Abstract
Marker-assisted breeding serves as a potent tool for screening target germplasm, assessing genetic diversity, and determining breeding potential of a crop. Therefore, inter primer binding site (iPBS)-retrotransposons marker system was employed to evaluate a collection of 33 Brassica genotypes, including 10 Brassica juncea, 5 B. oleracea, 7 Sinapis alba, 5 B. nigra, and 6 B. rapa, were utilized to evaluate their genetic diversity and variations 10 polymorphic primers that generated a total of 144 bands. Various diversity indices were calculated in the studied germplasm, including polymorphism information content (0.13–0.30), effective number of alleles (1.217–1.689), Shannon’s information index (0.244–0.531), and gene diversity (0.148–0.370). These indices collectively affirmed substantial genetic variations within the germplasm. Molecular variance analysis revealed that the majority (62%) of genetic variations were present within populations. The Brassica accessions were categorized into three populations utilizing a model-based structure algorithm. Evaluation of diversity indices based on the structure indicated that populations III and II exhibited higher diversity. Principal coordinate analysis and neighbor-joining analysis further corroborated the three distinct populations, confirming the reliability of the STRUCTURE analysis. Notably, the genetic distance assessment identified BN1 and BN3 from B. nigra species and the genotypes BO1 and BO3 from B. oleracea as genetically diverse mustard accessions. The extensive genetic diversity observed within the Brassica germplasm underscores its significance as a valuable genetic resource for comprehensive Brassica breeding programs. Moreover, these accessions hold promise as suitable candidates for heterosis breeding initiatives aimed at improving mustard production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Brassica is characterized by a diverse array of plant species, and are extensively used for over 5000 years BCE, (Campbell et al. 2016; Watson and Preedy 2010). This family comprises 338 genera and 3709 species, displaying significant morphological diversity encompassing various crop species with commercial importance, including industrial oilseeds, spices, vegetables, and fodder crops (Li, et al. 2017), which offer a diverse array of end products due to their varied range of use.
Brassica rapa L. is a source of vegetable oil for cooking, food processing, and industrial applications (Chew 2020). The seeds of Brassica arvensis L. could be used for biofuel production (Gidik, et al. 2023). Brassica nigra L. is a popular spice used in many cuisines as well as its paste is used as a condiment (Kayacetin 2020). Brassica juncea L. is another popular spice used in South Asian cuisines (Kumar, et al. 2011). Brassica napus L. is a hybrid of B. rapa L. and B. oleracea L. and is a source of vegetable oil, which is used for animal feed and industrial purposes (Das, et al. 2022). Following U's triangle model, the interspecific hybridization of B. rapa × B. nigra produces the amphidiploid species B. juncea [AABB (n: 18)], derived from two fundamental diploid species, B. rapa [AA (n: 10)] and B. nigra [BB (n: 8)].
Mustard species, including Sinapis alba (white mustard or yellow mustard), B. nigra (black mustard), B. juncea (brown mustard), B. rapa (also known as B. campestris, field mustard, or turnip), represent a distinguished germplasm, which serve as primary raw materials for numerous agriculture-based industries globally including Turkey (Kayacetin 2020). S. alba is commonly utilized as a seasoning agent in its dry form, while B. nigra holds significance not only as a condiment but also for its medicinal purposes (Aslan and Eryilmaz 2020; Kayacetin 2023a, b; Tomar and Shrivastava 2014). B. juncea is employed as a biofuel feedstock and spice, whereas B. rapa finds widespread use as a biofuel feedstock (Kayacetin 2023a, b; Kwon, et al. 2020).
Mustard and rape breeding programs require information on the diversity and relationships within and among landraces to facilitate germplasm recognition, preservation, and application (Rabbani et al. 1998; Ilyas et al. 2018). Previous research (Ali, et al. 2003; Azam, et al. 2013; Jan, et al. 2017; Naheed, et al. 2016; Neeru, et al. 2016) indicated that breeders extensively assess this germplasm exploring both morphological and genetic diversity before incorporating it into breeding programs. Nevertheless, molecular markers offer a more robust approach, representing a highly efficient, reproducible, and resource-saving strategy for selecting diverse genotypes.
