Abstract
Maize is one of the three major crops worldwide based on its yield and quality. Starch is crucial to both the yield and quality of maize as it accounts more than 60% of the seed weight, and its structure influences the quality of the crop. Starch synthase I (SSI) contributes to the majority of the starch synthase activity in the maize endosperm. An in-depth understanding of the starch synthesis regulatory mechanism would provide opportunities for improving the yield and quality of maize. In this study, ZmPLATZ2, a plant AT-rich sequence and zinc-binding protein (PLATZ) transcription factor related to starch synthesis, was selected based on co-expression analysis. The semiquantitative RT-PCR and qRT-PCR assays revealed that ZmPLATZ2 had a high expression in the endosperm, and reached the peak at 12 days after pollination (DAP). Different treatments demonstrated that ZmPLATZ2 was downregulated by the presence of sucrose. Subsequent transactivation and subcellular localization analyses showed that ZmPLATZ2 was localized in the nuclei without transactivation. Yeast one-hybrid and transient expression in maize endosperm indicated that ZmPLATZ2 could bind to the promoters of ZmSSI, ZmISA1, and ZmISA2 and increase their gene expression. After ZmPLATZ2 overexpression in rice, four starch synthesis genes were significantly upregulated in the transgenic plant, including the OsSSI gene. In vitro DAP-seq data showed that ZmPLATZ2 could bind to the CAAAAAAA element. In conclusion, our data support that ZmPLATZ2 binds to the CAAAAAAA element in the ZmSSI promoter and mediates the Glu signal pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Maize (Zea mays) is an important crop that contributes to solving the food security problem (Godfray et al. 2010). Starch accounts for more than half the weight of the seed; therefore, increasing the starch content is an essential measure for improving the yield. Nevertheless, the mechanism of starch biosynthesis is unclear as it includes a series of complex metabolic processes. Four types of enzymes are involved in the starch biosynthesis reaction: adenosine diphosphate glucose pyrophosphorylase (AGPase), starch synthase (SS), starch branching enzyme (SBE), and starch debranching enzyme (DBE) (Leterrier et al. 2008; Zhou et al. 2016; Hennen-Bierwagen et al. 2008). AGPase is the first key enzyme in starch synthesis that involved in ADP-glucose production (Huang et al. 2014; Li et al. 2018). SS mainly participates in amylopectin synthesis. IbSSI, a soluble starch synthase I gene, is involved in the synthesis of amylopectin chains and alters the content and structure of starch in the sweet potato (Wang et al. 2017). SBE is an essential factor in generate different heteromorphous starch granules (Wang et al. 2018a, b). The maize sbe2b (amylose-extender, ae) mutant shown that the amylose and resistant starch content increase 60% and 40% respectively (Li et al. 2008). DBE is responsible for generated linear starch fraction (Reddy et al. 2018). These four enzymes, during the seed filling period, influence the starch content and accumulation rate.
Transcription factors (TFs) regulate the expression patterns of target genes by binding to the cis-elements and play a vital role in starch synthesis. One example is that of ZmMYB14, which directly binds to the Bt1 promoter through a TAACTG motif of promote BT1 expression (Xiao et al. 2017). ZmABI4 binds to the CACCG sites of the ZmSSI promoter to mediate the ABA signal pathway (Hu et al. 2012). ZmEREB94 and ZmEREB156 are induced by sucrose and ABA, and are involved in the starch biosynthetic and metabolic pathways (Li et al. 2017b; Huang et al. 2016).
Nowadays, several TFs appear at one branch of the evolution tree. For example, the nuclear receptor family is absent in plants and yeast but is found in animals (Bennett et al. 2017), whereas the homeobox family has been found in animals and plants but not in yeast. The plant AT-rich sequence and zinc-binding protein (PLATZ) TF family is present only in plants and was first found in peas (Nagano et al. 2001). Previous studies of the PLATZ family are rare. However, it was recently established that PLATZ1 is significant in seed germination and seedling establishment and mediates the ABA, GA, and ethylene signaling pathways in cotton (Zhang et al. 2018). In addition, ORESARA15, a number of the PLATZ TF family, functions as a coordinator in leaf growth and senescence in Arabidopsis (Kim et al. 2018). The PLATZ TF family is also involved in grain development. The rice GL6, belonging to the PLATZ family, participates in rice grain development by interacting with C53 to negatively regulate grain numbers (Wang et al. 2019). Mutation in ZmPLATZ12 results in a floury endosperm, which indicates that ZmPLATZ12 plays a crucial role in endosperm development and storage reserve filling (Li et al. 2017a).
