Abstract
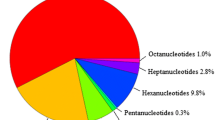
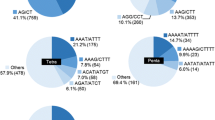
Linseed (Linum usitatissimum L.) is regarded as a cash crop of tomorrow because of the presence of nutraceutically important α-linolenic acid (ALA) and lignan. However, only limited breeding progress has been made in this crop, mainly due to the lack of sufficient genetic and genomic resources. Among these, simple sequence repeats (SSR) are useful DNA markers for diversity analysis, genetic mapping and tagging traits because of their co-dominant and highly polymorphic nature. In order to develop SSR markers for linseed, we used three microsatellite isolation methods, viz., PCR Isolation of Microsatellite Arrays (PIMA), 5′-anchored PCR method, and Fast Isolation by AFLP of Sequences COntaining repeats (FIASCO). The amplified products from these methods were pooled and sequenced using the 454 GS-FLX platform. A total of 36,332 reads were obtained, which assembled into 2,183 contigs and 2,509 singlets. The contigs and the singlets contained 1,842 microsatellite motifs, with dinucleotide motifs as the most abundant repeat type (54%) followed by trinucleotide motifs (44%). Based on this, 290 SSR markers were designed, 52 of which were evaluated using a panel of 27 diverse linseed genotypes. Among the three enrichment methods, the 5′-anchored PCR method was most efficient for isolation of microsatellites, while FIASCO was most efficient for developing SSR markers. We show the utility of next-generation sequencing technology for efficiently discovering a large number of microsatellite markers in non-model plants.



Similar content being viewed by others
References
Aggarwal RK, Hendre PS, Varshney RK, Bhat PR, Krishnakumar V, Singh L (2007) Identification, characterization and utilization of EST-derived genic microsatellite markers for genome analyses of coffee and related species. Theor Appl Genet 114:359–372
Bennett MD, Leitch IJ (2005) Nuclear DNA amounts in angiosperms: progress, problems and prospects. Ann Bot 95:45–90
Bickel CL, Gadani S, Lukacs M, Cullis CA (2011) SSR markers developed for genetic mapping in flax (Linum usitatissimum L.). Res Rep Biol 2011:23–29
Cardle L, Ramsay L, Milbourne D, Macaulay M, Marshall D, Waugh R (2000) Computational and experimental characterization of physically clustered simple sequence repeats in plants. Genetics 156:847–854
Cavagnaro PF, Senalik DA, Yang LM, Simon PW, Harkins TT, Kodira CD, Huang SW, Weng YQ (2010) Genome-wide characterization of simple sequence repeats in cucumber (Cucumis sativus L.). BMC Genomics 11:569–570
Cloutier S, Niu ZX, Datla R, Duguid S (2009) Development and analysis of EST-SSRs for flax (Linum usitatissimum L.). Theor Appl Genet 119:53–63
Cloutier S, Ragupathy R, Niu Z, Duguid S (2011) SSR-based linkage map of flax (Linum usitatissimum L.) and mapping of QTLs underlying fatty acid composition traits. Mol Breed (In press) doi:10.1007/s11032-010-9494-1
Collard BCY, Jahufer MZZ, Brouwer JB, Pang ECK (2005) An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142:169–196
da Maia LC, Palmieri DA, Souza VQ, Kopp MM, Carvalho FIF, Costa de Oliveira A (2008) SSR Locator: tool for simple sequence repeat discovery integrated with primer design and PCR simulation. Int J Plant Genomics, Article ID 412696, p 9
Deng X, Long SH, He DF, Li X, Wang YF, Liu J, Chen XB (2010) Development and characterization of polymorphic microsatellite markers in Linum usitatissimum. J Plant Res 123:119–123
