Abstract
Landraces can be defined as locally adapted cultivated forms of domesticated plant species that have distinctive traits resulting from both environmental adaptation and farmer selection. In tomato (Solanum lycopersicum L.), they are important for preserving genetic diversity and as a potential source of adaptive traits that are not found in contemporary commercial varieties. In this work, we analyzed the genetic variability within and between 16 tomato landraces originating in Southern Italy, a secondary center of diversity. We used the highly discriminant (GATA)4 DNA fingerprinting to assess genetic homogeneity, relationship, differentiation, and structure, as well as to infer the occurrence of possible contaminant or admixed genotypes. Within-landrace variability was found in a third of the landraces. The population structure (without using predefined groups) indicated in different instances, a correspondence between fruit type and landraces’ clusters. The results of multivariate and Bayesian analyses showed a low frequency of possible admixed or contaminant genotypes. Moreover, the quantification of intra-landrace variability allowed to estimate the potential loss of diversity in storing tomato landraces as pure lines. Finally, issues related to possible contamination of the germplasm also imply that more thorough assessment of variability is required not only to store genuine diversity but also to exploit tomato landraces in breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
While landraces are universally recognized as a highly valuable component of plant genetic resources for food and agriculture, their definition has been the subject of extensive discussion (Casañas et al. 2017; Villa et al. 2005; Zeven 1998). Broadly speaking, a crop landrace is a cultivated form of a domesticated plant species that is locally adapted mainly because of geographical isolation. More recently, greater attention has been given to the cultural value of a landrace, for example in terms of visible characteristics of identity, gastronomic value, and specific transformation processes and uses. This is to underline that the distinctive traits of a landrace are due not only to environmental adaptive traits but also to a deliberate selection by farmers (Pusadee et al. 2009), thus allowing them to be distinguished from ecotypes.
In the past, every agricultural production system was based on landraces, but they have been progressively replaced by more specialized germplasm since the nineteenth century. Currently, landraces play a role especially in traditional production systems of developing countries whereas in high-income countries, they are associated with amateur or traditional farming (Galluzzi et al. 2010; Gibson 2009). In advanced economies, landraces usually receive attention as niche products, because of the consumer perception of higher quality, and as a resource for crop improvement yet to be fully untapped. For instance, landraces are considered important germplasm to improve yield stability in the face of climate change, compliance to low input agriculture, resilience to stress, nutritional value, and quality (Bellon and van Etten 2014; Corrado and Rao 2017; Newton et al. 2011; Petropoulos et al. 2018).
The tomato (Solanum lycopersium L.) is among the most widespread and economically important vegetable crops worldwide (https://www.fao.org/faostat/). Following domestication in the Americas, this species was imported to Europe in the sixteenth century to later spread across the globe (McCue 1952). Italy and Spain are considered secondary centres of diversity because of the presence of a morphological variation that has not been documented elsewhere (Kulus 2022). In the cultivated tomato germplasm, genetic diversity has been mostly reduced because of founder effect and breeding (Miller and Tanksley 1990; Williams and Clair 1993), expect for the more recent introgression of variability from wild relatives (Bergougnoux 2014; Liang et al. 2017). Tomato landraces are therefore a valuable genetic material for their yield stability, resilience to stress, adaptability to organic farming, and fruit quality (Andreakis et al. 2004; Boziné-Pullai et al. 2021; Rouphael et al. 2021; Sumalan et al. 2020; Villareal et al. 1978). Moreover, tomato landraces are of increasing commercial importance because of the promotion of local food systems and short food supply chains as leverage towards more sustainable and resilient agricultural systems (Enthoven and Van den Broeck 2021; Samoggia et al. 2021).
