Abstract
The presence of microparticle viruses significantly impacts the quality of silkworm seeds for domestic sericulture, making their exclusion from detection in silkworm seed production crucial. Traditional methods for detecting microparticle viruses in silkworms, such as manual microscopic observation, molecular biology, and immunological approaches, are cumbersome and unable to achieve intelligent, batch real-time detection. To address this challenge, we employ the YOLOv8 algorithm in this paper. Firstly, NAM attention is introduced in the original algorithm’s Backbone component, allowing the model to extract more generic feature information. Secondly, ODConv replaces Conv in the Head component of the original algorithm, enhancing the model’s ability to identify microparticle viruses. Finally, NWD-LOSS modifies the CIoU loss of the original algorithm to obtain a more accurate prediction box. Experimental results demonstrate that the NN-YOLOv8 model outperforms mainstream detection algorithms in detecting silkworm microparticle diseases. With an average detection time of 22.6 milliseconds per image, the model shows promising prospects for future applications. This model improvement enhances detection efficiency and reduces human resource costs, effectively realizing detection intelligence.
Similar content being viewed by others
Explore related subjects
Find the latest articles, discoveries, and news in related topics.Avoid common mistakes on your manuscript.
1 Introduction
1.1 Motivation
As a natural fiber, silk combines lightness, softness, and fineness, making it ideal for industrial production, medical testing, and manufacturing processes. High-quality silkworm seeds can improve silk production and cocoon quality. However, the presence of microparticle viruses in the silkworm Bombyx mori seriously affects the quality of silkworm species. This not only leads to a decline in the survival rate of silkworms and a reduction in silk production but also results in substantial economic losses to the silk industry. Since silk is the primary raw material for silk production, infection by microparticle viruses can reduce the quality and quantity of silk, impacting the output and quality of silk products. Consequently, this can decrease enterprise income and competitiveness in the silk market. Moreover, as silk is an important trade commodity, the issue of microparticle viruses may also affect a country’s trade income and adversely impact the national economy. Therefore, achieving intelligent detection of microparticle viruses in domestic silkworms is essential. Microsporidium domestica is a specialized intracellular parasite of the domesticated silkworm, commonly found in the mulberry silkworm breeding industry as a pathogen of microsporidiosis. If silkworm seeds are infected with microparticle disease, symptoms such as black spots, a swollen abdomen, developmental delays, and other issues may arise, seriously affecting silk production quality. Traditional methods for detecting microparticle viruses in domestic silkworms, such as manual microscopic observation, molecular biology, and immunology, are cumbersome, slow, and incapable of achieving intelligent, batch real-time detection. Hence, there is an urgent need for a sensitive and intelligent detection method for rapid real-time detection of microparticle viruses in silkworm species.
1.2 Contributions
In 1857, Nageli [1], in his research, first discovered microsporidia inside the cells of silkworm pupae, which is the primary pathogen of infection in silkworm species. Pasteur [2] found that oviposition could transmit the microsporidia from the mother moth to the offspring. Based on this research finding, examining mother moths using microscopic means were first developed in 1870 by Pasteur, which was based on the use of the naked eye through tiny observation of Nb spore morphology to make a judgment. However, this detection method requires experienced laborers to carry out the identification, which is time-consuming, labor-intensive, subjective to human factors, and unsuitable for large-scale detection in the sericulture industry.
As research into mulberry microsporidiosis has progressed further, various detection methods have been tried. Many scholars have used molecular biology and immunology in a laboratory setting to determine detection, such as Polymerase Chain Reaction [3] (PCR), Quantitative Real-time PCR [4], Loop-mediated isothermal amplification [5] (LAMP). The above detection methods reduce the wastefulness of detecting moths by judging the infection based on the offspring rather than sampling and observing them from the mother. This type of detection provides a more reliable and effective method for detecting microsporidiosis in the mulberry silkworm. Hatakeyama and Hayasaka [6] and Liu et al. [7] developed PCR assays and LAMP methods, respectively, in the examination of silkworm eggs for microparticle disease based on the DNA sequence of microparticle disease, and these two types of method techniques are based on DNA information with specificity, and high sensitivity has been rapidly developed. Rahul et al. [8] observed the whole process of microsporidia sampling and summarized the standard protocols for operation in different samples (silkworm eggs, larvae). Fu et al. [9] developed a Quantitative Real-time PCR-based molecular assay with high accuracy and high-throughput screening capability. He et al. [10] developed an assay system that combines PCR and Nucleic Acid Lateral Flow Strip (NAFLS), which is highly sensitive and easy to use. Li et al. [11] proposed an ALMS-qPCR technique combining rapid and straightforward DNA extraction and Quantitative Real-time PCR, which can be applied to screening pathogenic molecules. Methods based on molecular biology are susceptible and specific but require special instruments and cumbersome operational steps.
Wang et al. [12] developed two monoclonal antibodies, 2G10 and 2B10, to study the localization of spore wall proteins of microparticle viruses using Immunological Fluorescence Assay (IFA) and protein blotting tests. An experimental study by Li et al. [13] successfully identified silkworm microsporidia by DALDI-TOF-MS, named SWP26, which showed that it could be used for diagnostic and drug detection studies. Immunological methods are highly feasible and faster to detect. However, there are several drawbacks to this method, including cross-contamination, limited applications in practical production, as well as extended time requirements, and incapacity to handle large numbers of samples.