In previous study, we identified a gene, GRMZM2G31165, via co-expression analysis. This gene encoded a TF containing a PLATZ domain, named ZmPLATZ2. The assays used determined that ZmPLATZ2 was able to bind to the SSI, SSIIa, SSIV, SSV, ISA1, and ISA2 promoters. In addition, ZmPLATZ2 increased the promoter activities of Sh2, Bt2, GBSSI, SSI, and SSIIIa. Meanwhile, we found that ZmPLATZ2 expression was stimulated when treated with glucose. In addition, the transcription factor identified in our study could regulate maize starch synthesis. Our study will contribute to elucidate the starch synthesis mechanism and improve yield by molecular technology.
Materials and methods
Co-expression analysis of the ZmPLATZ2 gene and starch synthesis related gene
Co-expression analysis of the data that include seven different period endosperms from Stelpflug et al. (2016) was used to select a candidate gene (GRMZM2G311656) coding a PLATZ protein. Twenty-seven maize starch synthesis related genes, which possessed a high expression in maize seed, were considered as the guide genes (Crofts et al. 2017). The Pearson correlation coefficient (PCC) between ZmPLATZ2 transcription factor and starch synthesis related genes were calculated by excel, and cytoscape_3.7.2 was use to display the co-expression analysis results.
Plant materials
Maize (Z. mays) inbred line 18-599 was grown during the summer field season at the Sichuan Agricultural University Farm. Developing kernels from self-pollinated plants were collected at specified days after pollination (DAP) for quantitative real-time PCR. The roots, stems, leaves, embryos, endosperms, and seeds were harvested at 15 DAP for semiquantitative RT-PCR. All samples were frozen in liquid N2 and stored at − 80 °C for further use. Endosperms that used in transient expression and ABA, sucrose and glucose treatment were harvested 9 or 10 DAP, as described previously (Xiao et al. 2017).
Transcript level measurement by RT-PCR and semiquantitative RT-PCR
Total RNA was extracted from the developing kernels by using TRIzol (Invitrogen, China) and RNase-free DNase to remove the genomic DNA. The 2 ng DNase treated RNA was converted into cDNA using reverse transcription PCR (Takara, Dalian, China). The transcription level of ZmPLATZ2 in various tissues was measured by semiquantitative RT-PCR and the temporal expression of ZmPLATZ2 in the seed was analyzed by qRT-PCR. For RT-PCR, the reaction mixture was composed of cDNA first-strand template, primer mix, and SYBR Green mix (Takara, Dalian, China). ZmACTIN was used as the internal control. Gene specific primers sets used for qRT-PCR are list in Supplementary Table S1.
Gene cloning and bioinformatics analysis of ZmPLATZ2
ZmPLATZ2 was cloned from the maize inbred line Mo17 based on the B73 reference genome using a primer designed by Primer 5 software. The ZmPLATZ2 gene comprised 930 base pairs (bp) and encodes 309 amino acids. SMART (http://smart.embl-heidelberg.de/) analysis was used to predict the protein domain. PSIPRED analysis (http://bioinf.cs.ucl.ac.uk/psipred/) was used to predict the secondary domain. Primers were list in Supplementary Table S1.
Influence of ABA, sucrose, and glucose treatment on the expression level of ZmPLATZ2
At 9–10 DAP, the maize kernels were surface sterilized with 75% ethanol and split in half under sterile conditions. Murashige and Skoog (MS) liquid medium was used to analyze ZmPLATZ2 expression in the maize endosperm in response to sucrose (200 μM), ABA (100 μM), glucose (200 μM), and 100 μM ABA + 200 μM sucrose, as described previously (Li et al. 2017b; Acevedo-Hernández et al. 2005). The treated samples were harvested at different times postexposure: 0.5, 1, 3, 6, 12, 24, and 48 h, and ZmPLATZ2 expression was analyzed using qRT-PCR.
Subcellular localization
The full-length ORF of ZmPLATZ2 was cloned into the transient expression vector p2300-eGFP at the BamHI/XbaI site using a ClonExpress II one-step cloning kit (Vazyme, Nanjing, China). Onion epidermal cells were peeled and cultured on Hypertonic MS medium for 4 h. A 0.9 µg sample of plasmid per vector was used to coat the gold powder, which was bombarded into the cells by a helium biolistic gun transformation system. The bombarded samples were then cultured for 24 to 48 h at 28 °C, as described in a previous study (Zhang et al. 2016). The samples were observed under BX61 fluorescent microscopy. The p2300-eGFP vector was used as the control. Primers were list in Supplementary Table S1.
Transient expression assays in maize endosperm
For the qualitative transactivation assay, the test vector was generated by inserting the starch synthesis promoter into the PstI and BamHI sites of the pBI221 vector. The ubi-ZmPLATZ2 effector construct was created by cloning its coding sequences into the BamHI and SacI sites. Gus, driven by a ubiquitin promoter, was used as the internal control. For analysis of the promoter activity in the endosperm, the test construct and internal control plasmid were combined at a molar ratio of 2:1. For analysis of the effect of ZmPLATZ2 on the expression of different types of starch synthesis, the molar ratio of the test construct, effector construct, and internal control plasmid was 2:1:1. Endosperm at 9 or 10 DAP were sterilized with 75% (v/v) enthanol and cultivated on MS medium containing 8.5‰ agar and 12% sucrose for 4 h prior to bombardment using a helium biolistic gun transformation system. The bombarded endosperms were cultivated for 24 h to analyze LUC and GUS reporter gene expression. The independent experiments consisting of three replicates were conducted. Primers were list in Supplementary Table S1.