Although several authors agree that landraces are dynamic, non-uniform populations, works documenting their genetic intra-variability are limited in comparison with those related to the study of their relationships or comparison with contemporary cultivars. Limiting our attention to a few predominantly selfing-species, the DNA analysis of a collection of 12 barley landraces of the Sardinia region of Italy indicated that intra-landraces variability accounted for 11% of the total variance (Papa et al. 1998). In a regional collection of 25 common bean landraces, the within landraces genetic variability was estimated at 32%, with almost all landraces composed of slightly different genotypes (Scarano et al. 2014). Genetic variability could be higher when confronting landraces from different nations, as demonstrated for Triticum aestivum (Dreisigacker et al. 2005), or from different regions, as in Lens culinaris (Fikiru et al. 2007) and Triticum durum (Pagnotta et al. 2005). In tomato, intra-landrace molecular variability has been found in secondary centers (García-Martínez et al. 2006, 2013; Mazzucato et al. 2010) and non-centers of origin for tomato (Alzahib et al. 2021).
On-farm conservation of landraces refers to the practice of maintaining and preserving traditional crop varieties by active cultivation where they have been traditionally grown and developed over generations, and provides multiple benefits (Bellon and van Etten 2014). Nonetheless, it has been also pointed out that farming cam also associates with contamination of landraces, either with other landraces or cultivars, for instance, because of the own seed production (Hagenblad et al. 2012; Kyratzis et al. 2019; Zeven and Waninge 1989).
The aim of this work was to analyze the genetic variability among and within 16 tomato landraces collected in the Campania region (Italy), representing different fruit shapes. Specifically, our aims included the quantification of the levels of genetic homogeneity, and the assessment of genetic differentiation and structure, considering that previous works indicated the possible presence of spurious and/or contaminant genotypes (Corrado et al. 2014; Rao et al. 2006). Among the different molecular markers available, we used the (GATA)4 minisatellite because of its ability to detect high levels of polymorphism. In tomato this marker type successfully distinguished cultivars and landraces that are otherwise difficult to differentiate (García-Martínez et al. 2013; Kaemmer et al. 1995; Rao et al. 2006).
Material and methods
Plant material
The work was carried out on sixteen tomato (Solanum lycopersicum L.) landraces present in the Campania region of Italy. Table 1 reports the code and the main characteristic of the fruits along with the farmer’s classification. A definition of the terms used in Table 1 to describe the overall fruit shape (e.g., circular, cordate, etc.) can be found in the UPOV guidelines for S. lycopersicum TG/44/11 (https://www.upov.int; accessed the 1 of February 2023). In this work, fruits with a narrow (elongated) cylindric (parallel) shape in lateral view are also defined as “San Marzano type”. Fruits with an average weight below 25 g were classified as "small".
DNA analysis
We analysed four plants per landraces, for a total of 64 genotypes. DNA was isolated from the first pair of true leaves of plants germinated in a growth chamber. Leaves were harvested, immediately frozen in liquid nitrogen, and stored at − 80 °C until use. Total DNA was isolated using the GenElute Plant Genomic DNA Miniprep Kit (Sigma Aldrich, Milan, Italy) and quantified using an agarose gel procedure (Sambrook et al. 1989). Five microg of total DNA was digested with 40 units of Taq I (Promega, Milan, Italy) overnight in a final volume of 450 µl following the manufacturer instructions. After ethanol precipitation and washing with 70% ethanol, the DNA was resuspended, and resolved in a 0.8% agarose gel in 1xTAE buffer for 18 h at 30 V (Sambrook et al. 1989). Following a 30-min denaturation in a 1,5 M NaCl and 0,5 M NaOH solution, and a 45 min neutralization in a 1,5 M NaCl and 1 M TrisHCl (pH 7,4) solution, the DNA was transferred overnight by Southern blotting to a Hybond N membrane (Amersham, Milan, Italy) using a 20 X SSC solution (Sambrook et al. 1989), and then crosslinked with UV (150 mJoule) using the GS Gene Linker (Bio-Rad, Milan, Italy). γ32PdATP 5' terminal labeling (Perkin Elmer, Milan, Italy), probe purification, hybridization with the (GATA)4 probe, washing and visualization on Kodak Bio-Max X-ray films were carried out as previously described (García-Martínez et al. 2013).