Among the various molecular markers, retrotransposons stand out as versatile, reproductive, and efficient tools for determining genetic diversity in plant species. Retrotransposons, also known as jumping elements, constitute between 50 and 90% of plant genomes (SanMiguel, et al. 1996). There are two main types of retrotransposons: long terminal repeat (LTR) and non-LTR, serving as predominant type markers in plant genomes. ParadoxicallyKalendar, et al. (2010) has proposed a universal DNA fingerprinting technique applicable to both plants and animals, termed as the "inter primer binding site (iPBS)," introducing a new marker system. This technique allows for the analysis of both LTR and non-LTR retrotransposons. Inter primer binding site primers are designed for reverse transcription during the retrotransposon replication cycle, based on primer binding site (PBS) sequences with conserved tRNA segments (Guo, et al. 2014; Kalendar, et al. 2010).
The efficiency of the iPBS-retrotransposon marker system has been demonstrated in a wide range of crops including laurel (Karık, et al. 2019), chickpea (Andeden, et al. 2013), pea (Baloch, et al. 2015b, a), lens (Baloch, et al. 2015b, a), quinoa (Barut, et al. 2020), Turkish okra (Yıldız, et al. 2015), safflower (Ali, et al. 2019), pepper (Yildiz and Arbizu 2022), Turkish wild and cultivated Emmer (Arystanbekkyzy, et al. 2019), Turkish bread wheat (Nadeem 2021), common bean (Aydın and Baloch 2019) and grapevine (Güler, et al. 2024). Despite the extensive use of SSR markers to differentiate for species (Thakur, et al. 2017), genetic diversity (Hobson and Rahman 2016; Raza, et al. 2019; Yin, et al. 2023), yield potential (Wolko, et al. 2022), and breeding (Singh, et al. 2022); the iPBS marker system have not yet been employed for genetic diversity assessment.
As far as we know, this study constitutes the initial efforts to utilize iPBS markers for evaluating genetic diversity within Brassica species.
Materials and methods
Plant growth conditions
The Brassica accessions (shown in Table 1) were surface sterilized following the procedure as described previously (Aslam, et al. 2022). The seeds were washed in running tap water followed by surface sterilization using 20% (v/v) commercial bleach (Domestos) for 15 min, followed by 5 × 3 rinses with sterile distilled water. Subsequently, the seeds were placed in Petri plates containing sterile filter paper moistened with sterile water for germination of the seedlings. The growth chamber conditions were 16 h light/8 h dark, 25 °C temperature, and 65% humidity.
Plant genomic DNA extraction from Brassica plant
Brassica seedlings were crushed into a fine powder using a mortar and pestle that had been pre-chilled with liquid nitrogen. The genomic DNA was isolated following the plant genomic DNA kit protocol (Exgene™ Plant SV, GeneAll co. LTD, Seoul, South Korea. The DNA was stored at -80 °C for future use.
Genomic DNA gel electrophoresis
Thegel electrophoresis was performed using TAE buffer at 100 V for 60 min in an agarose gel (1%) to determine the purity and quantity of the isolated genomic DNA. The gel was stained with ethidium bromide solution for 5 min and rinsed with distilled water to remove excess of the dye. The gel was visualized under the gel documentation system (Bio-Rad) and photographe.
iPBS-retrotransposon analysis and agarose gel electrophoresis
The primers utilized were derived from the work of Kalendar, et al. (2010) in this study. Initially, 80 iPBS-retrotransposon primers were tested on a randomly selected group of 10 Brassica accessions. Thereafter, 10 most polymorphic primers were identified and subsequently employed for PCR amplification across all 33 Brassica accessions (refer to Table 2 for details). The amplification conditions of iPBS-PCR were adapted based on the protocol outlined by Kalendar et al. (2010), with minor modifications (Saba, et al. 2020). The PCR reaction mixture included 30 ng of template DNA, 1 × Taq reaction buffer, 200 µM of dNTPs, 0.2 µM primer for 11–15 length oligonucleotides or 18-bases long oligonucleotides, and 1.25 U Taq polymerase (Promega, Life Technologies, NEB) in a 25 μL reaction volume.