Transcription activation
The yeast transactivation assay, using Yeast two-hybrid system, was performed as described in a previous study (Chen et al. 2016). The full-length ORF of ZmPLATZ2 was cloned into a DNA-binding domain vector (pGBKT7) with NdeI and BamHI sites and transformed into yeast AH109, harboring the LacZ and Trp reporter genes. The empty vector, pGBKT7 was used as the negative control. Primers were list in Supplementary Table S1.
Generation of rice transgenic plants
The ORF of ZmPLATZ2 was cloned into an overexpression vector, pCAMBIA1301 with HindIII and EcoRI restriction enzymes. This recombination construct, pCAMBIA1301-ZmPLATZ2, harbored a Gus reporter gene and overexpressed a full-length cDNA of ZmPLATZ2 driven by ubiquitin promoter and was subsequently transformed into rice using Agrobacterium-mediated transformation. Nipponbare was used as the wide-type (WT) plant and ZmPLATZ2 as the transgenic (TG) plant, which were confirmed by histochemical staining of Gus activity and PCR. Then, qRT-PCR was used to test the expression levels of the starch synthesis genes in the TG and WT rice.
Yeast one-hybrid assay
Yeast one-hybrid assays were performed as described in a previous study (Chen et al. 2016). The full-length ZmPLATZ2 was cloned into the NdeI and BamHI sites of the pGADT7-Rec2 (Takara, Dalian, China) vector. The promoters of Bt2, Sh2, GBSSI, SSI, SSIIa, SSIIIa, SSIV, SSV, ISA1, ISA2, and pullulanase were cloned into pHis2 with RcoRI and SacI using a ClonExpress II one-step cloning kit (Vazyme, Nanjing, China). Primers were list in Table S1.
Protein expression in E. coli and purification
The full-length cDNA sequence of ZmPLATZ2 was cloned into the pET32a vector between the BamHI and HindIII sites to generate His:PLATZ2. Then, the fusion vectors were transformed into BL21(DE3) in E. coli. Bacteria, containing the construct, were grown at 37 °C to OD600 = 0.6, then the fusion protein was induced by 0.5 mM isopropyl b-d-1-thiogalactopyranoside at 16 °C for 8 h. The His-PLATZ2 was purified using Nickle-IDA agarose beads following the manufacturer’s instructions. Primers were list in Table S1.
DAP-seq and DAP-qPCR
The total DNA of Mo17 and purified His-PLATZ2 was used for the DAP assays. The total DNA was broken into 300–500 bp fragments using an ultrasonic crusher. The His fusion protein was purified on Nickle-IDA agarose beads (GenStar, Beijing, China). His-PLATZ2 and DNA fragments were co-incubated for 12 h. The incubation buffer included 50 mM Tris, 1 mM EDTA, and 100 mM KCl, adjusted to a pH of 7.0 by HCl. 5% glycerol, 0.1% Triton X-100, 1 mM DTT. After incubation, the Nickle-IDA agarose beads were washed three to five times using the incubation buffer. Then, 100 µl of 5 M NaCl was added to the bead solution and incubated for 2 h to relieve the crosslinking of Nickle-IDA agarose beads and DNA fragments. The phenol–chloroform method was used to extract the DNA fragments. The fragments were sequenced with a Hiseq2500 system (Novogene, Beijing, China). An empty carrier vector served as the negative control. The DNA fragments prepared in DAP-seq were used for qPCR using SSI-CH1 primer pairs. Three replications were performed for each sample and the input sample served as the negative control in DAP-qPCR.
Results
Co-expression of ZmPLATZ2 with maize starch synthesis genes
Co-expression analysis between starch synthesis related genes and ZmPLATZ2 is shown in Fig. 1. The results shown that SBE1, AGPLS4, Sh2, Bt2, SS1, GBSS1, SS3b-1 and SS2c have a PCC absolute value greater than 0.83 with ZmPLATZ2. The PCC between other genes and ZmPLATZ2 is small than 0.8.
