Abstract
This study investigated genotype × environment interactions for the stability of expression of four productivity traits (cobs yield, cobs I class trade share, lend of cobs and fulfilment of cobs) of sweet maize hybrids (Zea mays L.). The additive main effects and multiplicative interaction (AMMI) model was employed to assess genotype × environment interaction. AMMI stability value was used to evaluate both stability and genotype. The genotype selection index was calculated for each hybrid, incorporating both the average trait value and the stability index. Ten sweet maize hybrids were evaluated: Golda, GSS 1453, GSS 3071, GSS 5829, GSS 8529, Overland, Noa, Shinerock, Sindon, and Tessa. Trials were ran conducted over four vegetative seasons at a single location in the Wielkopolska region using replicated field experiments. The AMMI model revealed significant genotypic and environmental effects for all analyzed traits. Based on their superior stability and favorable average trait values, both the Golda cultivar and the GSS 3071 hybrid are recommended for further breeding program inclusion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sweet maize is a widely cultivated and adaptable crop holding a distinct position within both agricultural and culinary fields. Its unique flavor and nutritional value make it a staple ingredient in many diets around the world (Swapna et al. 2020). Over the years, different sweet maize varieties have been established, catering to range of agronomic requirements and consumer preferences (Revilla et al. 2021; Marenya et al. 2022).
Sweet maize varieties come in a variety of flavor profiles, ranging from traditional sweet and buttery flavors to more unique and exotic flavors (Yactayo-Chang et al. 2022; Zhang et al. 2023). This variety allows consumers to choose the maize that suits their palate, making it a very versatile crop for a variety of culinary applications. Sweet maize varieties are prized for their ability to retain their freshness and quality when harvested fully ripe (Shirke and Pinjarkar 2023; Jiménez-Viveros and Valiente-Banuet 2023). This characteristic makes them an ideal choice for both home gardeners and commercial cultivation, ensuring consistent delivery of high-quality and flavorful maize to consumers.
Sweet maize varieties are available in early, medium and late types. This extended harvest season guarantees a continuous supply of fresh maize throughout the growing season, reducing the need for long-term transportation and storage (Amjath-Babu et al. 2020). Sweet maize is a good source of essential nutrients such as dietary fiber, vitamins (e.g., vitamin C, B vitamins) and minerals (e.g., potassium) (Palacios-Rojas et al. 2020; Garg et al. 2021; Galani et al. 2022). Some varieties of maize are even rich in antioxidants, which contributes to overall health and well-being (Crupi et al. 2023). Sweet maize varieties are extremely versatile in cooking (Vukadinović et al. 2022). They can be eaten raw, grilled, steamed, boiled or used in a variety of dishes such as soups, salads and baked dishes. This versatility adds flavor and nutritional value to a wide variety of meals.
Continuing research and breeding programs have resulted in the development of enhanced disease and pest resistance, increased yields, and improved adaptability to diverse growing environments in sweet maize varieties (Yadav et al. 2015). The breeding of sweet maize cultivars remain an essential component of modern agriculture, holding significant value for both producers and consumers. Sweet maize, cultivated worldwide, is prized for its distinctive flavor profile and well-established nutritional content (Nuss and Tanumihardjo 2010).
Breeding sweet maize varieties is a complex process that involves many steps (Maqbool et al. 2021). The main steps in breeding sweet maize are variety selection (selection of parental varieties with desirable traits), crossing, progeny selection (progeny price to select plants that best meet breeding criteria) and growth tests (evaluation of performance under different soil and climate conditions) (Paterniani 1990; Shelton and Tracy 2015).
