Abstract
MADS-box family proteins play an important role in grain formation and flower development; however, the molecular mechanisms by which transcription factors regulate the starch metabolism pathway are unclear in maize. Here, we report a transcription factor, ZmMADS1a, that controls starch biosynthesis in maize (Zea mays L.). We demonstrate the expression of ZmMADS1a in tassel, silk, and endosperm, and show that the protein is localized to the cell nucleus. Compared with the control, seeds of overexpressing ZmMADS1a increased starch content (especially amylose content), had smaller starch granules and altered chemical structure. Meanwhile, overexpression of ZmMADS1a resulted in increases in the contents of soluble sugars and reducing sugars in maize. ZmMADS1a plays a positive regulatory role in the starch biosynthesis pathway by up-regulating several starch biosynthesis related genes. We also show that ZmMADS1a has a similar adjustment mechanism of starch biosynthesis in rice. Collectively, our study suggests that ZmMADS1a functions as a positive regulator of starch biosynthesis by regulating the expression of key starch metabolism genes during seed development.
Similar content being viewed by others
Introduction
Based on domain structure, rates of evolution, developmental function, and the degree of functional redundancy, the MADS-box transcription factor family can be divided into two types of MADS-box proteins; Type I (the SRF superfamily) and Type II (the MEF2 superfamily), which contain three additional sequence domains and differ in their DNA binding properties compared to Type I1. The biological function of Type I proteins, which are weakly expressed in all species’ tissues, may participate in male and male gametophyte development and early development of the embryo and endosperm2,3,4; Four Arabidopsis of Type I genes have been cloned, including AGL23, AGL28, AGL61, and AGL62, which play important roles in embryonic development, flowering time, female gamete differentiation, and formation of the endosperm cells, respectively5,6,7,8. Studies describing the biological function of Type II genes are fairly common, and these proteins are known to participate in different flowering regulation pathways which promote or delay flowering time by regulating the expression of genes involved in flower morphological differentiation, and they also participate in the regulation of fruit and seed development9. Type II MADS-box domain usually have only one exon10 and, similar to the bZIP transcription factors, can form dimers11. Dimers of MADS-box transcription factors can bind to CArG-boxes, conserved DNA elements with the consensus sequence 5′-CC[A/T]6GG-3′12, which are recognized by the MADS-box domain.
Functional studies in maize and rice have shown that Type II MADS-box genes play important roles in the processes of grain formation and flower development. In maize, the bearded-ear gene can regulate the normal development of the corn flower13, the ZmMADS1 and ZmMADS3 genes affect the growth of the maize ear14, and ZmMADS2 and ZmMADS4 regulate the development of the inflorescence and the formation of pollen, respectively15. Loss of the ZmMADS19 protein leads to significantly larger glumes and androgynous corn16; the ZmMADS47 gene can regulate maize prolamine directly to affect endosperm storage activity by interactions with Opaque217. Type II MADS-box genes have also been well studied in rice. OsMADS57 plays a negative regulatory role in regulating the expression of DWARF14 (D14) through the interaction of miR444 with OsTb1 in rice shoots18; RNAi silencing of the OsMADS29 gene can lead to the severe shrunken seed phenotype, and this regulatory factor can also bind targets directly involved in programmed cell death (PCD)19.
In our study, we identified and characterized the ZmMAD1a gene, which encodes a protein that belongs to the MIKCC-group of TypeII. MADS-box proteins. MIKCc-group genes control various aspects of sporophyte development20,21. The MIKCc-group genes are classified by phylogenetic analysis into 12 major subfamilies that include AG, AGL2, AGL6, AGL12, AGL15, AGL17, AP3/PI, GGM13, STMADS11, TM3, AP1/SQUA, and FLOWERING LOCUS C (FLC). Expression of some MIKCC-group genes from rice, tomato, and A. thaliana, among others, is affected by stress treatment, indicating that they are involved in regulating flowering time in response to stress22. For instance, the AGL12 gene affects flowering time in A. thaliana23, and GmNMH7 is expressed in different periods of flower development in soybean, and expression in root nodules was also found to be controlled by photoperiod24. The OsMADS29 gene is associated with the degradation of the nucellus and nucellus swelling in rice19. Some genes in the MIKCc-group have been cloned such as ZAG1, ZAG2, ZAG3-ZAG5, and ZMM3-ZMM825. ZAG3 is expressed in the inflorescence and can affect the growth of the meristem in maize13. These genes are especially prominent on almost all levels of the gene regulatory network that controls reproductive development in flowering plants, but seed growth regulation has not been extensively studied.
In the present study, we identified and characterized the ZmMAD1a gene, which encodes a MADS-box transcription factor. Overexpression of the ZmMAD1a gene resulted in changes to the structures and contents of starch, and changes in sugar content, as well as changes in the expression levels of starch biosynthesis genes in maize and rice. Our study will clarify the mechanism underlying the molecular regulation of starch synthesis by ZmMAD1a, and will lay the foundation for further analyses of the starch metabolic pathways in maize.