Data analysis
Bands were scored for presence (1) or absence (0) and treated as dominant markers. The Shannon index was calculated with the vegan package (Dixon 2003). AMOVA was carried out with the poppr package, using 9999 permutations for testing statistical significance (Kamvar et al. 2014). Hierarchical classification of the individuals and of the populations was performed with the ape package (Paradis and Schliep 2019), using the Provesti distance and the Unweighted Pair Group Method with Arithmetic (UPGMA) clustering algorithm. The cophenetic correlation coefficient between the distance matrix and the corresponding cophenetic matrix was calculated using the Mantel’s test (with 9999 permutations) implemented in the ape package.
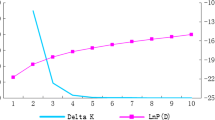
Possible population structure was estimated using the software STRUCTURE v2.3 (Pritchard et al. 2000). The analysis was carried out using a burning period of 25,000 iterations and a run length of 250,000 MCMC replications. We tested a continuous series of K, from 1 to 12, in 10 independent runs each to verify the consistency across runs. The most informative K was identified using the ΔK method (Evanno et al. 2005) with the aid of Structure harvester (Earl and VonHoldt 2012). The estimated cluster membership coefficient matrices of each run were permuted (n = 1,000) so that all replicates have as close a match as possible, and then averaged across the 10 runs using the Largely Greedy algorithm of the software CLUMPP (Jakobsson and Rosenberg 2007).
Results
Genetic diversity among and within landraces
(GATA)4 DNA fingerprinting of the 16 landraces revealed a total of 29 reproducible scorable bands (Fig. 1).
Main genetic parameters of the scored markers calculated on the whole population (n = 64) are reported in Supplementary Table 1. There were 27 polymorphic bands (93% of the total) considering all the individuals. The number of bands per landrace ranged from a maximum of 18 to a minimum of 8, for an average value of 12.4 ± 2.8 (mean ± s.d.). In total, we detected 22 multilocus genotypes. Eleven landraces were genetically homogeneous (i.e., they display a single DNA profile), while five had intra-landraces variability (i.e., bands were polymorphic within the landrace) (Table 2).
Two different genotypes were detected for the CB and the SMC landraces, while only one plant of the CI and TM was different. The most variable landrace was the CB, with three different profiles (Shannon Index = 1.04). The hierarchical classification of the individuals indicated that larger intra-landrace difference was present for the SMC, a San Marzano type. Specifically, while two plants clustered along with landraces with similar fruit type (i.e., SMMU and SMMO), the other two SMC plants were in a distinct position, joining the tree at the last node (Figure S1). The number of private alleles was low, with only one specific band for the CB landrace and another specific for the homogeneous landrace TD. The number of less common bands among landraces (i.e., with a frequency less or equal to 25%) was also low, on average 14% of the total number of bands (Table 2).
As landraces were collected and named according to the information provided by farmers, we employed an analysis of molecular variance (AMOVA) based on the polymorphic molecular markers to identify a possible genetic differentiation. This analysis considered the pre-defined landraces as hierarchical level for partitioning genetic variation. There was a significant (p < 0.001) differentiation among populations (Table 3). Specifically, most of the variance was attributed to differences among populations, with 14.4% of the molecular variance occurring among samples within populations.
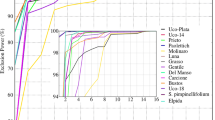
To categorize landraces based on levels of genetic relatedness, we calculated pairwise genetic distances between pre-defined populations. Distances were employed to build a dendrogram using the UPGMA algorithm (Fig. 2).
The distribution of the landraces indicated that all the pre-defined populations were well separated. The high Fst values indicated a significant pairwise differentiation between landraces, which was not surprising given the level of homogeneity observed within landraces (Table S2). The agreement between the genetic distance and the hierarchical classification was evaluated with the Mantel’s test. The cophenetic correlation coefficient was 0.81 and significant (p < 0.001), also suggesting the presence of a genetic structure in the data.