2 Related work
As computer technology has developed rapidly, researchers are exploring machine vision-based approaches to detect microparticle viruses in silkworm species. Xu [14] applied digital technology to create a microphotography system that solved the reproduction problem in examining microparticle disease, improved examination ergonomics, reduced costs, and eased work intensity. Zhou et al. [15] addressed the inefficiency of manual detection of microparticle viruses in silkworms by experimenting with electronic techniques for image analysis of microparticle virus pathogens. A CIAS image analyzer was used to test the method on stained images, and the results indicated that the technique could detect microparticle viruses effectively. Hu [16] proposed a microparticle image segmentation technique based on the HSI model for complex backgrounds. This technique separated only the target image from the non-target image and improved the adaptability of the two-dimensional Ostu segmentation method. However, machine vision-based detection methods are cumbersome in their steps, require high experimental instrumentation, are poorly resistant to interference, and detection algorithms need to provide better detection of complex microparticle virus images. Table 1 provides a comparative summary of related work based on references, type of analysis, and methodology.
A growing number of neural network-based target detection algorithms have been developed and used in a variety of fields in recent decades, including image classification [17], industrial inspection [18], autonomous driving [19], intelligent visual charging technology [20,21,22], the fields of traffic signs and road object detection [23, 24]. Modern target detection algorithms fall into two major categories. One is the two-stage algorithm, like Region-CNN [25] (RCNN), Fast Region-based CNN (Fast RCNN) [26], Faster Region-based CNN (Faster-RCNN) [27]. The other is the one-stage algorithm, for instance, CenterNet [28], Single Shot Multibox Detector (SSD) [24], You Only Look Once (YOLO) [29,30,31,32,33,34]. This algorithm eliminates the need for complicated processes such as image pre-processing and image feature extraction, which is highly operational and can be adapted to detect complex scenes.
Silva et al. [35] developed the original RetinaNet model to address the issue of rapid detection and counting of leukocyte counts in an In vivo microscope (IVM), which transforms the image randomly and uniformly by RGB and HSV color schemes, and then expands the data by simulating sample motion with different Point spread functions (PSF) as well as variability transformations. The method has been shown to be more effective in detecting white blood cell counts. Wang et al. [36] proposed an SSD-KD skin cancer detection model. The model uses a weighted loss function of cross-entropy for improving disease information extraction and incorporates a self-supervised assisted learning strategy to aid training. Experiments showed that the model could better detect skin diseases. Fourier laminar microscopy is used to detect small white cells. Wang et al. [37] developed the SO-YOLO model. The method combines multi-resolution features with grids with higher resolutions to detect small targets. Li et al. [38] proposed an MTC-YOLOv5 algorithm for the detection of cucumber plant diseases, which added Coordinate attention (CA) to reduce background interference, combined a Multi-scale (MS) training to improve the detection accuracy of small targets, and experimental results showed that the algorithm had high detection accuracy and speed. Zhu et al. developed a faster and more accurate SE-YOLOV5 model [39] proposed an improved YOLOv5 model for fast sperm detection, which added a Shuffle attention (SA) mechanism to enhance the detection performance of sperm and used DWConv to enhance the speed of convergence. Experiments showed that the model effectively reduced sperm leakage and improved detection accuracy. Zhang et al. [40] proposed an enhanced algorithm for YOLOv5 and random forest, which introduced HSV and Res-Net to extract features and used the random forest to classify and calculate wheat ears for Fusarium head blight (FHB) infection in wheat, which cannot be detected with high accuracy. Experiments show that the algorithm can quickly and efficiently assess the damage caused by FHB to wheat.
In summary, deep learning-based intelligent detection algorithms hold significant promise for cell detection. However, there has been limited exploration of such algorithms for detecting microparticle viruses. The current detection methods, which rely on manual observation, molecular biology, and machine vision, are labor-intensive and unsuitable for large-scale microparticle virus detection. Traditional deep learning detection tasks typically rely on publicly available datasets, which often feature distinct and large-scale features. However, microparticle virus samples are rare and require observation and sampling using an electron microscope. Moreover, microparticle viruses exhibit small and inconspicuous features, posing challenges for accurate feature extraction and detection. These factors increase the difficulty of detection and accuracy in microparticle virus detection tasks. In light of this research status, this paper develops a microparticle intelligence algorithm for silkworm species based on YOLOv8l named NN-YOLOv8, which has the following main contributions:
-
(1)
Using NAM attention as a mechanism introduced into the Backbone component in the model with the intention of enhancing its ability to extract generic features.
-
(2)
In this study, ODConv is used instead of Conv in the Head part of the original model, which enhances its ability to recognize small targets.
-
(3)
In this study, by using NMS-LOSS as opposed to the CIoU loss in the original model, one will obtain a more accurate prediction box.
3 Materials and methods
3.1 Dataset construction
Currently, there needs to be more research on the detection of silkworm microparticle viruses using deep learning methods at home and abroad, and there are no publicly available datasets. Therefore, a silkworm species microparticle virus dataset was constructed in this work. The dataset was sampled from a silkworm breeding farm in the Guangxi Zhuang Autonomous Region, from which 244 specimens containing microparticle viruses were sampled and magnified 400 times by electron microscopy, with 7–10 images saved for each specimen at random, yielding 2180 images, the number of labeled viruses is about 26,000, all with a pixel size of 3840 \(\times\) 2160. They were randomly divided into Train, Val and Test in the ratio of 8:1:1. Microparticle viruses are oblong and green in color, measuring (3.6–3.8) \(\upmu \hbox {m}\) \(\times\) (2.0–2.3) \(\upmu \hbox {m}\), and occupy only \(15 \times 25\) pixel values in the image. In addition, the dataset images contain other impurities, such as silkworm moth fragments, air bubbles, and nematodes. Figure 1 shows the microparticle virus sample data.