Results
ZmMADS1a structural characteristics and gene expression analyses
ZmMADS1a is predicted to encode a polypeptide of 269 amino acids with a calculated molecular weight of 30.23 kDa (Fig. 1I). Like other MADS-box family proteins, ZmMADS1a contains the MADS box (2–58aa) and K box (84–172aa) domains at the N-terminus that are key to the functions of MADS-box proteins. Sequence comparisons indicate that ZmMADS1a codes for a putative Type II MIKCC transcription factor. The protein also has a MADS_MEF2 like (Myocyte Enhance Factor 2-like) domain which can bind DNA and exist as hetero- and homo-dimers, and has an important effect in homeotic regulation in plants. The protein also has an ARG80 region which is a regulator of arginine metabolism and related MADS-box-containing transcription factors (Fig. 1I). The deduced amino acid sequences were used in BLAST searches and the best hits, four AG-like genes from Zea mays (ZAG1, ZAG2, ZAG3, ZAG5), are highly conserved with respect to their MADS and K domains (Fig. 1II). Phylogenetic analysis indicates that ZmMADS1a is closely related to ZAG2 (zag2), ZmMADS1 and other proteins that function in the regulation of flower and seed development (Fig. 1III). The functional properties of ZmMADS1a were analysed, and the results show that ZmMADS1a is localized to the cell nucleus (Fig. 2).
Protein sequence analysis and alignment of ZmMADS1a with other maize MADS-box proteins. (I) Protein structure analysis of ZmMADS1a. Sequence domains are MADS (2–58 aa), MCM1, Agamous, Deficiens, and SRF; MADS_MEF2_like (2–73 aa), MEF2-like/Type II subfamily of MADS box family of eukaryotic transcriptional regulators; K-box (84–172aa), which is commonly found associated with SRF-type transcription factors. (II) Homology analysis of amino acid sequences. ZmMADS1a was aligned with ZAG1, GRMZM2G052890_P01; ZAG2 GRMZM2G160687_P03; ZAG3, GRMZM2G160565_P01; ZAG5, GRMZM2G003514_P01. Different colors represent different conservativeness. The black box denotes the MADS box, the red box represents K-box. (III) Phylogenetic analysis based on an alignment of the ZmMADS1a sequence with MADS-box proteins from other plant species. The ZmMADS1a is indicated by the red triangle. The accession numbers are listed as follows: TaAGL2, ABF57921; TaWM3B, CAM59043; TuMADS13, EMS56192; TaMAF1, AHA51692; TaAGL31, AHA49652; TaWM3A, CAM59042; OsMADS13, AF151693_1; PeMADS13, AHM92069; SiMADS13, XP_004960786; SbMADS13, XP_002441989.1; ZAG2, XP_020405641; ZMM1, CAA57073; ZmMADS1, AQK40454; MuMADS1, DQ060444; ZAG1, NP_001105321.1; ZmMADS2, AAO85643;ZAG3, NP_001105332.1; ZAG5, NP_001105692.2; CaRIN, DQ999998; LeMADS-RIN, AF448522_1; FaMADS9, AF484683; VvMADS39, XM_002263003.
To understand the expression pattern of ZmMADS1a, we performed quantitative qPCR using RNA extracted from tissues at different developmental stages. ZmMADS1a was found to be expressed at a relatively high level in the tassel, silk, and endosperm (Fig. 3A). We sampled embryos and endosperm every two days from 6- to 20-DAP (days after pollination). We found that the relative expression of ZmMADS1a increased gradually in the endosperm until 14-DAP, after which it increased by 3-fold at 16-DAP, peaked at 18-DAP, and then decreased to approximately the 10-DAP level in 20-DAP endosperm (Fig. 3B).
Expression analysis of ZmMADS1a in maize by qRT-PCR. (A) Expression of ZmMADS1a in eight different maize tissues (B) Expression of ZmMADS1a in embryo (Em) and endosperm (En) at eight stages of seed development. R, root; S, stem; L, leaf; T, tassel; C, comsilk, S, stegophyll; En6, 6-DAP endosperm; Em6, 6-DAP embryo; En8, 8-DAP endosperm; Em8, 8-DAP embryo; En10, 10-DAP endosperm; Em10, 10-DAP embryo; En12, 12-DAP endosperm; Em12, 12-DAP embryo; En14, 14-DAP endosperm; Em14, 14-DAP embryo; En16, 16-DAP endosperm; Em16, 16-DAP embryo; En18, 18-DAP endosperm; Em18, 18-DAP embryo; En20, 20-DAP endosperm; Em20, 20-DAP embryo. All data are presented as mean ± SD from three replicates. Relative expression was calculated by setting the expression of ZmMADS1a in the stem as 1.0.
Overexpression of ZmMADS1a regulates starch metabolism genes
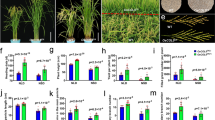
To investigate the function of ZmMADS1a at the whole plant level, we generated a transformation vector containing the full-length ZmMADS1a sequence driven by the CaMV 35S promoter (Pro35S:ZmMADS1a), and transformed the maize line ‘ZZC01’. Sixteen starch biosynthesis genes were selected for analysis of gene expression levels in the maize endosperm between the transgenic ZmMADS1a-overexpressing lines and the wild-type at different days after pollination. Transcription of ZmAGPL1, ZmAGPS1a, ZmBEIIb, ZmGBSSIa (wx), ZmISA1, and ZmSSIIa showed significant increases in the T3-generation transgenic lines. The expression levels of ZmAGPL2, ZmBEI, and ZmBEIIa genes were lower than the wild-type in the whole stage. The gene ZmAGPL2 and ZmBEI were down-regulated at 15d. The expression of ZmPUL and ZmGBSSIIa was low at 20 DAP but increased rapidly at 25 DAP, while ZmGBSSIIb, ZmISA2 and ZmSSI showed almost no change in expression over the course of the experiment compared to wild-type (Fig. 4). We also transformed the Pro35S:ZmMADS1a construct into the japonica rice cultivar ‘Zhonghua 11’. Transcription of OsAGPS2b, OsAGPS1, OsBEIIb, OsISA1, and OsISA2 was significantly increased in the T3-generation transgenic lines. The expression level of OsGBSSIa (wx) increased rapidly after 15 DAP, peaked at 20 DAP, and then fell to the control level by 25 DAP. The expression levels of OsAGPL2, OsBEI and OsSSIIIa were significantly down-regulated compared to the control. OsAGPL1 and OsSSI showed almost no change in relative expression from 3 to 25 DAP compared to the wild-type control (Fig. S1). Comparing the gene expression levels of ZmMADS1a in maize and rice, we found that many of the gene homologs (BEIIb, ISA1, SSIIa, GBSSIa, AGPL2, BEI, PUL, and SSI) had similar expression patterns, and have an important influence on the regulation of starch biosynthesis in both species. This indicates that the ZmMADS1a gene is extensively involved in the regulation of starch and glucose metabolism in maize endosperm.
