Abstract
Background
Machine learning (ML) has seen an increase in application, and is an important element of a digital evolution. The role of ML within upper gastrointestinal surgery for malignancies has not been evaluated properly in the literature. Therefore, this systematic review aims to provide a comprehensive overview of ML applications within upper gastrointestinal surgery for malignancies.
Methods
A systematic search was performed in PubMed, EMBASE, Cochrane, and Web of Science. Studies were only included when they described machine learning in upper gastrointestinal surgery for malignancies. The Cochrane risk-of-bias tool was used to determine the methodological quality of studies. The accuracy and area under the curve were evaluated, representing the predictive performances of ML models.
Results
From a total of 1821 articles, 27 studies met the inclusion criteria. Most studies received a moderate risk-of-bias score. The majority of these studies focused on neural networks (n = 9), multiple machine learning (n = 8), and random forests (n = 3). Remaining studies involved radiomics (n = 3), support vector machines (n = 3), and decision trees (n = 1). Purposes of ML included predominantly prediction of metastasis, detection of risk factors, prediction of survival, and prediction of postoperative complications. Other purposes were predictions of TNM staging, chemotherapy response, tumor resectability, and optimal therapy.
Conclusions
Machine Learning algorithms seem to contribute to the prediction of postoperative complications and the course of disease after upper gastrointestinal surgery for malignancies. However, due to the retrospective character of ML studies, these results require trials or prospective studies to validate this application of ML.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Artificial intelligence (AI) has been introduced within healthcare recently, therefore its role has become increasingly important. As the application of AI is escalating, it may be apparent that a digital evolution is ongoing in healthcare [1].
Artificial intelligence is described as the capacity of machines to mimic intelligent human behavior [2]. Artificial intelligence has proven its potential of using large datasets to perform specific tasks, such as image recognition [3]. Machine learning (ML) can be defined as an analytic approach in which models are trained on databases to make predictions on new unseen data [2]. Relevant examples of ML involve support vector machines, decision trees, and gradient boosting. Deep learning is an important subdiscipline of ML, in which multiple datasets are simultaneously included, these datasets are evaluated and modified until the next turn of evaluation. These evaluations are represented as layers and are based on the output of the preceding layer. These processes of evaluation are continued until a final output has been reached. Radiomics are a separate subdiscipline of AI and are able to extract textural features by analyzing medical images [4]. To develop a better understanding, all subdisciplines of AI are depicted in Fig. 1 and explained in Table 1.
Since the datasets within electronic medical records are expanding, the abilities of humans to analyze data have been exceeded. As a result, the human role is causing problems such as diagnostic errors, inefficiencies in workflow, and inappropriate treatments in the healthcare system [5]. To eliminate these problems, AI is currently being applied due to its great capacity for analyzing large quantities of data and gaining more experience in a relatively shorter amount of time.
Esophageal and gastric cancer have a high prevalence with poor outcomes, keeping the treatment procedure challenging [6,7,8,9]. The introduction of multimodality treatments and minimally invasive techniques have resulted in better treatment outcomes [10]. To further minimize postoperative complication rates, AI tools may be helpful. Another important issue concerns the selection of patients who could optimally benefit from neoadjuvant chemoradiotherapy (nCRT) [11]. Artificial intelligence could contribute to upper gastrointestinal surgery by identifying patients with a complete response to nCRT. Such pre-operative differentiations could be performed if large surgical datasets would be analyzed by ML. Eventually, using ML could provide personalized medicine to optimize treatment outcomes after upper gastrointestinal surgery.
Despite the potential benefits of AI, the scope of ML applications in upper gastrointestinal surgery for malignancies is barely described in literature. However, to support ML implementations in daily practice, it is vital to fill this gap to understand the role and progress of ML within upper gastrointestinal surgery properly. Therefore, this systematic review will provide a comprehensive overview of ML applications in upper gastrointestinal surgery for malignancies.
Materials and methods
Search strategy
This study was reported in accordance with the Cochrane Handbook for Systematic Reviews of Interventions version 6.0 and PRISMA guidelines. A systematic search was performed in the databases: PubMed, Embase.com, Clarivate Analytics/Web of Science Core Collection, and the Wiley/Cochrane Library. The timeframe within the databases was from inception to the 7th of July 2021 and conducted by M.B. and G.L.B. The search included keywords and free text terms for (synonyms of) 'machine learning' combined with (synonyms of) 'digestive system surgical procedures'. A full overview of the search terms per database can be found in the supplementary information (see appendix 1).
