare. Within oesophageal cancer (OC) care, ML techniques have already been applied with early success to the analyses of histological samples and radiology imaging; however, it has not yet been applied to the MDT itself where such models are likely to benefit from incorporating information-rich, diverse datasets to increase predictive model accuracy.
Methods
This review discusses the current role the MDT plays in modern UGI cancer care as well as the utilisation of ML techniques to date using histological and radiological data to predict treatment response, prognostication, nodal disease evaluation, and even resectability within OC.
Results
The review finds that an emerging body of evidence is growing in support of ML tools within multiple domains relevant to decision-making within OC including automated histological analysis and radiomics. However, to date, no specific application has been directed to the MDT itself which routinely assimilates this information.
Conclusions
The authors feel the UGI MDT offers an information-rich, diverse array of data from which ML offers the potential to standardise, automate, and produce more consistent, data-driven MDT decisions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oesophageal cancer (OC) is the 14th most common cancer in the UK yet the 7th commonest cause of cancer death.1 Only 39% of patients enter a curative pathway and less than 15% are alive at 5 years.2,3 Adenocarcinoma (OAC) of the oesophagus, in particular, has seen a 400% increase over the last 2 decades in part owing to the increased prevalence of gastro-oesophageal reflux and Barrett’s oesophagus and is now more prevalent than squamous cell carcinoma (OSCC) in some world regions including North America, Northern Europe, and Oceania.4
Gold standard management of OC remains curative resection, stage-permitting. Patients presenting with nodal disease also require neoadjuvant therapy (NAT) either as chemotherapy (NACT) or chemoradiotherapy (NACRT).5 Both have been shown to offer a survival advantage over surgery alone although to date, debate remains over which regime offers the better outcome.5,6,7,8,9 The Neo-AEGIS trial was intended to answer this very question, and yet 3-year follow-up data remains equivocal (despite a noticeably higher incidence of tumour regression grade (TRG) 1–2 within the CRT arm).10 Longer follow-up data is still awaited. The survival benefit from NAT, however, may not be conferred universally. A multicentre study of 1293 patients by Noble et al. demonstrated that a meaningful local response to NACT was only seen in those with TRG 1–2 (14.8% of the cohort) deemed “responders”. Overall survival in this group was 7.68 years versus 2.22 years in those with TRG 3–5 (85.2%).11 A major challenge is therefore predicting responders before starting NAT, although some groups have found modest success modelling variables available prior to surgery.12,13 Reliable predictive tools might then permit early triaging of non-responders directly to surgery in a bid to reduce NAT-associated morbidity and mortality for potentially little gain as it is recognised that NAT can decondition patients prior to surgery, potentially even rendering them inoperable.14,15,16
OC patients are consequently reliant on high-quality decision-making in often complex clinical contexts, with significant implications for their outcomes and quality of life.17 Currently, their treatment decisions are made by a multidisciplinary team (MDT), which is shown to improve patient outcomes.18,19,20 However, these services face ever-growing caseloads and clinical complexity, potentially leading to inconsistent and sometimes suboptimal decisions.21 Individual experience, perception, and bias can also lead to discordance within that decision-making consistency, effectively a form of “noise” in the process.22
Data-driven clinical decision tools are increasingly commonplace within medicine. The National Emergency Laparotomy Audit (NELA), for instance, has achieved widespread use for more objective operative risk stratification and the need for higher levels of care following emergency laparotomy.23,24 The domain of machine learning (ML) and by extension deep learning (a subset of ML which uses unstructured data, processing this through multiple “hidden layers” between the input and output layer to form a “neural network” designed to approximate human neural networks)25 offers huge potential to take ML a step further by standardising, optimising, and streamlining decision-making for cancer patients. Thus far, ML has been applied to decision-making with cardiac patients,26 breast cancer therapies,27 lung cancer,28 pancreatic cancer,29 and dermatological cancers.30 To date, no such approach has been made to the OC MDT. The purposes of this review are twofold: to contextualise the MDT’s role within OC and to discuss the applications of ML techniques within OC to date. This includes predicting treatment response using both histopathological and radiological data, as well as the emerging potential for radiomics for prognostication, nodal disease evaluation, and even resectability.
Methods
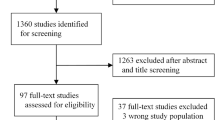
Studies were selected on their use of, or discussion of artificial intelligence–based techniques on the UGI MDT as a whole or data types used by the MDT to determine treatment decisions for oesophageal cancer patients.
Studies will be further discussed by the modality of data they apply their machine learning approaches to. Within the MDT framework, the two main data sources outside of standard clinical patient information are histopathological and imaging based. This review will therefore discuss each of these separately.
Studies were obtained by a systematic search of PubMed using a combination of key terms including “Machine Learning”, “Artificial Intelligence”, “Oesophageal Cancer”, “Oesophagogastric Cancer”, “Esophageal”, “Esophagogastric”, “Upper Gastrointestinal Cancer”, “Upper Gastrointestinal Multidisciplinary team”, “Multidisciplinary team”, “Radiomics”, and “Predicting response”. Additional relevant studies were also identified through bibliographic examination of articles retrieved through the initial literature searches.
The Multidisciplinary Team (MDT)
The clinical management of all cancer patients within the UK was centralised through MDTs following the Calman-Hine report in 1995.31 This brought together experts from all aspects of a patient’s care to focus on rapid, nuanced, complex, and above all-shared decision-making from the outset. MDTs comprise a variety of healthcare professionals: surgeons, physicians, oncologists, radiologists, histopathologists, specialist nurses, physiotherapists, occupational therapists, palliative care teams, and administrative staff. Centralisation also ensured adequate caseload to maintain clinical and operative skills. MDTs assess cancer site, stage, resectability, fitness for surgery, and necessary oncological adjuncts to formulate a treatment plan within the context of the patient’s wishes.