Ectopic expression of ZmMADSla results in grain morphology
Nineteen maize T3 lines were randomly selected for the analysis of grain morphology. Panicle length, panicle width, panicle circumference, and 100-seed weight showed no obvious changes in ZmMADSla-overexpressing transgenic maize plants compared to the wild-type ‘ZZC01’ (Fig. S2). We further field tested the homozygous T3 rice lines and compared them with wild type (‘Zhonghua 11’) and the empty vector (1301a) transgenic plants. Seeds of all of the ZmMADSla transfomants were slightly longer than the controls (Fig. S3I,III), but there were no significant changes in the 1000-seed weights (Fig. S3II). We also compared other phenotypes in the maize and rice T3 families. In maize, both plant height and ear height were reduced compared to wild-type plants, but leaf length and leaf width were not significantly different (Tables S4, S5). These results indicate that overexpression of the ZmMADSla gene has no significant effects on grain morphology.
Effects of ectopic expression of ZmMADS1a on starch granule size and structure
To investigate the possible effects of ZmMADS1a overexpression on starch granules in the endosperm, transverse sections of seeds were observed using scanning electron microscopy (SEM). Three regions were compared in the maize seed sections (Fig. 5). No changes were observed in the first region due to the compacted starch granules (Fig. 5a). The second region from the three lines showed noticeable changes in granule size, with the granules from the transgenic seeds being smaller than in the wild-type (Fig. 5IIb,IIIb), while granules from the third region of the endosperm were only slightly smaller than in wild-type (Fig. 5IIc,IIIc).
Comparisons of starch granule size and structure in maize endosperm. WT, wild-type plants (‘ZZC01’); Om-1, Om-6, and Om-18 are three ZmMADS1a-overexpressing maize lines; a-c, position of endosperm; (I), Bar = 1 mm; (II), Bar = 30 μm; (III) Starch granules size of b and c. To investigate the possible effects of ZmMADS1a overexpression on starch granules in the endosperm, transverse sections of seeds from T3-generation plants were observed by scanning electron microscopy (SEM). Three regions were compared in the maize seed sections. No changes were observed in the first region due to the compacted granules (a). There were obvious changes in the second regions in the three lines, with the granules from the transgenic seeds being smaller than in wild-type (b). The third areas of the seeds from the three lines showed only slight changes in granule size compared to wild-type (c). The asterisks indicate that the correlation coefficients were highly significantly different (*P < 0.05, **P < 0.01).Values represent the mean ± SD of triplicates.
The 13C {1H} CP/MAS spectra for starch granules isolated from rice and maize endosperm are shown in Fig. S4. Signals from 94–105 ppm were assigned to C1, signals from 68–78 ppm were assigned to C2, C3, and C5, signals at 58–65 ppm were associated with C6, and the signal at 82.0 was assigned to C4. The chemical shifts of each major peak are presented in Tables S7, S9. From Fig. S4I we can see that all of the chemical shifts in the ZmMADS1a-overexpressing maize lines are moved to the left about 0.5 ppm compared with wild-type. To calculate peak area in maize, the peak area from 52.1–112 was taken as 100%, and was also divided into four parts (C1, 112–88.5 ppm; C4, 88.5–78.4 ppm; C2, C3, C5, 78.4–65.1 ppm; C6, 65.1–52.1 ppm). Compared with the wild-type, the peak areas of C1 and C4 increased slightly, the area of C6 decreased slightly, and the area of the C2, C3, C5 peak was almost unchanged in the ZmMADS1a-overexpressing maize lines (Table S8). The variation in peak areas showed similar results in rice (Fig. S4II, Table S10). These results indicate that overexpression of the ZmMADS1a gene can change the chemical structure of starch in both maize and rice.