Inference of population structure
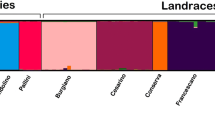
To classify the individuals in populations without using a priori knowledge, we used the model-based clustering method developed by Pritchard et alii (Pritchard et al. 2000). This analysis was also carried out to clarify the features of divergent samples identified in the hierarchical clustering approach, as they may be admixed genotypes (e.g., deriving from the cross of two landraces populations) or contaminants genotypes (e.g., not attributable to a particular landrace population). For the whole set of markers, the Evanno’s test indicated that the most informative number of populations was nine (Figure S2). The Q-matrix with the individual membership coefficients indicated that the samples were strongly assigned to the clusters (Table S3). At K = 9, a biological interpretation of the sub-division can be performed considering the fruit shape and folk taxonomy (this being also dependant on the morphological traits of the plants and its use). As illustrated in Fig. 3, the landraces with a small, circular, pointed fruits (PB and PBI) grouped together (C1, brown). The landraces with an elongated, San Marzano-type fruit (namely SMMO, SMMU, and two plants of the SMC) also clustered together (C2, grey). The other two plants of the SMC were considered a different population from all the others (C3, light green). The largest population was made of landraces with a medium size (around 40–50 g) circular fruit (TL, VD, CI, and GD) (C7, orange), with one plant of the CI (CI3) which appears to be admixed with another landraces (TI; C6, azure) with circular fruits. Another cluster was made of two landraces sharing the same fruits shape in lateral view (C8, blue). Nonetheless, the plants of the C9 group (dark blue) did not have a clear similarity considering fruit shape and size.
Estimated population structure of the tomato genotypes. Each genotype (reported on the X-axis) is represented by a horizontal line, which is partitioned into colored segments that represent the estimated membership fractions in the 9 clusters (C). See Figure S2 for the determination of the most informative K and Table S3 for the membership coefficient
Discussion
The appreciation of landraces has gained momentum in tomato because of their possible use in breeding as a source of adaptive and quality traits (Dwivedi et al. 2016). One of the main issues is that pedigree information, or more generally (co-)ancestry, is usually not available, and collections are usually made on the sole basis of farmers’ knowledge. The origin, differentiation, and relationships among and within landraces are often undetermined but this information is necessary to rationalize germplasm management and to guide future users in their sampling (Engels 2003; Spooner 2005).
This work evaluated the molecular diversity among and within locally cultivated tomato accessions using a highly informative DNA marker (Caramante et al. 2009). Many landraces were genetically uniform, as expected considering the predominantly selfing nature of tomato and the farmers’ maintenance selection (Zeven 2000). Variability within landraces was found in around 31% of the germplasm, although this estimation is based on markers scored as dominant.
The hierarchical classification of the individual suggested that, when present, the less frequent DNA profile of one landrace was not identical to those of others, expressing true variability rather than possible contamination or seed mixture. It was noteworthy the distinct position of two genotypes of the SMC landrace, because they joined the dendrogram at the last node, while the other different profiles were scattered within the dendrogram.
Considering the a priori knowledge (i.e., the predefined groups), the genetic analysis indicated that all the landraces' populations could be separated by the (GATA)4 analysis. Hierarchical classification based on population distances is useful to understand the molecular diversity of plants because it does not require assumptions (e.g., Hardy–Weinberg equilibrium) and can be applied to a wide range of dataset regardless of the biological properties of the species (being solely based on the chosen pairwise distance and clustering algorithm). At the high node level, some landrace groups (e.g., those with the San Marzano type fruits, or some with small round fruits) clustered together. Moreover, the high correlation coefficient between genetic distance and the cophenetic matrices is also considered a clue of the presence of a population structure (Odong et al. 2011).