World production of maize has shown a light but steady increase over the years, but human consumption of the grain has remained steady (Ranum et al. 2014). It is thought that the majority of the increase in production has corresponded to an increase in the use of maize for animal feed. Newly developed maize varieties with a very short growing season produce a softer, smaller kernel of white maize (Ranum et al. 2014). Important sweet maize characteristics for growers are earliness, disease resistance, yield, and ear characteristics (Fekonja et al. 2011). Desirable attributes for consumers are sweetness, texture, and flavor (Leksrisompong et al. 2012; Alemu 2023). New varieties of sweet maize have been develop with improved consistency taste and shelf life (Becerra-Sanchez and Taylor 2021). The adoption of never “high sugar or sweeter” varieties of longer shelf life and new sweet maize products have increase sweet maize consumption and have helped to further expand the marked. Sweet maize eating quality of fresh and processed whole-kernel, canned or frozen, is determined by it’s unique combination of flavor, texture, and aroma. Sweetness is the most important factor in consumer satisfaction with sweet maize (Lertrat and Pulam 2007). The results of conducted research indicate that the yield of cobs and length of vegetation season of maize varieties depends on air temperature and rainfall total (Waligóra et al. 2010). In maize breeding programs, the research for genotypes with high grain yield adapted in the most varied environments is one of the most important objectives for breeders. For that, the choice of populations that show good genetic homeostasis is essential for yield increases (Balestre et al. 2009).
In breeding sweet maize varieties, understanding the genotype-environment interaction (GEI) is a key factor in the success of the crop and in achieving maximum yields (Bocianowski et al. 2018, 2019a, 2019b; Katsenios et al. 2021; Nowosad et al. 2023). This interaction refers to how a plant's genotypes respond to changing environmental conditions, such as soil, climate and biotic factors. The genotype-environment interaction allows growers to select varieties of sweet maize that are more resistant to specific diseases or pests found in specific areas. This increases crop yields and reduces the need for crop protection chemicals. Varieties that are adapted to specific climatic conditions can survive extremes of temperature, drought or rainfall, which is extremely important in regions with variable climates. By matching varieties of sweet maize to a specific soil and climate, maximum yields can be achieved, resulting in higher agricultural productivity and greater food availability. Genotype-environment interaction allows sugar maize varieties to vary their genotypes depending on the conditions in which they are grown. This means that breeders can select and develop varieties that will best adapt to specific regions or cropping systems. This avoids yield losses caused by an unsuitable genotype for a given environment.
Breeding sweet maize varieties that show stable yields under a variety of environmental conditions is critical to securing the food supply (Ekpa et al. 2018). Genotype-environment interaction (in our case, hybrid-year interaction) can help identify and develop varieties that not only achieve good yields under typical conditions, but also perform well under more difficult circumstances. For farmers, understanding genotype-environment (or genotype-year or genotype-location) interaction can mean reducing risks associated with unpredictable weather changes or other factors affecting crops (de Leon et al. 2016). Being able to choose varieties adapted to specific conditions can help minimize losses. Multiple statistical methods have been established to elucidate GEI effects within multi-environmental trials. Among these, the Additive Main Effects and Multiplicative Interaction (AMMI) model (Gauch 1992) and Genotype + Genotype × Environment (GGE) biplot analysis (Yan et al. 2007) are robust and widely employed techniques for GEI analysis. The AMMI model leverages a combination of Analysis of Variance (ANOVA) to separate main effects of genotypes (G) and environments (E), and Principal Component Analysis (PCA) incorporating multiplicative terms to capture GEI effects. Both methods rely on PCA, which offers a unique approach to assess interrelationships between environments, genotypes, and their interaction. Additionally, these methods provide a visual representation of the results, facilitating a deeper understanding of how test genotypes respond across diverse environments (Linus et al. 2023). AMMI and GGE biplot methods serve as complementary tools, enabling breeders to not only comprehend GEI effects but also identify suitable environments and superior genotypes. Deciphering GEI is a crucial step in determining genotype adaptation and stability (Luo et al. 2015). These methods have demonstrably facilitated the untangling of complex GEI patterns and the identification of high-yielding, stable genotypes within multi-environmental trials, as evidenced by numerous studies (Wang et al. 2023; Vaezi et al. 2019; Bocianowski and Prażak 2022; Esan et al. 2023; Pour-Aboughadareh et al. 2023a, b). Beyond AMMI and GGE biplot approaches, Best Linear Unbiased Prediction (BLUP) also plays a role in MET GEI analysis. BLUP provides high-efficiency estimations of mean genotype yield within mixed models (Smith et al. 2005). Recently, Olivoto et al. (2019) proposed procedures to address limitations of the AMMI model by integrating its functionalities with BLUP. The Weighted Average of Absolute Scores from the Singular Value Decomposition (SVD) of the BLUP matrix for GEI effects (WAASB) and the WAASB incorporating response variable (WAASBY) support researchers in making informed decisions regarding genotype selection or recommendation, especially when balancing average performance and stability considerations.