Study selection
In the first step, to avoid missing studies with overlapping content, articles were included when they described ML within general surgery. Afterwards, studies were only approved if they met the following criteria: (1) describing ML in upper gastrointestinal surgery for malignancies, (2) clinical study, (3) conducted on adults. This review focused only on the most commonly used ML algorithms with adaptive learning abilities, therefore regression models have been excluded as these have been considered traditional statistical approaches. Articles were excluded if they: (1) did not describe the use of ML, (2) the involvement of upper gastrointestinal surgery and malignancies was absent, (3) were not written in English, (4) were publications reporting on reviews, histological analysis, study abstracts, conference proceedings, book chapters, editorials, errata, letters, notes, surveys or tombstones. No peculiar study design was chosen as an inclusion criterium. The title and abstract screening was independently performed by two reviewers (M.B. and G.L.B.) according to the inclusion and exclusion criteria. Studies were included for full-text evaluation when both reviewers agreed on inclusion. Disagreements were solved by discussions between two reviewers, resulting in consensus.
Quality assessment and risk of bias
The methodological quality of the included randomized controlled trials was assessed using the Revised Cochrane risk-of-bias tool for randomized trials [22]. This tool determines the overall risk of bias based on five bias domains: the randomization process, deviations from intended interventions, missing outcome data, measurement of outcomes, and selection of reported results. The ROBINS-I assessment tool was used to determine the methodological quality of non-randomized studies [23], which determines the overall risk of bias based on seven bias domains; confounding, participant selection, intervention classification, deviations from intended interventions, missing outcome data, measurement of outcomes, and selection of reported results.
Data synthesis and outcome assessment
After the full-text assessment, the following data from included studies were independently extracted by two reviewers (M.B. and G.L.B.): first author, year, country, number of patients, mean age, study design, carcinoma type, surgical procedure, type of ML, purpose of ML, outcome measurements, and predictive performance. Subsequently, studies were categorized based on surgical procedures and ML subdisciplines, and results of ML use were described within these categories. To summarize the results of studies in quantitative data, the mean accuracy (ACC) and area under the curve (AUC) were calculated as a representation of predictive performances. Conflicts among reviewers were solved by consensus.
Results
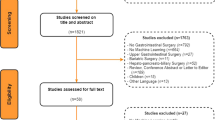
The search strategy provided a total of 1821 studies after the removal of duplicates (Fig. 2). Therefore, 1821 studies were screened for eligibility based on the title and abstract screening. Afterwards, 164 studies remained for the full-text assessment, resulting in the inclusion of 27 studies.
In all studies, various ML models have been applied within upper gastrointestinal surgery. The majority of these studies focused on neural networks (n = 9), multiple machine learning (n = 8), and random forests (n = 3). Remaining studies involved radiomics (n = 3), support vector machines (n = 3), and decision trees (n = 1). Eleven studies involved esophagectomy procedures, whereas sixteen studies concerned gastrectomy procedures. Purposes of ML included predominantly prediction of metastasis (n = 5), detection of risk factors (n = 5), prediction of survival (n = 5), and prediction of postoperative complications (n = 4). Other purposes were predictions of TNM staging (n = 2), chemotherapy response (n = 2), tumor resectability (n = 2), and optimal therapy (n = 2).
In most studies, patients were randomly divided into a training set and a test set. Training sets were used for the development of ML models, afterwards test sets were utilized to determine the accuracy of developed models.
An overview of the study characteristics is presented in Table 2.
Methodological quality assessment
All included articles were retrospective cohort studies, therefore only the ROBINS-I assessment tool was applied (Table 3). Due to the nature of ML, domains such as bias due to confounding and bias in outcome measurements received low risk-of-bias scores. However, because of the retrospective study design of these studies, moderate risk-of-bias scores were given for bias in the intervention classification domain.
Esophagectomy
For patients undergoing esophagectomy for malignant esophageal cancer, the following ML models were used: decision trees (n = 1), random forests (n = 3), neural networks (n = 3), radiomics (n = 1), and multiple machine learning (n = 3).
Decision tree
Shao et al. developed a decision tree model to predict the occurrence of anastomotic leakage after surgery in patients with esophageal tumors [24]. During the training phase of the model, univariate analysis indicated that the CRP to albumin ratio, postoperative CRP, lymphocytes, surgical duration, postoperative albumin, pre-operative red blood cells, tumor size, TNM, and ASA score were major predictive factors for anastomotic leakage. This model predicted 55 anastomotic leakage cases with an accuracy of 98% and an AUC of 0.95.
Random forest
Bolourani et al. trained a random forest model to predict patients with early readmissions in 30 days due to severe complications after esophagectomy [25]. Additionally, the most important risk factors for early readmission were determined. Reintubation, prolonged intubation, tracheostomy, aspiration pneumonitis, pyothorax, anastomotic leak, pneumonia, and acute kidney failure appeared to be the essential risk factors for early readmission. After application, the random forest predicted 383 early readmitted patients with an AUC of 0.74. Furthermore, Rice et al. developed a random forest model to detect essential factors associated with lymph node metastasis in patients with esophageal tumors [26]. This model discovered that the increasing cancer size, increasing depth of cancer invasion, and decreasing cancer differentiation are the most important factors for lymph node metastasis after esophagectomy. Additionally, another random forest model was trained to predict the optimal therapy for patients with esophageal tumors [27]. Therapy options included only esophagectomy and esophagectomy combined with adjuvant chemoradiotherapy or neoadjuvant chemoradiotherapy. Optimal therapy was determined as the treatment with a maximum mean survival time. For all patients that received an esophagectomy procedure only, this therapy was optimal in 61% of the cases. In 20% of the remaining patients, the survival time would have increased by 7% if adjuvant therapy was applied, and another 19% would also have benefitted if neoadjuvant therapy was used.