We then assessed the possible population structure without using predefined groups. This allowed us not only to understand the genetic diversity and relationships among different landraces but also to infer the origin and nature of the detected intra-landrace variability. The data analysis indicated that in different instances there is a correspondence between fruit type and clusters, which is consistent with previous studies that have primarily focused on cultivated tomato varieties. These studies have suggested that tomato cultivars can be grouped based on their fruit shape and market classes, which are largely fixed by breeding in contemporary cultivars (Corrado et al. 2013; Sim et al. 2012, 2011).
The number of identified populations was high and this was supported by the high value of pairwise genetic differentiation between landraces (measured as Fst). Although Fst also depends on the sample size, natural plant populations tend to have lower Fst values compared to cultivated plant populations, especially predominantly autogamous crops like S. lycopersicum. This indirectly suggests that the presence of the Bayesan groups with a similar fruit shape is likely to be indicative of a common origin because gene flow among tomato landraces should be limited. Nonetheless, as the genotypes have been collected in a culturally, gastronomically, and geographically homogenous area, an effect of the environment (including selection for common traits) over subpopulations cannot be ruled out. We used neutral DNA markers, which have limited (if any) obvious function and should not be under adaptive selection pressure. (GATA)4 tandem repeats are expected to be inherited independently of traits under selection because of their genomic organization, hence they should provide an unbiased estimate of genetic variation (Vosman and Arens 1997). However, further studies, based for instance on more extensive genome scans, will have to determine whether the sub-populations of landraces identified by the Bayesian analysis mostly reflect an evolutionary relationship related to a single founder population or are mainly due to a common human selection pressure.
The identity of a plant landrace can be altered by cross-pollination, seed mixing, or accidental or intentional introduction of unrelated genotypes (Zeven 1999). This may be due to the proximity to other varieties or the farmers’ seed production from hybrid cultivars (Zeven 1999). The latter is likely to occur more frequently for species like the tomato, whose seed production is dominated by hybrid genotypes. It has been discussed the possibility that in Greece tomato landraces were initially derived from introduced germplasm and later, hybrid material (Gonias et al. 2019). Moreover, the landrace from the Italian region Umbria called ‘Conserva’ is suspected to derive from recent crosses between genetically diverse lines because of the highly negative Fixation Index, while the Observed Heterozygosity was close to zero as the other five landraces analyzed (Castellana et al. 2020). On the other hand, the DNA analysis of the Spanish landrace ‘Pera’ revealed the presence of two related sub-populations, which may represent true intra-landrace variability (García-Martínez et al. 2013). The hierarchical clustering of the individuals and the Bayesian analysis of the subpopulations indicated the presence of genotypes that can be considered contaminant (e.g., highly unrelated to the alleged and other landraces), as in the case of the SMC, and those deriving from admixture, as in the case of the CI landraces.
In conclusion, this study offers valuable insights for the characterization and conservation of cultivated tomato genetic resources in a secondary centre of diversity, indicating the strong differentiation of the landraces and the possible presence of landrace groups based on fruit shape. The result highlights the importance of the molecular characterization of traditional tomato varieties that are still being grown as a tool to enhance the credibility of conservation strategies, facilitate sound decision-making (especially at regional level), and promote further actions for a genetic-based selection and purification. To protect the genetic variability of this species, our work also underlines the importance to introduce strategies that encourage the cultivation and marketing of landraces that are adapted to local conditions, while aligning with traditional gastronomy. Finally, our study also highlights the need for comprehensive and rigorous testing of tomato landraces in secondary centers of diversity, to understand the role and impact of intra-landrace variability, identify undesired genotypes, preserve appropriate genetic diversity and ultimately, develop improved plant material.
Ackwoledgments
This study was partly carried out within the Agritech National Research Center and received funding from the European Union Next-GenerationEU (PIANO NAZIONALE DI RIPRESA E RESILIENZA (PNRR)–MISSIONE 4 COMPONENTE 2, INVESTIMENTO 1.4–D.D. 1032 17/06/2022, CN00000022). This manuscript reflects only the authors’ views and opinions, neither the European. Union nor the European Commission can be considered responsible for them.