Co-expression analysis of ZmPLATZ2 and starch synthesis related genes based on the expression data of Stelpflug et al. (2016). The vertical distance from gene to Y axis indicates the size of PCC value
Cloning and primary sequence analysis
ZmPLATZ2 was cloned from the maize inbred line Mo17 based on the B73 reference genome. The primary sequence analysis showed that the ZmPLATZ2 gene comprised 894 bp and encoded a 33.679 kDa protein composed of 297 amino acids (Fig. 2a). Analysis using the PSIPRED software revealed that the n-termini of the ZmPLATZ2 protein consists of an α-helix (Fig. 2a). Using domain analysis with SMART software, we found that the ZmPLATZ2 protein included the single conserved PLATZ domain (Fig. 2b). In conclusion, the results confirmed that the ZmPLATZ2 protein belongs to the PLATZ family.
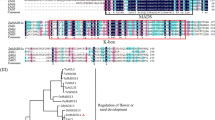
The primary sequence of ZmPLATZ2. a Secondary structure predicted by PSIPRED analysis (http://bioinf.cs.ucl.ac.uk/psipred/). b Conserved domain predicted by SMART analysis (http://smart.embl-heidelberg.de/)
Expression pattern
To determine the expression pattern of ZmPLATZ2, semiquantitative RT-PCR and RT-PCR were performed. The different organs of ZmPLATZ2 expression were examined by semiquantitative RT-PCR. ZmPLATZ2 was expressed mainly in the seed, with higher expression in the endosperm (Fig. 3a). RT-PCR was performed with Actin as the internal reference gene to determine expression pattern of the different stages of ZmPLATZ2. The expression of ZmPLATZ2 was highest at 12 DAP and decreased rapidly post 12 DAP (Fig. 3b).
Analysis of the expression pattern of ZmPLATZ2. a Expression analysis of ZmPLATZ2 in different organs of maize. R root, St stem, L leaf, 10ds 10-day seed, F filament, An anther, Em embryo, En endosperm, SC seed coat. b Expression analysis of ZmPLATZ2 at different stages of seed development. The data represents the mean ± standard deviation (SD) of three biological replicate experiments. c Treatment of glucose. d Treatment of ABA, sucrose, and ABA + sucrose. The treated samples were harvested at different time points postexposure: 0.5, 1, 3, 6, 12, 24, and 48 h. The data are given as the means ± SD of three replicates
To analyze the expression regulation of ZmPLATZ2 in greater detail, a series of treatments were performed. Maize endosperms, collected at 9 or10 DAP, were exposed to glucose (200 μM), sucrose (200 μM), ABA (100 μM), and 100 μM ABA + 200 μM sucrose. The concentration used in each treatment was based on those reported in previous studies (Acevedo et al. 2005; Hu et al. 2011).
As shown in Fig. 3c, ZmPLATZ2 expression was upregulated more than twofold after 0.5 h with glucose treatment and the expression of ZmPLATZ2 decreased by approximately 50% after 6 h when exposed to ABA and sucrose separately, whereas the treatment of ABA together with sucrose remitted this decrease. Therefore, we hypothesized that the expression of ZmPLATZ2 was strongly upregulated by glucose and temporarily downregulated by sucrose, which was complemented by adding ABA.
ZmPLATZ2 binds to the starch synthetic genes promoter
To analyze the interaction of ZmPLATZ2 with the promoters of the genes involved in the synthesis of starch, a yeast one-hybrid experiment was performed. The constructs of pGADT7-ZmPLATZ2 and pHis2, harboring the promoter of the key gene in starch synthesis, were co-transformed into the yeast strain Y187 using. ZmPLATZ2 bound strongly to the promoters of SSI and SSIIa and showed binding to the promoters of SSIV, ISA2, and SSV (Fig. 4). Therefore, ZmPLATZ2 can bind to the promoters of SSI, SSIIa, SSIV, ISA2, and SSV genes and may regulate their transcription.
Transient expression of ZmPLATZ2 in maize endosperm
Transient expression analysis was undertaken to analyze the regulatory role of the ZmPLATZ2 TFs in starch synthesis gene expression. Luc/(4-0) H Gus was used to detect whether ZmPLATZ2 affected promoter activity. The promoter activities of the four starch synthetic genes were significantly upregulated by at least a two-fold threshold (Fig. 5). These upregulated genes included Sh2 (AGPase large subunit), Bt2 (AGPase small subunit), SSI (starch synthesis I), and SSIIIa (starch synthesis IIIa) which was more than twofold. Meanwhile, WX, ISA1, and ISA2 promoter activities were significantly upregulated by ZmPLATZ2, although these were less than two-fold.
ZmPLATZ2 promotes the promoter activities of starch synthesis genes. a Schematic of the vector constructs used for transient expression. b–j The results of the transient expression in maize endosperms. The data are reported as the means ± standard errors (SEs) of three replicates. A one-sided paired t-test (**P < 0.01) was used to analyze the significance of the differences between with and without effector vectors
Transcription factor characterization
PLATZ2 is a novel class of plant-specific zinc-dependent DNA-binding TF; therefore, how it regulates the downstream processes was investigated via the analysis of the subcellular location of ZmPLATZ2 by the transient expression of the ZmPLATZ2-GFP protein in onion epidermal cells. The results showed that PLATZ2 was localized in the nuclei, which is consistent with the role of ZmPLATZ2 being a TF (Fig. 6d).