Deep learning algorithms rely on datasets with labeled data for effective training. In this process, the targets to be detected are manually outlined and assigned corresponding labels. The annotated data are then used to train the algorithm. In this document, the LabelImg software is used for data annotation. Sample frames within the regions of interest are annotated, and labeling information is added. Once the annotations are complete, the software generates a TXT file for each labeled sample box, containing position information and corresponding labels. These files serve as the training dataset for the YOLOv8 model. The data labeling process used in this paper is illustrated in Fig. 2 below.
3.2 NN-YOLOv8
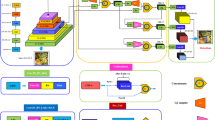
Among the current mainstream algorithms for target detection, the YOLOv8 algorithm is widely utilized in various fields. Comprising Backbone, Neck, and Head components, YOLOv8 serves as a cornerstone in numerous applications. In this study, we introduce enhancements to the NN-YOLOv8 model, which is built upon the YOLOv8 framework. The structural diagram is depicted in Fig. 3. Firstly, to enhance the model’s capability in extracting generic features, we integrate the NAM attention mechanism into the end of the Backbone. Secondly, aiming to improve the model’s ability to detect microparticle viruses, we replace Conv with ODConv in the Head section to enhance performance. Lastly, we recalibrate the losses using NWD-Loss instead of CIoU to obtain more accurate prediction boxes.
3.3 NAM attention module
For improved extraction of generic feature information from the model, attention mechanisms are often incorporated to highlight the key features of the image, reducing the focus on other factors, and thus improving detection speed and accuracy [41,42,43]. Liu et al. [44] proposed a Normalization-based Attention Module (NAM), which redesigned the spatial and channel attention modules using the idea of weights. According to Eq. (1), the channel attention module uses a scale factor based on Batch Normalization (BN).
Where \(\textrm{BN}\) denotes batch normalization, \(B_{\text {in }}\) and \(B_{\text {out }}\) as the input and output of image features. \(\mu _{B}\) is the mean, and \(\sigma _{B}\) is the standard deviation. \(\varepsilon\) for errors, \(\gamma\) and o are trainable affine transformation parameters (scale and displacement). The diagram of the channel attention submodule can be found in Fig. 4 and is calculated as shown in Eqs. (2) and (3).
Where \(M_{c}\) is used as the output of the image features, for each channel, \(\gamma\) is the scale factor, and \(W_{\lambda }\) is the corresponding weight of the module.
Pixel importance can be measured using scale factors. The corresponding spatial attention module is shown in Fig. 5 and is calculated as shown in Eq. (4).
\(M_{s}\) is used as the output of the image features, and \(W_{\gamma }\) is the corresponding mass of the module. In Eqs. (3) and (4), the higher the \(\gamma\), the richer the information contained in the channel, the higher the weight given through the sigmoid function, and the lower the \(\gamma\), the more homogeneous the information contained in the channel, the lower the weight given, and the lower the channel with lower feature information is suppressed through the idea of weight.
To reduce other weight interference, Eq. (5) is amended with a regularization term. x, y represents the input and output, and W represents the weights of the network.
where l(x) is the loss function and g(x) is the L1 parametrization. \(\textrm{p}\) is a function of the equilibrium \(g(\gamma )\) and \(g(\lambda )\). Adding L1 regularization compresses some weight parameters to minimal values to achieve feature selection and sparsity. This approach can help the model learn important features more efficiently and improve the generalization ability and performance.
3.4 ODConv
Recently, many scholars have enhanced their models’ ability to extract small target objects by utilizing convolutional kernels of varying sizes. Omni-dimensional dynamic convolution (ODConv) achieves dynamism by parallelizing four dimensions: kernel size, number of kernels, input channels, and output channels [45]. By allowing the convolution operation to integrate these four dimensions for output, ODConv can better extract features from images and effectively apply them to objects with complex backgrounds and irregular shapes. Additionally, after the input image undergoes feature vector compression through Global Average Pooling (GAP), it is mapped to a new feature space following the Fully Connected (FC) layer and ReLu activation function. Furthermore, the convolutional feature extraction capability is enhanced through parallel complementary superposition, achieved by four independent branches in four dimensions and four independent Sigmoid functions.” ODConv belongs to the category of plug-and-play dynamic convolution. In this study, Backbone’s Conv is replaced by ODConv to enhance the model’s ability to recognize small-sized targets. The model’s operational diagram is illustrated in Fig. 6.
Calculated as shown in Eq. (6), x and y are the output of the image features, and \(\mathrm {W_i}\) is the i-th convolutional kernel. \(\alpha _{\omega i}\) is the attention scalar for the i-th convolutional kernel, \(\alpha _{s i}, \alpha _{c i}, \alpha _{f i}\) are the attention scalars along the space, input, and output channels. Across the various dimensions of the kernel space, \(\odot\) is the multiplication operation.