Neural networks
Chen et al. developed artificial neural networks (ANNs) to predict lymph node metastasis in patients with T1 esophageal tumors undergoing surgery. Multivariate analysis identified six essential factors for developing lymph node metastasis: alcohol use, tumor size, invasion depth, histologic grade, lymph-vessel invasion, and positive imaging results [28]. The ANN model appeared to have an accuracy of 91% for predicting 133 patients with lymph node metastasis. Another study trained an ANN model to accurately predict N staging in patients with T1–T2 esophageal carcinomas that underwent surgery [29]. Univariate analysis indicated that tumor invasion depth, tumor length, tumor differentiation, lymph-vessel invasion, and dysphagia were important factors for predicting positive lymph nodes after surgery. By using this ANN model, 148 patients with N1, 65 patients with N2, and three patients with N3 positive lymph nodes were predicted, demonstrating an AUC of 0.85 for this prediction. Another ANN model was developed by Mofidi et al. to predict 1- and 3- year disease-free survival of patients that underwent surgery for esophageal carcinomas [30]. For the 1-year disease-free survival, the ANN model showed an overall accuracy of 88%, whereas an accuracy of 92% was demonstrated for the 3-year disease-free survival.
Radiomics
Rishi et al. developed a radiomics model for PET and CT scans to predict the response of esophageal tumors to neoadjuvant chemoradiotherapy [31]. Based on multiple radiomic features, patients with a low radiomic score showed a significant improvement in response to neoadjuvant chemoradiotherapy (nCRT). A complete response to nCRT was predicted in 34 patients with an accuracy of 77% and an AUC of 0.87.
Multiple machine learning
Ou et al. extracted radiomic features and combined them with multiple ML models to predict the resectability of esophageal carcinomas [32]. The resectability of tumors was defined according to the National Comprehensive Cancer Network (NCCN) [33]. A total of 270 patients with resectable carcinomas was predicted by the included ML algorithms. The gradient boosting model showed an accuracy of 79% with an AUC of 0.84. The SVM model indicated an accuracy of 79% with an AUC of 0.82. An accuracy of 69% and an AUC of 0.66 were discovered for the decision tree model. Finally, the random forest model included an accuracy of 67% with an AUC of 0.67 for this prediction. A combined model of the random forest and gradient boosting techniques was developed by Rahman et al. to predict early recurrence of esophageal cancer after surgery [34]. Univariate statistical analysis signified the number of positive lymph nodes and lymphovascular invasion as the most important predictors for early recurrence. Early recurrence after 1 year occurred in a total of 236 patients. Furthermore, for the discriminative ability, this combined model showed an AUC of 0.81 for the interval validation. Application on external databases resulted in an AUC of 0.80. Wang et al. trained a model by combining radiomics with the random forest technique to predict the overall survival of patients that have received surgery for esophageal carcinomas [35]. The index of concordance (C-index) was measured for this model, this index represented the fraction of patients whose survival was correctly predicted from 0 to 1. For this model, the C-index was discovered to be 0.54.
Gastrectomy
For patients that received a gastrectomy procedure for malignancies, several ML models were applied: SVM (n = 3), neural networks (n = 6), radiomics (n = 2), and multiple machine learning (n = 5).
Support vector machine
Dai et al. developed an SVM model to detect the resectability of gastric cancer [36]. The accuracy of this SVM model was discovered to be 87% in performing a classification between tumor and healthy tissue. Liu et al. trained an SVM model to decide between D1 and D2 lymphadenectomy in patients with gastric cancer [37]. Statistical analysis showed that the T stage, N1 lymph nodes, maximum length of tumor, parenchymal enhancement of tumor, and diameter of lymph node were essential factors for this clinical decision. This SVM model proved an AUC of 0.94 for this classification. Another SVM model was developed to predict postoperative complications after gastrectomy for gastric cancer [38]. The most important clinical features for postoperative complications were discovered to be age, tumor size, comorbidities, hemoglobin, and total protein levels. A total of 100 patients with postoperative complications was predicted by this model with an accuracy of 78%.