Data availability
The data supporting the findings of this study that are not already included within the article and its supplementary material are available from the corresponding author (G.C.) upon reasonable request.
References
Alzahib RH, Migdadi HM, Ghamdi AAA, Alwahibi MS, Afzal M, Elharty EH, Alghamdi SS (2021) Exploring genetic variability among and within hail tomato landraces based on sequence-related amplified polymorphism markers. Diversity 13(3):135
Andreakis N, Giordano I, Pentangelo A, Fogliano V, Graziani G, Monti LM, Rao R (2004) DNA fingerprinting and quality traits of Corbarino cherry-like tomato landraces. J Agric Food Chem 52(11):3366–3371
Bellon MR, van Etten J (2014) Climate change and on-farm conservation of crop landraces in centres of diversity. Plant genetic resources and climate change, 137–150
Bergougnoux V (2014) The history of tomato: from domestication to biopharming. Biotechnol Adv 32(1):170–189
Boziné-Pullai K, Csambalik L, Drexler D, Reiter D, Tóth F, Tóthné Bogdányi F, Ladányi M (2021) Tomato landraces are competitive with commercial varieties in terms of tolerance to plant pathogens—a case study of Hungarian gene bank accessions on organic farms. Diversity 13(5):195
Caramante M, Rao R, Monti LM, Corrado G (2009) Discrimination of ‘San Marzano’accessions: a comparison of minisatellite, CAPS and SSR markers in relation to morphological traits. Sci Hortic 120(4):560–564
Casañas F, Simó J, Casals J, Prohens J (2017) Toward an evolved concept of landrace. Front Plant Sci 8:145
Castellana S, Ranzino L, Beritognolo I, Cherubini M, Luneia R, Villani F, Mattioni C (2020) Genetic characterization and molecular fingerprint of traditional Umbrian tomato (Solanum lycopersicum L.) landraces through SSR markers and application for varietal identification. Genet Resour Crop Evolut 67(7):1807–1820
Corrado G, Rao R (2017) Towards the genomic basis of local adaptation in landraces. Diversity 9(4):51
Corrado G, Piffanelli P, Caramante M, Coppola M, Rao R (2013) SNP genotyping reveals genetic diversity between cultivated landraces and contemporary varieties of tomato. BMC Genom 14:1–14
Corrado G, Caramante M, Piffanelli P, Rao R (2014) Genetic diversity in Italian tomato landraces: Implications for the development of a core collection. Sci Hortic 168:138–144
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14(6):927–930
Dreisigacker S, Zhang P, Warburton M, Skovmand B, Hoisington D, Melchinger A (2005) Genetic diversity among and within CIMMYT wheat landrace accessions investigated with SSRs and implications for plant genetic resources management. Crop Sci 45(2):653–661
Dwivedi SL, Ceccarelli S, Blair MW, Upadhyaya HD, Are AK, Ortiz R (2016) Landrace germplasm for improving yield and abiotic stress adaptation. Trends Plant Sci 21(1):31–42
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Engels J (2003) A guide to effective management of germplasm collections. Bioversity International
Enthoven L, Van den Broeck G (2021) Local food systems: reviewing two decades of research. Agric Syst 193:103226
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14(8):2611–2620
Fikiru E, Tesfaye K, Bekele E (2007) Genetic diversity and population structure of Ethiopian lentil (Lens culinaris Medikus) landraces as revealed by ISSR marker. Afr J Biotechnol 6(12):1460–1468
Galluzzi G, Eyzaguirre P, Negri V (2010) Home gardens: neglected hotspots of agro-biodiversity and cultural diversity. Biodivers Conserv 19:3635–3654
García-Martínez S, Andreani L, Garcia-Gusano M, Geuna F, Ruiz JJ (2006) Evaluation of amplified fragment length polymorphism and simple sequence repeats for tomato germplasm fingerprinting: utility for grouping closely related traditional cultivars. Genome 49(6):648–656
García-Martínez S, Corrado G, Ruiz JJ, Rao R (2013) Diversity and structure of a sample of traditional Italian and Spanish tomato accessions. Genet Resour Crop Evol 60:789–798