The objective of this study was to assess different maize (Zea mays L.) variety by years interaction for cobs yield, lends of cobs and genotype selection index by the AMMI model.
Material and methods
Field experiment
Field experiment carried out in years 2015–2018 in the experimental and didactic department in Złotniki (52°29′ N, 16°49′ E), belonging to Poznań University of Life Sciences as the one factor, randomized complete blocks design in four replications. They were located on soil of bonitation class IVa belonging to the group of fawn, typical formed from light clay sands shallowly lying on light clay. The size of the plots was 24.5 m2 (2.4 m × 8.75 m). In the years of research sowing was carried out in the second half of May, while in October the cobs were harvested by hand in the milky maturity of the grain. Before sowing maize mineral fertilization was applied with nitrogen in the form of urea in the amount of 200 kg ha–1 and phosphorus using 400 kg ha–1 Polifoska M (5-16-21). After sowing maize, the herbicide Lumax 535.5 SE at the dose of 3.5 l ha–1 was applied. For sweet maize grown in 2015 the previous crop was maize, while in the years 2016–2018 was winter wheat.
Plant material
The study used 10 hybrids of sweet maize: Golda, GSS 1453, GSS 3071, GSS 5829, GSS 8529, Overland, Noa, Shinerock, Sindon and Tessa. Cobs yield, lend and fulfilment of cobs, cobs I class trade share pertentage were analyzed. The share of first class was assessed on a sample of all collected cobs. I grades of cobs fully matured were counted.
Weather conditions
Weather conditions were favorable to maize vegetation (Table 1). During the research average temperature in several months where similar to each other but higher than the long term average from 1957–2018. The total precipitation in the period May–September was lower than sum of many years in 2015 by 45 mm and in 2018 by 118.4 mm. In several years the rainfall deficit occurred already in May (only in year 2017 they exceeded the average long term total) and in 2018 it also concern June. July apart from 2018 was more abundant in rainfall than it usually was in many years. August was rain-poor month in 2018, but September in 2016. Analyzing the course of temperature and precipitation, it can be concluded that the most optimal year for maize vegetation was 2016, and the least 2018.
Statistical analysis
Obtained data were analysed using additive main effects and multiplicative interaction (AMMI) model (Gauch and Zobel 1990) for each trait, independently. The AMMI model first fits additive effects for the main effects of genotypes-hybrids (G) and years (Y) followed by multiplicative effects for GYI by principal component analysis. Results of AMMI analysis were presented by biplot graphs. The AMMI model (Nowosad et al. 2016) is given by:
where yg,e,k is the trait mean of genotype (hybrid) g in year e for replicate k, μ is the grand mean, αg is the genotypic mean deviations, βe is the year mean deviations, N is the number of PCA axis retained in the adjusted model, λn is the eigenvalue of the PCA axis n, γg,n is the genotype score for PCA axis n, δe,n is the score eigenvector for PCA axis n, Qg,e is the residual containing all multiplicative terms not included in the model, \({\epsilon }_{g,e,k}\) is the experimental error. The AMMI stability value (ASV) was used to compare the stability of genotype as described by Purchase et al. (2000):
where SSIPCA1 is the sum of squares for IPCA1, SSIPCA2 – the sum of squares for IPCA2, the IPCA1 and IPCA2 scores are the genotypic scores in the AMMI model. Lower ASV score indicate a more stable genotype across years (Nowosad et al. 2017).
Genotype selection index (GSI) was calculated for each hybrid which incorporates both mean of trait and ASV index in single criteria (GSIi) as (Farshadfar and Sutka 2003).