Overexpression of ZmMADS1a improves starch and sugar contents
To understand the effects of overexpressing ZmMADS1a on starch synthesis, we analyzed the starch content and related biochemical indicators in T3-generation transgenic maize seeds. The total starch, amylose, and amylopectin contents in the four ZmMADS1a-overexpressing maize lines were increased (Fig. 6a–c), and the expression level of the four lines were increased (Fig. S5). In these transgenic lines, the increases in total starch and amylopectin contents were 3.26% and 5.08% respectively, while the relative increase in the amylose content was considerably higher than wild-type, up to 11.42%. Comparing the physiological and biochemical characteristics of maize endosperm in the ZmMADSla-overexpressing lines with wild-type, we found that the contents of soluble sugars and reducing sugars were significantly increased, but that the crude protein, crude fat, and carbon contents were not significantly different than in the control line (Fig. 7, Table S6). We also compared the rice ZmMADSla-overexpressing T4-generation seeds with the non-transgenic control. The results showed that the total starch and amylose contents followed similar trends in maize and rice, especially for amylose content, which had a maximum increase of 26.62% in the transgenic rice lines Or-8 and Or-12 (Figs 6 and S6). We then examined the maize seeds under transmitted light, which showed that the shaded areas of the kernels from the ZmMADS1a-overexpressing lines were slightly larger than in wild-type seeds (Fig. 6D). All of the above results indicate that overexpressing the ZmMADS1a gene can affect starch (especially amylose) and sugar contents in the endosperm.
Endosperm starch content in seeds of ZmMADSla-overexpressing transgenic maize lines and wild-type. (A) total starch content; (B) amylose content; (C) amylopectin content; (D) light transmission; Om-1, Om-6, Om-12, and Om-18 are four overexpressing lines of ZmMADSla seeds; WT, wild-type plants (‘ZZC01’). The dashed blue lines indicate the average for the overexpressing lines. The asterisks indicate that the correlation coefficients were highly significantly different (*P < 0.05, **P < 0.01).Values represent the mean ± SD of triplicates.
Physiological and biochemical characteristics of endosperm in ZmMADSla-overexpressing maize lines and wild-type. CP, Crude protein content; CF, Crude fat content; C, Carbon content; SS, Reducing sugar content; RS, Soluble sugar content; WT, wild-type plants (‘ZZC01’). Om-1, Om-6, Om-12, and Om-18 are four ZmMADSla-overexpressing lines. The dashed blue lines indicate the average values in the overexpressing lines. The asterisks indicate that the correlation coefficients were highly significantly different (*P < 0.05, **P < 0.01).Values represent the mean ± SD of triplicates.
Discussion
ZmMADS1a, a typical MADS-box transcription factor, belongs to the MIKCc-subgroup of the Type II MADS-box proteins (Fig. 1I). MADS-box TF genes are especially prominent on almost all levels of the gene regulatory networks that control reproductive development in flowering plants22,23. In recent years, research into seed and embryo development has become increasingly focused on MADS-box transcription factors in species such as rice (Oryza sativa L)26, soybean (Glycine max (Linn.) Merr.)27, tomato (Lycopersicum esculentum)28, Arabidopsis thaliana8, and strawberry (Fragaria × ananassa Duch.)29.
In our study, we found that ZmMAD1a is expressed at a relatively high level in the maize flower (tassel and silk) and endosperm (Fig. 3A). This expression pattern is similar to that found in other plants in which seed or fruit development is regulated by MADS-box genes, which are mainly expressed in the reproductive organs. OsMADS6, which is important for ovule development, was highly expressed in developing seeds and flowers, but was not detected in roots, leaves, and suspension cells30. In tomato, the genes TAG1, TAGL1, TAGL2, and TAGL11, which are related to fruit development, were only expressed in flowers and fruits31, and the seed growth genes, SHP1 and SHP2, are expressed mainly in the carpels and control pod shattering in Arabidopsis32. The level of expression of ZmMAD1a in the endosperm increases with the increase in days after pollination until 18-DAP, after which it declines by >10-fold in 20-DAP endosperm (Fig. 3B). Similar phenomena have been observed in other species; in rice, OsMADS6 is expressed abundantly in the ovule during meiosis, and expression in the endosperm peaks at 10 DAP and then declines slightly at 20 DAP30. In soybean, GmAGL15 is expressed in the seeds during development; its expression level peaked initially at 15 d after flowering in the young embryo, reached the highest level at 30 DAP, then dropped rapidly27. Transcription of the tomato SlMADS1 gene occurs mainly in sepals and green fruits, and the mRNA levels decrease significantly with fruit ripening3.