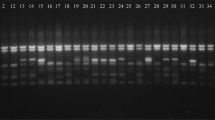
The PCR reaction procedure included an initial denaturation phase at 94 °C for 3 min, succeeded by 40 denaturation cycles at 94 °C for 15 s, an annealing step at a temperature ranging from 49 °C to 65 °C, based on the oligonucleotides used, for 1 min, and a final extension at 72 °C for 10 min (Kalendar et al. 2010). The amplified retrotransposon bands were subjected to analysis using 3% (w/v) agarose gel electrophoresis at 120 V. The resulting amplified bands were observed and documented using a gel documentation system (Bio-Rad GDS, California, USA).
Genetic diversity analysis
Retrotransposons were employed as markers to assess the relationships among individual plant genotypes within the collection. An iPBS marker-based neighbor-joining tree was constructed. The dissimilarity matrix was calculated using the Manhattan index in TASSEL software (Bradbury et al. 2007) to construct both the tree and principal coordinate analyses (PCoA). Different diversity parameters were also computed using GenAlEx V6.5 (Peakall and Smouse 2006) to investigate genetic differences between landraces and cultivars.
Results
Following PCR amplification of retrotransposon markers, the resulting products were separated on agarose gels and visualized using a gel documentation system. Thereafter the bands were scored, and the genetic diversity analysis was performed on the Brassica genotypes listed in Table 1. On average, each primer produced 14 bands, and the mean count of polymorphic bands was 13.5. Notably, iPBS 2376 exhibited the maximum rate of polymorphism (94.12%), followed by 88.24% bands by iPbS 2376 and minimum percentage of 64.17 was noted for iPBS 2074 primers. The values for Polymorphic Information Content (PIC) varied from 0.13 for iPBS 2074 to 0.30 for iPBS 2081. Additional details regarding the primer performance metrics are provided in Table 2. Across all primers, recessive alleles (q) were more frequent. The primers iPBS2376, and iPBS2081 exhibited the greatest allele count (Na), had values in close proximity to 1.882 and 1.824 correspondingly. Although iPBS 2074 exhibited the lowest values of Na at 1.294, the Shannon’s information index (I) was the highest for iPBS 2081 and the lowest for iPBS 2074. The effective alleles (Ne) were the highest for iPBS 2081 and the lowest for iPBS 2074. The values of He (expected heterozygosity) in this study ranged 0.148 (iPBS 2074) to 0.370 (iPBS 2081).
Similarly, the values of uHe (unbiased expected heterozygosity) for the same set of primers varied between 0.150 to 0.376. The Brassica genotypes were classified into three clades according to the Neigbor-joining clustering analysis. The first clade consisted of 11 genotypes all of which were from B. juncea species except one BO1 from B. oleracea clustered with BJ10. The second group consisted of 10 genotypes both from B. napus and rapa. There were 6 B. rapa and 4 B. napus accessions in this clade. Out of total 12 members in the third clade, 7 genotypes belonged to S. alba and one to B. nigra. Rest of four from B. oleracea showed diverse nature of genotypes that were grouped under the same clade (Fig. 1) (Table 3).
STRUCTURE analysis using the Evanno method (Fig. 2) identified that three populations (K = 3) had the optimal number of genetic clusters. This conclusion was supported by the maximum ΔK value of 302.71 observed at K = 3. The FST values, indicative of genetic differences, were computed as 0.64, 0.65, and 0.40 for the Brassica populations q1 to q3, correspondingly. Furthermore, the structure analysis depicted the proportions of genetic diversity within these three populations (Fig. 3). Notably, clustering analysis provided similar results in identifying distinct populations. The structural analysis offered the added advantage of visualizing the proportional contributions of genetic diversity.