Cellular localization and transcription activity assays of ZmPLATZ2 in yeast strain AH109. a Schematic diagram of the yeast two-hybrid assay. b Yeast screened on SD/–Trp medium in accordance with the structure outlined in A; Yeast screened on SD/–Trp–His–Ura medium with X-α-gal in accordance with the structure outlined in A. c Schematic diagram of the subcellular localization assay. d Results of ZmPLATZ2-eGFP and 35S-eGFP under GFP field, bright, and in the merged image
ZmPLATZ2 localizes to the nucleus and acts as a transcription factor. To examine its transcription activity, a fused vector, pGBKT7-ZmPLATZ2, was generated and transformed into the yeast strain AH109, with the transformants of pGBKT7, pGBKT7-Lam, and pGBKT7-53 as the negative controls and pGBKT7-Gal as the positive control. Each of the transformants was confirmed by PCR. All the yeast cells grew well on the SD/–Trp medium. Only the positive control had α-galactosidase activity on the SD/–Trp/–His/–Ura/X-α-GAL medium (Fig. 6b); therefore, ZmPLATZ2 had no transcription activity in the yeast cells.
Analysis of ZmPLATZ2 influences the starch synthesis gene in transgenic rice
To further study the effect of ZmPLATZ2 on starch synthesis in vivo, overexpressing rice plants were obtained using Agrobacterium-mediated transformation and a stable overexpression line was obtained. The transgenic rice was confirmed by Gus staining (Fig. 7a), DNA detection (Fig. 7b), and transcript detection (Fig. 7c). The main starch synthesis genes were measured in transgenic rice using qRT-PCR. The expression levels of SSI, BEIIb, BT1, SSIIIa, and Wx were significantly upregulated 3- to 10-fold in TG rice compared to those in WT rice (Fig. 7d). These results indicate that ZmPLATZ2 could regulate the starch synthesis in rice.
Overexpression of ZmPLATZ2 in rice. a Gus staining analysis of transgenic (TG) and wild-type (WT). b DNA detection of transformed plants. c Transcript detection of transformed plants. d qRT-PCR analysis of starch synthesis expression in WT and TG rice. The data are reported as the means ± SEs of four replicates. A one-sided paired t-test (**P < 0.01) was used to analyze the significance of the differences between TG and WT
DAP-Seq identifies genomic sites are directly bound by ZmPLATZ2
To assess the target genes of the ZmPLATZ2 protein, we performed a DAP-seq experiment using His antibodies against His-tagged ZmPLATZ2 protein (Fig. 8h) and chromatin extracted from Mo17 (Fig. 8i). The empty vector was used as the negative control.
Profiles of ChIP-seq analysis. a Distribution of ZmPLATZ2 binding regions in the maize genome. b Peak width of ZmPLATZ2 in the ChIP-seq. c Distribution of ZmPLATZ2 binding sites corresponding to the − 5000 to + 3000 bp regions flanking the TSS. d GO classification of DEGs between ZmPLATZ2 and PET32a. e The most prominent motif was identified as ZmPLATZ2 binding motif. f In vitro pull down of target DNA by ZmPLATZ2. His-ZmPLATZ2 or His alone were incubated with DNA for 12 h, pulled down, and qPCR was performed for the ZmSSI gene. g OsSSI and ZmSSI has the same CAAAAAC/GTTTTTG motif in promoter
The results of the distribution of binding peaks in the ZmPLATZ2 DAP-seq experiment showed that approximately 91% of the ZmPLATZ2 binding sequences were located in the intergenic. Of these, 7% were located in the promoter regions (− 2000 bp to the TSS) (Fig. 8a). We analyzed the peak width and distance between each peak summit and the TSS of its nearest gene to survey the promoter regions of ZmPLATZ2 binding. The peak widths of ZmPLATZ2 were mainly concentrated in the 300 to 400 bp region (Fig. 8b) and there was a peak in the gene promoter regions (Fig. 8c). Therefore, the ZmPLATZ2 peaks were highly focused in the 300 to 400 bp region, upstream of the TSSs.
By GO (Gene Ontology) classification of DEGs (Differentially Expressed Gene), we found that ZmPLATZ2 was mostly bound to the nucleic acid activity and cellular nitrogen compound biosynthesis (Fig. 8d). CAAAAAAA was identified as a statistically defined motif (P-value = 1E-138; Fig. 8e). We identified the same motif with DAP-seq results from the promoter of ZmSSI, and then DAP-qPCR showed that the ZmSSI promoter was pulled down 15-fold compared to the control (Fig. 8f). Meanwhile, the promoter of OsSSI and ZmSSI has the same CAAAAAC/GTTTTTG motif in same site of promoter (Fig. 8g). These results suggested that CAAAAAAA was a highly conserved binding motif for ZmPLATZ2.
