Abstract
Landscape disturbances can affect reproductive performance of animal-pollinated trees. We verified the effects of the loss and fragmentation of natural forests caused by the creation of coniferous plantations on fruit and seed production as well as mating patterns of animal-pollinated trees. We investigated 146 and 134 individual flowering trees of Prunus verecunda (Koidz.) Koehne in 2012 and 2013, respectively, at 12 sites. These sites were at least 1.2 km apart from each other in an 8 × 15-km forestry region, and the composition and configuration of natural forests varied in the study area. The mean outcrossing rate was >0.99 across the sites. Among trees within the sites, the number of fruits per inflorescence was positively correlated with the basal stem area and leaf chlorophyll density of the trees. Among the sites, the mean number of fruits per inflorescence was positively correlated with the site elevation, and the correlated paternity was positively correlated with the mean distance between trees. The sound-seed rate was positively correlated with the natural-forest area among the sites. These results suggest that environments and resources of trees influence their fruit production, that a loss of natural forests increases in embryo mortality, and that a reduction in tree density decreases pollen donor diversity in P. verecunda. Thus, a landscape disturbance may decrease seed production, whereas outbreeding is maintained, and fruit production is not likely to be dependent on the landscape disturbance in this species.
Similar content being viewed by others
Introduction
Landscape disturbances, including loss and fragmentation of natural habitats, is likely to affect animal-mediated pollination, which may lead to changes in fruit and seed production of flowering plants (Aizen and Feinsinger 1994; Ghazoul 2005; Hadley and Betts 2012). A meta-analysis of studies on animal-pollinated plants demonstrated negative effects of landscape disturbances on fruit and seed production (Aguilar et al. 2006). Landscape disturbances also negatively affect pollination success, as measured by the frequency of pollinator visits to a flower, the amount of pollen loads on a stigma, and the number of pollen tubes in a style, suggesting that pollen limitation underlies the reduced production of fruits and seeds (Aguilar et al. 2006). The negative effects on fruit and seed production are larger in self-incompatible plants than in self-compatible plants, because the limited availability of outcrossed pollen has more direct effects on fruit and seed production in self-incompatible plants (Aguilar et al. 2006). In self-compatible plants, landscape disturbances decrease the outcrossing rate (Aguilar et al. 2008; Eckert et al. 2010). Therefore, loss and fragmentation of habitats tend to reduce fruit and seed production while shifting mating patterns toward inbreeding.
Mating patterns and pollen dispersal are also related to landscape properties (Sork and Smouse 2006), which are described by the composition and configuration of habitats (patches) and non-habitats (the matrix) of plants and pollinators (Hadley and Betts 2012). Although the matrix is expected to impede pollen flow, substantial pollen immigration has been observed in isolated patches, particularly in those with reduced plant density (Sork and Smouse 2006; Kramer et al. 2008). Pollen dispersal distance sometimes increases in patches with reduced plant density (Llorens et al. 2012; Côrtes et al. 2013) and between fragmented patches because of changes in pollinator movements in the matrix (Kamm et al. 2010). In constrast, landscape disturbances decrease mate diversity (Breed et al. 2013). Not only the size but also the shape of patches affects mating patterns (Llorens et al. 2013). Overall, these findings suggest that loss and fragmentation of habitats either increase or decrease mate diversity because of various consequences of the landscape disturbances on pollen dispersal.
Anthropogenic landscape disturbances typically occur with the conversion of intact habitats, such as natural forests, grasslands, and wetlands, to lands used for human activities, such as plantations, farmlands, and residential as well as industrial areas. The effects of converting natural habitats to farmlands in agricultural landscapes on pollination have been studied frequently, because of the importance of pollination services from those natural habitats, forests in particular, for farmland crop plants (Klein et al. 2007; Taki et al. 2007). However, the effects of converting natural forests to artificial plantations in silvicultural landscapes on pollination of forest trees have rarely been investigated, despite the fact that monocultured plantations of wind-pollinated conifers have lower species richness and population density of pollinators than natural forests (Taki et al. 2013).
Loss and fragmentation of habitats often occur simultaneously after a landscape disturbance event and tend to occur interactively (Smith et al. 2009). Furthermore, the landscape disturbance usually leads to reduced plant density in the disturbed habitats (Eckert et al. 2010). Therefore, the landscape disturbance complicates loss and fragmentation of habitats as well as reduced plant density. However, only a few studies have tried to disentangle these confounding factors (Hadley and Betts 2012).
Fruit and seed production as well as mate diversity (reproductive performance) of flowering trees have been compared between an intact habitat, such as a large continuous forest, and a disturbed habitat, such as a small fragmented forest. These comparisons often lack replication; thus, the findings cannot be generalized. However, independent comparisons between continuous and fragmented forests in multiple regions have revealed a consistent effect of fragmentation on reproductive performance in these regions (Aizen and Feinsinger 1994; Mimura et al. 2009). Patterns observed at multiple sites with different landscape properties within a region also demonstrate the effects of landscape properties on reproductive performance, which can be estimated using statistical models (Llorens et al. 2012; Breed et al. 2013). The spatial independence of sites and the separation of factors confounded with the sites are crucial when fitting the models to observed patterns.
We compared landscape properties and reproductive performance among multiple sites. Using this approach, we investigated the wild cherry species, Prunus verecunda (Koidz.) Koehne, because it occurs in various forest patches with different landscape properties and depends on generalist pollinators in natural forest habitats. This species commonly occurs in broadleaf forests in the semi-mountainous ranges of central Japan. Most Prunus species have gametophytic self-incompatibility and predominantly reproduce by outcrossing (Ushijima et al. 2004; Shuri et al. 2012). Various animals, birds, bees, flies, and beetles, are the main pollinators of P. verecunda and are more abundant in natural forests than in coniferous plantations in central Japan (Taki et al. 2013).
The aim of this study was to examine the effects of loss and fragmentation of natural forests on fruit and seed production and mate diversity of P. verecunda. We predicted that these factors would have negative effects on the reproductive performance of P. verecunda. To verify this prediction, we observed the reproductive performance during two years at 12 sites in a forestry region, where coniferous plantations have fragmented the natural broadleaf forests, and used statistical models to detect potential effects of landscape properties on the reproductive performance of P. verecunda.
Materials and methods
Study sites and trees
We located 12 sites (site j from 1 to 12) in a 14.9 × 7.8 km area (36.800–36.938°N; 140.570–140.657°E) in a forestry region in central Japan (Fig. 1). We measured elevation (m) at each site. We categorized the vegetation into natural broadleaf forests and others, mainly including coniferous (Cryptomeria japonica and Chamaecyparis obtusa) plantations (Fig. 1). The distance between the sites ranged from 1.2 to 15.9 km, which was beyond the radius of the foraging ranges of most pollinators. The pollinators of P. verecunda consist of Japanese white-eyes (Zosteros japonicus), bees dominated by honeybee (Apis cerana) workers and bumblebee (Bombus) queens, flies dominated by syrphid flies, and beetles dominated by cerambycid beetles. The maximum daily movements of the Japanese white-eyes ranged from 0.2 to 1.1 km (Aslan et al. 2014). The foraging radii of honeybee workers from their nests ranged from 1.0 to 5.5 km (Beekman and Ratnieks 2000). Bumblebee queens flew up to 1.3 km within a few days (Hagen et al. 2011) and dispersed by at least 3 km (Lepais et al. 2010). The flies and beetles were likely to have much smaller flight ranges than the birds and bees. Therefore, the sites seemed to be spatially independent because of little overlap between the pollinator communities.
Location of sites and landscape patterns of natural forests (dark gray), coniferous plantations (pale gray), and other land (white) in a forestry region in central Japan. Numbers indicate sites. Thick lines indicate observation trails at the sites. Circles indicate 500-ha buffer areas around the sites. An inset map shows the location of the study area within Japan
We established a trail (152–1061 m long) at each site as a survey route for walking through the natural forests. We recorded the locations of all (12–44) flowering P. verecunda trees within a 20 m width along each site of the trail at each site during 2012 and 2013. We calculated the mean distance (d j m) between neighboring flowering trees along the trail at site j to evaluate the local density of flowering trees at each site. We selected a mean of 12.2 (8–15) trees in 2012 and 11.2 (8–13) trees in 2013 as maternal trees from the flowering trees at each site. We observed 124 maternal trees in both years among 146 and 134 maternal trees selected in 2012 and 2013, respectively. We measured the diameters of all stems of each maternal tree at breast height (1.3 m). Based on the diameters, we calculated the basal stem area (h i m2) of the ith maternal tree. We calculated the distances to the first nearest (df i m) and 10th nearest (dt i m) flowering trees from maternal tree i along the trail to evaluate the spatial distribution of flowering trees around each maternal tree. Because the effective number of pollen donors for each maternal tree was approximately 10 (see Results), dt i represented the radius of the local mating range of maternal tree i.
We calculated the area (f j m2) and edge length (g j m) of natural forest patches at site j in six buffers around the midpoint of the trail with radii of 126, 178, 399, 564, 798, and 1262 m, corresponding to buffer areas of 5, 10, 50, 100, 200, and 500 ha, respectively, using the vegetation map and ArcGIS (ESRI Japan, Tokyo). These spatial scales corresponded to different foraging ranges of the various P. verecunda pollinators. We calculated the relative area (a jk = f jk /b k ) and relative edge length (e jk = 0.5g jk p−1/2 f −1/2 jk ) of natural forests at site j, where the buffer area was b k (m2) at spatial scale k. The relative edge length was 0, where there was no forest edge within a buffer, because the size of forest patch exceeded the buffer width.
Measurements
The fruits of P. verecunda mature and are dispersed in July in central Japan. Thus, we collected branches with immature fruits from maternal trees at each site in June 2012 and 2013, before the fruits were dispersed. We sampled one leaf from each of five shoots on the branches collected from each maternal tree. First, we measured the chlorophyll density of each of the five leaves from the shoots using a SPAD-502 Plus portable chlorophyll meter (Konica Minolta, Tokyo, Japan) and averaged them (c i ) for maternal tree i. Second, we measured the number of fruits per inflorescence on sampled branches as an index of fruit production. Fallen inflorescences leave vestiges on 1-year-old shoots in P. verecunda. Thus, we estimated the number of inflorescences (l i ) by adding the number of vestiges of fallen inflorescences and infructescences with immature fruits on the branches of maternal tree i. We counted the number of immature fruits (u i ) on the branches and calculated the number of fruits per inflorescence (m i = u i /l i ) for maternal tree i. Third, we measured wet weight (mg) of five immature fruits randomly sampled from the branches and averaged them (w i mg) for maternal tree i.
A single seed matures in each fruit, and a single embryo develops in each seed in P. verecunda. We cut the immature fruits and examined the embryos in their seeds. We categorized the seeds as those with viable or dead embryos. The viable embryos had a pair of white globular cotyledons and a hypocotyl. We classified seeds with the hollow or jelly-like material inside and seeds with withered black embryos as seeds with dead embryos. We counted the number of examined seeds (x i ) until the number of sound seeds with viable embryos (v i ) was ≥16 to obtain the sound-seed rate (s i = v i /x i ) for maternal tree i. We stored approximately 16 viable embryos and a leaf of each maternal tree at −20 °C for later DNA extraction.
Genotyping and genetic analysis
We extracted DNA from the leaves of 146 and 134 maternal trees in 2012 and 2013, respectively, using a DNeasy Plant Mini Kit (Qiagen, Hilden, Germany) and their 2,237 and 2,037 viable embryos using PrepMan Ultra reagent (Life Technologies, Foster City, CA, USA).
We tested 12 primer pairs of BPPCT001, 002, 005, 012, 014, 026, 028, 034, 037, 038, 040, and 041 (Dirlewanger et al. 2000); five of UDP96-001, 96-008, 97-401, 97-402, and 97-403 (Cipriani et al. 1999); and six of UDP98-025, 98-405, 98-408, 98-410, 98-412, and 98-416 (Testolin et al. 2000). Among them, primer pairs for eight loci successfully amplified the polymorphic DNA fragments in P. verecunda. Among the eight loci, BPPCT034 is on the linkage group two (LG2), UDP97-403 is on LG3, BPPCT005 and BPPCT040 on LG4, BPPCT014 and BPPCT037 are on LG5, and UDP96-001 and UDP98-412 are on LG6 (Dirlewanger et al. 2004; Clarke et al. 2009). We conducted polymerase chain reactions (PCRs) using a forward primer labeled with a fluorescent dye (FAM, HEX, or NED) and a reverse primer assigned to each locus. We used two multiplex sets of four loci, set 1: BPPCT034 (FAM), UDP96-001 (FAM), BPPCT005 (HEX), and UDP98-412 (NED), and set 2: BPPCT014 (FAM), BPPCT037 (FAM), UDP97-403 (HEX), and BPPCT040 (NED). The two primer pairs labeled with FAM in each set produced PCR products with different ranges in size. We performed multiplex PCRs in a 12 µL mixture containing 6 µL Multiplex PCR Master Mix (Qiagen), 0.2 µM of each primer, and 1 µL of template DNA solution using a GeneAmp 9700 thermal cycler (Life Technologies). The thermal cycler was programmed for 15 min at 95 °C, followed by 32 cycles, consisting of 30 s at 94 °C, 90 s at 60 °C, and 60 s at 72 °C, and finished by 30 min at 60 °C. The sizes of the PCR products were measured using an ABI PRISM 3100-Avant Genetic Analyzer with POP-7 in 36-cm capillaries and GeneScan analysis software (Life Technologies).
We determined genotypes of the maternal trees and their embryos. We estimated null alleles in putative homozygous genotypes from genotype mismatches between the maternal trees and their embryos. We described genetic parameters in the maternal tree population in 2012. First, we examined the deviation from Hardy–Weinberg equilibrium at each locus and linkage disequilibrium among the eight loci using GenePop 4.2 (Rousset 2008). Second, we estimated the Fisher’s F statistics, F IS, F ST, and pairwise F ST, in the maternal tree population consisting of 12 subpopulations corresponding to the sites using GenePop 4.2. Third, we tested isolation by distance among the 12 subpopulations with the Mantel test based on the genetic distance (F ST/[1 − F ST]) and the log-transformed spatial distance using GenePop 4.2.
We estimated mating pattern indices for each maternal tree. First, we estimated the multilocus outcrossing rate (tm i ), which is the proportion of outcrossed embryos, for maternal tree i using MLTR 3.4 (Ritland 2002). Second, we estimated the biparental-inbreeding rate (p i = tm i − ts i ), which is a measure of parent relatedness, for maternal tree i, where ts i was the outcrossing rate estimated from individual loci using MLTR 3.4. Third, we estimated the multilocus correlated paternity (r i ) within maternal tree i, which is the probability that two embryos of maternal tree i share the same father, using KINDIST in POLDISP 1.0c (Robledo-Arnuncio et al. 2007). Similarly, we estimated r ii’ between maternal trees i and i’, which is the probability that two embryos of different maternal trees i and i’ share the same father.
Statistical analysis
We predicted each of the five response variables in maternal tree i: the number of fruits per inflorescence (m i ), fruit weight (w i mg), sound-seed rate (s i ), biparental-inbreeding rate (p i ), and correlated paternity (r i ), from a linear model. The model included the nine explanatory variables of maternal tree i: basal stem area (h i m2), leaf chlorophyll density (c i SPAD value), and distances to the first nearest (df i m) and 10th nearest (dt i m) flowering trees, as well as m i , w i , empty-seed rate (1 − s i ), selfing rate (1 − tm i ), p i , and r i , except for the predicted variable itself. We included the latter variables in the models, because fruit and seed production as well as mating patterns could affect themselves and each other. The explanatory variables were log10-transformed, except for c i (SPAD value) and w i (mg), to reduce their skewness. We replaced zero and negative values with 0.001 to prevent the transformed values from being infinite. We fitted the data of maternal tree i in each year to the model using glm [family = Gamma (link = “log”)] for m i , w i , p i , and r i and glmmML [family = binomial (link = “logit”), cluster = “maternal tree”] for s i in R 3.0.2 (R Core Team 2013). We reduced the influence of overdispersion on the model fitting using a gamma distribution with shape and rate parameters as well as a binomial distribution with random effects of maternal trees. All models had fixed effects of the explanatory variables and 11 fixed effects of the sites (site 1 as an intercept). Thus, we examined the variations in the response variables among maternal trees within the sites using these so-called tree models.
We calculated the mean values of the five response variables at site j from those in maternal tree i as follows: m j = (Σ i∈j m i )/n j , w j = (Σ i∈j w i )/n j , s j = (Σ i∈j v i )/(Σ i∈j x i ), p j = (Σ i∈j p i )/n j , and r j = (Σ i∈j r i )/n j , where n j is the number of maternal trees at site j. We also obtained the six explanatory variables at site j: elevation (m), relative area (a jk ) and relative edge length (e jk ) of natural forests at spatial scale k, mean basal stem area (h j = [Σi∈j h i ]/n j m2) and mean leaf chlorophyll density (c j = [Σi∈j c i ]/n j SPAD value) of maternal trees, and mean distance (d j m) between neighboring flowering trees. We performed redundancy analyses (RDA) using rda() in R 3.0.2 (Legendre and Legendre 2012) to summarize the variations in the explanatory and response variables among the sites. We examined the variables for each year (2012 or 2013) and the relative area and edge length of natural forests at an intermediate spatial scale (100-ha buffer area). We plotted the sites and drew vectors for the explanatory and response variables on the coordinates of the first and second axes of the RDA plot.
We examined the variations in the response variables among the sites using so-called site models. The site model for each response variable included the six explanatory variables. We could not add reproductive performance variables to the explanatory variables, because there were too many explanatory variables to fit the data based on the small sample size (12 sites). Among the explanatory variables, h j (m2) and d j (m) were log10-transformed. We fitted the data of site j at spatial scale k for each year to the site model using glm [family = Gamma (link = “log”)] for m j , w j , p j , and r j and glmmML [family = binomial (link = “logit”), cluster = “site”] for s j in R 3.0.2.
After fitting the data to the site models, we selected the most predictable spatial scale for each response variable in each year. We compared the lowest Akaike’s Information Criterion (AIC) values of the site models at spatial scale k and selected the k value with the lowest AIC value among the six spatial scales. We selected a site model with the lowest AIC value at the most predictable spatial scale. We also selected a tree model with the lowest AIC value for each response variable in each year. In these selected models, we estimated the parameter of every fixed effect and calculated its effect size from the parameter estimate divided by the estimated standard error of each parameter. We examined the statistical significance (P < 0.05) obtained from the Wald test using summary() of the output of glm() or glmmML() in R 3.0.2. We obtained predictions from the explanatory variables with significant effects that were consistent between the years.
Results
Genetic structure
The genotypes of the maternal trees and their embryos are recorded in Online Resources 1a, b. No missing genotypes were detected in the maternal trees, and the proportions of missing genotypes ranged from 0.54 to 14.02 % among the loci and 4.00 % on an average across the loci in the embryos.
Null alleles were found at five loci, BPPCT005, BPPCT014, BPPCT034, BPPCT040, and UDP97-403 in 2012 and 2013. The proportions of null alleles were 0.54 and 0.28 % at BPPCT005, 0.20 and 0.26 % at BPPCT014, 0.20 and 0.09 % at BPPCT034, 3.75 and 3.21 % at BPPCT040, and 1.79 and 1.64 % at UDP97-403 in the maternal trees and their embryos, respectively. The genetic parameters were estimated in a population of 146 maternal trees observed in 2012. No significant deviation from Hardy–Weinberg equilibrium was found at any loci (P ≥ 0.386), except BPPCT040 (P < 0.001), and no significant linkage disequilibrium was detected among eight loci (P ≥ 0.282), except between BPPCT005 and BPPCT040 (P < 0.001). Consequently, BPPCT040 was removed from the following analyses.
At the seven selected loci, the fixation indices were low (F IS = 0.001, F ST = 0.019) in the maternal tree population that consisted of 12 subpopulations corresponding to the sites. The pairwise F ST values between the subpopulations ranged from −0.009 to 0.065 (Online Resource 1c). No significant correlation was found between the spatial and genetic distances (P = 0.972).
Site properties and reproductive performance
The elevations of the 12 sites varied from 361 to 823 m. The range of the relative area of natural forests among the sites decreased as the spatial scale increased. The relative edge length of natural forests increased as the spatial scale increased. The properties of tree populations varied among the sites. The mean basal stem area ranged from 0.045 to 0.162 m2, the mean leaf chlorophyll density ranged from 37.1 to 41.8 SPAD values, and the mean distance between neighboring flowering trees along the trail ranged from 10.8 to 53.1 m.
Fruit and seed production and mating patterns of maternal trees differed between 2012 and 2013 (Fig. S1). The maternal trees produced significantly fewer (0.483 per inflorescence) and heavier (165 mg) fruits in 2013 than those in 2012 (0.515 and 149, respectively; Wilcoxon test, P < 0.003; Fig. S1a, b). The sound-seed rate was relatively high, and the mean value (0.969) in 2013 was significantly higher than that (0.930) in 2012 (P < 0.001; Fig. S1c). The seeds were almost completely outbred, because the mean outcrossing rate (tm) was 0.990 in 2012 and 0.997 in 2013 (Fig. S1d). We excluded tm from the response variables, because tm had little variation. The parents were nearly unrelated, because the mean biparental-inbreeding rate was 0.024 in 2012 and 0.025 in 2013 (Fig. S1e). The mean correlated paternity (r) was 0.104 in 2012 and 0.076 in 2013 (Fig. S1f). The significantly lower r in 2013 than in 2012 (P = 0.015) indicated that the pollen donors were more diverse in 2013 than in 2012. About 0.1 r value would indicate approximately 10 pollen donors per maternal tree, if each donor contributed equally to seed production. Because the r value between maternal trees was not negatively correlated with the spatial distance between maternal trees at some sites, we were unable to estimate the pollen dispersal distance at every site.
Variations among maternal trees within sites
The selected tree models with the highest predictability for the number of fruits per inflorescence (m i ) in maternal tree i were similar between 2012 and 2013 (Table 1). In both years, m i increased significantly as the basal stem area and leaf chlorophyll density of the maternal trees increased (Fig. 2a, b). In both years, m i decreased as the empty-seed rate increased, and it had a significant effect on m i in 2012 (Fig. 2c). In 2013, m i significantly increased as the correlated paternity increased (Table 1). In both years, m i significantly varied among the sites (Table 1). The selected tree model for the fruit weight did not have any explanatory variables with significant effects in either year (Table 1).
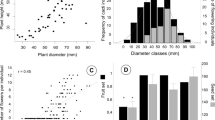
Relationships between explanatory and response variables among maternal trees in 2012 (circles) and 2013 (crosses). Number of fruits per inflorescence predicted from tree models with explanatory variables with significant effects (P < 0.05) that are consistent between years, a basal stem area, b leaf chlorophyll density, and c empty-seed rate of maternal trees, are shown by solid (2012) and broken (2013) lines. Leaf chlorophyll density was measured using a SPAD-502 Plus portable chlorophyll meter (Konica Minolta)
The selected tree model for the sound-seed rate (s i ) had no explanatory variable with a significant effect in either year, although s i significantly varied among the sites in 2013 (Table 1).
The selected tree model for the biparental-inbreeding rate (p i ) had two and no explanatory variables with significant effects in 2012 and 2013, respectively (Table 1). As the number of fruits per inflorescence and correlated paternity increased, p i increased significantly in 2012 (Table 1). In 2013, p i varied significantly among the sites (Table 1). Although the selected tree model for the correlated paternity (r i ) had no explanatory variable with a significant effect in 2013, the leaf chlorophyll density had a significantly positive effect on r i in 2012 (Table 1). In 2012, s i varied significantly among the sites (Table 1).
The data for fruit and seed production as well as mating patterns of the maternal trees and their properties are listed in Online Resource 1d.
Variations among sites
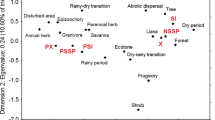
The variations in the explanatory and response variables among the sites were summarized with RDA. The first (53.0 % in 2012 and 42.9 % in 2013) and second (29.7 % in 2012 and 24.8 % in 2013) RDA axes explained most of the variations in both years (Fig. 3). Some patterns were consistent between the years as follows. (1) Sites 1, 2, and 3 were located in a negative region on the first and second axes, where the natural-forest area was large, and the biparental-inbreeding rate and correlated paternity were low. (2) Sites 10, 11, and 12 were plotted in a positive region on the first axis, where the elevation was low, and the number of fruits per inflorescence was small. (3) Site 7 was characterized by high values on the second axis, where the mean basal stem area of maternal trees was small, the distance between flowering trees was large, and the biparental-inbreeding rate or correlated paternity was high.
Variations in explanatory variables (gray arrows; elv: elevation, nfa: relative area of natural forests, nfe: relative edge length of natural forests, bsa: mean basal stem area of maternal trees, spd: mean leaf chlorophyll density of maternal trees, and dst: mean distance between flowering trees) and response variables (black arrows; FN: number of fruits per inflorescence, FW: fruit weight, SR: sound-seed rate, BI: biparental-inbreeding rate, and CP: correlated paternity) on coordinates of the first and second axes of redundancy analyses (RDA) in (a) 2012 and (b) 2013. Percentages in the axis labels are proportions of variations explained by two RDA axes
Spatial scales with the highest predictability (lowest AIC value) differed among the five response variables in the site models (Fig. 4). The AIC value for the number of fruits per inflorescence was independent of the spatial scales in 2012 (Fig. 4a) but was lower at larger spatial scales and lowest at the 200-ha scale in 2013 (Fig. 4f). The AIC value for the fruit weight was lowest at the 500-ha scale in both years (Fig. 4b, g), whereas the AIC value at the 5-ha scale was also low in 2013 (Fig. 4g). The AIC value for the sound-seed rate was lower in smaller spatial scales and lowest at the 5-ha scale in both years (Fig. 4c, h), although the AIC value at the 500-ha scale was also low in 2013 (Fig. 4h). The AIC value for the biparental-inbreeding rate was lower in larger spatial scales and lowest at the 500-ha scale in 2012 (Fig. 4d) and at the 200-ha in 2013 (Fig. 4i). The AIC value for the correlated paternity was lowest at the 500-ha scale in both years (Fig. 4e, j).
Scale dependency for predictability of site models with the five lowest Akaike’s Information Criterion (AIC) values in 2012 (upper row) and 2013 (lower row). Spatial scales represent buffer areas (ha). Site models for a, f the number of fruits per inflorescence, b, g fruit weight, c, h sound-seed rate, d, i biparental-inbreeding rate, and e, j correlated paternity are shown
The selected site models with the highest predictability for the number of fruits per inflorescence (m j ) at site j had significantly positive effects of the elevation on m j in both years (Table 2; Fig. 5a). In 2013, the relative edge length of natural forests at the 200-ha scale and the mean leaf chlorophyll density of the maternal trees had significantly negative effects on m j (Table 2). The selected site models for the fruit weight (w j mg) differed between 2012 and 2013 (Table 2). The relative area of natural forests at the 500-ha scale had a significantly positive effect on w j in 2012 but had a significantly negative effect on w j in 2013 (Table 2). In 2013, the elevation had a significantly positive effect on w j , and the mean basal stem area of the maternal trees had a significantly negative effect on w j (Table 2). Thus, no explanatory variables had a consistent trend in significant effects on w j between 2012 and 2013 (Table 2).
Relationships between explanatory and response variables among sites in 2012 (circles) and 2013 (crosses). a Number of fruits per inflorescence, b sound-seed rate, and c correlated paternity predicted from tree models with explanatory variables with significant effects that are consistent between years, elevation, relative area of natural forests, and mean distance between flowering trees, respectively, are shown as solid (2012) and broken (2013) lines
The selected site models for the sound-seed rate (s j ) had significantly positive effects of the relative area of natural forests at the 5-ha scale on s j in both years (Table 2, Fig. 5b). The mean distance between flowering trees had a significantly positive effect on s j in 2012, and the mean basal stem area and mean leaf chlorophyll density of the maternal trees had significantly negative effects on s j in 2013 (Table 2).
The selected site models for the biparental-inbreeding rate (p j ) had significantly negative effects of the relative area of natural forests at the 500-ha scale and the mean basal stem area of the maternal trees on p j in 2012 (Table 2). The selected site models for the correlated paternity (r j ) had positive effects of the mean distance between flowering trees on r j in both years (Table 2, Fig. 5c). The relative area of natural forests at the 500-ha scale had negative but non-significant effects on r j in both years (Table 2). In 2012, the mean basal stem area of the maternal trees had a significantly negative effect on r j (Table 2).
The data for fruit and seed production as well as mating patterns at the sites and their properties are listed in Online Resource 1e.
Discussion
General patterns
The reproductive performance in P. verecunda depended on various factors, such as the properties of forest landscapes, tree populations, and individual trees. Thus, the processes that determine the reproductive performance seem to be complex. Therefore, we have to interpret the selected models and the estimated effects carefully.
Pollen dispersal distance indicates the spatial scale, at which the properties of landscapes and populations affect the reproductive performance (Sork and Smouse 2006; Ashley 2010). However, we were unable to estimate the pollen dispersal distance in P. verecunda from the correlated paternity between the maternal trees. Pollen dispersal and mating patterns have been intensively studied in a congeneric species, P. avium L. The mean distance of pollen dispersal estimated in P. avium ranged from 20 to 145 m, although these values were underestimated, because pollen immigration from outside the study sites was ignored (Cottrell et al. 2009; Jolivet et al. 2011). These estimates of the pollen dispersal distance are shorter than the length (152–1061 m) of our observation trail at each site. The mean correlated paternity observed in P. avium ranged from 0.089 to 0.142 (Cottrell et al. 2009; Jolivet et al. 2011), which is similar to those (0.076 ≤ r ≤ 0.104) observed in P. verecunda. These values indicate that the effective number of fathers is approximately 10, which is smaller than the number (12–44) of flowering trees observed at each site. Therefore, we seemed to have surveyed available mates and mating neighborhoods at each site.
The reproductive performance at the sites depended on the landscape properties at various spatial scales. At the maximum scale (1.26-km radius corresponding to 500-ha buffer area), the buffers slightly overlapped among some sites. Some P. verecunda pollinators, such as honeybee workers and bumblebee queens, have the foraging ranges of several kilometers (Beekman and Ratnieks 2000; Lepais et al. 2010; Hagen et al. 2011). Thus, the pollinators potentially overlapped among the sites. Therefore, some sites may not be spatially independent in our statistical models. Thus, we are cautious when interpreting the effects of the landscape properties on the reproductive performance in the models.
Some reproductive performance variables differed between 2012 and 2013. Flower abundance within trees, flowering-tree density, pollinator availability, and environmental conditions during flowering and fruiting in P. verecunda are likely to vary between years. The higher sound-seed rate and lower correlated paternity in 2013 than in 2012 suggest that resource availability, pollination service, and/or mate diversity were higher in 2013 than in 2012. The fewer but heavier fruits in 2013 than in 2012 suggest a trade-off between the number and size of fruits due to limited resources for fruit production. Such fluctuations between years may have influenced the effects of the landscape properties on the reproductive performance. We attempt to identify effects that are robust against such annual fluctuations. Thus, we regard the explanatory variables with significant effects that are consistent between years as the robust effects in our models.
Fruit production
The basal stem area and leaf chlorophyll density in the maternal trees had positive effects on the number of fruits per inflorescence in both 2012 and 2013, suggesting that fruit production depends on the availability of resources. A larger basal stem area indicates a larger canopy mass and better light conditions. A larger tree size and higher chlorophyll content allow trees to achieve more photosynthesis and larger resource allocation to fruit production. These findings suggest that resource availability but not pollen availability would limit fruit production. Landscape disturbances are expected to reduce fruit production because of the limited availability of outcrossed pollen in self-incompatible species, such as P. verecunda (Aizen and Feinsinger 1994; Aguilar et al. 2006). The pollen availability limited the fruit-set rate in P. virginiana L. in fragmented forests surrounded by farmlands more than in continuous forests (Suarez-Gonzalez and Good 2014). However, forest vegetation was maintained in our study area, because coniferous plantations surrounded natural forests. The richness and abundance of bee communities in coniferous plantations at young forest ages were similar to those of natural forests (Taki et al. 2013). Thus, the forest vegetation in our study area may harbor pollinator communities to a certain extent, preventing pollen limitation in fruit production.
The number of fruits per inflorescence in the maternal trees increased as the empty-seed rate decreased. This result is reasonable because embryo mortality may cause fruit abortion, resulting in reduced fruit production.
The number of fruits per inflorescence increased as the elevation increased among the sites in both 2012 and 2013, suggesting that certain environmental conditions vary at different elevations and affect fruit production. In our study area, P. verecunda is more abundant at higher elevations (>500 m), and another cherry species, P. jamasakura Sieb. ex Koidz., is more abundant at lower elevations (<500 m), suggesting that higher elevations are more suitable for P. verecunda in our study area. This habitat suitability may result in higher fruit production at the higher elevations, although the environmental conditions responsible for the higher fruit production remain unclear.
Our results suggest that differences in resource availability among trees and environmental variability at different elevations mainly cause the variations in fruit production. In contrast to our expectations from previous studies (Aguilar et al. 2006), pollen limitation and landscape disturbances may not affect fruit production in P. verecunda in natural forests that have been fragmented by coniferous plantations
Seed production and mating patterns
The sound-seed rate increased among the sites as the natural-forest area increased in both 2012 and 2013. Loss of forest habitats has frequently been associated with decreased plant density and increased selfing rate, biparental-inbreeding rate, and correlated paternity in animal-pollinated plants (Breed et al. 2013). The forest loss resulted in increases in both biparental-inbreeding rate and correlated paternity in P. virginiana, because relatives were more aggregated spatially in fragmented forests than in a continuous forest (Suarez-Gonzalez and Good 2014). These findings are consistent with our results indicating that the biparental-inbreeding rate and correlated paternity tended to increase as the natural-forest area decreased. The loss of natural forests may limit mating availability as a result of reduced tree populations, infrequent pollinator movements, and poor pollinator communities (Ghazoul 2005; Winfree et al. 2009). Mate availability restricted to the neighborhood, where relatives are frequent, seems to increase both biparental-inbreeding rate and correlated paternity.
Biparental inbreeding may cause embryo mortality as a result of inbreeding depression. The magnitude of inbreeding depression is relatively high in outcrossing woody perennials (Duminil et al. 2009). Most Prunus species predominantly outcross because of their gametophytic self-incompatibility that prevents self-fertilization (Ushijima et al. 2004; Shuri et al. 2012). The observed outcrossing rate (0.994 ≤ tm i ≤ 0.998) confirms the self-incompatibility in P. verecunda. The nearly zero fixation index (F IS = 0.001) observed in the maternal trees indicates the exclusion of inbred progeny until maturity, suggesting a high magnitude of inbreeding depression. The fruit-set rate in P. virginiana was lower in fragmented forests, where the biparental-inbreeding rate was higher, than in a continuous forest (Suarez-Gonzalez and Good 2014).
The most predictable scale for the effect of the natural-forest area on the sound-seed rate was the 5-ha buffer area with the 126-m radius. Relatives were aggregated at such a small scale (<60 m radius) in P. virginiana (Suarez-Gonzalez and Good 2014). Thus, biparental inbreeding that occurs between the relatives at this scale may result in embryo mortality due to inbreeding depression. On the other hand, the most predictable scale for the effect of the natural-forest area on the biparental-inbreeding rate and correlated paternity was the 500-ha buffer area with the 1.26-km radius. Thus, long-distance pollen dispersal occurring such a large scale may affect the overall mating patterns (Kramer et al. 2008; Ashley 2010). This spatial scale corresponds to the foraging ranges of some P. verecunda pollinators, such as birds and bees (Beekman and Ratnieks 2000; Lepais et al. 2010; Hagen et al. 2011; Aslan et al. 2014). Thus, the abundance and activity of pollinators may be a primary factor that is responsible for reduced mate diversity resulted from the loss of natural forests (Klein et al. 2007; Winfree et al. 2009).
The correlated paternity increased among the sites as the mean distance between flowering trees increased in both 2012 and 2013. This result indicates that flowering-tree density, an inverse of the distance between the trees, affects mate diversity (Eckert et al. 2010; Breed et al. 2013). The flowering-tree density seems to be positively related to the number of mates within a mating range. A positive effect of tree density on mate diversity was also detected in insect-pollinated Eucalyptus species, although this effect was confounded with the effect of forest loss (Breed et al. 2013). Our findings in P. verecunda demonstrate that the flowering-tree density affects mate diversity independently of landscape properties.
Conclusions
Our comparison among multiple sites in a forestry region revealed relationships between forest landscapes and reproductive performance in P. verecunda. Our findings suggest that habitat loss reduces seed production in natural forests that have been fragmented by coniferous plantations as well as by farmlands (Klein et al. 2007; Taki et al. 2007). Our analyses discriminated among various effects of landscape, population, and tree properties on reproductive performance. The effects of landscape properties were likely to vary among spatial scales. Because our sampling sites may not have been completely statistically independent, particularly at the larger spatial scales, the effects of the natural-forest area on mating patterns at the 500-ha scale should be confirmed by further evidence. Our observational study implies that complicated factors are involved in fluctuating reproductive performance between years. Experimental studies can effectively verify the effects of each potential factor that is responsible for reproductive performance.
Our findings support three conclusions. First, our results demonstrate that landscape properties affect seed production in P. verecunda. The loss of natural forests may increase embryo mortality. On the other hand, resource and environmental factors seem to limit fruit production. Second, population properties may also affect mating patterns in P. verecunda. The local density of flowering trees may have an effect on mate availability. Third, our study reveals the robustness of fruit production and outbreeding despite a landscape disturbance. Even if changes in mating patterns result in reduced viability of offspring, P. verecunda trees seem to stably produce outbred progeny in fragmented natural forests in forestry regions.
References
Aguilar R, Ashworth L, Galetto L, Aizen MA (2006) Plant reproductive susceptibility to habitat fragmentation: review and synthesis through a meta-analysis. Ecol Lett 9:968–980. doi:10.1111/j.1461-0248.2006.00927.x
Aguilar R, Quesada M, Ashworth L, Herrerias-Diego Y, Lobo J (2008) Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Mol Ecol 17:5177–5188. doi:10.1111/j.1365-294X.2008.03971.x
Aizen MA, Feinsinger P (1994) Forest fragmentation, pollination, and plant reproduction in a Chaco dry forest, Argentina. Ecology 75:330–351. doi:10.2307/1939538
Ashley MA (2010) Plant parentage, pollination, and dispersal: how DNA microsatellites have altered the landscape. CRC Crit Rev Plant Sci 29:148–161. doi:10.1080/07352689.2010.481167
Aslan A, Hart P, Wu J, Aslan CE (2014) Evaluating the qualitative effectiveness of a novel pollinator: a case study of two endemic Hawaiian plants. Biotropica 46:732–739. doi:10.1111/btp.12159
Beekman M, Ratnieks F (2000) Long-range foraging by the honey-bee, Apis mellifera L. Funct Ecol 14:490–496
Breed MF, Ottewell KM, Gardner MG, Marklund MHK, Dormontt EE, Lowe AJ (2013) Mating patterns and pollinator mobility are critical traits in forest fragmentation genetics. Heredity. doi:10.1038/hdy.2013.48
Cipriani G, Lot G, Huang W-G, Marrazzo MT, Peterlunger E, Testolin R (1999) AC/GT and AG/CT microsatellite repeats in peach [Prunus persica (L) Batsch]: isolation, characterisation and cross-species amplification in Prunus. Theor Appl Genet 99:65–72. doi:10.1007/s001220051209
Clarke JB, Sargent DJ, Bošković RI, Belaj A, Tobutt KR (2009) A cherry map from the inter-specific cross Prunus avium ‘Napoleon’ × P. nipponica based on microsatellite, gene-specific and isoenzyme markers. Tree Genet Genom 5:41–51
Côrtes MC, Uriarte M, Lemes MR, Gribel R, Kress JW, Smouse PE, Bruna EM (2013) Low plant density enhances gene dispersal in the Amazonian understory herb Heliconia acuminata. Mol Ecol 22:5716–5729. doi:10.1111/mec.12495
Cottrell JE, Vaughan SP, Connolly T, Sing L, Moodley DJ, Russell K (2009) Contemporary pollen flow, characterization of the maternal ecological neighbourhood and mating patterns in wild cherry (Prunus avium L.). Heredity 103:118–128. doi:10.1038/hdy.2009.39
Dirlewanger E, Cosson P, Tavaud M, Aranzana J, Poizat C, Zanetto A, Arús P, Laigret F (2000) Development of microsatellite markers in peach [Prunus persica (L.) Batsch] and their use in genetic diversity analysis in peach and sweet cherry (Prunus avium L.). Theor Appl Genet 105:127–138. doi:10.1007/s00122-002-0867-7
Dirlewanger E, Graziano E, Joobeur T, Garriga-Calderé F, Cosson P, Howad W, Arús P (2004) Comparative mapping and marker-assisted selection in Rosaceae fruit crops. PNAS 101:9891–9896
Duminil J, Hardy OJ, Petit RJ (2009) Plant traits correlated with generation time directly affect inbreeding depression and mating system and indirectly genetic structure. BMC Evol Biol 9:177. doi:10.1186/1471-2148-9-177
Eckert CG, Kalisz S, Geber MA, Sargent R, Elle E, Cheptou P-O, Goodwillie C, Johnston MO, Kelly JK, Moeller DA, Porcher E, Ree RH, Vallejo-Marín M, Winn AA (2010) Plant mating systems in a changing world. Trends Ecol Evol 25:35–43. doi:10.1016/j.tree.2009.06.013
Ghazoul J (2005) Pollen and seed dispersal among dispersed plants. Biol Rev 80:413–443
Hadley AS, Betts MG (2012) The effects of landscape fragmentation on pollination dynamics: absence of evidence not evidence of absence. Biol Rev 87:526–544. doi:10.1111/j.1469-185X.2011.00205.x
Hagen M, Wikelski M, Kissling WD (2011) Space use of bumblebees (Bombus spp.) revealed by radio-tracking. PLoS ONE 6:e19997. doi:10.1371/journal.pone.0019997
Jolivet C, Höltken AM, Liesebach H, Steiner W, Degen B (2011) Mating patterns and pollen dispersal in four contrasting wild cherry populations (Prunus avium L.). Eur J For Res 131:1055–1069. doi:10.1007/s10342-011-0576-3
Kamm U, Gugerli F, Rotach P, Edwards P, Holderegger R (2010) Open areas in a landscape enhance pollen-mediated gene flow of a tree species: evidence from northern Switzerland. Landsc Ecol 25:903–911. doi:10.1007/s10980-010-9468-z
Klein A-M, Vaissière BE, Cane JH, Steffan-Dewenter I, Cunningham SA, Kremen C, Tscharntke T (2007) Importance of pollinators in changing landscapes for world crops. Proc Biol Sci 274:303–313. doi:10.1098/rspb.2006.3721
Kramer AT, Ison JL, Ashley MV, Howe HF (2008) The paradox of forest fragmentation genetics. Conserv Biol 22:878–885. doi:10.1111/j.1523-1739.2008.00944.x
Legendre P, Legendre L (2012) Numerical ecology, 3rd edn. Elsevier, Amsterdam
Lepais O, Darvill B, O’Connor S, Osborne JL, Sanderson RA, Cussans J, Goffe L, Goulson D (2010) Estimation of bumblebee queen dispersal distances using sibship reconstruction method. Mol Ecol 19:819–831. doi:10.1111/j.1365-294X.2009.04500.x
Llorens TM, Byrne M, Yates CJ, Nistelberger HM, Coates DJ (2012) Evaluating the influence of different aspects of habitat fragmentation on mating patterns and pollen dispersal in the bird-pollinated Banksia sphaerocarpa var. caesia. Mol Ecol 21:314–328. doi:10.1111/j.1365-294X.2011.05396.x
Llorens TM, Yates CJ, Byrne M, Nistelberger HM, Williams MR, Coates DJ (2013) Complex interactions between remnant shape and the mating system strongly influence reproductive output and progeny performance in fragmented populations of a bird-pollinated shrub. Biol Conserv 164:129–139. doi:10.1016/j.biocon.2013.05.011
Mimura M, Barbour RC, Potts BM, Vaillancourt RE, Watanabe KN (2009) Comparison of contemporary mating patterns in continuous and fragmented Eucalyptus globulus native forests. Mol Ecol 18:4180–4192. doi:10.1111/j.1365-294X.2009.04350.x
R Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Ritland K (2002) Extensions of models for the estimation of mating systems using n independent loci. Heredity 88:221–228. doi:10.1038/sj/hdy/6800029
Robledo-Arnuncio JJ, Austerlitz F, Smouse PE (2007) POLDISP: a software package for indirect estimation of contemporary pollen dispersal. Mol Ecol Notes 7:763–766. doi:10.1111/j.1471-8286.2007.01706.x
Rousset F (2008) Genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8:103–106. doi:10.1111/j.1471-8286.2007.01931.x
Shuri K, Saika K, Junko K, Michiharu K, Nagamitsu T, Iwata H, Tsumura Y, Mukai Y (2012) Impact of negative frequency-dependent selection on mating pattern and genetic structure: a comparative analysis of the S-locus and nuclear SSR loci in Prunus lannesiana var. speciosa. Heredity 109:188–198. doi:10.1038/hdy.2012.29
Smith AC, Koper N, Francis CM, Fahrig L (2009) Confronting collinearity: comparing methods for disentangling the effects of habitat loss and fragmentation. Landsc Ecol 24:1271–1285. doi:10.1007/s10980-009-9383-3
Sork VL, Smouse PE (2006) Genetic analysis of landscape connectivity in tree populations. Landsc Ecol 21:821–836. doi:10.1007/s10980-005-5415-9
Suarez-Gonzalez A, Good SV (2014) Pollen limitation and reduced reproductive success are associated with local genetic effects in Prunus virginiana, a widely distributed self-incompatible shrub. Ann Bot 113:595–605. doi:10.1093/aob/mct289
Taki H, Kevan PG, Ascher JS (2007) Landscape effects of forest loss in a pollination system. Landsc Ecol 22:1575–1587. doi:10.1007/s10980-007-9153-z
Taki H, Okochi I, Okabe K, Inoue T, Goto H, Matsumura T, Makino S (2013) Succession influences wild bees in a temperate forest landscape: the value of early successional stages in naturally regenerated and planted forests. PLoS ONE 8:e56678. doi:10.1371/journal.pone.0056678
Testolin R, Marrazzo T, Cipriani G, Quarta R, Verde I, Dettori MT, Pancaldi M, Sansavini S (2000) Microsatellite DNA in peach (Prunus persica L. Batsch) and its use in fingerprinting and testing the genetic origin of cultivars. Genome 43:512–520
Ushijima K, Yamane H, Watari A, Kakehi E, Ikeda K, Hauck NR, Iezzoni AF, Tao R (2004) The S haplotype-specific F-box protein gene, SFB, is defective in self-compatible haplotypes of Prunus avium and P. mume. Plant J 39:148–161. doi:10.1111/j.1365-313X.2004.02154.x
Winfree R, Aguilar R, Vázquez D, LeBuhn G, Aizen MA (2009) A meta-analysis of bees’ responses to anthropogenic disturbance. Ecology 90:2068–2076
Acknowledgments
The Japanese Ministry of Education, Science, Sports, and Culture financially supported this study (Grant-in-Aid No. 24380085). We thank the staff of the Kanto District Forest Office for their permission to study the national forests, Yoko Osone, Takuya Ishigane, Kosuke Maeda, and Shotaro Mine for their assistance in field measurement, and Shoko Hisamatsu and Yasuko Kawamata for their help with data arrangement.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Nagamitsu, T., Shuri, K., Taki, H. et al. Effects of converting natural forests to coniferous plantations on fruit and seed production and mating patterns in wild cherry trees. Ecol Res 31, 239–250 (2016). https://doi.org/10.1007/s11284-015-1331-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11284-015-1331-x