GSIi = RMi + RAi,
where RMi is rank of trait mean (from maximal to minimal) for i-th hybrid, RAi is rank of the ASV for the i-th hybrid. Finally, total genotype selection index (TGSI) was calculated for each hybrid as sum of GSIs for all four traits of study. All the analyses were conducted using the GenStat v. 23.1 statistical software package (GenStat 2023).
Results
The three (genotype, year, GY interaction) sources of variation were highly significant for all four observed traits (Table 2) of maize.
Cobs yield
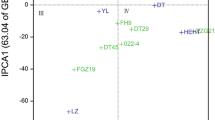
The sum of squares for year main effect represented 65.93% of the total cobs yield variation. The differences between genotypes (hybrids) explained 12.57% of the total cobs yield variation, while the effects of GYI explained 19.27% (Table 2). Values for the two principal components were also significant and accounted jointly for 92.16% of the whole effect it had on the variation of cobs yield. The first principal component (IPCA 1) accounted for 63.96% of the variation caused by interaction, while IPCA 2 accounted for 28.20% (Table 2, Fig. 1). Cobs yield of the tested hybrids varied from 7.8 (for Golda in 2018) to 25 t ha–1 (for GSS 8529 in 2016) throughout the four years, with an average of 14.11 t ha–1 (Table 3). The hybrid GSS 8529 had the highest average cobs yield (16.36 t ha–1), and the hybrid GSS 1453 had the lowest (10.55 t ha–1). The average cobs yield per years also varied from 9.83 t ha–1 in 2017, to 18.26 t ha–1 in 2016 (Table 3). The stability of tested genotypes can be evaluated according to biplot for cobs yield (Fig. 1). Maize hybrids interacted differently with climate conditions in the observed years. The hybrids Overland, Sindom, Tessa and Noa interacted positively with the 2015 (Fig. 1). Hybrids GSS 8529 and Golda interacted positively with 2016, however hybrid GSS 1453 with the 2017 and 2018. The analysis showed that some hybrids have high adaptation; however, most of them have specific adaptability. AMMI stability values (ASV) revealed variations in cobs yield stability among the ten hybrids (Table 3). According to Purchase et al. (2000), a stable variety is defined as one with ASV value close to zero. Consequently, the hybrids GSS 3071 and GSS 5829 with ASV of, respectively, 0.157 and 0.431, were the most stable, while the hybrid GSS 1453 (6.629) was the least stable (Table 3). The hybrid GSS 3071 with high average cobs yield (16.11 t ha–1) and the best ASV equal to 0.157 is hybrids with the best genotype selection index (3).
Cobs I class trade share
In the ANOVA, the sum of squares for year main effect represented 62.79% of the total, and this factor had the highest effect on the cobs I class trade share. The differences between genotypes explained 15.71% of the total cobs I class trade share variation, while the effects of GYI explained 21.11% (Table 2). Values for the two principal components were also significant and accounted jointly for 90.02% of the whole effect it had on the variation of cobs I class trade share (Fig. 2, Table 2). The first principal component (IPCA 1) accounted for 53.35% of the variation caused by interaction, while IPCA 2 accounted for 36.67% (Fig. 2). Cobs I class trade share of the tested genotypes varied from 10 (for Shinerock in 2015) to 97.6% (for GSS 3071 in 2018) throughout the four years, with an average of 63.46% (Table 4). The hybrid GSS 3071 had the highest average cobs I class trade share (77.91%), and the Shinerock had the lowest (49.88%). The average cobs I class trade share per years also varied from 32.18% in 2015, to 78.04% in 2016 (Table 4). The stability of tested genotypes can be evaluated according to biplot for cobs I class trade share (Fig. 2). The Tessa interacted positively with the 2015, but negatively with the 2017 (Fig. 2). Genotypes Shinerock, GSS 8529 and Sindon interacted positively with 2016 and 2017. The GSS 1453 and Noa interacted positively with the 2018, but negatively with the 2016 (Fig. 2). The Overland with ASV of 0.370 was the most stable, while the GSS 8529 (5.108) and Tessa (5.081) were the least stable (Table 4). The genotypes Golda, Overland and Noa with high average cobs I class trade share (69.82%, 67.19% and 74.97%, respectively) and ASV equal to, respectively, 1.149, 0.370 and 2.243 are genotypes with the best genotype selection index (5).