PCoA further confirmed the clustering observed with the model-based structure algorithm, as evidenced by the separation of the 33 Brassica accessions into three populations (Fig. 4).
Genetic variation among the genotypes was assessed through Analysis of Molecular Variance (AMOVA). As presented in Table 4, approximately 38% of the total variation was attributed to disparities among populations accounted for, while the remaining 62% of the variation was observed within the populations.
Discussion
Genetic variation within plant populations is a crucial resource for breeders. It allows them to develop improved cultivars with traits desired by farmers, industry, and consumers (Govindaraj, et al. 2015). Numerous research endeavors have explored relationships within Brassicaceae family, employing SSR markers (El-Esawi, et al. 2016; Singh, et al. 2018; Thakur, et al. 2017). Similarly, several investigations have utilized ISSR markers for assessing genetic associations within the Brassicaceae family (Khalil and El-Zayat 2019; Safari, et al. 2013; Shen, et al. 2016; Wang, et al. 2017). Nevertheless, to the best of our information, this study marks the first instance where the iPBS-retrotransposon marker system has been employed to evaluate genetic diversity in 5 different Brassica species populations.
A set of 10 highly polymorphic iPBS-retrotransposon primers was employed to investigate the genetic variability and structure of populations of Brassica species. These iPBS primers generated a total of 144 bands, with 135 of them identified as polymorphic (refer to Table 2). The total and polymorphic bands identified in this study surpassed those reported by Raza, et al. (2019) using SSR markers. However, it is worth noting that the PIC value was lower in the case of iPBS.
In contrast to another study utilizing ISSR markers with a set of 7 primers, where the number of polymorphic bands was higher (20) compared to iPBS (13), this discrepancy could be attributed to the limited number of primers and the marker system employed. Similarly, in another study of ISSR, the average number of polymorphic bands in B. rapa var. chinensis was 6.3, which is lower than the findings in the present study (Shen, et al. 2016). Furthermore, a study applying ISSR markers to Amphidiploid lines of B. juncea (2n = 4x = 36), reported a lower PIC value (0.18) compared to our iPBS system, highlighting the influence of species and marker choice on genetic diversity estimates.
This findings of current study on genetic diversity are comparable to those obtained by Wang et al. (2017) using ISSR markers. The average diversity index (I), effective allele number (Ne), and average genetic diversity (He) were 0.4045, 1.4557, and 0.2684, respectively. While a study by Thakur et al. (2018) reported a higher number of alleles (Na) using SSR markers for B. napus, these values were close to those obtained with iPBS markers in this study. However, the lowest allele count observed in the SSR study exceeded compared to iPBS markers. These variations likely stem from differences in both plant species analyzed and the marker systems employed.
The observed variations in scores for different diversity indices in this investigation may have arisen from disparities in germplasm and the inherent characteristics of the utilized molecular markers. The iPBS-retrotransposons marker system, known for its high reproducibility and universal applicability, has been validated in several investigations (Shimira, et al. 2021; Yildiz and Arbizu 2022). Hence, the marker system could be the adopted choice for molecular profiling of Brassica genotypes over co-dominant and dominant marker systems.
Importantly, AMOVA findings emphasized that the highest genetic variances in Brassica germplasm exist within populations. These findings align with previous research that also reported higher genetic variations within populations (Jozová, et al. 2023; Tesfaye, et al. 2023).
Nei's genetic distance analysis suggests that B. nigra accessions BN1 and BN3 belong to a distant clade. This genetic distance makes them potential parents for cross-pollination breeding programs aimed at improving mustard quality. For the accessions of species B. oleracea such as BO1 and BO3 could be desired candidate for heterosis research. These combinations could enhance mustard pungency and glucosinolate contents and yield potential. The use of iPBS in marker-assisted breeding is particularly advantageous for gaining insights into genetic potential and diversity among species or cultivars, expediting molecular-based breeding.