Strengths of the MDT
Numerous studies have shown a benefit to managing oesophageal cancer via an MDT framework (Fig. 1) over surgeons managing such cases independently.18,19,20,32 They have been shown to reduce the incidence of open-and-close laparotomies or thoracotomies (from 21 and 5, respectively, to 13% and 0%, p = 0.02). Operative mortality is lower (5.7% vs 26%, p = 0.004), and 5-year survival is significantly higher (52% vs 10%, p = 0.0001). On multi-variate analysis, MDT management, lymph node metastases, and American Society of Anaesthesiologist (ASA) grade were all found to be independently associated with survival.18 Freeman et al. reported that a formal thoracic MDT for OC improved the rate of complete staging from 67 to 97% (p < 0.0001) and increased the percentage assessment by MDT from 72 to 98% (p < 0.0001) and adherence to national guidelines for management from 83 to 98% (p < 0.0001).19 Van Hagen and colleagues found that over one-third of management plans pre-conceived by individual clinicians as the “best course of action” for potentially curative upper gastrointestinal (UGI) cancer cases were still changed after MDT discussion20.
These benefits are not restricted to curative cases. A Dutch study of 948 palliative oesophagogastric (OG) patients found a significantly shorter time from diagnosis to commencement of palliative therapy in the MDT group (20 days vs 30 days, p < 0.001), a higher incidence of palliative external beam radiotherapy (EBRT) (38% vs 21%, OR 2.7), higher incidence of systemic therapy (30% vs 23%, OR 1.6,), fewer patients treated with palliative stents (4% vs 12%, OR 0.3), and greater duration of survival (169 days vs 107 days, HR 1.3).32 The authors attributed at least part of this improved survival to the greater usage of tumour-specific palliative therapies such as EBRT and systemic therapy in the MDT group.
Vulnerabilities of the MDT
Despite the multitude of strengths of the MDT system, it is also vulnerable to clinical, inter-personal, and logistic challenges. Rising caseloads, reduced dedicated MDT time, missing data, patient complexity, and inter-member disagreement all lead to inconsistent and suboptimal decision-making with potentially life-limiting consequences for a patient’s health and quality of life.33 The dedicated preparation time required and associated financial cost are also considerable. Each hour of an MDT has been estimated to take 2 h for a radiologist and 2.4 h for a histopathologist to prepare for.34 A systematic review in 2011 exploring clinical, social, and technological factors influencing MDT decision-making found that definitive plans were only reached at first discussion in 47.6–73% of cases owing to time pressures or inadequacy of available information at the time of discussion (e.g., imaging, staging, pathology review, or patient comorbidities).21 A failure to implement MDT decisions was seen in 1–16% of cases owing to differing patient wishes or inappropriate management plans when factoring in patient comorbidities. General surgical, urological, and soft tissue cancer MDTs were found to have clinician-made decisions based almost entirely on clinical information. The review noted that physicians drove the decision-making, often ignoring nurse-led input usually at the detriment of the overall efficacy of the MDT.
Patient-centred decision-making varies within MDTs. Another study by Lamb et al. determined that patient wishes were infrequently considered at MDT unless nurses present could, and felt empowered to, speak up.35 Furthermore, essential social data such as a patient’s social position, attitude, values, and preferences often be missing, incomplete, or selectively presented in order to influence the discussion in a particular direction.36
Leadership and personal biases are salient factors. A study of breast cancer MDTs found that while a lack of clarity and conflict over leadership were negative predictors for effective internal communication, team effectiveness, and resource efficiency, a single strong leader also harmed innovation.37 Their results further highlighted that perceptions of team effectiveness could vary significantly by role, noting that breast surgeons and breast care nurses consistently rated their MDT’s performance higher than their radiology and histopathology counterparts.
Such vulnerabilities can have clinical impact on OC patients. A small observational multicentre Danish study investigated inter-observer variability between MDT decisions at four major UGI cancer units in Denmark.22 The study presented 20 OSCC cases as new referrals to each of the four centres to determine resectability, curability, and treatment strategy. The authors reviewed the frequency by which disagreement between MDTs resulted in a different treatment recommendation and whether this had a clinical impact. Moderate concordance was seen on classifying T-stage, M-stage, resectability, and curability, while N-stage and operability only reached “fair” concordance. The authors traced much of the disagreement back to classifying “Mx” and consideration of “further investigations”. The biggest impact of their findings was however that MDT disagreement led to a clinical impact in 60% of cases. The authors reported that while operability was crucial to determining an accurate treatment strategy, it was most vulnerable to inter-observer differences. Yet given the clinical information available at MDT, it remained difficult if not almost impossible to determine accurately at the time of the meeting. The authors reinforced the importance of being able to establish operability either pre-MDT or with additional data variables available at the time of discussion.
Similarly, comorbidity is inadequately presented or integrated into cancer MDTs, despite having a substantial impact on the quality of its judgements. A 2015 systematic review found that comorbidities were; not well considered (meaning MDTs were less likely to reach a treatment decision); were often the reason given for deviating from treatment guidelines; and where a treatment recommendation was given, was usually the reason it was not implemented.33
Decision-Making Within OC
Table 1 outlines the 2018 National Institute for Clinical Excellence (NICE) guidelines for the management of OC.38 Notably while some authors categorise T2N0 disease as early and amenable to endomucosal resection (EMR), NICE supports the use of NAT in this cohort, likely to minimise local recurrence risk from micro-metastases.39,40 It can be readily appreciated that histology, TNM staging, and an assessment of patient fitness (commonly quantified by the WHO Performance Status classification) account for the bulk of decision critical parameters. While the concept of comorbidity is acknowledged, especially when determining suitability for palliative chemotherapy, such guidelines remain simplistic, rarely factoring in dimensions such as high-risk comorbidities, social variables, or even ease of patient access to CRT centres.