Lend of cobs
The sum of squares for year main effect represented 52.62% of lend of cobs variation. The differences between genotypes explained 17.90% of lend of cobs variation, while the effects of GYI explained 20.52% (Table 2). Values for the two principal components were also highly significant and accounted jointly for 80.63% of the whole effect it had on the variation of lend of cobs. The IPCA 1 and IPCA 2 accounted for 51.94 and 28.70%, respectively, of the variation caused by interaction (Fig. 3). Lend of cobs of the tested hybrids varied from 14.3 cm (for GSS 8529 in 2015) to 22.6 cm (for Golda in 2017) throughout the four years, with an average of 19.37 cm (Table 5). The hybrid GSS 1453 had the highest average lend of cobs (20.38 cm), and the hybrid GSS 5829 the lowest (17.96 cm). The average lend of cobs per years also varied from 17.33 cm in 2015 to 21.06 cm in 2017. The GSS 5829 and Overland interacted positively with the 2015 (Fig. 3). The Tessa and GSS 8529 interacted positively with the 2016 and 2017, however Noa interacted positively with the 2018. The Overland with ASV of 0.254 was were the most stable, while the Sindon (2.383) was the least stable (Table 5). The Overland with the average lend of cobs equal to 19.79 cm and the best ASV equal to 0.254 is genotypes with the best genotype selection index (5).
Fulfilment of cobs
The sum of squares for year main effect represented 72.03% of fulfilment of cobs variation. The differences between genotypes explained 4.28% of fulfilment of cobs variation, while the effects of GYI explained 9.62% (Table 2). Values for the two principal components were also highly significant and accounted jointly for 90.42% of the whole effect it had on the variation of fulfilment of cobs. The IPCA 1 accounted for 46.76% of the variation caused by interaction, while IPCA 2 accounted for 43.66%. Fulfilment of cobs of the tested hybrids varied from 4.7 for Shinerock (2015) to 8.0 (for Golda, GSS 1453, GSS 3071, GSS 5829, Noa and Tessa in 2018) throughout the four years, with an average of 6.69 (Table 6). The Golda had the highest average fulfilment of cobs (6.925), and the Shinerock the lowest (6.3). The average fulfilment of cobs per years also varied from 5.43 in 2015 to 7.8 in 2018. The genotypes GSS 3071, Tessa and Overland interacted positively with the 2015 (Fig. 4). The GSS 1453 interacted positively with the 2016, however the Shinerock interacted positively with the 2017. The cultivar Golda with ASV of 0.047 was the most stable, while the hybrid GSS 1453 (0.999) was the least stable (Table 6). The cultivar Golda with the best the average fulfilment of cobs equal to 6.925 and the best ASV equal to 0.047 is genotypes with the best genotype selection index (2).
Total genotype selection index
The smallest total genotype selection index (for all four traits jointly) were observed for Golda (TGSI = 28) GSS 3071 (TGSI = 32), and Overland (TGSI = 35), while the worst for hybrid GSS 8529 (TGSI = 63). The cultivar Golda and hybrid GSS 3071 are recommended for further inclusion in the breeding programs because of their better stability than other lines and good average values of observed traits.
Discussion
Modern agriculture and food production present many environmental challenges (Soria-Lopez et al. 2023). Sweet maize varieties play an important role in the context of these challenges, both in terms of breeding and impact on ecosystems (Revilla et al. 2021). Some varieties of sweet maize, such as those with biomass-rich root systems, can help protect the soil from erosion (Yadav et al. 2022). Dense roots stabilize soil structure and prevent soil runoff during rainfall (Bodner et al. 2021). Growing different varieties of sweet maize can help increase agricultural biodiversity (Schulz et al. 2020). Different varieties can attract different insect species, which benefits pollinators and other organisms, supporting agricultural ecosystems. Some varieties of sweet maize can help maintain proper water balance in the soil. This is especially important in drought-prone areas, where properly selected varieties can use less water.