The model-based structure algorithm categorized 33 Brassica genotypes into three distinct populations primarily based on their species level (refer to Fig. 3). Population III, comprising 36.36% (12) accessions, is a mixed population of S. alba and B. oleracea. Population I, accounting for 33.33% (11) accessions, mainly consists of B. juncea, except for one that is from B. oleracea. Population II was the smallest and included 30.30% (10) genotypes, encompassing B. rapa and B. nigra. The presence of B. nigra and B. rapa in Population II, and S. alba and B. oleracea in Population III, contributed to the high diversity of these populations. Neighbor-joining analysis, as depicted in Fig. 1, aligns with the population structure (Fig. 3), showing congruence and support the diversity pattern with multiple lines of evidence. Furthermore, PCoA echoes the distribution pattern of genotypes seen in the neighbor-joining tree and structure analysis, underscoring the consistency and reproducibility of the analyses.
STRUCTURE analysis revealed higher genetic variations within populations III and II, which was further confirmed by AMOVA analysis. Importantly, AMOVA analysis confirms that most genetic variation resides within Brassica germplasm populations. This high within-population diversity holds promise for future breeding efforts aimed to improve mustard paste quality and yield. PCoA analysis further supports the clustering observed by the model-based structure algorithm, segregating Brassica accessions into three populations (refer to Fig. 4).
Conclusion
The current investigation offers a comprehensive understanding of genetic variations within a germplasm consisting of 5 Brassica species utilizing the iPBS-retrotransposons marker system. The study identified BN1, BN3, BO1, and BO3 as genetically diverse accessions within the germplasm, making them promising putative candidates for breeding programs. AMOVA findings emphasized increased genetic variations within populations compared to variations among populations. Notably, population II and III, as identified through structure clustering, exhibited higher diversity, suggesting that accessions within these populations should be prioritized for future Brassica breeding initiatives. The structure algorithm based on a model and PCoA successfully separated the examined germplasm into distinct populations, primarily attributed to their diverse genetic potential and increased heterozygosity. This study further corroborated the practicality and widespread applicability of the retrotransposons markers, positioning them as valuable tools for exploring genetic diversity in Brassica crops.
References
Ali N, Javidfar F, Elmira JY, Mirza M (2003) Relationship among yield components and selection criteria for yield improvement in winter rapeseed (Brassica napus L.). Pak J Bot 35:167–174
Ali F, Yılmaz A, Nadeem MA, Habyarimana E, Subaşı I, Nawaz MA, Chaudhary HJ, Shahid MQ, Ercişli S, Zia MAB (2019) Mobile genomic element diversity in world collection of safflower (Carthamus tinctorius L.) panel using iPBS-retrotransposon markers. PLoS ONE 14:e0211985
Andeden EE, Baloch FS, Derya M, Kilian B, Özkan H (2013) iPBS-Retrotransposons-based genetic diversity and relationship among wild annual Cicer species. J Plant Biochem Biotechnol 22:453–466
Arystanbekkyzy M, Nadeem MA, Aktas H, Yeken MZ, Zencirci N, Nawaz MA, Ali F, Haider MS, Tunc K, Chung G (2019) Phylogenetic and taxonomic relationship of turkish wild and cultivated emmer (Triticum turgidum ssp. dicoccoides) revealed by iPBSretrotransposons markers. Int J Agric Biol 21:155–163