Neural networks
Bollschweiler et al. trained an ANN model with the aim of predicting lymph node metastasis in patients with gastric cancer [39]. An accuracy of 93% was detected for this ANN model in predicting 38 patients with lymph node metastasis. Additionally, Jiang et al. developed an ANN model to predict peritoneal metastasis after surgery [40]. For predicting peritoneal metastasis, tumor type and tumor differentiation appeared to be the key predictors. For this model, an AUC of 0.92 was found for predicting peritoneal metastasis. In another study, a CNN model was trained to predict lymph node metastasis in gastric cancer patients [41]. For lymph node metastasis, tumor size, tumor location, differentiation, depth of invasion, and tumor stage appeared to be the most important predictors. After external validation, an AUC of 0.88 was discovered for this CNN model.
Li et al. developed an ANN model to predict 5-year survival in cohort groups of patients from three different institutes that only underwent a gastrectomy procedure [42]. Essential factors for a 5-year survival were the depth of invasion, lymph node metastasis, age, and tumor size. The AUC for this model appeared to be 0.80, 0.84, and 0.85 in these three groups. Another ANN model was trained to predict 5-year survival after radical gastrectomy [43]. For this model, the AUC was observed to be 0.81 for this prediction. To determine risk factors for the survival of gastric cancer patients, Zhu et al. developed an ANN model for patients that underwent radical gastrectomy [44]. According to this model, tumor stage, radical surgery, serum CA19-9 levels, peritoneal dissemination, and BMI were significant predictors of gastric cancer survival. An AUC of 0.89 was found for this ANN model.
Radiomics
By training a radiomics model, Chen et al. aimed to predict response to nCRT in patients with advanced gastric cancer [45]. Out of all clinical factors, only tumor differentiation was significant for this response. This model identified 61 patients with complete response to nCRT, and this prediction was performed with an AUC of 0.74. Additionally, Li et al. developed a radiomics model to detect risk factors for advanced gastric cancer [46]. Multivariate analysis indicated that the tumor stage, number of lymph nodes, lymphatic vascular infiltration, and histological grade were key radiomic features for advanced gastric cancer. This radiomics model detected these features with an AUC of 0.87, 0.73, 0.68, and 0.78, respectively.
Multiple machine learning
Multiple machine learning models were trained by Akcay et al., who aimed to predict overall survival for patients that received chemoradiation therapy after gastrectomy [47]. After multivariate statistical analysis, the most important factors for overall survival were discovered to be the number of removed lymph nodes, the number of metastatic lymph nodes, the lymph node ratio, and the neoadjuvant CT history. The median overall survival was predicted to be 23 months. Gradient boosting showed an AUC of 0.64, whereas the random forest model was observed to have an AUC of 0.59. The SVM model had an AUC of 0.50, and the ANN model had an AUC of 0.45 in predicting the overall survival.
Celik et al. developed an ANN, decision tree, and a random forest model to predict anastomotic leakage after gastrectomy [48]. For postoperative leakage, the random forest and decision tree model both showed a sensitivity of 9% and a specificity of 91%. The ANN model had a sensitivity of 8% and a specificity of 92%. Furthermore, a prediction model was developed based on radiomics and support vector machine techniques. This model was trained to predict lymph node metastasis in gastric cancer patients [49]. 297 patients with lymph node metastasis were identified. External validation showed an AUC of 0.76 and an accuracy of 71% for this model.
Random forest and decision tree models were developed by Huang et al. to predict risk factors and diagnosis for lymph node metastasis in gastric cancer [50]. Random forest showed that tumor size, CT findings, histological grade, hemoglobin, and carcinoembryonic antigen were key factors for the development of gastric lymph node metastasis. The decision tree model was discovered to have an accuracy of 76% for the diagnosis of lymph node metastasis in gastric cancer. Qiao et al. extracted radiomic features and combined them with SVM, decision tree, and random forest algorithms to predict TNM staging in gastric cancer patients [51]. The support vector machine model appeared to have an AUC of 0.79 in predicting TNM staging. The random forest model showed an AUC of 0.78, whereas the decision tree algorithm demonstrated an AUC of 0.63.
Discussion
From this systematic review, it can be concluded that ML has shown its potential in predicting the course of disease and postoperative complications after upper gastrointestinal surgery for malignancies. Machine learning models have predicted the course of disease with accuracies up to 93% and AUCs up to 0.94, whereas postoperative complications have been predicted with accuracies up to 98% and AUCs up to 0.95.
By using ML, surgeons could be able to pre-operatively identify patients at high risk of postoperative complications. Theoretically, this insight could aid the surgeon in deciding the type of operation and considering prophylactic measures to minimize postoperative complications. For many years, several risk factors for esophagogastric cancer and metastasis have been reported, but as many risk factors are varying from demographical to genetic characteristics, these reports remain controversial [52,53,54]. Machine learning could be used to select the most important risk factors out of all variables that have been inserted into the model. Studies within this review have already revealed the tumor size, depth of cancer invasion, tumor location, and histologic grade to be essential predictors for the course of disease concerning upper gastrointestinal cancer. Recognizing the most relevant risk factors could be very important for achieving early diagnosis and improving the prognosis of patients with esophagogastric cancer. In addition, ML could also predict response to nCRT in patients with upper gastrointestinal cancer. Based on these predictions, surgeons could reconsider surgical treatment strategies to optimize the treatment outcomes.