Discussion
PLATZ1 is the first reported member of the PLATZ TF family, which was identified from the pea (Pisum sativum) (Nagano et al. 2001). The first reported member in maize was ZmPLATZ12(fl3), which was specifically expressed in the maize endosperm starchy cell and regulated many tRNAs and 5S RNA by interactions with RPC53 (Wang et al. 2018a, b).
Semiquantitative RT-PCR showed that ZmPLATZ2 was mainly expressed in the endosperm and further series experiments showed that ZmPLATZ2 harbored the PLATZ domain and was located in the nucleus. These results showed that ZmPLATZ2 performs its function in the endosperm as a TF. However, no transcription activity was found in ZmPLATZ2. Thus, ZmPLATZ2 interacted with other proteins to regulate the target gene, such as ZmPLATZ12 (fl3). ZmPLATZ proteins are usually involved in the adjustment of RNAPIII transcription, whereas ZmPLATZ2 could not interact with RPC53 or TFC1, which is similar to most of the ZmPLATZ proteins (Wang et al. 2018a, b). We attempted to discover the key protein that interacted with ZmPLATZ2 using yeast two-hybrid assays, but no results were obtained.
Yeast one-hybrid experiment of the ZmPLATZ2 bound promoter of starch synthesis genes revealed centrally that SSIIa, SSI, SSIV, ISA2, and SSV promoters had a ZmPLATZ2 binding motif. Our analysis of the transient expression in endosperm supported the role of regulating the Sh2, Bt2, SSI, and SSIIIa genes. Therefore, we speculate that ZmPLATZ2 can bind to the ZmSSI promoter, and thus promote ZmSSI expression. The analysis of the starch synthesis gene expression in TG rice revealed that the OsSSI gene was significantly upregulated. Meanwhile, the transcript levels of the OsSBEIIb, OsSSIIIa, and OsWX genes had markedly higher overexpression in ZmPLATZ2 TG rice than that of WT rice. The same effect has been reported for the overexpression of ZmbZIP22 in maize and rice (Dong et al. 2019); therefore, these results in rice are credible. SSI has no isoforms in rice, unlike the other types of SS (Fujita et al. 2003); therefore, SSI might play an important role in starch synthesis. Meanwhile, we found that the promoter of OsSSI and ZmSSI has the same CAAAAAC/GTTTTTG motif with the same flanking sequence and the distance of the motif to TSS is 157 bp in rice and 115 bp. Therefore, ZmPLATZ2 could bind to the same highly conservative motif in maize and rice. The mechanism by which ZmPLATZ2 regulates starch synthesis is different from that of the other PLATZ proteins bound with RPC53 or TFC1. Therefore, ZmPLATZ2 might regulate the downstream gene as a TF by binding with the promoter (Supplementary Table S2).
Transcription factor participate in hormone signal pathway (Lian et al. 2020; Chen et al. 2019). Different treatments of endosperms showed that ZmPLATZ2 was positively induced by glucose and negatively regulated by ABA and sucrose. In the present study, we found that the glucose-induced ZmPLATZ2 acted as a positive regulator of starch synthesis. ZmPLATZ2 was downregulated by ABA and sucrose, whereas the expression of ZmPLATZ2 after treatments was the same with CK. However, further studies are required to determine the mechanism underlying ZmPLATZ2 induced by glucose and the role that ZmPLATZ2 plays in the ABA and sucrose pathways.
Starch synthesis is crucial for grain weight. Many previous studies have shown that hormones can induce gene expression by regulating the TF. For example, ZmABI4 could bind to the CACCG motif of ZmSSI, and thus mediate the ABA pathway (Hu et al. 2012). In conclusion, we identified a new gene, ZmPLATZ2, which functions in the SSI transcription machinery mediated in glucose. In addition, we observed that ZmPLATZ2 is induced by glucose and binds to the CAAAAAC/GTTTTTG motif of ZmSSI to promote expression. Although the detailed mechanism as to how ZmPLATZ2 regulates the expression of the starch synthesis genes and what role it plays in the ABA and sucrose pathways remains unclear, our data contribute toward the discovery of this process. Therefore, the exploitation of ZmPLATZ2 is a potential approach for regulating the expression of ZmSSI involved in the glucose pathway.
Abbreviations
- AGPase:
-
Adenosine diphosphate glucose pyrophosphorylase
- DAP:
-
Days after pollination
- DAP-qPCR:
-
DNA affinity purification sequencing-quantitative real-time polymerase chain reaction
- DAP-seq:
-
DNA affinity purification sequencing
- DBE:
-
Starch debranching enzyme
- DEGs:
-
Differentially expressed genes
- GO:
-
Gene ontology
- MS:
-
Murashige and Skoog medium
- SBE:
-
Starch branching enzyme
- SS:
-
Starch synthase
- TFs:
-
Transcription factors
- TG:
-
Transgenic
- WT:
-
Wide-type
References
Acevedo-Hernández GJ, León P, Herrera-Estrella LR (2005) Sugar and ABA responsiveness of a minimal RBCS light-responsive unit is mediated by direct binding of ABI4. Plant J 43:506–519. https://doi.org/10.1111/j.1365-313X.2005.02468.x
Bennett RL, Swaroop A, Troche C, Licht JD (2017) The role of nuclear receptor-binding SET domain family histone lysine methyltransferases in cancer. Cold Spring Harbor Perspect Med 7:a026708. https://doi.org/10.1101/cshperspect.a026708
Chen J, Yi Q, Cao Y et al (2016) ZmbZIP91 regulates expression of starch synthesis-related genes by binding to ACTCAT elements in their promoters. J Exp Bot 67:1327–1338. https://doi.org/10.1101/10.1093/jxb/erv527
Chen Z, Liu Y, Yin Y et al (2019) Expression of AtGA2ox1 enhances drought tolerance in maize. Plant Growth Regul 89:203–215. https://doi.org/10.1007/s10725-019-00526-x
Crofts N, Nakamura Y, Fujita N (2017) Critical and speculative review of the roles of multi-protein complexes in starch biosynthesis in cereals. Plant Sci 262:1–8. https://doi.org/10.1016/j.plantsci.2017.05.007
Dong Q, Xu QQ, Kong JJ et al (2019) Overexpression of ZmbZIP22 gene alters endosperm starch content and composition in maize and rice. Plant Sci 283:407–415. https://doi.org/10.1016/j.plantsci.2019.03.001
Fujita N (2003) Antisense inhibition of isoamylase alters the structure of amylopectin and the physicochemical properties of starch in rice endosperm. Plant Cell Physiol 44:607–618. https://doi.org/10.1093/pcp/pcg079
Godfray HC, Beddington JR, Crute IR, Haddad L, Lawrence D, Muir JF, Pretty J, Robinson S, Thomas SM, Toulmin C (2010) Food security: the challenge of feeding 9 billion people. Science 327:812–818. https://doi.org/10.1126/science.1185383
Hennen-Bierwagen TA, Liu F, Marsh RS et al (2008) Starch biosynthetic enzymes from developing maize endosperm associate in multisubunit complexes. Plant Physiol 146:1892–1908. https://doi.org/10.1104/pp.108.116285
Hu YF, Li YP, Zhang JJ et al (2011) PzsS3a, a novel endosperm specific promoter from maize (Zea mays L.) induced by ABA. Biotechnol Lett 33:1465–1471. https://doi.org/10.1007/s10529-011-0582-z
Hu YF, Li YP, Zhang JJ, Liu HM, Tian ML, Huang YB (2012) Binding of ABI4 to a CACCG motif mediates the ABA-induced expression of the ZmSSI gene in maize (Zea mays L.) endosperm. J Exp Bot 63:5979–5989. https://doi.org/10.1093/jxb/ers246
Huang B, Hennen-Bierwagen TA, Myers AM (2014) Functions of Multiple Genes Encoding ADP-glucose pyrophosphorylase subunits in maize endosperm, embryo, and leaf. Plant Physiol 164(2):596–611. https://doi.org/10.1104/pp.113.231605
Huang HH, Xie SD, Xiao QL et al (2016) Sucrose and ABA regulate starch biosynthesis in maize through a novel transcription factor. ZmEREB156. Sci Rep 6:27590. https://doi.org/10.1007/10.1038/srep27590
Kim JH, Kim J, Jun SE et al (2018) ORESARA15, a PLATZ transcription factor, mediates leaf growth and senescence in Arabidopsis. New Phytol 220:609–623. https://doi.org/10.1007/10.1111/nph.15291
Leterrier M, Holappa LD, Broglie KE, Beckles DM (2008) Cloning, characterisation and comparative analysis of a starch synthase IV gene in wheat: functional and evolutionary implications. BMC Plant Biol 8:98. https://doi.org/10.1007/10.1186/1471-2229-8-98
Li L, Jiang H, Campbell M et al (2008) Characterization of maize amylose-extender (ae) mutant starches. Part I: Relationship between resistant starch contents and molecular structures. Carbohydr Polym 74:396–404. https://doi.org/10.1016/j.carbpol.2008.03.012
Li Q, Wang J, Ye J, Zheng XX, Xiang XL, Li CS, Fu MM, Wang Q, Zhang ZY, Wu YR (2017a) The maize imprinted gene floury3 encodes a PLATZ protein required for tRNA and 5S rRNA transcription through interaction with RNA polymerase III. Plant Cell 29:2661–2675. https://doi.org/10.1007/10.1186/10.1105/tpc.17.00576