Gibson RW (2009) A review of perceptual distinctiveness in landraces including an analysis of how its roles have been overlooked in plant breeding for low-input farming systems. Econ Bot 63(3):242–255
Gonias ED, Ganopoulos I, Mellidou I, Bibi AC, Kalivas A, Mylona PV, Osanthanunkul M, Tsaftaris A, Madesis P, Doulis AG (2019) Exploring genetic diversity of tomato (Solanum lycopersicum L.) germplasm of genebank collection employing SSR and SCAR markers. Genet Resour Crop Evol 66:1295–1309
Hagenblad J, Zie J, Leino MW (2012) Exploring the population genetics of genebank and historical landrace varieties. Genet Resour Crop Evol 59:1185–1199
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23(14):1801–1806
Kaemmer D, Weising K, Beyermann B, Börner T, Epplen J, Kahlm G (1995) Oligonucleotide fingerprinting of tomato DNA. Plant Breeding 114(1):12–17
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281
Kulus D (2022) Genetic diversity for breeding tomato. Cash Crops: Genetic Diversity, Erosion, Conservation and Utilization 505–521
Kyratzis AC, Nikoloudakis N, Katsiotis A (2019) Genetic variability in landraces populations and the risk to lose genetic variation. The example of landrace ‘Kyperounda’and its implications for ex situ conservation. PLoS ONE 14(10):e0224255
Liang S, Jie C, Kai X, Wencai Y (2017) Origin of the domesticated horticultural species and molecular bases of fruit shape and size changes during the domestication, taking tomato as an example. Hortic Plant J 3(3):125–132
Mazzucato A, Ficcadenti N, Caioni M, Mosconi P, Piccinini E, Sanampudi VRR, Sestili S, Ferrari V (2010) Genetic diversity and distinctiveness in tomato (Solanum lycopersicum L.) landraces: the Italian case study of ‘A pera Abruzzese.’ Sci Hortic 125(1):55–62
McCue GA (1952) The history of the use of the tomato: an annotated bibliography. Ann Mo Bot Gard 39(4):289–348
Miller J, Tanksley S (1990) RFLP analysis of phylogenetic relationships and genetic variation in the genus Lycopersicon. Theor Appl Genet 80:437–448
Newton AC, Akar T, Baresel JP, Bebeli PJ, Bettencourt E, Bladenopoulos KV, Czembor JH, Fasoula DA, Katsiotis A, Koutis K (2011) Cereal landraces for sustainable agriculture. Sustain Agric 2:147–186
Odong T, Van Heerwaarden J, Jansen J, van Hintum TJ, Van Eeuwijk F (2011) Determination of genetic structure of germplasm collections: are traditional hierarchical clustering methods appropriate for molecular marker data? Theor Appl Genet 123:195–205
Pagnotta MA, Impiglia A, Tanzarella OA, Nachit MM, Porceddu E (2005) Genetic variation of the durum wheat landrace Haurani from different agro-ecological regions. Genet Resour Crop Evol 51:863–869
Papa R, Attene G, Barcaccia G, Ohgata A, Konishi T (1998) Genetic diversity in landrace populations of Hordeum vulgare L. from Sardinia, Italy, as revealed by RAPDs, isozymes and morphophenological traits. Plant Breeding 117(6):523–530
Paradis E, Schliep K (2019) ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35(3):526–528
Petropoulos SA, Fernandes Â, Ntatsi G, Petrotos K, Barros L, Ferreira IC (2018) Nutritional value, chemical characterization and bulb morphology of Greek garlic landraces. Molecules 23(2):319
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Pusadee T, Jamjod S, Chiang Y-C, Rerkasem B, Schaal BA (2009) Genetic structure and isolation by distance in a landrace of Thai rice. Proc Natl Acad Sci 106(33):13880–13885
Rao R, Corrado G, Bianchi M, Di Mauro A (2006) (GATA)4 DNA fingerprinting identifies morphologically characterized ‘San Marzano’tomato plants. Plant Breeding 125(2):173–176
Rouphael Y, Corrado G, Colla G, De Pascale S, Dell’Aversana E, D’Amelia LI, Fusco GM, Carillo P (2021) Biostimulation as a means for optimizing fruit phytochemical content and functional quality of tomato landraces of the San Marzano area. Foods 10(5):926
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold spring harbor laboratory press.