Sugar maize cultivation often uses a lot of water, which can be a problem in areas where water resources are limited (Asgharnejad et al. 2021). Over-intensive cultivation can lead to a drop in groundwater levels and contribute to environmental problems related to water resources (Patel 2023). In some cases, the cultivation of varieties of sweet maize may require intensive use of pesticides and chemical fertilizers (Revilla et al. 2021). This can lead to soil and groundwater contamination, and negatively affect insect populations and other organisms in the ecosystem. If sugar maize growing areas are monocultures, meaning that one crop is grown there in a large range, this can lead to a loss of biodiversity and an increased risk of pests and diseases (Tayyab et al. 2021). In the context of ecology, a key challenge is to promote sustainable sugar maize cultivation. This means choosing varieties and practices that minimize negative environmental impacts, such as using greener production methods, reducing water and chemical use, and preserving genetic diversity.
In recent years several methodologies have been proposed with the intention of describing in an easy and precise manner the performance of genotypes in diverse environments (Balestre et al. 2009). Crop yields are influenced by genetic potential and weather conditions (Szempliński and Dubis 2011). In the present study, both main factors (genotypes and years) and their interaction statistically significantly determined all considered quantitative traits. A significant effect of years of research (2017–2020) on the expression of observed traits of sweet maize was observed by Yu et al. (2023) studying whether sweet maize intercropped with soybean improves crops’ fresh grain yield and carbon and nitrogen accumulation compared to sweet maize monoculture. In the results presented here, a statistically significant effect of genotype-environment interaction on all observed traits was obtained. Similar results were obtained by other researchers analyzing sweet maize (Bachireddy et al. 1992; Zystro et al. 2021; Adham et al. 2022; Stansluos et al. 2023; Ye et al. 2023).
The result obtained from AMMI analyses are very important for developing and recommending best maize variety for production in a specific year. This method allows the analysis of unbalanced data sets naturally and is an efficient method for multi-environment studies (Bernardo Júnior et al. 2018). Babić et al. (2006) in their researches state that many authors showed the possibility of developing highly yielding and stable hybrids. The assumption was that commercial maize hybrids were characterized not only by the level of average yield but also by their stability. The authors state that they are significant differences in yield and some yield components between genotypes, environments and interaction G × L and there was a significant share of the genotype of the components and interaction G × L and G × Y in total phenotypic for yield and some of the components of the yield. The use of the AMMI model allowed an accurate analysis of GEI and the selection of the most stable varieties in all years of the study. Genotypes specific to conditions in particular years were also identified. Similar results in studies on sweet corn using the AMMI model have been obtained by other researchers. Choudhary et al. (2019) studied 13 hybrids in eight environments. Identification of stable (out of 45) sweet maize genotypes for breeding purposes through GEI analysis of green cob yield in a wide range of environments in western India (three localities in two years) was studied by Patel et al. (2023). Stability and adaptability of yield among earliness 18 sweet maize hybrids in six environments in West Java, Indonesia studied Ruswandi et al. (2020). Mousavi et al. (2023) used the AMMI model to evaluate GE interactions of eight genotypes in 16 environments in Debrecen, Hungary.
Conclusions
The analysis of AMMI stability values showed that the most stable in terms of the effect of environmental conditions on cob yield were the GSS 3071 and GSS 5829 varieties, while the least stable was the GSS 1453 variety. The Overland variety, showed the best genotype selection index for cob length stability. On the other hand, the Golda, Overland and Noa varieties had the highest share of first commercial grade cobs in yield, and the Golda variety had the highest stability of cob filling with maize grain.
Data availability
The datasets generated during and analysed during the current study are available from the corresponding author on reasonable request.