3.5 NWD-LOSS
In order to address the issue of slow convergence of the loss function and low detection accuracy for some objects, leading to a reduction in the Intersection over Union (IoU), several scholars have proposed alternative calculations to replace the original \({\textrm{CIoU}}\) [46,47,48]. In the context of small target detection, using \({\textrm{CIoU}}\) may result in a higher loss of confidence, as the predicted anchor box may not precisely enclose the target object in a standard rectangle. Therefore, this section adopts the Normalized Gaussian Wasserstein Distance Loss (NWD-Loss), which recalculates the loss using the Wasserstein distance [49]. This approach involves modeling the bounding box as a Gaussian distribution and computing the difference between two Gaussian distributions using the Wasserstein distance to measure the similarity between the detected bounding box and the actual target. Subsequently, this similarity metric is normalized to a range between [0,1] for loss calculation. A higher similarity metric indicates greater resemblance between the detected bounding box and the actual target, whereas a lower similarity metric suggests less resemblance.
The box is modeled as a two-dimensional Gaussian distribution. The equation for its inner connection ellipse is shown in Eq. (7).
In this case, \(\mu _{x}\) and \(\mu _{y}\) represent the coordinates of the ellipse’s center. Similarly, \(\sigma _{x}\) and \(\sigma _{y}\) represent the semi-axes along \(\textrm{x}\) and \(\textrm{y}\), respectively.
Equation (8) illustrates a two-dimensional Gaussian probability density function.
Where \(x, \mu\) and \(\Sigma\) denotes the coordinates, vectors of the Gaussian distribution when Eq. (9) are satisfied:
An ellipse in Eq. (7) indicates the densities profile of a 2D Gaussian distribution so that its border resembles a 2D Gaussian distribution, in which the values are shown in Eq. (10).
At this point, it is possible to translate border A’s similarity to border B’s similarity into a Gaussian distribution distance. In the case of two Gaussian distributions \(\mu _{1}\left( m_{1}, \Sigma _{1}\right)\) and \(\mu _{2}\left( m_{2}, \Sigma _{2}\right)\), the two-dimensional Gaussian distance between them is calculated as shown in Eq. (11).
In the following example, F stands for Frobenius parametrization. The Gaussian distribution for modeling border \(\textrm{A}\left( c x_{a}, c y_{a}, w_{a}, h_{a}\right)\) and border \(\textrm{B}\left( c x_{b}, c y_{b}, w_{b}, h_{b}\right)\) is \(\mathcal {N}_{a} \cdot \mathcal {N}_{b}\) can be further simplified to Eq. (12).
Normalizing this yields Eq. (13).
Constant C. Using NWD-LOSS instead of the original loss function IoU improves accuracy by maintaining scale invariance while better detecting small targets.
4 Experimental environment and evaluation criteria
4.1 Experimental environment
This study conducts all experiments under Windows 10. During the experiment, the hardware environment is Intel(R) i7-11700KF@3.60GHz 8CPU, NVIDIA RTX3080Ti GPU, 64 G RAM. Programmed in Python 3.7. Deep learning frameworks are CUDA 11.0 and CUDNN 11.1. A Stochastic Gradient Descent (SGD) method was employed to optimize the model. During the training process, we utilized input images with dimensions of \(3840\times 2160\), and the official yolov8l.pt file served as the pretrained model for training. We determined the epoch to be 150 based on early stopping criteria and the observation that the training loss stabilized without further decrease. Considering the performance of the GPU graphics card, we set the batch size to 16. Empirically, we assigned the initial learning rate to 0.001, the mosaic enhancement to 1.0, and the mix-up enhancement to 0.243. These hyper-parameters will remain consistent for subsequent experiments.
4.2 Evaluation criteria
In this experiment, the following evaluation criteria were used: Precision (P), Recall (R), Average Precision (AP), and mean Average Precision (mAP).
The model’s correct detection rate is True Positive (TP), and the incorrect detection rate is False Positive (FP). P is the model’s accuracy; R is the regression rate. The formulae are shown in Eqs. (14) and (15).
AP is expressed as the average precision of the prediction based on the following Eq. (16).
The mAP is an ideal metric for assessing models and is calculated as shown in Eq. (17):
In Eq. (18), F1 is calculated as the summed average of precision and recall.
5 Experimental results
5.1 Model validation
Based on the model’s width and depth, the YOLOv8 algorithm can be classified into five different versions: YOLOv8n, YOLOv8s, YOLOv8m, YOLOv8l, and YOLOv8x. In this section, we experiment with the constructed silkworm microparticle virus dataset using each of these five versions, and the comparative results are presented in Table 2.
Compared to the YOLOv8n model, the YOLOv8l model exhibits a 3.4% higher precision, a 1.4% higher recall, and a 5.3% higher mAP, with an average single image detection time increase of 3.5ms. In comparison with the YOLOv8s model, the YOLOv8l model demonstrates a 2.4% higher precision, a 0.1% higher recall, and a 3% higher mAP, with an average single image detection time increase of 3ms. When compared to the YOLOv8m model, the YOLOv8l model shows a 1.1% higher precision, a 1.4% higher mAP, and an increase in average single image detection time of 1.8ms. Finally, compared to the YOLOv8x model, the YOLOv8l model exhibits a 2.3% higher precision, a 0.2% higher recall, and a 2.2% higher mAP, with the average single image detection time reduced by 0.4ms.