Conventional statistics have been applied for many years in surgery. Traditional regression models have been used to predict several aspects concerning the course of disease, such as lymph node metastasis, distant metastasis, survival, and remnant cancer after upper gastrointestinal surgery for malignancies. Predictions were performed with AUCs generally ranging between 0.7 and 0.8 [55,56,57,58]. This might be partially overlapping with ML algorithms, as these ranged from 0.7 to 0.98 within this review. However, this review shows that the AUCs of ML algorithms were usually above 0.8, indicating superior predictive capabilities of ML in predicting the course of disease of upper gastrointestinal cancer. Both methods could be used for data analysis and predictions, but there are important differences. Conventional statistics are mainly used to find relationships between usually two variables, whereas ML focuses on recognizing patterns within the data, not being restricted by the number of variables [59]. As patient databases contain a vast amount of clinical variables, ML would be preferred due to its ability to analyze high numbers of variables and complex data.
This review has some limitations. More articles could have been included if more technical databases were searched for ML studies. In addition, due to the presence of inconsistencies in reported accuracies, a meta-analysis could not be performed.
Within this review, it has been discovered that ML models have been only applied retrospectively, therefore prospective studies should be prioritized in the future to gain clinical validation for ML. Gaining this clinical validation could enable patients to receive a personalized treatment plan based on ML predictions. However, to implement ML applications in the field of upper gastrointestinal surgery, a few obstacles need to be overcome. Even though ML models are already applied, clinicians and medical residents still experience difficulties in understanding how the model exactly functions, this is also called “the black box” issue [60]. Additionally, data scientists and engineers have issues with understanding the concepts and interpretations of clinical data, this might impede the development of ML models [61]. To overcome these problems, interdisciplinary collaborations are key to implementing ML applications successfully. To produce high-quality data, data scientists and engineers should be responsible for data preparations, data processing, and feature selections during the training phase. Additionally, clinicians should label clinical data clearly and provide accurate segmentation of esophagogastric cancer on medical images. Clinicians should also focus on feedback from patients to discover additional objectives that could be predicted with ML. However, before these steps, it is essential to agree on study protocols and patient schedules to ensure a systematic and transparent approach to the development of ML models [62]. Furthermore, stakeholders in the hospital should establish strict policies for patient data protection.
Conclusion
Machine learning has demonstrated accurate predictive capabilities for patients with upper gastrointestinal malignancies undergoing surgery. These capabilities became apparent for short-term outcomes such as prediction of postoperative complications, and long-term outcomes such as prediction of metastasis and survival. However, the current performances of ML are based on retrospective studies only. Therefore, these results require trials and prospective studies to gain clinical validation for ML.
References
Briganti G, Le Moine O (2020) Artificial intelligence in medicine: today and tomorrow. Front Med 7:27
Davenport T, Kalakota R (2019) The potential for artificial intelligence in healthcare. Future Healthc J 6(2):94–98
Kulkarni S, Seneviratne N, Baig MS, Khan AHA (2020) Artificial intelligence in medicine: where are we now? Acad Radiol 27(1):62–70
Mintz Y, Brodie R (2019) Introduction to artificial intelligence in medicine. Minim Invasive Ther Allied Technol 28(2):73–81
Topol EJ (2019) High-performance medicine: the convergence of human and artificial intelligence. Nat Med 25(1):44–56
Uhlenhopp DJ, Then EO, Sunkara T, Gaduputi V (2020) Epidemiology of esophageal cancer: update in global trends, etiology and risk factors. Clin J Gastroenterol 13(6):1010–1021
Karimi P, Islami F, Anandasabapathy S, Freedman ND, Kamangar F (2014) Gastric cancer: descriptive epidemiology, risk factors, screening, and prevention. Cancer Epidemiol Biomark Prev 23(5):700–713
Yeung JC (2020) Management of complications after esophagectomy. Thorac Cardiovasc Surg 30(3):359–366
Rawicz-Pruszyński K, van Sandick JW, Mielko J, Ciseł B, Polkowski WP (2018) Current challenges in gastric cancer surgery: European perspective. Surg Oncol 27(4):650–656
van Hagen P, Hulshof MC, van Lanschot JJ, Steyerberg EW, van Berge Henegouwen MI, Wijnhoven BP, Richel DJ, Nieuwenhuijzen GA, Hospers GA, Bonenkamp JJ, Cuesta MA, Blaisse RJ, Busch OR, ten Kate FJ, Creemers GJ, Punt CJ, Plukker JT, Verheul HM, Spillenaar Bilgen EJ, van Dekken H, CROSS Group (2012) Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med 366(22):2074–2084
Kim JY, Hofstetter WL (2012) Esophagectomy after chemoradiation: who and when to operate. Semin Thorac Cardiovasc Surg 24(4):288–293
Ramesh AN, Kambhampati C, Monson JR, Drew PJ (2004) Artificial intelligence in medicine. Ann R Coll Surg Engl 86(5):334–338
El Naqa I, Murphy M (2015) What is machine learning? Springer, Cham
Zhang L, Zhou W, Jiao L (2004) Wavelet support vector machine. IEEE Trans Syst Man Cybern B Cybern 34(1):34–39
Song YY, Lu Y (2015) Decision tree methods: applications for classification and prediction. Shanghai Arch Psychiatry 27(2):130–135
Breiman L (2001) Random forests. Mach Learn 45(1):5–32
Friedman JH (2002) Stochastic gradient boosting. Comput Stat Data Anal 38(4):367–378
Rusk N (2016) Deep learning. Nat Methods 13(1):35–35
Abraham A (2005) Artificial neural networks. Wiley, Chichester
Albawi S, Mohammed TA, Al-Zawi S (2017) Understanding of a convolutional neural network. In: International conference on engineering and technology, pp 1–6
Kumar V, Gu Y, Basu S, Berglund A, Eschrich SA, Schabath MB, Forster K, Aerts HJ, Dekker A, Fenstermacher D, Goldgof DB, Hall LO, Lambin P, Balagurunathan Y, Gatenby RA, Gillies RJ (2012) Radiomics: the process and the challenges. Magn Reson Imaging 30(9):1234–1248
Sterne JAC, Savović J, Page MJ, Elbers RG, Blencowe NS, Boutron I, Cates CJ, Cheng HY, Corbett MS, Eldridge SM, Emberson JR, Hernán MA, Hopewell S, Hróbjartsson A, Junqueira DR, Jüni P, Kirkham JJ, Lasserson T, Li T, McAleenan A, Reeves BC, Shepperd S, Shrier I, Stewart LA, Tilling K, White IR, Whiting PF, Higgins JPT (2019) RoB 2: a revised tool for assessing risk of bias in randomised trials. BMJ 366:l4898
Sterne JA, Hernán MA, Reeves BC, Savović J, Berkman ND, Viswanathan M, Henry D, Altman DG, Ansari MT, Boutron I, Carpenter JR, Chan AW, Churchill R, Deeks JJ, Hróbjartsson A, Kirkham J, Jüni P, Loke YK, Pigott TD, Ramsay CR, Regidor D, Rothstein HR, Sandhu L, Santaguida PL, Schünemann HJ, Shea B, Shrier I, Tugwell P, Turner L, Valentine JC, Waddington H, Waters E, Wells GA, Whiting PF, Higgins JP (2016) ROBINS-I: a tool for assessing risk of bias in non-randomised studies of interventions. BMJ 355:i4919
Shao CY, Liu KC, Li CL, Cong ZZ, Hu LW, Luo J, Diao YF, Xu Y, Ji SG, Qiang Y, Shen Y (2019) C-reactive protein to albumin ratio is a key indicator in a predictive model for anastomosis leakage after esophagectomy: application of classification and regression tree analysis. Thorac Cancer 10(4):728–737
Bolourani S, Tayebi MA, Diao L, Wang P, Patel V, Manetta F, Lee PC (2021) Using machine learning to predict early readmission following esophagectomy. J Thorac Cardiovasc Surg 161(6):1926–1939
Rice TW, Ishwaran H, Hofstetter WL, Schipper PH, Kesler KA, Law S, Lerut EM, Denlinger CE, Salo JA, Scott WJ, Watson TJ, Allen MS, Chen LQ, Rusch VW, Cerfolio RJ, Luketich JD, Duranceau A, Darling GE, Pera M, Apperson-Hansen C, Blackstone EH (2017) Esophageal cancer: associations with (pN+) lymph node metastases. Ann Surg 265(1):122–129
Rice TW, Lu M, Ishwaran H, Blackstone EH (2019) Precision surgical therapy for adenocarcinoma of the esophagus and esophagogastric junction. J Thorac Oncol 14(12):2164–2175
Chen H, Zhou X, Tang X, Li S, Zhang G (2020) Prediction of lymph node metastasis in superficial esophageal cancer using a pattern recognition neural network. Cancer Manag Res 12:12249–12258