References
Adham A, Ghaffar MBA, Ikmal AM, Shamsudin NAA (2022) Genotype × environment interaction and stability analysis of commercial hybrid grain corn genotypes in different environments. Life 12(11):1773. https://doi.org/10.3390/life12111773
Alemu T (2023) Texture profile and design of food product. J Agri Horti Res 6(2):272–281
Amjath-Babu TS, Krupnik TJ, Thilsted SH, McDonald AJ (2020) Key indicators for monitoring food system disruptions caused by the COVID-19 pandemic: Insights from Bangladesh towards effective response. Food Sec 12:761–768. https://doi.org/10.1007/s12571-020-01083-2
Asgharnejad H, Nazloo EK, Larijani MM, Hajinajaf N, Rashidi H (2021) Comprehensive review of water management and wastewater treatment in food processing industries in the framework of water-food-environment nexus. Compr Rev Food Sci Food Saf 20:1–37. https://doi.org/10.1111/1541-4337.12782
Babić V, Babić M, Delić N (2006) Stability parameters of commercial maize (Zea mays L.) hybrids. Genetika Belgrade 38(3):235–240
Bachireddy VR, Payne R Jr, Chin KL (1992) Conventional selection versus methods that use genotype × environment interaction in sweet corn trials. HortScience 7(5):436–438
Balestre M, Von Pinho RG, Souza JC, Oliveira RL (2009) Genotypic stability and adaptability in tropical maize based on AMMI and GGE biplot analysis. Genet Mol Res 8(4):1311–1322
Becerra-Sanchez F, Taylor G (2021) Reducing post-harvest losses and improving quality in sweet corn (Zea mays L): challenges and solutions for less food waste and improved food security. Food Energy Secur 10:e277. https://doi.org/10.1002/fes3.277
Bernardo Júnior LAY, de Silva CP, de Oliveira LA, Nuvunga JJ, Pires LPM, Von Pinho RG, Balestre M (2018) AMMI Bayesian models to study stability and adaptability in maize. Agron J 110:1765–1776. https://doi.org/10.2134/agronj2017.11.0668
Bocianowski J, Prażak R (2022) Genotype by year interaction for selected quantitative traits in hybrid lines of Triticum aestivum L. with Aegilops kotschyi Boiss. and Ae. variabilis Eig using the additive main effects and multiplicative interaction model. Euphytica 218:11
Bocianowski J, Szulc P, Nowosad K (2018) Soil tillage methods by years interaction for dry matter of plant yield of maize (Zea mays L.) using additive main effects and multiplicative interaction model. J Integr Agric 17(12):2836–2839. https://doi.org/10.1016/S2095-3119(18)62085-4
Bocianowski J, Nowosad K, Tomkowiak A (2019a) Genotype—environment interaction for seed yield of maize hybrids and lines using the AMMI model. Maydica 64:M13
Bocianowski J, Nowosad K, Szulc P (2019b) Soil tillage methods by years interaction for harvest index of maize (Zea mays L) using additive main effects and multiplicative interaction model. Acta Agric Scand Sect B 69(1):75–81. https://doi.org/10.1080/09064710.2018.1502343
Bodner G, Mentler A, Keiblinger K (2021) Plant roots for sustainable soil structure management in cropping systems. In: Rengel Z, Djalovic I (eds) The root systems in sustainable agricultural intensification. Wiley, New York. https://doi.org/10.1002/9781119525417.ch3
Choudhary M, Kumar B, Kumar P, Guleria SK, Singh NK, Khulbe R, Kamboj MC, Vyas M, Srivastava RK, Puttaramanaik Swain D, Mahajan V, Rakshit S (2019) GGE biplot analysis of genotype × environment interaction and identification of mega-environment for baby corn hybrids evaluation in India. Indian J Genet 79(4):658–669. https://doi.org/10.31742/IJGPB.79.4.3
Crupi P, Faienza MF, Naeem MY, Corbo F, Clodoveo ML, Muraglia M (2023) Overview of the potential beneficial effects of carotenoids on consumer health and well-being. Antioxidants 12(5):1069. https://doi.org/10.3390/antiox12051069
de Leon N, