Abstract
The aim of this study is to review the current state of and highlight the challenges in the production of microbial nitrilases as catalysts for the mild hydrolysis of industrially important nitriles. Together with aldoxime dehydratase, the nitrile-hydrolyzing enzymes (nitrilase, nitrile hydratase) are key enzymes in the aldoxime–nitrile pathway which is widely distributed in bacteria and fungi. The availability of nitrilases has grown significantly over the past decade due to the use of metagenomic and database-mining approaches. Databases contain plenty of putative enzymes of this type, whose overproduction may improve the spectrum and the industrial utility of nitrilases. By exploiting this resource, the number of experimentally verified nitrilases has recently increased to several hundred. We especially focus on the efficient heterologous expression systems that are applicable for the overproduction of wild-type nitrilases and their artificial variants. Biocatalyst forms with industrial potential are also highlighted. The potential industrial applications of nitrilases are classified according to their target products (α-hydroxy acids, α- and β-amino acids, cyano acids, amides). The emerging uses of nitrilases and their subtypes (cyanide hydratases, cyanide dihydratases) in bioremediation is also summarized. The integration of nitrilases with other enzymes into artificial multienzymatic and chemoenzymatic pathways is considered a promising strategy for future applications.
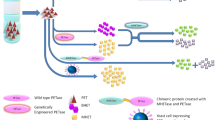
Graphical Abstract





Similar content being viewed by others
Abbreviations
- Approx.:
-
Approximately
- CHT:
-
Cyanide hydratase
- CLEAs:
-
Cross-linked enzyme aggregates
- CynD:
-
Cyanide dihydratase
- dcw:
-
Dry cell weight
- E.e.:
-
Enantiomeric excess
- IPTG:
-
Isopropyl-β-d-thiogalactopyranoside
References
Basile LJ, Willson RC, Sewell BT, Benedik MJ (2008) Genome mining of cyanide-degrading nitrilases from filamentous fungi. Appl Microbiol Biotechnol 80(3):427–435. doi:10.1007/s00253-008-1559-2
Baum S, van Rantwijk F, Stolz A (2012) Application of a recombinant Escherichia coli whole-cell catalyst expressing hydroxynitrile lyase and nitrilase activities in ionic liquids for the production of (S)-mandelic acid and (S)-mandeloamide. Adv Synt Catal 354(1):113–122. doi:10.1002/adsc.201100391
Bayer S, Birkemeyer C, Ballschmiter M (2011) A nitrilase from a metagenomic library acts regioselectively on aliphatic dinitriles. Appl Microbiol Biotechnol 89(1):91–98. doi:10.1007/s00253-010-2831-9
Bordier F, Stam M, Darii E, Tricot S, Fossey A, Rohault J, Debard A, Mariage A, Pellouin V, Petit JL, Perret A, Vallenet D, Salanoubat M, Weissenbach J, Vergne-Vaxelaire C, de Berardinis V, Zaparucha A (2014) Large alpha-aminonitrilase activity screening of nitrilase superfamily members: access to conversion and enantiospecificity by LC–MS. J Mol Catal B Enzym 107:79–88. doi:10.1016/j.molcatb.2014.05.019
Brady D, Beeton A, Zeevaart J, Kgaje C, van Rantwijk F, Sheldon RA (2004) Characterisation of nitrilase and nitrile hydratase biocatalytic systems. Appl Microbiol Biotechnol 64(1):76–85. doi:10.1007/s00253-003-1495-0
Cai WW, Su EZ, Zhu SJ, Ren YH, Wei DZ (2014) Characterization of a nitrilase from Arthrobacter aurescens CYC705 for synthesis of iminodiacetic acid. J Gen Appl Microbiol 60(6):207–214. doi:10.2323/jgam.60.207
Chen HY, Zhang TX, Sun TY, Ni Z, Le YL, Tian R, Chen Z, Zhang CX (2015) Clostridium thermocellum nitrilase expression and surface display on Bacillus subtilis spores. J Mol Microbiol Biotechnol 25(6):381–387. doi:10.1159/000441642
Chen HY, Chen Z, Ni Z, Tian R, Zhang TX, Jia JR, Chen KP, Yang SL (2016) Display of Thermotoga maritima MSB8 nitrilase on the spore surface of Bacillus subtilis using out coat protein CotG as the fusion partner. J Mol Catal B Enzym 123:73–80. doi:10.1016/j.molcatb.2015.11.002
Chmura A, Rustler S, Paravidino M, van Rantwijk F, Stolz A, Sheldon RA (2013) The combi-CLEA approach: enzymatic cascade synthesis of enantiomerically pure (S)-mandelic acid. Tetrahedron Asymmetry 24(19):1225–1232. doi:10.1016/j.tetasy.2013.08.013
Detzel C, Maas R, Jose J (2011) Autodisplay of nitrilase from Alcaligenes faecalis in E. coli yields a whole cell biocatalyst for the synthesis of enantiomerically pure (R)-mandelic acid. ChemCatChem 3(4):719–725. doi:10.1002/cctc.201000382
Gao BJ, Chen LL, Li YB (2016) Preparation of surface imprinted material of single enantiomer of mandelic acid with a new surface imprinting technique and study on its chiral recognition and resolution properties. J Chromatogr A 1443:10–20. doi:10.1016/j.chroma.2016.03.018
Gong JS, Li H, Zhu XY, Lu ZM, Wu Y, Shi JS, Xu ZH (2012a) Fungal His-tagged nitrilase from Gibberella intermedia: gene cloning, heterologous expression and biochemical properties. PLOS ONE 7(11):e50622. doi:10.1371/journal.pone.0050622
Gong JS, Lu ZM, Li H, Shi JS, Zhou ZM, Xu ZH (2012b) Nitrilases in nitrile biocatalysis: recent progress and forthcoming research. Microb Cell Fact 11:142. doi:10.1186/1475-2859-11-142
Gong JS, Lu ZM, Li H, Zhou ZM, Shi JS, Xu ZH (2013) Metagenomic technology and genome mining: emerging areas for exploring novel nitrilases. Appl Microbiol Biotechnol 97(15):6603–6611. doi:10.1007/s00253-013-4932-8
Han C, Yao PY, Yuan J, Duan YT, Feng JH, Wang M, Wu QQ, Zhu DM (2015) Nitrilase-catalyzed hydrolysis of 3-aminopropionitrile at high concentration with a tandem reaction strategy for shifting the reaction to β-alanine formation. J Mol Catal B Enzym 115:113–118. doi:10.1016/j.molcatb.2015.02.007
Hann EC, Sigmund AE, Hennessey SM, Gavagan JE, Short DR, Ben-Bassat A, Chauhan S, Fallon RD, Payne MS, DiCosimo R (2002) Optimization of an immobilized-cell biocatalyst for production of 4-cyanopentanoic acid. Org Proc Res Dev 6(4):492–496. doi:10.1021/op025515k
Kaplan O, Veselá AB, Petříčková A, Pasquarelli F, Pičmanová M, Rinágelová A, Bhalla TC, Pátek M, Martínková L (2013) A comparative study of nitrilases identified by genome mining. Mol Biotechnol 54(3):996–1003. doi:10.1007/s12033-013-9656-6
Kato Y, Ooi R, Asano Y (2000) Distribution of aldoxime dehydratase in microorganisms. Appl Environ Microbiol 66(6):2290–2296. doi:10.1128/AEM.66.6.2290-2296.2000
Kiziak C, Conradt D, Stolz A, Mattes R, Klein J (2005) Nitrilase from Pseudomonas fluorescens EBC191: cloning and heterologous expression of the gene and biochemical characterization of the recombinant enzyme. Microbiology 151:3639–3648. doi:10.1099/mic.0.28246-0
Li CY, Yue ZL, Feng FZ, Xi CW, Zang HL, An XJ, Liu KR (2016) A novel strategy for acetonitrile wastewater treatment by using a recombinant bacterium with biofilm-forming and nitrile degrading capability. Chemosphere 161:224–232. doi:10.1016/j.chemosphere.2016.07.019
Liu ZQ, Dong LZ, Cheng F, Xue YP, Wang YS, Ding JN, Zheng YG, Shen YC (2011) Gene cloning, expression, and characterization of a nitrilase from Alcaligenes faecalis ZJUTB10. J Agric Food Chem 59(21):11560–11570. doi:10.1021/jf202746a
Liu ZQ, Baker PJ, Cheng F, Xue YP, Zheng Y-G, Shen Y-C (2013) Screening and improving the recombinant nitrilases and application in biotransformation of iminodiacetonitrile to iminodiacetic acid. PLOS ONE 8(6):e67197. doi:10.1371/journal.pone.0067197
Liu ZQ, Zhang XH, Xue YP, Xu M, Zheng YG (2014) Improvement of Alcaligenes faecalis nitrilase by gene site saturation mutagenesis and its application in stereospecific biosynthesis of (R)-(−)-mandelic acid. J Agric Food Chem 62(20):4685–4694. doi:10.1021/jf405683f
Martínková L, Chmátal M (2016) The integration of cyanide hydratase and tyrosinase catalysts enables effective degradation of cyanide and phenol in coking wastewaters. Water Res 102:90–95. doi:10.1016/j.watres.2016.06.016
Martínková L, Křen V (2010) Biotransformations with nitrilases. Curr Opin Chem Biol 14(2):130–137. doi:10.1016/j.cbpa.2009.11.018
Martínková L, Uhnáková B, Pátek M, Nešvera J, Křen V (2009) Biodegradation potential of the genus Rhodococcus. Environ Int 35(1):162–177. doi:10.1016/j.envint.2008.07.018
Ni KF, Wang HL, Zhao L, Zhang MJ, Zhang SY, Ren YH, Wei DZ (2013) Efficient production of (R)-(−)-mandelic acid in biphasic system by immobilized recombinant E. coli. J Biotechnol 167(4):433–440. doi:10.1016/j.jbiotec.2013.07.024
O’Reilly C, Turner PD (2003) The nitrilase family of CN hydrolysing enzymes - a comparative study. J Appl Microbiol 95(6):1161–1174. doi:10.1046/j.1365-2672.2003.02123.x
Pai O, Banoth L, Ghosh S, Chisti Y, Banerjee UC (2014) Biotransformation of 3-cyanopyridine to nicotinic acid by free and immobilized cells of recombinant Escherichia coli. Proc Biochem 49:655–659. doi:10.1016/j.procbio.2014.01.023
Petříčková A, Sosedov O, Baum S, Stolz A, Martínková L (2012a) Influence of point mutations near the active site on the catalytic properties of fungal arylacetonitrilases from Aspergillus niger and Neurospora crassa. J Mol Catal B Enzym 77:74–80. doi:10.1016/j.molcatb.2012.01.005
Petříčková A, Veselá AB, Kaplan O, Kubáč D, Uhnáková B, Malandra A, Felsberg J, Rinágelová A, Weyrauch P, Křen V, Bezouška K, Martínková L (2012b) Purification and characterization of heterologously expressed nitrilases from filamentous fungi. Appl Microbiol Biotechnol 93(4):1553–1561. doi:10.1007/s00253-011-3525-7
Qiu J, Su EZ, Wang HL, Cai WW, Wang W, Wei DZ (2014a) Cloning, overexpression, and characterization of a high enantioselective nitrilase from Sphingomonas wittichii RW1 for asymmetric synthesis of (R)-phenylglycine. Appl Biochem Biotechnol 173(2):365–377. doi:10.1007/s12010-014-0845-y
Qiu J, Su EZ, Wang W, Wei DZ (2014b) High yield synthesis of D-phenylglycine and its derivatives by nitrilase mediated dynamic kinetic resolution in aqueous-1-octanol biphasic system. Tetrahedron Lett 55(8):1448–1451. doi:10.1016/j.tetlet.2014.01.044
Rinágelová A, Kaplan O, Veselá AB, Chmátal M, Křenková A, Plíhal O, Pasquarelli F, Cantarella M, Martínková L (2014) Cyanide hydratase from Aspergillus niger K10: Overproduction in Escherichia coli, purification, characterization and use in continuous cyanide degradation. Proc Biochem 49(3):445–450. doi:10.1016/j.procbio.2013.12.008
Robertson DE, Chaplin JA, DeSantis G, Podar M, Madden M, Chi E, Richardson T, Milan A, Miller M, Weiner DP, Wong K, McQuaid J, Farwell B, Preston LA, Tan XQ, Snead MA, Keller M, Mathur E, Kretz PL, Burk MJ, Short JM (2004) Exploring nitrilase sequence space for enantioselective catalysis. Appl Environ Microbiol 70(4):2429–2436. doi:10.1128/AEM.70.4.2429-2436.2004
Schreiner U, Hecher B, Obrowsky S, Waich K, Klempier N, Steinkellner G, Gruber K, Rozzell JD, Glieder A, Winkler M (2010) Directed evolution of Alcaligenes faecalis nitrilase. Enzyme Microb Technol 47(4):140–146. doi:10.1016/j.enzmictec.2010.05.012
Seffernick JL, Samanta SK, Louie TM, Wackett LP, Subramanian M (2009) Investigative mining of sequence data for novel enzymes: a case study with nitrilases. J Biotechnol 143(1):17–26. doi:10.1016/j.jbiotec.2009.06.004
Sohoni SV, Nelapati D, Sathe S, Javadekar-Subhedar V, Gaikaiwari RP, Wangikar PP (2015) Optimization of high cell density fermentation process for recombinant nitrilase production in E. coli. Bioresour Technol 188:202–208. doi:10.1016/j.biortech.2015.02.038
Sonbol SA, Ferreira AJS, Siam R (2016) Red Sea Atlantis II brine pool nitrilase with unique thermostability profile and heavy metal tolerance. BMC Biotechnol 16:14. doi:10.1186/s12896-016-0244-2
Sosedov O, Stolz A (2014) Random mutagenesis of the arylacetonitrilase from Pseudomonas fluorescens EBC191 and identification of variants, which form increased amounts of mandeloamide from mandelonitrile. Appl Microbiol Biotechnol 98(4):1595–1607. doi:10.1007/s00253-013-4968-9
Sosedov O, Matzer K, Bürger S, Kiziak C, Baum S, Altenbuchner J, Chmura A, van Rantwijk F, Stolz A (2009) Construction of recombinant Escherichia coli catalysts which simultaneously express an (S)-oxynitrilase and different nitrilase variants for the synthesis of (S)-mandelic acid and (S)-mandelic amide from benzaldehyde and cyanide. Adv Synth Catal 351(10):1531–1538. doi:10.1002/adsc.200900087
Sosedov O, Baum S, Bürger S, Matzer K, Kiziak C, Stolz A (2010) Construction and application of variants of the Pseudomonas fluorescens EBC191 arylacetonitrilase for increased production of acids or amides. Appl Environ Microbiol 76(11):3668–3674. doi:10.1128/AEM.00341-10
Sun HH, Gao WY, Fan HY, Wang HL, Wei DZ (2015a) Cloning, purification and evaluation of the enzymatic properties of a novel arylacetonitrilase from Luminiphilus syltensis NOR5-1B: a potential biocatalyst for the synthesis of mandelic acid and its derivatives. Biotechnol Lett 37(8):1655–1661. doi:10.1007/s10529-015-1830-4
Sun HH, Wang HL, Gao WY, Chen LF, Wu K, Wei DZ (2015b) Directed evolution of nitrilase PpL19 from Pseudomonas psychrotolerans L19 and identification of enantiocomplementary mutants toward mandelonitrile. Biochem Biophys Res Commun 468(4):820–825. doi:10.1016/j.bbrc.2015.11.038
Thuku RN, Brady D, Benedik MJ, Sewell BT (2009) Microbial nitrilases: versatile, spiral forming, industrial enzymes. J Appl Microbiol 106(3):703–727. doi:10.1111/j.1365-2672.2008.03941.x
Vergne-Vaxelaire C, Bordier F, Fossey A, Besnard-Gonnet M, Debard A, Mariage A, Pellouin V, Perret A, Petit JL, Stam M, Salanoubat M, Weissenbach J, De Berardinis V, Zaparucha A (2013) Nitrilase activity screening on structurally diverse substrates: providing biocatalytic tools for organic synthesis. Adv Synt Catal 355(9):1763–1779. doi:10.1002/adsc.201201098
Veselá AB, Petříčková A, Weyrauch P, Martínková L (2013) Heterologous expression, purification and characterization of arylacetonitrilases from Nectria haematococca and Arthroderma benhamiae. Biocatal Biotrans 31(1):49–56. doi:10.3109/10242422.2012.758117
Veselá AB, Křenková A, Martínková L (2015) Exploring the potential of fungal arylacetonitrilases in mandelic acid synthesis. Mol Biotechnol 57(5):466–474. doi:10.1007/s12033-015-9840-y
Veselá AB, Rucká L, Kaplan O, Pelantová H, Nešvera J, Pátek M, Martínková L (2016) Bringing nitrilase sequences from databases to life: the search for novel substrate specificities with a focus on dinitriles. Appl Microbiol Biotechnol 100(5):2193–2202. doi:10.1007/s00253-015-7023-1
Wang L, Watermeyer JM, Mulelu AE, Sewell BT, Benedik MJ (2012) Engineering pH-tolerant mutants of a cyanide dihydratase. Appl Microbiol Biotechnol 94(1):131–140. doi:10.1007/s00253-011-3620-9
Wang HL, Sun HH, Wei DZ (2013) Discovery and characterization of a highly efficient enantioselective mandelonitrile hydrolase from Burkholderia cenocepacia J2315 by phylogeny-based enzymatic substrate specificity prediction. BMC Biotechnol 13:14. doi:10.1186/1472-6750-13-14
Wang HL, Fan HY, Sun HH, Zhao L, Wei DZ (2015a) Process development for the production of (R)-(−)-mandelic acid by recombinant Escherichia coli cells harboring nitrilase from Burkholderia cenocepacia J2315. Org Proc Res Dev 19(12):2012–2016. doi:10.1021/acs.oprd.5b00269
Wang HL, Gao WY, Sun HH, Chen LF, Zhang LJ, Wang XD, Wei DZ (2015b) Protein engineering of a nitrilase from Burkholderia cenocepacia J2315 for efficient and enantioselective production of (R)-o-chloromandelic acid. Appl Environ Microbiol 81(24):8469–8477. doi:10.1128/AEM.02688-15
Xue YP, Wang YP, Xu Z, Liu ZQ, Shu XR, Jia DX, Zheng YG, Shen YC (2015) Chemoenzymatic synthesis of gabapentin by combining nitrilase-mediated hydrolysis with hydrogenation over Raney-nickel. Catal Commun 66:121–125. doi:10.1016/j.catcom.2015.03.035
Yoshida T, Mitsukura K, Mizutani T, Nakashima R, Shimizu Y, Kawabata H, Nagasawa T (2013) Enantioselective synthesis of (S)-2-cyano-2-methylpentanoic acid by nitrilase. Biotechnol Lett 35(5):685–688. doi:10.1007/s10529-012-1131-0
Yusuf F, Rather IA, Jamwal U, Gandhi SG, Chaubey A (2015) Cloning and functional characterization of nitrilase from Fusarium proliferatum AUF-2 for detoxification of nitriles. Funct Integr Genom 15(4):413–424. doi:10.1007/s10142-014-0430-z
Yutthalekha T, Warakulwit C, Limtrakul J, Kuhn A (2015) Enantioselective recognition of DOPA by mesoporous platinum imprinted with mandelic acid. Electroanalysis 27(9):2209–2213. doi:10.1002/elan.201500145
Zhang ZJ, Xu JH, He YC, Ouyang LM, Liu YY, Imanaka T (2010) Efficient production of (R)-(-)-mandelic acid with highly substrate/product tolerant and enantioselective nitrilase of recombinant Alcaligenes sp. Proc Biochem 45(6):887–891. doi:10.1016/j.procbio.2010.02.011
Zhang ZJ, Pan JA, Liu JF, Xu JH, He YC, Liu YY (2011) Significant enhancement of (R)-mandelic acid production by relieving substrate inhibition of recombinant nitrilase in toluene–water biphasic system. J Biotechnol 152(1–2):24–29. doi:10.1016/j.jbiotec.2011.01.013
Zhang CS, Zhang ZJ, Li CX, Yu HL, Zheng GW, Xu JH (2012) Efficient production of (R)-o-chloromandelic acid by deracemization of o-chloromandelonitrile with a new nitrilase mined from Labrenzia aggregata. Appl Microbiol Biotechnol 95(1):91–99. doi:10.1007/s00253-012-3993-4
Zhang XH, Liu ZQ, Xue YP, Zheng YG (2014) Activity improvement of a regioselective nitrilase from Acidovorax facilis and its application in the production of 1-(cyanocyclohexyl) acetic acid. Proc Biochem 49(12):2141–2148. doi:10.1016/j.procbio.2014.08.018
Zhang ZJ, Yu HL, Imanaka T, Xu JH (2015) Efficient production of (R)-(−)-mandelic acid by isopropanol-permeabilized recombinant E. coli cells expressing Alcaligenes sp nitrilase. Biochem Eng J 95:71–77. doi:10.1016/j.bej.2014.12.009
Zhu DM, Mukherjee C, Biehl ER, Hua L (2007) Discovery of a mandelonitrile hydrolase from Bradyrhizobium japonicum USDA110 by rational genome mining. J Biotechnol 129(4):645–650. doi:10.1016/j.jbiotec.2007.02.001
Acknowledgements
This study was funded by the Ministry of Education, Youth and Sports of the Czech Republic (project COST LD15107) and the Institute of Microbiology of the Academy of Sciences of the Czech Republic, v.v.i. (project RVO61388971).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Martínková, L., Rucká, L., Nešvera, J. et al. Recent advances and challenges in the heterologous production of microbial nitrilases for biocatalytic applications. World J Microbiol Biotechnol 33, 8 (2017). https://doi.org/10.1007/s11274-016-2173-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-016-2173-6