The average single image detection time has been reduced by 0.4ms. Precision and Recall are often negatively correlated in a model, while mAP serves as the most important evaluation metric for measuring model performance. In summary, the YOLOv8l model demonstrates superior detection performance for silkworm microparticle viruses. Additionally, to provide a more visual representation of the YOLOv8l model’s performance, we compare the F1_Curve with the PR_Curve for the YOLOv8 model in its four different versions. The results clearly indicate that the YOLOv8l model outperforms the other models in both the F1_Curve and PR_Curve. Figure 7 illustrates the comparison results.
5.2 Ablation experiments
In order to verify the effectiveness of the module used in this paper in the model, we replace the mainstream attention mechanisms such as ECA [50], SimAM [42] and CBAM [51] and compare them with the original YOLOv8l model in terms of Precision, Recall, and mAP, and the comparison of the data is shown in Table 3:
As can be seen from Table 3, YOLOv8l+CBAM performs best in Precision with 65.4%, which is 2.2% higher than the YOLOv8l model, and YOLOv8l+NAM has the best performance in Recall and mAP with 71.8% and 67.6%, which is higher than the YOLOv8l model with 3.7% and 3.1%, respectively, compared to the YOLOv8l model. The implementation results show that the NAM attention mechanism performs excellently in this paper’s dataset. Using different types of convolutions will give the model different features. When processing the features, we used three convolutions, DSConv [52], and GNConv [53] for validation. The comparison results are shown in Table 4:
As can be seen from Table 4, in terms of Precision, YOLOv8l+DSConv performs better with 68.3%, which is 5.1% higher compared to the YOLOv8l model, and in terms of Recall and mAP, the YOLOv8l+ODConv model performs the best with 72.1% and 67.7%, which is 4% higher compared to the YOLOv8l model and 3.2%. The experimental results show that ODConv improves the expressiveness of the model in the dataset used in this paper and is more effective than other convolutions.
Adjusting the loss of the model can recalculate the loss to the model, which in turn improves the accuracy of the prediction; we used four loss functions: SIoU [47], WIoU [48], GIoU [54] recalculation and validation. The comparison results are shown in Table 5:
As can be seen from Table 5, YOLOv8l+NWD-LOSS has the best performance in terms of Precision, Recall, and mAP, reaching 69.1%, 70.3%, and 68.5%, respectively, which are 5.9%, 2.2%, and 4% higher than the YOLOv8l model compared to the YOLOv8l model, respectively. The implementation results show that NWD-LOSS can minimize the loss in training and guide the model to converge toward the optimal solution.
To validate the proposed model, this section conducts a comparative experiment for each improvement step. Table 6 provides a concise overview of the outcomes of their comparison at each stage.
Incorporating the NAM attention mechanism into the YOLOv8l model improved Precision and Recall by 1.5% and 3.7%, respectively, while increasing mAP by 3.1%. This enhancement came at the cost of a 6.6ms increase in average single image detection time. Subsequently, integrating ODConv further improved Precision and Recall by 0.3% and 2.3%, respectively, and mAP by 2.5%, with a slight increase in detection time by 1.2ms compared to YOLOv8l+NAM. Finally, implementing NWD-LOSS led to a 2.1% increase in Precision, a 2.4% increase in mAP, and only a marginal 0.1ms increase in detection time compared to YOLOv8l+NAM+ODConv.
In addition, to give a more visual representation of the NN-YOLOv8 model’s performance, this section compares the F1_Curve and PR_Curve of the YOLOv8l and NN-YOLOv8 models with the actual inspection plots. Figure 8 shows that the modified NN-YOLOv8 model performs much better in F1_Curve and PR_Curve. Figure 9a shows the actual position of the microparticle virus in the picture to be detected, Fig. 9b depicts a comparison of model heat maps, and Fig. 9c compares the level of detection of the model. Comparing the NN-YOLOv8 model to the YOLOv8l model indicates that the NN-YOLOv8 algorithm is more accurate and has a lower miss detection rate. However, compared to all the microparticle viruses present in Fig. 9a, it can be observed that there is still a certain status quo of missed detections. NN-YOLOv8 More detection diagrams are shown in Fig. 10. A comparison of Figs. 9 and 10 reveals that, despite the strong performance of our proposed NN-YOLOv8 model in detection, there are still instances of missed detections. This is attributed to the smaller size and less prominent features of the undetected microparticle viruses.
5.3 Comparative analysis of different algorithms
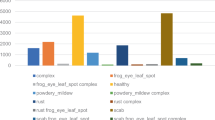
In this section, a comprehensive comparison between the proposed NN-YOLOv8 algorithm and mainstream algorithms, including YOLOv3, YOLOv4, SSD, Fast-RCNN, and YOLOv8l, is presented to further validate its performance. Figure 11 depicts a comparison of their mAP values. Introducing error bars in the bar chart image data analysis enhance its credibility. Error bars are used to represent the uncertainty or variability of the data, visually indicating the range of the data and assisting in evaluating the credibility of prediction results. Compared to the YOLOv3 algorithm, the proposed NN-YOLOv8 model achieves a remarkable 27.65% improvement in mAP. Similarly, in comparison with the YOLOv4 algorithm, the NN-YOLOv8 model exhibits a substantial improvement of 25.15% in mAP. Moreover, when compared to the YOLOv8l model, the NN-YOLOv8 model demonstrates an 8% improvement in mAP, signifying its superior performance. Additionally, when compared to the Faster-RCNN algorithm, the NN-YOLOv8 model outperforms with a significant improvement of 32.68% in mAP. Furthermore, compared to the SSD algorithm, the NN-YOLOv8 model shows a remarkable enhancement of 45.89% in terms of mAP. In summary, the proposed NN-YOLOv8 algorithm, as presented in this study, exhibits higher mAP and detection performance compared to mainstream detection algorithms.
6 Conclusion and future studies
In this paper, we propose NN-YOLOv8l as an enhanced version of the YOLOv8l model. The enhancements include the incorporation of the NAM attention mechanism into the Backbone component to improve generic feature extraction. Additionally, ODConv is utilized instead of Conv in the Head component to enhance the model’s capability in detecting small target objects. Lastly, NMS-LOSS is employed to mitigate the CIoU loss, aiming for more accurate prediction boxes. According to experimental results, the NN-YOLOv8 model achieves a remarkable mAP of 72.5%. Compared to mainstream target detection algorithms, NN-YOLOv8 demonstrates superior performance in detection. In practical applications, researchers only need to collect samples of the specimens, as the model can swiftly determine the location and density of microparticle viruses in the image. This capability fulfills real-time detection requirements, eliminating the necessity for experienced validators to make judgments and effectively addressing the issue of high manual labor intensity.
Although the model exhibits a superior mean Average Precision (mAP) compared to other mainstream algorithms, several shortcomings still require attention. These include the relatively lower mAP of the model, the excessive number of layers in the improved model leading to a high number of parameters, which results in a large model size unsuitable for embedded device applications, and the presence of smaller microparticle viral microspores leading to detection leakage. Following team consultation, it was determined that excessive background impurities and motion blur in the captured images contribute to these issues. To address this, the next step involves processing the background impurities using image processing techniques such as top-hat arithmetic on the dataset. This will help lighten the model and improve its performance. Generative adversarial networks (GAN) will deblur the images, resulting in more precise and accurate pictures. The high-quality images generated through this process will be used to train the detection model further, leading to improved detection accuracy.
Data availability
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
References
Nageli C (1857) uber die neue krankheit der seidenraupe und verwandte organismen [abstract of report before 33. versamml. deutsch. naturf. u. aerzte. bonn, 21 sept.]. Bot. Ztg. 15:760–761
Pasteur L (1870) Etudes sur la maladie des vers à soie: 2.: Notes et documents. Gauthier-Villars, Paris. https://doi.org/10.5962/bhl.title.119544
Fu Z, He X, Cai S, Liu H, He X, Li M, Lu X (2016) Quantitative PCR for detection of Nosema bombycis in single silkworm eggs and newly hatched larvae. J Microbiol Methods 120:72–78. https://doi.org/10.1016/j.mimet.2015.12.003
Klee J, Tay WT, Paxton RJ (2006) Specific and sensitive detection of Nosema bombi (Microsporidia: Nosematidae) in bumble bees (Bombus spp.; Hymenoptera: Apidae) by PCR of partial rRNA gene sequences. J Invertebr Pathol 91(2):98–104. https://doi.org/10.1016/j.jip.2005.10.012
Yan W, Shen Z, Tang X, Xu L, Li Q, Yue Y, Xiao S, Fu X (2014) Detection of Nosema bombycis by FTA cards and loop-mediated isothermal amplification (LAMP). Curr Microbiol 69:532–540. https://doi.org/10.1007/s00284-014-0619-3
Hatakeyama Y, Hayasaka S (2003) A new method of pebrine inspection of silkworm egg using multiprimer PCR. J Invertebr Pathol 82(3):148–151. https://doi.org/10.1016/S0022-2011(03)00019-3
Liu J, Cheng W, Yan Y, Wei J, Yang J et al (2015) Detection of pebrine disease in Bombyx mori eggs with the loop-mediated isothermal amplification (LAMP) method based on EB1 gene. Acta Entomol Sin 58(8):846–855
Rahul K, Manjunatha GR, Sivaprasad V (2021) Pebrine monitoring methods in sericulture. Methods Microbiol 49:79–96. https://doi.org/10.1016/bs.mim.2021.04.003
Fu Z, He X, Cai S, Liu H, He X, Li M, Lu X (2016) Quantitative PCR for detection of Nosema bombycis in single silkworm eggs and newly hatched larvae. J Microbiol Methods 120:72–78. https://doi.org/10.1016/j.mimet.2015.12.003
He Z, Ni Q, Song Y, Wang R, Tang Y, Wu Y, Liu L, Bao J, Chen J, Long M et al (2019) Development of a nucleic acid lateral flow strip for rapid, visual detection of Nosema bombycis in silkworm eggs. J Invertebr Pathol 164:59–65. https://doi.org/10.1016/j.jip.2019.04.004
Li P, Mi R, Zhao R, Li X, Zhang B, Yue D, Ye B, Zhao Z, Wang L, Zhu Y et al (2019) Quantitative real-time PCR with high-throughput automatable DNA preparation for molecular screening of Nosema spp. in Antheraea pernyi. J Invertebr Pathol 164:16–22. https://doi.org/10.1016/j.jip.2019.04.003
Wang J-Y, Chambon C, Lu C-D, Huang K-W, Vivarès CP, Texier C (2007) A proteomic-based approach for the characterization of some major structural proteins involved in host-parasite relationships from the silkworm parasite Nosema bombycis (Microsporidia). Proteomics 7(9):1461–1472. https://doi.org/10.1002/pmic.200600825
Li Y, Wu Z, Pan G, He W, Zhang R, Hu J, Zhou Z (2009) Identification of a novel spore wall protein (SWP26) from microsporidia Nosema bombycis. Int J Parasitol 39(4):391–398. https://doi.org/10.1016/j.ijpara.2008.08.011
Xu G, Pan L (2002) Application and research of digital microscopy in the inspection of microparticle disease. Jiangsu Sericul Ture 1:14–16
Zhou Y, Zeng C, Xie J (1995) Analysis of the pathogenic image of microparticle disease of the silkworm. J Sichuan Univ Sci Ed 32(2):224–226. https://doi.org/10.1016/j.ijpara.2008.08.011
Hu X (2011) Research on micro-particle image recognition method of silkworm based on machine vision. Wuhan Institute of Technology, Wuhan
Mustafa HT, Zareapoor M, Yang J (2020) MLDNet: multi-level dense network for multi-focus image fusion. Signal Process Image Commun 85:115864. https://doi.org/10.1016/j.image.2020.115864
Kaya GU (2023) Development of hybrid optical sensor based on deep learning to detect and classify the micro-size defects in printed circuit board. Measurement 206:112247. https://doi.org/10.1016/j.measurement.2022.112247
Emin Güney C, Williams R, Shi X, Yuan Q, Trigg M (2022) Autonomous control of shore robotic charging systems based on computer vision. J Hydrol 612:128217. https://doi.org/10.1016/j.jhydrol.2022.128217
Güney E, Bayılmış C, Çakar S, Erol E, Atmaca Ö (2023) Autonomous control of shore robotic charging systems based on computer vision. Expert Syst Appl 238:122116. https://doi.org/10.1016/j.eswa.2023.122116
Güney E, Sahin IH, Cakar S, Atmaca O, Erol E, Doganli M, Bayilmis C (2022) Electric shore-to-ship charging socket detection using image processing and yolo. In: 2022 International Symposium on Multidisciplinary Studies and Innovative Technologies (ISMSIT), pp 1069–1073. https://doi.org/10.1109/ISMSIT56059.2022.9932841
Güney E, Bayilmiş C, Çakan B (2022) An implementation of real-time traffic signs and road objects detection based on mobile GPU platforms. IEEE Access 10:86191–86203. https://doi.org/10.1109/ACCESS.2022.3198954
Güney E, Bayilmiş C, Çakan B (2022) Corrections to “an implementation of real-time traffic signs and road objects detection based on mobile GPU platforms’’. IEEE Access 10:103587–103587. https://doi.org/10.1109/ACCESS.2022.3209832
Güney E, Bayılmış C (2022) An implementation of traffic signs and road objects detection using faster R-CNN. Sakarya Univ J Comput Inf Sci 5:216–224. https://doi.org/10.35377/saucis.05.02.1073355
Girshick R, Donahue J, Darrell T, Malik J (2014) Rich feature hierarchies for accurate object detection and semantic segmentation. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp 580–587. https://doi.org/10.1109/CVPR.2014.81
Girshick R (2015) Fast R-CNN. In: Proceedings of the IEEE International Conference on Computer Vision, pp 1440–1448. https://doi.org/10.1109/ICCV.2015.169
Ren S, He K, Girshick R, Sun J (2015) Faster R-CNN: towards real-time object detection with region proposal networks. In: Advances in Neural Information Processing Systems, vol 28.https://doi.org/10.48550/arXiv.1506.01497
Duan K, Bai S, Xie L, Qi H, Huang Q, Tian Q (2019) CenterNet: keypoint triplets for object detection. In: Proceedings of the IEEE/CVF International Conference on Computer Vision, pp 6569–6578. https://doi.org/10.48550/arXiv.1904.08189
Liu W, Anguelov D, Erhan D, Szegedy C, Reed S, Fu C-Y, Berg AC (2016) SSD: single shot multibox detector. In: Computer Vision–ECCV 2016: 14th European Conference, Amsterdam, The Netherlands, October 11–14, 2016, Proceedings, Part I 14. Springer, pp 21–37. https://doi.org/10.1007/978-3-319-46448-0_2
Redmon J, Divvala S, Girshick R, Farhadi A (2016) You only look once: unified, real-time object detection. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp 779–788. https://doi.org/10.48550/arXiv.1506.02640
Redmon J, Farhadi A (2017) YOLO9000: better, faster, stronger. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 7263–7271. https://doi.org/10.48550/arXiv.1612.08242
Redmon J, Farhadi A (2018) YOLOV3: an incremental improvement. arXiv preprint arXiv:1804.02767
Bochkovskiy A, Wang C-Y, Liao H-YM (2020) YOLOV4: optimal speed and accuracy of object detection. arXiv preprint arXiv:2004.10934
Wang C-Y, Bochkovskiy A, Liao H-YM (2023) YOLOV7: trainable bag-of-freebies sets new state-of-the-art for real-time object detectors. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp 7464–7475. https://doi.org/10.48550/arXiv.2207.02696
Silva BCG, Tam R, Ferrari RJ (2021) Detecting cells in intravital video microscopy using a deep convolutional neural network. Comput Biol Med 129:104133. https://doi.org/10.48550/arXiv.2207.02696
Wang Y, Wang Y, Cai J, Lee TK, Miao C, Wang ZJ (2023) SSD-KD: a self-supervised diverse knowledge distillation method for lightweight skin lesion classification using dermoscopic images. Med Image Anal 84:102693. https://doi.org/10.1016/j.media.2022.102693
Wang X, Xu T, Zhang J, Chen S, Zhang Y (2018) SO-YOLO based WBC detection with Fourier ptychographic microscopy. IEEE Access 6:51566–51576. https://doi.org/10.1109/ACCESS.2018.2865541
Li S, Li K, Qiao Y, Zhang L (2022) A multi-scale cucumber disease detection method in natural scenes based on YOLOv5. Comput Electron Agric 202:107363. https://doi.org/10.1016/j.compag.2022.107363
Zhu R, Cui Y, Huang J, Hou E, Zhao J, Zhou Z, Li H (2023) YOLOv5s-SA: light-weighted and improved YOLOv5S for sperm detection. Diagnostics 13(6):1100. https://doi.org/10.3390/diagnostics13061100
Zhang D-Y, Luo H-S, Wang D-Y, Zhou X-G, Li W-F, Gu C-Y, Zhang G, He F-M (2022) Assessment of the levels of damage caused by Fusarium head blight in wheat using an improved YOLOv5 method. Comput Electron Agric 198:107086. https://doi.org/10.1016/j.compag.2022.107086
Dai T, Cai J, Zhang Y, Xia S-T, Zhang L (2019) Second-order attention network for single image super-resolution. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp 11065–11074. https://doi.org/10.1109/CVPR.2019.01132
Yang L, Zhang R-Y, Li L, Xie X (2021) SimAM: a simple, parameter-free attention module for convolutional neural networks. In: International Conference on Machine Learning. PMLR, pp 11863–11874. https://doi.org/10.1109/CVPR.2019.01132
Liu Y, Shao Z, Hoffmann N (2021) Global attention mechanism: retain information to enhance channel-spatial interactions. arXiv preprint arxiv:2112.05561
Liu Y, Shao Z, Teng Y, Hoffmann N (2021) NAM: normalization-based attention module. arXiv preprint arXiv:2111.12419
Li C, Zhou A, Yao A (2022) Omni-dimensional dynamic convolution. arXiv preprint arxiv:2209.07947
Zheng Z, Wang P, Liu W, Li J, Ye R, Ren D (2020) Distance-IoU loss: faster and better learning for bounding box regression. In Proceedings of the AAAI conference on artificial intelligence, vol. 34, pp 12993–13000. https://doi.org/10.1609/aaai.v34i07.6999
Gevorgyan Z (2022) SIoU loss: more powerful learning for bounding box regression. arXiv preprint arxiv:2205.12740
Tong Z, Chen Y, Xu Z, Yu R (2023) Wise-IoU: bounding box regression loss with dynamic focusing mechanism. arXiv preprint arXiv:2301.10051
Wang J, Xu C, Yang W, Yu L (2021) A normalized Gaussian Wasserstein distance for tiny object detection. arXiv preprint arXiv:2110.13389
Wang Q, Wu B, Zhu P, Li P, Zuo W, Hu Q (2020) ECA-Net: efficient channel attention for deep convolutional neural networks. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp 11534–11542. https://doi.org/10.48550/arXiv.1910.03151
Woo S, Park J, Young LJ (2018) CBAM: convolutional block attention module. In: Proceedings of the European Conference on Computer Vision (ECCV), pp 3–19. https://doi.org/10.48550/arXiv.1807.06521
Gennari N, Fawcett R, Prisacariu VA (2019) DSConv: efficient convolution operator. In: Proceedings of the IEEE/CVF International Conference on Computer Vision, pp 5148–5157. https://doi.org/10.48550/arXiv.1901.01928
Rao Y, Zhao W, Tang Y, Zhou J, Lim SN, Lu J(2022) HorNet: efficient high-order spatial interactions with recursive gated convolutions. In: Advances in Neural Information Processing Systems. https://doi.org/10.48550/arXiv.2207.14284
Rezatofighi H, Tsoi N, Gwak J, Sadeghian A, Reid I, Savarese S (2019) Generalized intersection over union: a metric and a loss for bounding box regression. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp 658–666. https://doi.org/10.48550/arXiv.1902.09630
Acknowledgements
Not applicable.
Funding
Central Leading Local Science and Technology Development Funds Project, Hechi Science and Technology Planning No. [2022] 5 (2022HKJZ05), Hechi University 2021 High-level Talent Research Initiation Project (2021GCC014)
Author information
Authors and Affiliations
Contributions
Conceptualization was contributed by J.S., Y.Z. and T.W.; methodology was contributed by J.S., Y.Z. and T.W.; software was contributed by T.W.; data curation was contributed by C.X. and Y.A.; validation was contributed by T.W. and J.S.; formal analysis was contributed by C.X. and Y.Z.; investigation was contributed by J.S., Y.Z., T.W., C.X. and Y.A; funding acquisition was contributed by J.S.; resources were contributed by J.S.; writing—original draft preparation, was contributed by J.S; writing—review and editing, was contributed by J.S., Y.Z., T.W.; visualization was contributed by Y.Z; supervision was contributed by T.W; project administration was contributed by T.W.; all authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no Conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Y., Su, J., Wang, T. et al. Intelligent detection method of microparticle virus in silkworm based on YOLOv8 improved algorithm. J Supercomput 80, 18118–18141 (2024). https://doi.org/10.1007/s11227-024-06159-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11227-024-06159-w