Liu XL, Shao CY, Sun L, Liu YY, Hu LW, Cong ZZ, Xu Y, Wang RC, Yi J, Wang W (2020) An artificial neural network model predicting pathologic nodal metastases in clinical stage I-II esophageal squamous cell carcinoma patients. J Thorac Dis 12(10):5580–5592
Mofidi R, Deans C, Duff MD, de Beaux AC, Paterson-Brown S (2006) Prediction of survival from carcinoma of oesophagus and oesophago-gastric junction following surgical resection using an artificial neural network. Eur J Surg Oncol 32(5):533–539
Rishi A, Zhang GG, Yuan Z, Sim AJ, Song EY, Moros EG, Tomaszewski MR, Latifi K, Pimiento JM, Fontaine JP, Mehta R, Harrison LB, Hoffe SE, Frakes JM (2021) Pretreatment CT and 18F-FDG PET-based radiomic model predicting pathological complete response and loco-regional control following neoadjuvant chemoradiation in oesophageal cancer. J Med Imaging Radiat Oncol 65(1):102–111
Ou J, Li R, Zeng R, Wu CQ, Chen Y, Chen TW, Zhang XM, Wu L, Jiang Y, Yang JQ, Cao JM, Tang S, Tang MJ, Hu J (2019) CT radiomic features for predicting resectability of oesophageal squamous cell carcinoma as given by feature analysis: a case control study. Cancer Imaging 19(1):66
Pluchino LA, D’Amico TA (2020) National comprehensive cancer network guidelines: who makes them? What are they? Why are they important? Ann Thorac Surg 110(6):1789–1795
Rahman SA, Walker RC, Lloyd MA, Grace BL, van Boxel GI, Kingma BF, Ruurda JP, van Hillegersberg R, Harris S, Parsons S, Mercer S, Griffiths EA, O’Neill JR, Turkington R, Fitzgerald RC, Underwood TJ (2020) Machine learning to predict early recurrence after oesophageal cancer surgery. Br J Surg 107(8):1042–1052
Wang J, Wu LL, Zhang Y, Ma G, Lu Y (2021) Establishing a survival prediction model for esophageal squamous cell carcinoma based on CT and histopathological images. Phys Med Biol 66(14):145015
Dai H, Bian Y, Wang L, Yang J (2021) Support vector machine-based backprojection algorithm for detection of gastric cancer lesions with abdominal endoscope using magnetic resonance imaging images. Sci Program 2021:1–8
Liu C, Qi L, Feng QX, Sun SW, Zhang YD, Liu XS (2019) Performance of a machine learning-based decision model to help clinicians decide the extent of lymphadenectomy (D1 vs. D2) in gastric cancer before surgical resection. Abdom Radiol (NY) 44(9):3019–3029
Lu S, Yan M, Li C, Yan C, Zhu Z, Lu W (2019) Machine-learning-assisted prediction of surgical outcomes in patients undergoing gastrectomy. Chin J Cancer Res 31(5):797–805
Bollschweiler EH, Mönig SP, Hensler K, Baldus SE, Maruyama K, Hölscher AH (2004) Artificial neural network for prediction of lymph node metastases in gastric cancer: a phase II diagnostic study. Ann Surg Oncol 11(5):506–511
Jiang Y, Liang X, Wang W, Chen C, Yuan Q, Zhang X, Li N, Chen H, Yu J, Xie Y, Xu Y, Zhou Z, Li G, Li R (2021) noninvasive prediction of occult peritoneal metastasis in gastric cancer using deep learning. JAMA Netw Open 4(1):e2032269
Jin C, Jiang Y, Yu H, Wang W, Li B, Chen C, Yuan Q, Hu Y, Xu Y, Zhou Z, Li G, Li R (2021) Deep learning analysis of the primary tumour and the prediction of lymph node metastases in gastric cancer. Br J Surg 108(5):542–549
Li Z, Wu X, Gao X, Shan F, Ying X, Zhang Y, Ji J (2020) Development and validation of an artificial neural network prognostic model after gastrectomy for gastric carcinoma: an international multicenter cohort study. Cancer Med 9(17):6205–6215
Oh SE, Seo SW, Choi MG, Sohn TS, Bae JM, Kim S (2018) Prediction of overall survival and novel classification of patients with gastric cancer using the survival recurrent network. Ann Surg Oncol 25(5):1153–1159
Zhu L, Luo W, Su M, Wei H, Wei J, Zhang X, Zou C (2013) Comparison between artificial neural network and Cox regression model in predicting the survival rate of gastric cancer patients. Biomed Rep 1(5):757–760
Chen Y, Wei K, Liu D, Xiang J, Wang G, Meng X, Peng J (2021) A Machine learning model for predicting a major response to neoadjuvant chemotherapy in advanced gastric cancer. Front Oncol 11:675458
Li Q, Qi L, Feng QX, Liu C, Sun SW, Zhang J, Yang G, Ge YQ, Zhang YD, Liu XS (2019) Machine learning-based computational models derived from large-scale radiographic-radiomic images can help predict adverse histopathological status of gastric cancer. Clin Transl Gastroenterol 10(10):00079
Akcay M, Etiz D, Celik O (2020) Prediction of survival and recurrence patterns by machine learning in gastric cancer cases undergoing radiation therapy and chemotherapy. Adv Radiat Oncol 5(6):1179–1187