A Role for Machine Learning?
Machine learning (ML) has gained popularity within healthcare environments for its potential to assist clinical decision-making by detecting complex patterns within large datasets. Great promise has been shown even in OC, in predicting outcomes following oesophagectomy.41 However, while post-operative models have shown good discrimination and calibration, pre-operative models are more challenging.12 Despite this, the pre-treatment MDT discussion remains a key mile marker in the patient’s care pathway, and optimising the decision-making at this stage is vital. MDTs typically assimilate information from clinical, pathological, and radiological sources, each of which offers a potential focus for ML applications, yet surprisingly, this has not been exploited in UGI MDTs to date.
Machine learning is traditionally divided into supervised and unsupervised learning. Supervised learning requires the “labelling” of data (the ground truth is given to the machine). The machine is then able to compare the input and outcome data to determine the best fitting model which explains any underlying structure of the data. Supervised learning is thus well suited to smaller datasets where the ground truth is known—a prime example being the outcomes of historic MDTs where treatment decisions of patients are already known. By comparison, unsupervised learning algorithms identify patterns within datasets to extract features that may speak to their structure. Such techniques are useful when the ground truth is unknown, necessitating large volumes of data—a challenge frequently encountered in cancer datasets. Models are trained using data partitioned from the main dataset, by which the machine searches for patterns between the selected variables and the designated outcome. Ideal models learn from training data to make accurate predictions when fed new unseen data (testing datasets), minimising “under-” or “over-fitting”. Under-fitted models are too simplistic or inflexible to capture the underlying relationships leading to high error rates in both training and testing (bias). Over-fitting occurs when the model features are too numerous or complex resulting in high variance. These models perform well within training but struggle on test/validation sets.42 This may be mitigated by increasing the size of the training set available and the diversity of the observations themselves, making it more representative of the theoretical population distribution. In real-world settings however, this is often difficult with health data especially for rarer clinical scenarios under study. Table 2 summarises some common ML-based techniques.
ML Applications Within OC to Date
Histopathological Analysis
The application of ML to histopathology in order to augment decision-making in clinical care is gaining popularity.43,44,45 RNA and whole genome sequencing (WGS) offer detailed and individualised data for analysis at the cost of expensive tissue analytical processes.41 Computer vision–based ML promises comparatively low-cost, automated large-scale analysis in OC, although to date very few studies have applied such techniques to OC (Table 3).41,46 Pilot work using convolutional neural networks (CNN) to process unlabelled high-resolution digital OAC histology slides achieved good internal validation in predicting response to NAT (C-index 0.836).41 While these results are promising, validation over larger datasets and external data sources remains necessary before use in clinical practice, especially as the use of unsupervised learning creates a “black box” solution impeding transparency, “explainability”, and ultimately trust in the solution. An additional confounder in the Rahman et al. study was the use of both NACRT and NACT within the patient cohort. The training of the CNN in this instance utilised ImageNet (non-specific images from a vast online database of everyday images) in the form of transfer learning. This circumvented the need for the sheer volume of histology-specific training images otherwise needed to produce a sufficiently accurate model. Pre-trained networks have performed competitively against models trained from scratch.47 However, with academic collaborative projects such as the Northern Pathology Imaging Co-operative looking to accumulate large-scale digital pathology repositories, this challenge may become more achievable in the future, especially as transfer learning is unlikely to be sufficiently robust for routine clinical use.
With only a minority of OC patients benefitting from NAT, it is appealing for MDTs to be able to identify them as early as possible. Accurate prediction of tumour response from initial biopsies usually available at the beginning of a referral pathway would allow patients to be filtered towards the most beneficial therapy in the timeliest fashion.
Imaging-Based Approaches—Radiomics
Over the last two decades, advances in image processing and analysis have allowed the field of radiomics to flourish developing a substantial evidence base across numerous solid organ cancer types.48 Radiomics refers to the extraction of quantitative, clinically significant, high-dimensional imaging biomarkers from standard-of-care medical imaging to predict a range of clinical outcomes.42 Standard radiological assessments within MDTs for OC are traditionally largely qualitative, with some quantification of tumour size, number, and position of suspected lymphadenopathy and the presence of distant metastases. A human eye–based assessment however may struggle to pick out additional hidden data on a pixel/voxel level within the image stacks and inherently involves a degree of both selection bias as well as inter- and intra-observer variability.49 Radiomics seeks to mine this data for more tailored decision-making. Coupling this to the MDT infrastructure would benefit OC patients by achieving highly detailed assessment of their disease burden, resectability, and probable interval response to NAT at a very early stage.
Radiomic Workflow
The radiomic workflow (Fig. 2) can be summarised as image acquisition, image pre-processing, segmentation, feature extraction, data preparation, feature reduction, and model development.42,50 Image acquisition relates to the curating of imaging stacks containing regions of interest (ROI) under investigation. Features extracted from ROIs may mirror the tumour phenotype and its molecular fingerprint.49 Image pre-processing includes segmentation of ROIs which may be manual (considered gold standard but resource intensive), automatic, or hybridised. Automated segmentations while potentially error-prone offer workflow automation with reasonable accuracy.51 The next step is feature extraction which is the functional core of radiomics. Visual features embedded within images are extracted and converted into quantifiable vectors.42,49 Vectors may differ in scales; thus, data preparation includes feature scaling, data continuation, discretisation, and under- or over-sampling for class imbalances.52 The resultant features may be hundreds in number and counter-productive to a well-performing model.53 Dimensionality reduction and feature selection can minimise those redundant, non-relevant features which may slow a model for little gain.54,55,56 The final feature pool which forms the radiomic model is then used to classify groups of patients into one of several outcome classes, whether this is based on a perceived risk or intervention outcome. Finally, validation of the generated model must then be done internally and externally as it speaks to the generalisability of the final model.57
Radiomics in OC
An evolving body of evidence is now emerging for OC in predicting treatment response, prognosis, nodal status, and even resectability.16 Improving the diagnostic accuracy of each of these aspects in turn using radiomics can drive forward a large portion of the MDT’s weekly workflow. Table 4 summarises studies which have applied radiomics to the OC domain.