Samoggia A, Grillini G, Del Prete M (2021) Price fairness of processed tomato agro-food chain: the italian consumers’ perception perspective. Foods 10(5):984
Scarano D, Rubio F, Ruiz JJ, Rao R, Corrado G (2014) Morphological and genetic diversity among and within common bean (Phaseolus vulgaris L.) landraces from the Campania region (Southern Italy). Sci Hortic 180:72–78
Sim S, Robbins M, Van Deynze A, Michel A, Francis D (2011) Population structure and genetic differentiation associated with breeding history and selection in tomato (Solanum lycopersicum L.). Heredity 106(6):927–935
Sim S-C, Van Deynze A, Stoffel K, Douches DS, Zarka D, Ganal MW, Chetelat RT, Hutton SF, Scott JW, Gardner RG (2012) High-density SNP genotyping of tomato (Solanum lycopersicum L.) reveals patterns of genetic variation due to breeding.
Spooner DM (2005). Molecular markers for genebank management. Bioversity International
Sumalan RM, Ciulca SI, Poiana MA, Moigradean D, Radulov I, Negrea M, Crisan ME, Copolovici L, Sumalan RL (2020) The antioxidant profile evaluation of some tomato landraces with soil salinity tolerance correlated with high nutraceuticaland functional value. Agronomy 10(4):500
Villa TCC, Maxted N, Scholten M, Ford-Lloyd B (2005) Defining and identifying crop landraces. Plant Genetic Resources 3(3):373–384
Villareal R, Lai S, Wong S (1978) Screening for Heat Tolerance in the Genus Lycopersicon L. HortScience 13(4):479–481
Vosman B, Arens P (1997) Molecular characterization of GATA/GACA microsatellite repeats in tomato. Genome 40(1):25–33
Williams CE, Clair DAS (1993) Phenetic relationships and levels of variability detected by restriction fragment length polymorphism and random amplified polymorphic DNA analysis of cultivated and wild accessions of Lycopersicon esculentum. Genome 36(3):619–630
Zeven AC (1998) Landraces: a review of definitions and classifications. Euphytica 104:127–139
Zeven AC (1999) The traditional inexplicable replacement of seed and seed ware of landraces and cultivars: a review. Euphytica 110:181–191
Zeven AC (2000) Traditional maintenance breeding of landraces: 1. Data by Crop Euphytica 116:65–85
Zeven A, Waninge J (1989) The presence of three groups of Scalavatis and other hexaploid bread wheat plants contaminating durum wheat fields in Cyprus. Euphytica 43:117–124
Funding
Open access funding provided by Università degli Studi di Napoli Federico II within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
Conceptualization, G.C.; formal analysis, M.C., Y.R. and G.C.; investigation, M.C. and G.C.; writing—original draft preparation, G.C..; writing—review and editing, Y.R. and G.C. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Caramante, M., Rouphael, Y. & Corrado, G. Genetic diversity among and within tomato (Solanum lycopersicum L.) landraces grown in Southern Italy. Genet Resour Crop Evol 71, 157–166 (2024). https://doi.org/10.1007/s10722-023-01613-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01613-9