Celik S, Sohail A, Ashraf S, Arshad A (2019) Application of machine learning techniques to analyze anastomosis integrity after Total gastrectomy for prediction of clinical leakage. Health Technol (Berl) 9(5):757–763
Feng QX, Liu C, Qi L, Sun SW, Song Y, Yang G, Zhang YD, Liu XS (2019) An intelligent clinical decision support system for preoperative prediction of lymph node metastasis in gastric cancer. J Am Coll Radiol 16(7):952–960
Huang C, Hu C, Zhu J, Zhang W, Huang J, Zhu Z (2020) Establishment of decision rules and risk assessment model for preoperative prediction of lymph node metastasis in gastric cancer. Front Oncol 10:1638
Qiao X, Li Z, Li L, Ji C, Li H, Shi T, Gu Q, Liu S, Zhou Z, Zhou K (2021) Preoperative T2-weighted MR imaging texture analysis of gastric cancer: prediction of TNM stages. Abdom Radiol (NY) 46(4):1487–1497
Domper Arnal MJ, Ferrández Arenas Á, Lanas Arbeloa Á (2015) Esophageal cancer: risk factors, screening and endoscopic treatment in Western and Eastern countries. World J Gastroenterol 21(26):7933–7943
Jiang KY, Huang H, Chen WY, Yan HJ, Wei ZT, Wang XW, Li HX, Zheng XY, Tian D (2021) Risk factors for lymph node metastasis in T1 esophageal squamous cell carcinoma: A systematic review and meta-analysis. World J Gastroenterol 27(8):737–750
Yusefi AR, Bagheri Lankarani K, Bastani P, Radinmanesh M, Kavosi Z (2018) Risk factors for gastric cancer: a systematic review. Asian Pac J Cancer Prev 19(3):591–603
Kim SM, Lee H, Min BH, Kim JJ, An JY, Choi MG, Bae JM, Kim S, Sohn TS, Lee JH (2019) A prediction model for lymph node metastasis in early-stage gastric cancer: toward tailored lymphadenectomy. J Surg Oncol 120(4):670–675
Zhu C, You Y, Liu S, Ji Y, Yu J (2020) A nomogram to predict distant metastasis for patients with esophageal cancer. Oncol Res Treat 43(1–2):2–9
Xie SH, Santoni G, Mälberg K, Lagergren P, Lagergren J (2021) Prediction model of long-term survival after esophageal cancer surgery. Ann Surg 273(5):933–939
Lu J, Zheng ZF, Zhou JF, Xu BB, Zheng CH, Li P, Xie JW, Wang JB, Lin JX, Chen QY, Truty MJ, He QL, Huang CM (2019) A novel prognosis prediction model after completion gastrectomy for remnant gastric cancer: development and validation using international multicenter databases. Surgery 166(3):314–321
Bzdok D, Altman N, Krzywinski M (2018) Statistics versus machine learning. Nat Methods 15:233–234
Habli I, Lawton T, Porter Z (2020) Artificial intelligence in health care: accountability and safety. Bull World Health Organ 98(4):251–256
Martín-Noguerol T, Paulano-Godino F, López-Ortega R, Górriz JM, Riascos RF, Luna A (2021) Artificial intelligence in radiology: relevance of collaborative work between radiologists and engineers for building a multidisciplinary team. Clin Radiol 76(5):317–324
Kusters R, Misevic D, Berry H, Cully A, Le Cunff Y, Dandoy L, Díaz-Rodríguez N, Ficher M, Grizou J, Othmani A, Palpanas T, Komorowski M, Loiseau P, Moulin Frier C, Nanini S, Quercia D, Sebag M, Soulié Fogelman F, Taleb S, Tupikina L, Wehbi F (2020) Interdisciplinary research in artificial intelligence: challenges and opportunities. Front Big Data 3:577974
Funding
No funding was required for this systematic review.
Author information
Authors and Affiliations
Contributions
MB: participated in the design of the study, data collection and interpretation, wrote and submitted the manuscript. GLB: participated in the design of the study and wrote parts of the manuscript. HJB: participated in the design of the study and revised the manuscript critically. DLvdP: participated in the design of the study, revised the manuscript critically, and submitted the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Disclosures
Dr. Bektaş, Dr. Burchell, Professor van der Peet, Professor Bonjer have no conflicts of interest or financial ties to disclose.
Ethical Approval
None.
Research registration number
None.
Guarantor
M Bektaş.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bektaş, M., Burchell, G.L., Bonjer, H.J. et al. Machine learning applications in upper gastrointestinal cancer surgery: a systematic review. Surg Endosc 37, 75–89 (2023). https://doi.org/10.1007/s00464-022-09516-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00464-022-09516-z