Treatment Response Evaluation
Most studies predicting treatment response have focussed on NACRT rather than NACT, using OSCC primarily or mixed histology datasets.58,59,60,61 As many of these studies originate from China, where 90% of OC is the OSCC subtype, this is unsurprising. Nonetheless, it has long been appreciated that tumour heterogeneity on imaging is associated with aggressive tumour biology and impaired treatment response in OC leading to many ML techniques being applied to this very issue.62 As imaging is often one of the earliest potential sources of information on tumour biology for OC patients, accurate characterisation here can tailor the oncological plan even before histology has been returned.
Flurodeoxyglucose (18F)-positron emission tomography (FDG-PET) is used to assess for metastatic disease by uptake of FDG in metabolically active cells. Metabolic tumour volume (MTV) and standardised uptake value (SUV) on FDG-PET may variably predict response to NACRT in OC across serial imaging time points as well holding prognostic significance for survival.16,63,64 One PET study drew inspiration from DNA microarray analysis combining an extracted radiomic signature with a LASSO-logistic regression model to predict treatment response (AUC 0.835). While the authors contended with a class imbalance favouring responders and a radiomic signature derived from only 20 patients, the approach was nevertheless an intriguing one.65 A drawback to FDG-PET is its expense, time consumption, and lack of the complete molecular characterisation that one wishes to exploit when mining spatial heterogeneity in tissue architecture and metabolic activity.62 Contrast-enhanced CT is comparatively ubiquitous in day-to-day clinical practice for assessing treatment response; it is quick and easily accessible. In smaller case series, it has even successfully predicted response to NACRT using as few as five shape and histogram-based metrics (AUC 0.686–0.727).59
Studies combining multimodal data frequently show superior performance compared to single data streams alone. Zhang et al. predicted pathological tumour response to NACRT in OC patients applying both logistic regression (LR) and support vector machine (SVM) models finding that a combination of conventional PET/CT response measures, clinical data (TNM, histology, patient demographics), and spatial–temporal PET/CT features offered superior predictive performance over individual feature sets (AUC of 1.0 for SVM vs 0.9 for LR).66 However, the study did not factor in nodal disease and was small (N = 20), thus risking over-fitting in the absence of external validation. Another study combining clinical information, geometry, PET textural features, and CT textural features used a LASSO-regularised LR model to produce an AUC of 0.78 versus 0.58 for SUVmax alone.62
Prognostication
A number of studies have attempted to prognosticate in OC. Qiu et al., for instance, reported disease recurrence in one-third of patients who experienced a pathological complete response following NACRT and surgery for OSCC.67 Their CT-based nomogram combined clinical risk factors and a radiomic signature of eight features. This proved superior (C-index of 0.746) versus radiomic (0.685) and clinical (0.614) features alone (p < 0.001 in all cases). The model could effectively stratify patients into high and low risk categories potentially offering tailored adjuvant therapy post-resection.
One Dutch study predicted 3-year survival after NACRT using a random forest model comparing clinical and radiomic feature sets on pre-treatment CT. This study did include both OAC and OSCC, albeit heavily weighted towards the former.60 They reported an AUC of 0.61 on external validation for their radiomic model versus 0.62 for their clinical dataset. While the authors did show clear survival differences between TRG 1–2 and TRG 3–5 patients within the study cohort, this did not translate to a statistically significant difference in survival within validation sets when risk was stratified by the model again reflecting the Neo-AEGIS trial.5
Deep convolutional neural networks (CNN) have also proved capable of predicting 1-year survival in OSCC when trained on PET images. A Taiwanese study pre-trained a ResNet 3D CNN using a mixed set of 1,107 OSSC and lung cancer PET scans.68 Their best model attained an AUC of 0.738, outperforming clinical data alone. The authors found that CNN predictions themselves were significant on multivariable analysis for survival indicating that meaningful prognostic hidden data could be extricated. The authors did recognise that the extraction and selection of features was not transparent, i.e. a “black box” problem.
While accurate knowledge of operability and treatment response is vital for counselling patients of MDT treatment recommendations, precise prognostication allows them to contextualise the cost–benefit balance. The studies described above therefore highlight the significant role ML can play here.
Nodal Status
The prediction of lymph node (LN) disease conveys implications for prognosis and MDT treatment decisions. Tan and colleagues achieved a test set validation AUC of 0.773 using LASSO-LR when predicting LN metastases in resectable OSCC cases, outperforming size criteria alone on CT imaging.61 Another CT-based study reported near-identical performance in testing using an elastic net approach across what was implied to be a mixed histological cohort.69
Other Outcomes
Less conventional radiomic–based problems have also been explored. Resectability, for example, was predicted in one study of 591 OSCC patients. A LASSO-enhanced dimensionality reduction technique across multiple ML algorithms showed that multivariable logistic regression (MLR) offered the best performance (AUC 0.87, accuracy 0.86).58 Another study in radio-genomics used CT imaging to help predict microRNA-1246 expression, a biomarker linked with prognostic significance in OSCC.70 Correlation analysis extracted image features correlating with miR-1246 levels in 92 patients. Linear regression then separated patients into low and high expression correlating with survival. Unfortunately, while miR-1246 levels were significantly raised in stage 2 disease, no difference was seen between healthy controls and stage 1 disease, thereby limiting miR-1246’s potential for screening.