Aslam N, Sameeullah M, Yildirim M, Baloglu MC, Yucesan B, Lössl AG, Waheed MT, Gurel E (2022) Isolation of the 3β-HSD promoter from Digitalis ferruginea subsp. ferruginea and its functional characterization in Arabidopsis thaliana. Mol Biol Rep 49:7173–7183
Aslan V, Eryilmaz T (2020) Polynomial regression method for optimization of biodiesel production from black mustard (Brassica nigra L.) seed oil using methanol, ethanol, NaOH, and KOH. Energy 209:118386
Aydın MF, Baloch FS (2019) Exploring the genetic diversity and population structure of Turkish common bean germplasm by the iPBS-retrotransposons markers. Legume Res 42(1):18–24. https://doi.org/10.18805/LR-423
Azam SM, Farhatullah AN, Shah S, Iqbal S (2013) Correlation studies for some agronomic and quality traits in Brassica napus L. Sarhad J Agric 29:547–550
Baloch FS, Alsaleh A, de Miera LES, Hatipoğlu R, Çiftçi V, Karaköy T, Yıldız M, Özkan H (2015a) DNA based iPBS-retrotransposon markers for investigating the population structure of pea (Pisum sativum) germplasm from Turkey. Biochem Syst Ecol 61:244–252
Baloch FS, Derya M, Andeden EE, Alsaleh A, Cömertpay G, Kilian B, Özkan H (2015b) Inter-primer binding site retrotransposon and inter-simple sequence repeat diversity among wild Lens species. Biochem Syst Ecol 58:162–168
Barut M, Nadeem MA, Karaköy T, Baloch FS (2020) DNA fingerprinting and genetic diversity analysis of world quinoa germplasm usingiPBS-retrotransposon marker system. Turk J Agric for 44:479–491
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Campbell L, Rempel CB, Wanasundara JP (2016) Canola/rapeseed protein: Future opportunities and directions—Workshop proceedings of IRC 2015. Plants 5(2), 17; https://doi.org/10.3390/plants5020017
Chew SC (2020) Cold-pressed rapeseed (Brassica napus) oil: chemistry and functionality. Food Res Int 131:108997
Das GG, Malek MA, Shamsuddin AKM, Sagor GHM (2022) Development of high yielding early matured and shattering tolerant Brassica napus L. through interspecific hybridization between B. rapa L. and B. oleracea L. Genet Resour Crop Evol 69:1009–1018
El-Esawi MA, Germaine K, Bourke P, Malone R (2016) Genetic diversity and population structure of Brassica oleracea germplasm in Ireland using SSR markers. C R Biol 339:133–140
Gidik B, Volkan G, Önemli F, Girgel Ü (2023) Fatty acid composition and biodiesel quality of Brassica nigra, Brassica napus and sinapis arvensis seeds. Bahçe 52:1–6
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Genet Res Int 2015:431487
Güler E, Karadeniz T, Özer G, Uysal T (2024) Diversity and association mapping assessment of an untouched native grapevine genetic resource by iPBS retrotransposon markers. Genet Resour Crop Evol 71:679–690
Guo D-L, Guo M-X, Hou X-G, Zhang G-H (2014) Molecular diversity analysis of grape varieties based on iPBS markers. Biochem Syst Ecol 52:27–32
Hobson N, Rahman H (2016) Genome-wide identification of SSR markers in the Brassica A genome and their utility in breeding. Can J Plant Sci 96:808–818
Ilyas M, Ghulam S, Rabbani MA, Malik SI, Cheema NM, Muhammad A, JAN SA, (2018) Genetic divergence in Brassica napus L. germplasm as determined by quantitative attributes. Pak J Bot 50:1039–1045
Jan SA, Shinwari ZK, Rabbani MA, Niaz IA, Shah SH (2017) Assessment of quantitative agro-morphological variations among Brassica rapa diverse populations. Pak J Bot 49:561–567
Jozová E, Rost M, Rychlá A, Stehlíková D, Pudhuvai B, Hejna O, Beran P, Čurn V, Klíma M (2023) Microsatellite markers: a tool to assess the genetic diversity of yellow mustard (Sinapis alba L.). Plants 12:4026
Kalendar R, Antonius K, Smýkal P, Schulman AH (2010) iPBS: a universal method for DNA fingerprinting and retrotransposon isolation. Theor Appl Genet 121:1419–1430