Dutta S, Kumawat G, Singh BP, Gupta DK, Singh S, Dogra V, Gaikwad K, Sharma TR, Raje RS, Bandhopadhya TK, Datta S, Singh MN, Bashasab F, Kulwal P, Wanjari KB K, Varshney R, Cook DR, Singh NK (2011) Development of genic-SSR markers by deep transcriptome sequencing in pigeonpea (Cajanus cajan (L.) Millspaugh). BMC Plant Biol 11:17
FAOSTAT Database (2009) http://faostat.fao.org Accessed 10 Feb 2011
Fisher PJ, Gardner RC, Richardson TE (1996) Single locus microsatellites isolated using 5′-anchored PCR. Nucleic Acids Res 24:4369–4371
Gadaleta A, Mangini G, Mule G, Blanco A (2007) Characterization of dinucleotide and trinucleotide EST-derived microsatellites in the wheat genome. Euphytica 153:73–85
Ghosh R, Paul S, Ghosh SK, Roy A (2009) An improved method of DNA isolation suitable for PCR-based detection of begomoviruses from jute and other mucilaginous plants. J Virol Methods 159:34–39
Gimenes MA, Hoshino AA, Barbosa AVG, Palmieri DA, Lopes CR (2007) Characterization and transferability of microsatellite markers of the cultivated peanut (Arachis hypogaea). BMC Plant Biol 7:9
He GH, Meng RH, Newman M, Gao GQ, Pittman RN, Prakash CS (2003) Microsatellites as DNA markers in cultivated peanut. BMC Plant Biol 3:3
Hiremath PJ, Farmer A, Cannon SB, Woodward J, Kudapa H, Tuteja R, Kumar A, Bhanuprakash A, Mulaosmanovic B, Gujaria N, Krishnamurthy L, Gaur PM, Kavikishor PB, Shah T, Srinivasan R, Lohse M, Xiao Y, Town CD, Cook DR, May GD, Varshney RK (2011) Large-scale transcriptome analysis in chickpea (Cicer arietinum L.), an orphan legume crop of the semi-arid tropics of Asia and Africa. Plant Biotechnol J (In press) doi:10.1111/j.1467-7652.2011.00625.x
Huang XQ, Madan A (1999) CAP3: a DNA sequence assembly program. Genome Res 9:868–877
Jae-Woong Y, Anupam D, Ma KH, Jong-Wook C, Park YJ (2009) A study on relative abundance, composition and length variation of microsatellites in 18 underutilized crop species. Genet Resour Crop Evol 56:237–246
Joshi SP, Ranjekar PK, Gupta VS (1999) Molecular markers in plant genome analysis. Curr Sci India 77:230–240
Katti MV, Ranjekar PK, Gupta VS (2001) Differential distribution of simple sequence repeats in eukaryotic genome sequences. Mol Biol Evol 18:1161–1167
Kaur S, Cogan NO, Pembleton LW, Shinozuka M, Savin KW, Materne M, Forster JW (2011) Transcriptome sequencing of lentil based on second-generation technology permits large-scale unigene assembly and SSR marker discovery. BMC Genomics 12:265
Kolliker R, Jones ES, Drayton MC, Dupal MP, Forster JW (2001) Development and characterization of simple sequence repeat (SSR) markers for white clover (Trifolium repens L.). Theor Appl Genet 102:416–424
Li WZ, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22:1658–1659
Lunt DH, Hutchinson WF, Carvalho GR (1999) An efficient method for PCR-based isolation of microsatellite arrays (PIMA). Mol Ecol 8:891–893
McCouch SR, Chen XL, Panaud O, Temnykh S, Xu YB, Cho YG, Huang N, Ishii T, Blair M (1997) Microsatellite marker development, mapping and applications in rice genetics and breeding. Plant Mol Biol 35:89–99
Newcomb RD (2006) Analysis of expressed sequence tags from apple. Plant Physiol 141:147–166
Powell W, Machray GC, Provan J (1996) Polymorphism revealed by simple sequence repeats. Trends Plant Sci 1:215–222
Rabello E, de Souza AN, Saito D, Tsai SM (2005) In silico characterization of microsatellites in Eucalyptus spp.: Abundance, length variation and transposon associations. Genet Mol Biol 28:582–588