Li H, Xiao QL, Zhang CX et al (2017b) Identification and characterization of transcription factor ZmEREB94 involved in starch synthesis in maize. J Plant Physiol 216:11–16
Li JJ, Shi T, Huang LY, He DL et al (2018) Systematic transcriptomic analysis provides insights into lotus (Nelumbo nucifera) seed development. Plant Growth Regul 86:339–350. https://doi.org/10.1007/s10725-018-0433-1
Lian L, Xu H, Zhang H et al (2020) Overexpression of OsSPL14 results in transcriptome and physiology changes in indica rice ‘MH86.’ Plant Growth Regul 90:265–278. https://doi.org/10.1007/s10725-019-00569-0
Nagano Y, Furuhashi H, Inaba T, Sasaki Y (2001) A novel class of plant-specific zinc-dependent DNA-binding protein that binds to A/T-rich DNA sequences. Nucleic Acids Res 29:4097–4105. https://doi.org/10.1093/nar/29.20.4097
Reddy CK, Choi SM, Lee DJ, Lim ST (2018) Complex formation between starch and stearic acid: effect of enzymatic debranching for starch. Food Chem 244:136–142. https://doi.org/10.1016/j.foodchem.2017.10.040
Stelpflug SC, Sekhon RS, Vaillancourt B, Hirsch CN, Buell CR, Leon ND, Kaeppler SM (2016) An expanded maize gene expression atlas based on RNA sequencing and its use to explore root development. Plant Genome 9:1. https://doi.org/10.3835/plantgenome2015.04.0025
Wang Y, Li Y, Zhang H, Zhai H, Liu QC, He SZ (2017) A soluble starch synthase I gene, IbSSI, alters the content, composition, granule size and structure of starch in transgenic sweet potato. Sci Rep 7:2315. https://doi.org/10.1038/s41598-017-02481-x
Wang J, Hu P, Lin L, Chen ZC, Liu QQ, Wei CX (2018a) Gradually decreasing starch branching enzyme expression is responsible for the formation of heterogeneous starch granules. Plant Physiol 176:582–595. https://doi.org/10.1104/pp.17.01013
Wang JC, Ji C, Li Q, Zhou Y, Wu YR (2018b) Genome-wide analysis of the plant-specific PLATZ proteins in maize and identification of their general role in interaction with RNA polymerase III complex. BMC Plant Biol 18:221. https://doi.org/10.1186/s12870-018-1443-x
Wang AH, Hou QQ, Si LZ et al (2019) The PLATZ transcription factor GL6 affects grain length and number in rice. Plant Physiol 180:2077–2090. https://doi.org/10.1104/pp.18.01574
Xiao QL, Wang YY, Du J et al (2017) ZmMYB14 is an important transcription factor involved in the regulation of the activity of the ZmBT1 promoter in starch biosynthesis in maize. Febs J 284:3079–3099. https://doi.org/10.1111/febs.14179
Zhang JJ, Zhang SS, Li H, Du H, Huang HH, Li YP, Hu YF, Liu HM, Liu YH, Yu GW (2016) Identification of transcription factors ZmMYB111 and ZmMYB148 involved in phenylpropanoid metabolism. Front Plant Sci. https://doi.org/10.3389/fpls.2016.00148
Zhang SC, Yang R, Huo YQ, Liu SS, Yang GD, Huang JG, Zheng CC, Wu CG (2018) Expression of cotton PLATZ1 in transgenic Arabidopsis reduces sensitivity to osmotic and salt stress for germination and seedling establishment associated with modification of the abscisic acid, gibberellin, and ethylene signalling pathways. Plant Biol 18:218. https://doi.org/10.1073/10.1186/s12870-018-1416-0
Zhou H, Wang L, Liu G et al (2016) Critical roles of soluble starch synthase SSIIIa and granule-bound starch synthase Waxy in synthesizing resistant starch in rice. Proc Natl Acad Sci USA 113:12844–12849. https://doi.org/10.1073/pnas.1615104113
Acknowledgements
Y.H., J.Z., H.L., and Y.W designed the study. H.L. and Y.W. performed the experiments. L.L., C.M., J.D., T.L., Y.L., and C.Z. analyzed the data. Y.H., Q.X. J.Z., Y.C., Q.Y., Y.W., Y.L., H.H., H.L., Y.H., G.Y., and H.L. prepared and revised the manuscript and all the authors read and approved the manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (Grant No: 31771702) and National Major Project for Transgenic Organism Breeding (Grant No: 2016ZX08003-001).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by J. Celenza
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, H., Wang, Y., Xiao, Q. et al. Transcription factor ZmPLATZ2 positively regulate the starch synthesis in maize. Plant Growth Regul 93, 291–302 (2021). https://doi.org/10.1007/s10725-020-00687-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-020-00687-0