Karık Ü, Nadeem MA, Habyarimana E, Ercişli S, Yildiz M, Yılmaz A, Yang SH, Chung G, Baloch FS (2019) Exploring the genetic diversity and population structure of Turkish laurel germplasm by the iPBS-retrotransposon marker system. Agronomy 9:647
Kayacetin F (2020) Botanical characteristics, potential uses, and cultivation possibilities of mustards in Turkey: a review. Turk J Bot 44:101–127
Kayacetin F (2023a) Comparison of some species in genus Brassica cultivated on clay loamy soils under semi-arid agroecosystem for suitability to biodiesel production. Energy Sour, Part A 45:11963–11980
Kayacetin F (2023b) Influence of sowing dates and genotypes on phenology, morphology, yield and fatty acid compounds of Sinapis alba L. for the energy industry. Gesunde Pflanz 75:613–623
Khalil R, El-Zayat M (2019) Molecular characterization of some Brassica species. Adv Plants Agric Res 9:112–119
Kumar V, Thakur AK, Barothia ND, Chatterjee SS (2011) Therapeutic potentials of Brassica juncea: an overview. Cell Med 1(1):2–1
Kwon H-Y, Choi S-I, Park H-I, Choi S-H, Sim W-S, Yeo J-H, Cho J-H, Lee O-H (2020) Comparative analysis of the nutritional components and antioxidant activities of different Brassica juncea cultivars. Foods 9:840
Li P, Zhang S, Li F, Zhang S, Zhang H, Wang X, Sun R, Bonnema G, Borm TJ (2017) A phylogenetic analysis of chloroplast genomes elucidates the relationships of the six economically important Brassica species comprising the triangle of U. Front Plant Sci 8:111
Nadeem MA (2021) Deciphering the genetic diversity and population structure of Turkish bread wheat germplasm using iPBS-retrotransposons markers. Mol Biol Rep 48:6739–6748
Naheed H, Mohammad F, Sohail Q, Abid S, Khan N (2016) Genetic diversity of bread wheat lines based on agro-morphological traits. Int J Agric Biol 18:1049–1055
Neeru N, Avtar R, Singh A (2016) Evaluation and classification of Indian mustard (Brassica juncea L.) genotypes using principal component analysis. J Oilseed Brassica 1:167–174
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Rabbani MA, Iwabuchi A, Murakami Y, Suzuki T, Takayanagi K (1998) Genetic diversity in mustard (Brassica juncea L.) germplasm from Pakistan as determined by RAPDs. Euphytica 103:235–242
Raza A, Saher MS, Farwa A, Ahmad KRS (2019) Genetic diversity analysis of Brassica species using PCR-based SSR markers. Gesunde Pflanzen 71:1–7
Saba K, Sameeullah M, Asghar A, Gottschamel J, Latif S, Lössl AG, Mirza B, Mirza O, Waheed MT (2020) Expression of ESAT-6 antigen from Mycobacterium tuberculosis in broccoli: an edible plant. Biotechnol Appl Biochem 67:148–157
Safari S, Mehrabi A-A, Safari Z (2013) Efficiency of RAPD and ISSR markers in assessment of genetic diversity in Brassica napus genotypes. Int J Agric Crop Sci 5(273):279
SanMiguel P, Tikhonov A, Jin YK, Motchoulskaia N, Zakharov D, Melake-Berhan A, Springer PS, Edwards KJ, Lee M, Avramova Z, Bennetzen JL (1996) Nested retrotransposons in the intergenic regions of the maize genome. Science 274:765–768
Shen X, Zhang Y, Xue J, Li M, Lin Y, Sun X, Hang Y (2016) Analysis of genetic diversity of Brassica rapa var. chinensis using ISSR markers and development of SCAR marker specific for Fragrant Bok Choy, a product of geographic indication. Genet Mol Res 15(1):11
Shimira F, Boyaci HF, Çilesiz Y, Nadeem MA, Baloch FS, Taşkin H (2021) Exploring the genetic diversity and population structure of scarlet eggplant germplasm from Rwanda through iPBS-retrotransposon markers. Mol Biol Rep 48:6323–6333
Singh BK, Choudhary SB, Yadav S, Malhotra EV, Rani R, Ambawat S, Priyamedha PA, Kumar R, Kumar S, Sharma HK, Singh DK, Rai PK (2018) Genetic structure identification and assessment of interrelationships between Brassica and allied genera using newly developed genic-SSRs of Indian Mustard (Brassica juncea L.). Ind Crops Prod 113:111–120
Singh KP, Kumari P, Raipuria RK, Rai PK (2022) Development of genome-specific SSR markers for the identification of introgressed segments of Sinapis alba in the Brassica juncea background. 3 Biotech 12(12):332
Tesfaye M, Feyissa T, Hailesilassie T, Kanagarajan S, Zhu L-H (2023) Genetic diversity and population structure in ethiopian mustard (Brassica carinata A. Braun) as revealed by single nucleotide polymorphism markers. Genes 14:1757
Thakur AK, Singh KH, Singh L, Nanjundan J, Khan YJ, Singh D (2017) SSR marker variations in Brassica species provide insight into the origin and evolution of Brassica amphidiploids. Hereditas 155:6
Tomar RS, Shrivastava V (2014) Efficacy evaluation of ethanolic extract of Brassica nigra as potential antimicrobial agent against selected microorganisms. Int J Pharma Sci Health Care 3:117–123
Wang A, Zhou G, Lin C, Wang B, Huang X, Deng Y, Zhang R (2017) Genetic diversity study of Brassica campestris L. ssp. chinensis Makino based on ISSR markers. Caryologia 70:48–54
Watson RR, Preedy VR (2010) Bioactive foods and extracts: Cancer treatment and prevention. CRC Press
Wolko J, Łopatyńska A, Wolko Ł, Bocianowski J, Mikołajczyk K, Liersch A (2022) Identification of SSR Markers Associated with Yield-Related Traits and Heterosis Effect in Winter Oilseed Rape (Brassica napus L.). Agronomy 12:1544. https://doi.org/10.3390/agronomy12071544
Yıldız M, Koçak M, Baloch FS (2015) Genetic bottlenecks in Turkish okra germplasm and utility of iPBS retrotransposon markers for genetic diversity assessment. Genet Mol Res 14(3):10588–10602. https://doi.org/10.4238/2015.September.8.20
Yildiz M, Arbizu C (2022) Inter-primer binding site (iPBS) retrotransposon markers provide insights into thegenetic diversity and population structure of carrots (Daucus Apiaceae). Turk J Agri for 46(2):214–223
Yin J, Zhao H, Wu X, Ma Y, Zhang J, Li Y, Shao G, Chen H, Han R, Xu Z (2023) SSR marker based analysis for identification and of genetic diversity of non-heading Chinese cabbage varieties. Front Plant Sci 14:1112748. https://doi.org/10.3389/fpls.2023.1112748
Acknowledgements
The authors acknowledge the funding and suport by TÜBİTAK (The Scientific and Technological Research Council of Türkiye) to support this study through Project number 5190038.
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK). The Funding was provided by Türkiye Bilimsel ve Teknolojik Araştırma Kurumu, 5190038
Author information
Authors and Affiliations
Contributions
MS: Investigation, Methodology, Laboratory research, Data curation, Visualization, Software, Writing–original draft. FK: Resources supply, KMK: Supervision, writing, review and editing, AYP: Laboratory research and experimentation, SM: Laboratory research and experimentation, MTW: review & editing, VÇ: Resources supply.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known conflict of interests for the work reported in this paper.
Ethical approval
The research in this article does not involve any human or animal subjects therefore it does not need any ethical approval.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sameeullah, M., Kayaçetin, F., Khavar, K.M. et al. Decoding genetic diversity and population structure of Brassica species by inter primer binding site (iPBS) retrotransposon markers. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01986-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01986-5