We found that overexpressing the ZmMAD1a gene had no significant effects on maize or rice traits (Fig. S2). Phylogenetic analysis also showed that ZmMADS1a groups closely with ZAG2 (zag2), ZmMADS1, and other proteins that function in the regulation of flower and seed development (Fig. 1III). Our results show that ZmMAD1a affects the physiological and biochemical characteristics in the endosperm of maize and rice seeds. Overexpression of ZmMADS1a led to altered expression of starch biosynthesis genes (Figs 4 and S1) and changes in starch composition and structure in the endosperm (Figs 5, 6, S4, S6). The expression levels of AGPL1 (sh2) and AGPS1 (bt2) were increased, but AGPL2 was generally down-regulated during development of maize endosperm (Fig. 4). These genes encode subunits of the AGPase enzyme, which controls the rate-limiting step in starch production and is regulated by allosteric effectors, and can affect the total starch content33. AGPL1 (sh2) and AGPS1 (bt2) exhibited significantly higher expression levels than AGPL2, which led to increases in the total starch content (Figs 6A, S6B). GBSSI is primarily responsible for amylose biosynthesis34, and GBSSII does not show significant activity in endosperm, but may be essential in other non-storage tissues35. Our results indicate that the expression of GBSSI and GBSSII show different trends during endosperm development. The GBSSI (wx) gene was significantly up-regulated and GBSSII was down-regulated during maize endosperm development (Fig. 4). The increased level of GBSSI expression in response to overexpression of ZmMADS1a leads to an increase in amylose content (Fig. 6B). SS genes are exclusively involved in amylopectin biosynthesis, and their distribution within the plastid between the stroma and starch granules varies between species, tissue, and developmental stages36. The expression of the SSI and SSII genes showed very different patterns in seeds of maize and rice ZmMADS1a-overexpressing lines. SSI is the major SS in cereal endosperm, but expression of the SSI gene was not drastically changed in seeds of ZmMADS1a-overexpressing vs. wild-type seeds (Figs 4, S1). SSIIa is mainly responsible for the synthesis of branched starch of medium chain length37, and SSIIIa can control the branched structure through its interaction with DBE38. Expression of SSIIa was significantly up-regulated at 15 and 20 DAP, while the SSIIIa gene was up-regulated at 15 DAP compared to the control (Fig. 4); therefore, the resulting amylopectin contents were changed only slightly (Fig. 6C). Expression of the BEI gene was down-regulated, particularly at 15 and 20 DAP. BEIIa was down-regulated at 20 and 25 DAP, and BEIIb showed significant up-regulation from 10 to 25 DAP in maize endosperm (Fig. 4). Isoamylases can play vital roles in starch granule initiation33. The relative expression level of ISA can change starch granule structure in potato, and also in Arabidopsis mutants39. PUL mainly plays a role in starch degradation, but is also involved in starch synthesis, and mutations in this enzyme affect starch content and structure40,41. ISA1 (su1) expression was sharply up-regulated at 15 DAP, and PUL was down-regulated at 20–25 DAP compared to wild-type (Fig. 4). Changes in the expression levels of these genes can partially account for the observed reduction in the size of starch granules in the endosperm of ZmMADS1a-overexpressing transgenic maize lines (Fig. 5). The changes in the ZmMADS1a-overexpressing lines were not caused by a single starch gene, but rather by multiple genes which may interact and affect the expression of one another at different stages of seed development. These results indicate that ZmMADS1a mainly functions at the mid-late grain-filling period of kernel development in maize, and that it positively regulates the expression of endosperm starch biosynthesis genes.
Starch granules consist of ordered crystalline and disordered amorphous areas, and are composed of amylose and amylopectin in different proportions. Crystalline and amorphous structures are mainly the information reflected in the C1 signal zone in the 13C{1H}CP/MAS spectra41.The peak locations in our study were consistent with the results of previous studies42. The signal at 102.4 ppm is associated with the amorphous part of the granule. Signals at 100.3 and 101.2 ppm are associated with the crystalline structure. The signal at 82 ppm was attributed to C4 of the V-type starch, which is amylose in a single helix conformation, and the 82 ppm signal in the C4 region provides information on the amorphous components of the starch43. The chemical shifts are shifted to the left by ~0.5 ppm and the peak area of C4 increased noticeably in the ZmMADS1a-overexpressing lines (Fig. S4). Overexpression of ZmMADS1a had a significant impact on the disordered structures in the starch granules.
The ZmMADS1a gene is involved in the expression of multiple genes associated with enhanced sugar contents (reducing and soluble sugars) (Fig. 7), and the ZmMADS1a-overexpressing transgenic plants exhibited better growth under drought stress conditions. We provide candidate regulators and show that ZmMADS1a has a similar adjustment mechanism for starch biosynthesis in maize and rice grain and can enhance the expression of most starch biosynthesis genes, especially the expression of GBSSIa (wx), resulting in increased amylose content in mature seeds. The results presented here will further our understanding of the ZmMADS1a regulatory mechanism in endosperm starch synthesis in maize.
Materials and Methods
Plant material and growth conditions
Maize plants for laboratory analyses were grown in a greenhouse at a temperature of 28 ± 2 °C and a photoperiod of 14 hours light/10 hours dark. Rice seedlings were grown at 30 °C under a 16/8-h light/dark cycle in controlled environment chambers and were then transferred to the field and grown under normal rice cultivation conditions at the Experimental Station of Anhui Agricultural University, Hefei, China. Seeds were sown in a seed bed in mid-May and transplanted to the field in mid-June.
Isolation of the ZmMADS1a cDNA and DNA Sequence Analysis
The ZmMADS1a gene sequence was obtained from MaizeSequence (http://www.maizesequence.org/index.html, GRMZM2G160687_T01, 1473 bp). Protein sequence analysis was performed using Pfam (http://pfam.janelia.org/search/sequence) and EXPASY (http://web.expasy.org/compute_pi/). Amino acid sequences were predicted using the DNAMAN software package (Lynnon Biosoft). The full-length cDNA of ZmMADS1a was amplified by RT-PCR from RNA extracted from seeds of maize genotype B73 16 days after pollination with primers MA-F and MA-R.
Generation of DNA constructs and transformation
The amplified ZmMADS1a cDNA from the maize B73 inbred line was digested with KpnI and SmaI (primers: mMA-F-1 and mMA-R-1, Table S1) and then inserted into the p1 vector, which is a modied version of pCAMBIA130144, resulting in a vector called p1-ZmMADS1a. These two clones were digested with EcoRI and SmaI and inserted into the binary vector pCAMBIA1301 under the control of the Cauliflower mosaic virus 35S (CaMV-35S) promoter, and the NOS terminator was inserted between the SphI and HindIII sites.
The pZZ00005 vector (China National Seed Group Co., Ltd) was used for overexpression analysis in maize. The 35S::ZmMADS1a::NOS gene cassette was amplifed from p1-ZmMADS1a using primers mMA-F-2 and mMA-R-2 (Table S1) carrying terminal HindIII and PmeI sites and inserted into the pZZ00005 vector, giving p2-ZmMADS1a. Zea mays ‘ZZC01’ (China National Seed Group Co., Ltd) was the maize line used for transformation.
The p1-ZmMADS1a vector was introduced into the Agrobacterium strain EHA105, and transformed into the rice (Oryza sativa L. subsp. japonica) cultivar ‘Zhonghua 11’ as described previously45. Using hygromycin selection, 18 independent T0 transgenic lines were obtained; expression of the ZmMADS1a transgene was confirmed in these lines by RT-PCR, and homozygous T3 transgenic lines were obtained for further analysis.
The p2-ZmMADS1a vector was also introduced into the Agrobacterium strain EHA105. Timentin (100 mg/L) and bialaphos (5 mg/L) were added to the selective medium. Using a protein test strip for bialaphos selection, 20 independent T0 transgenic lines were obtained and the homozygous T3 transgenic lines were obtained for further analysis.
Gene expression analyses
The relative expression of each endosperm gene was analyzed in eight different maize tissues by qRT-PCR, and the α-tubulin gene (GRMZM2G152466_T01) was used as an internal control for normalization of gene expression levels. PCR primer pairs were designed with Primer Express 3.0 using the B73 genome sequence as a reference (MaizeGDB release 5b.60; http://www.maizegdb.org/). To study the expression pattern of ZmMADS1a, 16 different tissues, including stage-3 root, stage-3 stem, stage-3 leaf, tassel, cornsilk, stegophyl, 12 DAP_endosperm, 12 DAP_embryos, 14 DAP_endosperm, 14 DAP_embryos, 16 DAP_endosperm, 16 DAP_embryos, 18 DAP_endosperm, 18 DAP_embryos, 20 DAP_endosperm, and 20 DAP_embryos were collected (DAP = days after pollination).
To analyze the expression patterns of starch synthesis genes in seeds of transgenic and wild-type plants, developing seeds were harvested from maize (5 DAP, 10 DAP, 15 DAP, 20 DAP, and 25 DAP) and rice (3 DAP, 6 DAP, 10 DAP, 15 DAP, 20 DAP, and 25 DAP). All samples were frozen immediately in liquid nitrogen and stored at −80 °C prior to use. Total RNA was isolated using the E.Z.N.A. MagSi Plant RNA Kit (Omega Biotek), and was treated with DNaseI to remove contaminating genomic DNA. All PCR primers used in this study are given in Tables S2, S3. First-strand cDNAs were synthesized using the PrimeScript™ RT reagent Kit with gDNA Eraser (Perfect Real Time; Takara). Expression levels were calculated using the 2−ΔΔCT method of Livak and Schmittgen (2001)46 and statistically analyzed as described by Liu et al.47.
Subcellular localization
Three fusion vectors were constructed to study the localization of ZmMADS1a. The full-length cDNA sequence of ZmMADS1a without the terminator codon (TGA) was amplified from B73 RNA. The identity of the gene fragment was confirmed by DNA sequencing and was then inserted between the HindIII and BamHI sites of the pEZS-NL vector, which contains the enhanced green fluorescent protein (eGFP) reporter gene, to generate a fusion construct under the control of the CaMV 35S promoter, namely pEZS-MS-eGFP (the primer names were MA-SL-F and MA-SL-R, Table S1). The localization vector was transferred into protoplasts of Arabidopsis and maize by electroporation. The leaves of Arabidopsis were selected for 4 weeks after the leaves were not smoked about 90 pieces and the leaves of maize were selected 2 weeks of growth. The two kinds of leaves were cut into 1 mm wide strips.
Grain trait measurements
More than three randomly chosen, fully filled grains from each line or strain were aligned length-wise along a vernier caliper to measure seed length, after which they were rearranged to measure grain width and thickness48. The 1,000-grain weight was determined by weighing 10 replicates of 100-grain samples independently on an electronic balance.
Analysis of starch properties
Harvested paddy seeds were air-dried and stored at room temperature. Embryos and pericarps were removed before the experiment, and the endosperms were ground to a powder using a grinding mill. The starch and amylose contents were measured using starch assay kits following the manufacturer’s instructions (K-TSTA and K-AMYL; Megazyme). To determine the amylose content, the powder was soaked for 48 h in 0.4% NaOH (powder:NaOH = 1:3) at room temperature, washed several times with distilled water until no slimy liquid remained, and drained. To determine the amount of soluble sugars, 50 mg of endosperm powder was washed twice in 80% (v/v) ethanol at 80 °C for 40 min and assayed using anthrone reagent49.
Observation of endosperm starch granules
Seeds were dried completely under low pressure, cut across the short axis, and the surface was sputter coated with gold prior to observation with a scanning electron microscopy. To analyze the structure of starch granules, solid-state 13C CP/MAS NMR spectra was used. About 0.3 mg of starch samples were packed into a 4-mm diameter, cylindrical, partially stabilized zirconium oxide (PSZ) rotor (6 kHz, 54.7°) with a Kelf and cap. The experiments were conducted at a 13C frequency of 75.46 MHz on a Bruker MSL-300 spectrometer43,50,51.
References
Messenguy, F. & Dubois, E. Role of MADS box proteins and their cofactors in combinatorial control of gene expression and cell development. Gene 316, 1–21 (2003).
Portereiko, M. F. & Lloydm, A. Steff en, J. G., et al. AGL80 is required for central cell and endosperm development in Arabidopsis. Plant Cell 18, 1862–1872 (2006).
Tiwari, S. et al. Transcriptional profiles underlying parent-of-origin effects in seeds of Arabidopsis thaliana. Bmc Plant Biol. 10, 72 (2010).
Masiero, S. et al. The emerging importance of Type I MADS Box transcription factors for plant reproduction. Plant Cell 23, 865–872 (2011).
Yoo, S. K., Lee, J. S. & Ahn, J. H. Overexpression of AGAMOUS-LIKE 28 (AGL28) promotes flowering by upregulating expression of floral promoters within the autonomous pathway. Biochem. Biophys Res. Commun. 348, 929–936 (2006).
Bemer, M. et al. The MADS dDomain protein DIANA acts together with AGAMOUS-LIKE80 to specify the central cell in Arabidopsis Ovules. Plant Cell 20, 2088–2101 (2008).
Kang, I. H. et al. The AGL62 MADS domain protein regulates cellularization during endosperm development in Arabidopsis. Plant Cell 20, 635–647 (2008).
Colombo, M. et al. AGL23, a type I MADS-box gene that controls female gametophyte and embryo development in Arabidopsis. Plant J. 54, 1037–1048 (2010).
Ferrándiz, C., Gu, Q., Martienssen, R. & Yanofsky, M. F. Redundant regulation of meristem identity and plant architecture by FRUITFUL, APETALA1 and CAULIFLOWER. Development 127, 725–734 (2000).
Henschel, K. et al. Two ancient classes of MIKC-type MADS-box genes are present in the Moss Physcomitrella patens. Mol. Biol. Evol. 19, 801–814 (2002).
Pellegrini, L., Tan, S. & Richmond, T. J. Structure of serum response factor core bound to DNA. Nature 376, 490–498 (1995).
Kaufmann, K., Melzer, R. & Theissen, G. MIKC-type MADS-domain proteins: structural modularity, protein interactions and network evolution in land plants. Gene 347, 183–198 (2005).
Thompson, B. E. et al. Bearded-ear encodes a MADS box transcription factor critical for maize floral development. Plant Cell 21, 2578–2590 (2009).
Heuer, S. et al. The maize MADS box gene ZmMADS3 affects node number and spikelet development and is co-expressed with ZmMADS1 during flower development, in egg cells, and early embryogenesis. Plant Physiol. 127, 33 (2001).
Heuer, S., Lörz, H. & Dresselhaus, T. The MADS box gene ZmMADS2 is specifically expressed in maize pollen and during maize pollen tube growth. Sex. Plant Reprod. 13, 21–27 (2000).
Han, J. J. & Martienssen, R. Pod corn is caused by rearrangement at the Tunicate1 locus. Plant Cell 24, 2733–2744 (2012).
Qiao, Z. et al. ZmMADS47 regulates Zein gene transcription through interaction with Opaque2. Plos Genet. 12, e1005991 (2016).
Guo, S. et al. The interaction between OsMADS57 and OsTB1 modulates rice tillering via DWARF14. Nat. Commu. 4, 1566 (2013).
Yin, L. L. & Xue, H. W. The MADS29 transcription factor regulates the degradation of the nucellus and the nucellar projection during rice seed development. Plant Cell 24, 1049–1065 (2012).
Becker., A. & Theissen, G. The major clades of MADS-box genes and their role in the development and evolution of flowering plants. Mol. Phylogenet. Evol. 29, 464–489 (2003).
Melzer, R., Wang, Y. Q. & Theissen, G. The naked and the dead: The ABCs of gymnosperm reproduction and the origin of the angiosperm flower. Semin. Cell Dev. Biol. 21, 118–128 (2010).
Arora, R. et al. MADS-box gene family in rice: genome-wide identifi cation, organization and expression profi ling during reproductive development and stress. BMC Genomics 8, 242 (2007).
Tapia-López, R. & Alvarez-Buylla, E. R. An AGAMOUS-related MADS-box gene, XAL1 (AGL12), regulates root meristem cell proliferation and flowering transition in. Arabidopsis. Plant Physiol. 146, 1182–1192 (2008).
Wu, C. et al. In situ expression of the GmNMH7 gene is photoperiod-dependent in a unique soybean (Glycine max [L.] Merr.) flowering reversion system. Planta 223, 725–735 (2006).
Fischer, A. et al. Chromosomal mapping of the MADS-box multigene family in Zea mays reveals dispersed distribution of allelic genes as well as transposed copies. Nucleic Acids Res. 23, 1901 (1995).
Dreni, L. et al. The D-lineage MADS-box gene OsMADS13, controls ovule identity in rice. Plant J. 52, 690–699 (2010).
Wang, X. L. Expression of the MADS-Box gene GmAGL15 in seed development of soybean: expression of the MADS-Box gene GmAGL15 in seed development of Soybean. Acta Agronomica Sinica 34, 330–332 (2008).
Mazzucato, A. et al. Characterization of genes controlling stamen identity and development in a parthenocarpic tomato mutant indicates a role for the DEFICIENS ortholog in the control of fruit set. Physio.l Plant 132, 526–537 (2010).
Seymour, G. B. et al. A SEPALLATA gene is involved in the development and ripening of strawberry (Fragaria × ananassa Duch.) fruit, a non-climacteric tissue. J. Experim. Bot. 62, 1179–1188 (2011).
Zhang, J. et al. OsMADS6 plays an essential role in endosperm nutrient accumulation and is subject to epigenetic regulation in rice (Oryza sativa. L). Plant J. 64, 604–617 (2010).
Busi, M. V. et al. MADS-box genes expressed during tomato seed and fruit development. Plant Mol. Biol. 52, 801–815 (2003).
Eshed, Y. SHATTERPROOF MADS-box genes control seed dispersal in Arabidopsis. Nature 404, 766–770 (2000).
Kawagoe, Y. et al. Roles of isoamylase and ADP-glucose pyrophosphorylase in starch granule synthesis in rice endosperm. Plant J. 42, 164–174 (2010).
Hanashiro, I. et al. Granule-Bound Starch Synthase I is Responsible for Biosynthesis of Extra-Long Unit Chains of Amylopectin in Rice. Plant Cell Physiol. 49, 925–933 (2008).
Jongseong, J. et al. Starch biosynthesis in cereal endosperm. Plant Physiol. Bioch. 48, 383–392 (2010).
Commuri, P. D. & Keeling, P. L. Chain-length specificities of maize starch synthase I enzyme: studies of glucan affinity and catalytic properties. Plant J. 25, 475–486 (2001).
Vrinten, P. L. & Nakamura, T. Wheat granule-bound starch synthase I and II are encoded by separate genes that are expressed in different tissues. Plant Physiol. 122, 255–264 (2000).
Li, Z. et al. The structure and expression of the wheat starch synthase III gene. Motifs in the expressed gene define the lineage of the starch synthase III gene family. Plant Physiol. 123, 613–624 (2000).
Wattebled, F. et al. Mutants of arabidopsis lacking a chloroplastic isoamylase accumulate phytoglycogen and an abnormal form of amylopectin. Plant Physiol. 138, 184–195 (2005).
Dinges, J. R., Colleoni, C., James, M. G. & Myers, A. M. Mutational analysis of the pullulanase-type debranching enzyme of maize indicates multiple functions in starch metabolism. Plant Cell 15, 666–680 (2003).
Delval, F. et al. Characterization of crosslinked starch materials with spectroscopic techniques. Journal of Appl. Polym. Sci. 93, 2650–2663 (2004).
Fan, D. et al. Determining the effects of microwave heating on the ordered structures of rice starch by NMR. Carbohyd. Polym. 92, 1395–1401 (2013).
Tan, I. et al. A method for estimating the nature and relative proportions of amorphous, single, and double-helical components in starch granules by 13C CP/MAS NMR. Biomacromolecules 8, 885–891 (2007).
Yuan, B., Shen, X., Li, X., Xu, C. & Wang, S. Mitogen-activated protein kinase OsMPK6 negatively regulates rice disease resistance to bacterial pathogens. Planta 226, 953–960 (2007).
Ge, X., Chu, Z., Lin, Y. & Wang, S. A tissue culture system for different germplasms of indica rice. Plant Cell Rep 25, 392–402 (2006).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2012).
Liu, J. X., Srivastava, R., Che, P. & Howell, S. H. An endoplasmic reticulum stress response in Arabidopsis is mediated by proteolytic processing and nuclear relocation of a membrane-associated transcription factor. bZIP28. Plant Cell 19, 4111–4119 (2007).
Mao, H. et al. Linking differential domain functions of the GS3 protein to natural variation of grain size in rice. P. Natl. Acad.Sci. USA 107, 19579–19584 (2010).
Wang, J. C., Xu, H., Zhu, Y., Liu, Q. Q. & Cai, X. L. OsbZIP58, a basic leucine zipper transcription factor, regulates starch biosynthesis in rice endosperm. J. Exp. Bot. 64, 3453–3466 (2013).
Dong, T. et al. A tomato MADS-box transcription factor, SlMADS1, acts as a negative regulator of fruit ripening. Plant Physiol. 163, 1026–1036 (2013).
Fujita, N. et al. Characterization of pullulanase (PUL)-deficient mutants of rice (Oryza sativa L.) and the function of PUL on starch biosynthesis in the developing rice endosperm. J. Exp. Bot. 60, 1009–1023 (2009).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31801374), Anhui Natural Science Foundation (1808085QC89) and National key research and development plan (2016YFD0101002), National 973 project (2014CB 138200), President Youth Innovation Found of Anhui Academy of Agricultural Sciences (17B0915).
Author information
Authors and Affiliations
Contributions
Q.D. performed the majority of the experimental work. F.W. and J.K. assisted in most of the experiments. L.C. and Q.X. conducted part of the determination of transgenic material. H.J. and T.L. gave valuable advice. H.C. assisted the experiments material breeding. C.L. and B.C. revised the paper. B.C. conceived the project and supervised the work.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dong, Q., Wang, F., Kong, J. et al. Functional analysis of ZmMADS1a reveals its role in regulating starch biosynthesis in maize endosperm. Sci Rep 9, 3253 (2019). https://doi.org/10.1038/s41598-019-39612-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-39612-5
- Springer Nature Limited
This article is cited by
-
Genome-wide association analysis of kernel nutritional quality in two natural maize populations
Molecular Breeding (2023)
-
Comparative proteomic analyses of Tartary buckwheat (Fagopyrum tataricum) seeds at three stages of development
Functional & Integrative Genomics (2022)