Rajwade AV, Arora RS, Kadoo NY, Harsulkar AM, Ghorpade PB, Gupta VS (2010) Relatedness of Indian flax genotypes (Linum usitatissimum L.): An Inter-Simple Sequence Repeat (ISSR) primer assay. Mol Biotechnol 45:161–170
Rallo P, Dorado G, Martin A (2000) Development of simple sequence repeats (SSRs) in olive tree (Olea europaea L.). Theor Appl Genet 101:984–989
Roose-Amsaleg C, Cariou-Pham E, Vautrin D, Tavernier R, Solignac M (2006) Polymorphic microsatellite loci in Linum usitatissimum. Mol Ecol Notes 6:796–799
Sanguinetti CJ, Neto ED, Simpson AJG (1994) Rapid silver staining and recovery of PCR products separated on polyacrylamide gels. Biotechniques 17:914–921
Santana QC, Coetzee MPA, Steenkamp ET, Mlonyeni OX, Hammond GNA, Wingfield MJ, Wingfield BD (2009) Microsatellite discovery by deep sequencing of enriched genomic libraries. Biotechniques 46:217–223
Sharopova N, McMullen MD, Schultz L, Schroeder S, Sanchez-Villeda H, Gardiner J, Bergstrom D, Houchins K, Melia-Hancock S, Musket T, Duru N, Polacco M, Edwards K, Ruff T, Register JC, Brouwer C, Thompson R, Velasco R, Chin E, Lee M, Woodman-Clikeman W, Long MJ, Liscum E, Cone K, Davis G, Coe EH (2002) Development and mapping of SSR markers for maize. Plant Mol Biol 48:463–481
Singh M, Kumar R, Nagpure NS, Kushwaha B, Mani I, Murmu K, Chauhan UK, Lakra WS (2010) Nucleotide sequences and chromosomal localization of 45S and 5S rDNA in Neolissochilus hexagonolepis (Pisces, Cyprinidae), using dual-color fish. Zool Sci 27:709–716
Soto-Cerda BJ, Carrasco RA, Aravena GA, Urbina HA, Navarro CS (2011) Identifying novel polymorphic microsatellites from cultivated flax (Linum usitatissimum L.) following data mining. Plant Mol Biol Rep 29:753–759
Squirrell J, Hollingsworth PM, Woodhead M, Russell J, Lowe AJ, Gibby M, Powell W (2003) How much effort is required to isolate nuclear microsatellites from plants? Mol Ecol 12:1339–1348
Tang SJ, Liu ZZ, Tang WQ, Yang JQ (2009) A simple method for isolation of microsatellites from the mudskipper (Boleophthalmus pectinirostris), without constructing a genomic library. Conserv Genet 10:1957–1959
Toth G, Gaspari Z, Jurka J (2000) Microsatellites in different eukaryotic genomes: Survey and analysis. Genome Res 10:967–981
Varshney RK, Graner A, Sorrells ME (2005) Genic microsatellite markers in plants: features and applications. Trends Biotechnol 23:48–55
Weber JL, May PE (1989) Abundant class of human DNA polymorphisms which can be typed using the polymerase chain-reaction. Am J Hum Genet 44:388–396
Wiesner I, Wiesnerova D, Tejklova E (2001) Effect of anchor and core sequence in microsatellite primers on flax fingerprinting patterns. J Agric Sci 137:37–44
Zane L, Bargelloni L, Patarnello T (2002) Strategies for microsatellite isolation: a review. Mol Ecol 11:1–16
Acknowledgments
This research was financially supported partly by an in-house grant (MLP020226) of the National Chemical Laboratory, Pune, and partly by the Indo–Canadian project (GAP278426) sponsored by the Department of Biotechnology, India. The authors also acknowledge the Council of Scientific and Industrial Research (CSIR), India, for providing fellowships to SMK and VCP.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kale, S.M., Pardeshi, V.C., Kadoo, N.Y. et al. Development of genomic simple sequence repeat markers for linseed using next-generation sequencing technology. Mol Breeding 30, 597–606 (2012). https://doi.org/10.1007/s11032-011-9648-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-011-9648-9