Challenges and Future Directions for ML and the MDT
One of the main challenges facing ML tools designed for the MDT is inevitably the degree of noise within the datasets. This may be attributable to several factors such as variation in attendance of specific MDT members, the allocated time they possess to be present and discuss each case, clinical equipoise over treatment options, clinician preferences, and even social factors such as patient geography and their relationships to high-resource units.35 Incorporating some or all such factors into future model training may adjust for this noise. Trustworthiness and transparency remain another key issues for model deployment within healthcare settings. Patients, clinicians, and health regulators alike will likely require a degree of explainability for ML solutions. A route through this would be to focus on more simplistic and/or explainable models such as logistic regression and decision tree algorithms (a process which falls under explainable AI or “XAI” 71). However while XAI intuitively fits the perception of providing understanding of a system’s decisions, inherently explainable algorithms and post-hoc explainability tools may conversely reflect a misleading sense of true trustworthiness, with patient safety potentially better achieved through robust validation techniques instead.72 Once model performance is confirmed at a single unit, the tool may then be extended to other MDTs. This may be through tailoring a model to each unit individually or applying a single model to multiple units. The former approach is labour intensive yet minimises under-fitting or poor generalisability as we are no longer modelling noise and idiosyncrasies particular to one MDT and applying these “rules” to another. Alternatively, a one-size-fits-all model may be designed for generalisation across multiple provided the practices of each such unit follow a consistent pattern. To achieve this, the training data requires amalgamation and homogenisation from multiple sources which pose challenges such as data sharing agreements between centres, standardised patient data acquisition, and navigating the variation in imaging protocols associated with each individual hospital.50 Daramola et al. propose a multimodal AI framework for amalgamation, processing, and model development using similar data types in managing infectious diseases within sub-Saharan Africa.73 Through these approaches, ML allows OC MDTs to automate aspects of their workflow, potentially extract clinically meaningful information from imaging data, and streamline decision-making which has been learned from its historic decision-making framework. As UGI MDTs also manage gastric cancers, the concept is also transferrable to their gastric cancer patients and potentially other solid body cancers.
Conclusion
The OC MDT handles complex treatment decisions with potentially life-altering implications for its patients, increasingly under pressures of modern practice and caseloads. ML has shown great promise as an assistive tool in many clinical domains. While ML approaches have been applied to several data types relevant to the OC MDT, the MDT itself is as yet an unexplored arena. Future work should now look to integrate these techniques to streamline and assist the MDT’s own decision-making. This in turn may offer the capacity to offer data-driven solutions, reduce costs and help prioritise their caseload, and thereby positively impact patient cancer care.
Data Availability
Data availability was not applicable to this article.
References
Heartburn Cancer UK. Oesophageal cancer [Internet]. Available from: https://www.heartburncanceruk.org/oesophageal-cancer
Maynard N, Crosby T, Trudgill N, Varangunam M, Park MH, Sinha S. An audit of the care received by people with Oesophago-gastric Cancer in England and Wales [Internet]. Third Annual Report. 2010.
Chadwick G, Groene O, Cromwell D, Hardwick R, Riley S, Crosby T, et al. National Gastric Cancer Audit. Natl Oesophegao-Gastric Cancer Audit. 2013;1–69.
Arnold M, Ferlay J, Van Berge Henegouwen MI, Soerjomataram I. Global burden of oesophageal and gastric cancer by histology and subsite in 2018. Gut. 2020;69(9):1564–71.
Reynolds J V, Preston SR, O’Neill B, Lowery MA, Baeksgaard L, Crosby T, et al. Neo-AEGIS (Neoadjuvant trial in Adenocarcinoma of the Esophagus and Esophago-Gastric Junction International Study): Preliminary results of phase III RCT of CROSS versus perioperative chemotherapy (Modified MAGIC or FLOT protocol). (NCT01726452). J Clin Oncol [Internet]. 2021 May 20;39(15_suppl):4004.
Cunningham D, Allum WH, Stenning SP, Thompson JN, Van de Velde CJH, Nicolson M, et al. Perioperative chemotherapy versus surgery alone for resectable gastroesophageal cancer. N Engl J Med [Internet]. 2006 Jul 6;355(1):11–20.
Allum WH, Stenning SP, Bancewicz J, Clark PI, Langley RE. Long-term results of a randomized trial of surgery with or without preoperative chemotherapy in esophageal cancer. J Clin Oncol. 2009;27(30):5062–7.
Shapiro J, van Lanschot JJB, Hulshof MCCM, van Hagen P, van Berge Henegouwen MI, Wijnhoven BPL, et al. Neoadjuvant chemoradiotherapy plus surgery versus surgery alone for oesophageal or junctional cancer (CROSS): Long-term results of a randomised controlled trial. Lancet Oncol [Internet]. 2015;16(9):1090–8. A
Al-Batran SE, Homann N, Pauligk C, Goetze TO, Meiler J, Kasper S, et al. Perioperative chemotherapy with fluorouracil plus leucovorin, oxaliplatin, and docetaxel versus fluorouracil or capecitabine plus cisplatin and epirubicin for locally advanced, resectable gastric or gastro-oesophageal junction adenocarcinoma (FLOT4): a ra. Lancet (London, England) [Internet]. 2019;393(10184):1948–57.
Reynolds J V., Preston SR, O’Neill B, Lowery MA, Baeksgaard L, Crosby T, et al. Neo-AEGIS (Neoadjuvant trial in Adenocarcinoma of the Esophagus and Esophago-Gastric Junction International Study): Preliminary results of phase III RCT of CROSS versus perioperative chemotherapy (Modified MAGIC or FLOT protocol). (NCT01726452). J Clin Oncol [Internet]. 2021 May 20;39(15_suppl):4004–4004.
Noble F, Lloyd MA, Turkington R, Griffiths E, O’Donovan M, O’Neill JR, et al. Multicentre cohort study to define and validate pathological assessment of response to neoadjuvant therapy in oesophagogastric adenocarcinoma. Br J Surg. 2017;104(13):1816–28.
Goense L, van Rossum PSN, Xi M, Maru DM, Carter BW, Meijer GJ, et al. Preoperative Nomogram to Risk Stratify Patients for the Benefit of Trimodality Therapy in Esophageal Adenocarcinoma. Ann Surg Oncol [Internet]. 2018;25(6):1598–607.
Bott RK, George G, McEwen R, Zylstra J, Knight WRC, Baker CR, et al. Predicting response to neoadjuvant chemotherapy in patients with oesophageal adenocarcinoma. Acta Oncol (Madr) [Internet]. 2021;60(12):1629–36.
Jiang W, de Jong JM, van Hillegersberg R, Read M. Predicting Response to Neoadjuvant Therapy in Oesophageal Adenocarcinoma. Cancers (Basel). 2022;14(4):1–37.
Depypere L, Thomas M, Moons J, Coosemans W, Lerut T, Prenen H, et al. Analysis of patients scheduled for neoadjuvant therapy followed by surgery for esophageal cancer, who never made it to esophagectomy. World J Surg Oncol. 2019;17(1):1–9.
Findlay JM, Bradley KM, Wang LM, Franklin JM, Teoh EJ, Gleeson F V., et al. Predicting pathologic response of esophageal cancer to neoadjuvant chemotherapy: The implications of metabolic nodal response for personalized therapy. J Nucl Med. 2017;58(2):266–75.
Al-Batran SE, Ajani JA. Impact of chemotherapy on quality of life in patients with metastatic esophagogastric cancer. Cancer [Internet]. 2010 Jun 1;116(11):2511–8.
Stephens MR, Lewis WG, Brewster AE, Lord I, Blackshaw GRJC, Hodzovic I, et al. Multidisciplinary team management is associated with improved outcomes after surgery for esophageal cancer. Dis Esophagus. 2006;19(3):164–71.
Freeman RK, Van Woerkom JM, Vyverberg A, Ascioti AJ. The effect of a multidisciplinary thoracic malignancy conference on the treatment of patients with esophageal cancer. Ann Thorac Surg [Internet]. 2011;92(4):1239–43.
Van Hagen P, Spaander MCW, Van Der Gaast A, Van Rij CM, Tilanus HW, Van Lanschot JJB, et al. Impact of a multidisciplinary tumour board meeting for upper-GI malignancies on clinical decision making: A prospective cohort study. Int J Clin Oncol. 2013;18(2):214–9.
Lamb BW, Brown KF, Nagpal K, Vincent C, Green JSA, Sevdalis N. Quality of care management decisions by multidisciplinary cancer teams: a systematic review. Ann Surg Oncol [Internet]. 2011 Aug;18(8):2116–25.
Achiam MP, Nordsmark M, Ladekarl M, Olsen A, Loft A, Garbyal RS, et al. Clinically decisive (dis)agreement in multidisciplinary team assessment of esophageal squamous cell carcinoma; a prospective, national, multicenter study. Acta Oncol (Madr) [Internet]. 2021;60(9):1091–9.
Mak M, Hakeem AR, Chitre V. Pre-NELA vs NELA - has anything changed, or is it just an audit exercise? Ann R Coll Surg Engl. 2016;98(8):554–9.
Hunter Emergency Laparotomy Collaborator Group, Hunter Emergency Laparotomy Collaborator Group. High-Risk Emergency Laparotomy in Australia: Comparing NELA, P-POSSUM, and ACS-NSQIP Calculators. J Surg Res [Internet]. 2020;246:300–4.
IBM. Deep Learning [Internet]. IBM Cloud Education. 2020. Available from: https://www.ibm.com/cloud/learn/deep-learning#toc-deep-learn-md_Q_Of3
Diller GP, Kempny A, Babu-Narayan S V., Henrichs M, Brida M, Uebing A, et al. Machine learning algorithms estimating prognosis and guiding therapy in adult congenital heart disease: Data from a single tertiary centre including 10 019 patients. Eur Heart J. 2019;40(13):1069–77.
Lin FPY, Pokorny A, Teng C, Dear R, Epstein RJ. Computational prediction of multidisciplinary team decision-making for adjuvant breast cancer drug therapies: A machine learning approach. BMC Cancer [Internet]. 2016;16(1):1–10.
Wang Z, Sun J, Sun Y, Gu Y, Xu Y, Zhao B, et al. Machine Learning Algorithm Guiding Local Treatment Decisions to Reduce Pain for Lung Cancer Patients with Bone Metastases, a Prospective Cohort Study. Pain Ther [Internet]. 2021;10(1):619–33.
Bradley A, Van Der Meer R, McKay C. Personalized Pancreatic Cancer Management: A Systematic Review of How Machine Learning Is Supporting Decision-making. Pancreas. 2019;48(5):598–604.
Andrew TW, Hamnett N, Roy I, Garioch J, Nobes J, Moncrieff MD. Machine-learning algorithm to predict multidisciplinary team treatment recommendations in the management of basal cell carcinoma. Br J Cancer [Internet]. 2022;126(4):562–568.
Calman K, Hine D. A policy framework for commissioning cancer services. BMJ. 1995;310:1425.
Vermeulen BD, Bruggeman L, Bac DJ, Schrauwen RWM, Epping LSM, Scheffer RCH, et al. Impact of multidisciplinary tumor board discussion on palliation of patients with esophageal or gastro-esophageal junction cancer: a population-based study. Acta Oncol (Madr) [Internet]. 2020;59(4):410–6.
Stairmand J, Signal L, Sarfati D, Jackson C, Batten L, Holdaway M, et al. Consideration of comorbidity in treatment decision making in multidisciplinary cancer team meetings: A systematic review. Ann Oncol [Internet]. 2015;26(7):1325–32.
Taylor C, Munro AJ, Glynne-Jones R, Griffith C, Trevatt P, Richards M, et al. Multidisciplinary team working in cancer: what is the evidence? BMJ [Internet]. 2010 Mar 23;340(mar23 2):c951–c951.
Lamb BW, Sevdalis N, Arora S, Pinto A, Vincent C, Green JSA. Teamwork and team decision-making at multidisciplinary cancer conferences: Barriers, facilitators, and opportunities for improvement. World J Surg. 2011;35(9):1970–6.
Hamilton DW, Heaven B, Thomson RG, Wilson JA, Exley C. Multidisciplinary team decision-making in cancer and the absent patient : a qualitative study. BMJ Open. 2016;6(7):e012559
Haward R, Amir Z, Borrill C, Dawson J, Scully J, West M, et al. Breast cancer teams: the impact of constitution, new cancer workload, and methods of operation on their effectiveness. Br J Cancer [Internet]. 2003 Jul 7;89(1):15–22.
The National Institute for Health and Care Excellence (NICE). Oesophago-gastric cancer: Assessment and management in adults (NG83). NICE Guidel [Internet]. 2018;4(January 2018):970–6.
Smyth EC, Lagergren J, Fitzgerald RC, Lordick F, Shah MA, Lagergren P, et al. Oesophageal cancer. Nat Rev Dis Prim [Internet]. 2017 Jul 27;3:17048.
Lang CCJ, Lloyd M, Alyacoubi S, Rahman S, Pickering O, Underwood T, et al. The Use of miRNAs in Predicting Response to Neoadjuvant Therapy in Oesophageal Cancer. Cancers. 2022;14(5):1171
Rahman S, Early J, De Vries M, Lloyd M, Grace B, Ramchurn G, et al. Predicting response to neoadjuvant therapy using image capture from diagnostic biopsies of oesophageal adenocarcinoma. Eur J Surg Oncol [Internet]. 2021 Jan;47(1):e4.
Koçak B, Durmaz EŞ, Ateş E, Kılıçkesmez Ö. Radiomics with artificial intelligence: A practical guide for beginners. Diagnostic Interv Radiol. 2019;25(6):485–95.
Dimitriou N, Arandjelović O, Caie PD. Deep Learning for Whole Slide Image Analysis: An Overview. Front Med. 2019;6(November):1–7.
Gurcan MN, Boucheron LE, Can A, Madabhushi A, Rajpoot NM, Yener B. Histopathological Image Analysis: A Review. IEEE Rev Biomed Eng. 2009;2:147–71.
Komura D, Ishikawa S. Machine Learning Methods for Histopathological Image Analysis. Comput Struct Biotechnol J [Internet]. 2018;16:34–42.
Tomita N, Abdollahi B, Wei J, Ren B, Suriawinata A, Hassanpour S. Attention-Based Deep Neural Networks for Detection of Cancerous and Precancerous Esophagus Tissue on Histopathological Slides. JAMA Netw Open. 2019;2(11):1–13.
Kieffer B, Babaie M, Kalra S, Tizhoosh HR. Convolutional Neural Networks for Histopathology Image Classification : Training vs . Using Pre-Trained Networks. In: 2017 Seventh International Conference on Image Processing Theory, Tools and Applications (IPTA). Montreal, QC, Canada, 2017. pp. 1–6. https://doi.org/10.1109/IPTA.2017.8310149.
Bogowicz M, Vuong D, Huellner MW, Pavic M, Andratschke N, Gabrys HS, et al. CT radiomics and PET radiomics: Ready for clinical implementation? Q J Nucl Med Mol Imaging. 2019;63(4):355–70.
Varghese BA, Cen SY, Hwang DH, Duddalwar VA. Radiologists Need to Know. Ajr. 2019;(212):1–9.
Xie C yi, Pang C lap, Chan B, Wong EY yuen, Dou Q, Vardhanabhuti V. Machine Learning and Radiomics Applications in Esophageal Cancers Using Non-Invasive Imaging Methods—A Critical Review of Literature. Cancers (Basel) [Internet]. 2021 May 19;13(10):2469.
Jin J, Zhu H, Zhang J, Ai Y, Zhang J, Teng Y, et al. Multiple U-Net-Based Automatic Segmentations and Radiomics Feature Stability on Ultrasound Images for Patients With Ovarian Cancer. Front Oncol. 2021;10(February):1–8.
van Timmeren JE, Cester D, Tanadini-Lang S, Alkadhi H, Baessler B. Radiomics in medical imaging—“how-to” guide and critical reflection. Insights Imaging. 2020;11:91
Brunzell H, Eriksson J. Feature reduction for classification of multidimensional data. Pattern Recognit. 2000;33(10):1741–8.
Ringnér M. What is principal component analysis? Nat Biotechnol. 2008;26(3):303–4.
Balakrishnama S, Ganapathiraju A. Linear Discriminant Analysis - A Brief Tutorial [Internet]. 1995. Available from: https://datajobstest.com/data-science-repo/LDA-Primer-Balakrishnama-and-Ganapathiraju.pdf
Saeys Y, Inza I, Larrañaga P. A review of feature selection techniques in bioinformatics. Bioinformatics. 2007;23(19):2507–17.
Rhys HI. Machine Learning with R, the tidyverse, and mlr. 1st Edition. New York: Manning; 2020.
Ou J, Li R, Zeng R, Wu CQ, Chen Y, Chen TW, et al. CT radiomic features for predicting resectability of oesophageal squamous cell carcinoma as given by feature analysis: A case control study. Cancer Imaging. 2019;19(1):1–10.
Hou Z, Ren W, Li S, Liu J, Sun Y, Yan J, et al. Radiomic analysis in contrast-enhanced CT: Predict treatment response to chemoradiotherapy in esophageal carcinoma. Oncotarget. 2017;8(61):104444–54.
Larue RTHM, Klaassen R, Jochems A, Leijenaar RTH, Hulshof MCCM, Henegouwen MIVB, et al. Pre-treatment CT radiomics to predict 3-year overall survival following chemoradiotherapy of esophageal cancer. Acta Oncol (Madr) [Internet]. 2018;57(11):1475–81.
Tan X, Ma Z, Yan L, Ye W, Liu Z, Liang C. Radiomics nomogram outperforms size criteria in discriminating lymph node metastasis in resectable esophageal squamous cell carcinoma. Eur Radiol [Internet]. 2019 Jan;29(1):392–400.
Beukinga RJ, Hulshoff JB, Van Dijk L V., Muijs CT, Burgerhof JGM, Kats-Ugurlu G, et al. Predicting response to neoadjuvant chemoradiotherapy in esophageal cancer with textural features derived from pretreatment 18F-FDG PET/CT imaging. J Nucl Med. 2017;58(5):723–9.
Simoni N, Rossi G, Benetti G, Zuffante M, Micera R, Pavarana M, et al. F-FDG PET / CT Metrics Are Correlated to the Pathological Response in Esophageal Cancer Patients Treated With Induction Chemotherapy Followed by Neoadjuvant Chemo-Radiotherapy. Front Oncol. 2020;10:599907
Pan LL, Gu P, Huang G, Xue HP, Wu SQ. Prognostic significance of SUV on PET/CT in patients with esophageal cancer: A systematic review and meta-analysis. Eur J Gastroenterol Hepatol. 2009;21(9):1008–15.
Cao Q, Li Y, Li Z, An D, Li B, Lin Q. Development and validation of a radiomics signature on differentially expressed features of 18F-FDG PET to predict treatment response of concurrent chemoradiotherapy in thoracic esophagus squamous cell carcinoma. Radiother Oncol [Internet]. 2020;146:9–15.
Zhang H, Tan S, Chen W, Kligerman S, Kim G, D’Souza WD, et al. Modeling pathologic response of esophageal cancer to chemoradiation therapy using spatial-temporal 18F-FDG PET features, clinical parameters, and demographics. Int J Radiat Oncol Biol Phys [Internet]. 2014;88(1):195–203.
Qiu Q, Duan J, Deng H, Han Z, Gu J, Yue NJ, et al. Development and Validation of a Radiomics Nomogram Model for Predicting Postoperative Recurrence in Patients With Esophageal Squamous Cell Cancer Who Achieved pCR After Neoadjuvant Chemoradiotherapy Followed by Surgery. Front Oncol. 2020;10(August):1–10.
Yang CK, Yeh JCY, Yu WH, Chien LI, Lin KH, Huang WS, et al. Deep convolutional neural network-based positron emission tomography analysis predicts esophageal cancer outcome. J Clin Med. 2019;8(6):1–9.
Shen C, Liu Z, Wang Z, Guo J, Zhang H, Wang Y, et al. Building CT Radiomics Based Nomogram for Preoperative Esophageal Cancer Patients Lymph Node Metastasis Prediction. Transl Oncol [Internet]. 2018;11(3):815–24.
Hoshino I, Yokota H, Ishige F, Iwatate Y, Takeshita N, Nagase H, et al. Radiogenomics predicts the expression of microRNA-1246 in the serum of esophageal cancer patients. Sci Rep. 2020;10(1):1–8.
Holzinger A, Biemann C, Pattichis CS, Kell DB. What do we need to build explainable AI systems for the medical domain? 2017. ArXiv, abs/1712.09923
Ghassemi M, Oakden-rayner L, Beam AL. Viewpoint The false hope of current approaches to explainable artificial intelligence in health care. Lancet Digit Heal [Internet]. 2021;3(11):e745–50.
Daramola O, Nyasulu P, Mashamba-Thompson T, Moser T, Broomhead S, Hamid A, et al. Towards AI-Enabled Multimodal Diagnostics and Management of COVID-19 and Comorbidities in Resource-Limited Settings. Informatics [Internet]. 2021 Sep 23;8(4):63.
Acknowledgements
The authors wish to acknowledge the Institute for Life Sciences and University Hospital Southampton who jointly provide a funded studentship for NT.
Funding
NT receives a joint studentship from the Institute for Life Sciences (University of Southampton) and University Hospital Southampton.
Author information
Authors and Affiliations
Contributions
1) NT was involved in the conception of this work, its drafting, and revising for critical and important intellectual content, final approval, and agreement of accountability for accuracy.
2) GV was involved in the conception of this work, its drafting, and revising for critical and important intellectual content, final approval, and agreement of accountability for accuracy.
3) IB was involved in the conception of this work, its drafting, and revising for critical and important intellectual content, final approval, and agreement of accountability for accuracy.
4) TJU was involved in the conception of this work, its drafting, and revising for critical and important intellectual content, final approval, and agreement of accountability for accuracy. In addition, TJU acts in role of study supervision.
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thavanesan, N., Vigneswaran, G., Bodala, I. et al. The Oesophageal Cancer Multidisciplinary Team: Can Machine Learning Assist Decision-Making?. J Gastrointest Surg 27, 807–822 (2023). https://doi.org/10.1007/s11605-022-05575-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11605-022-05575-8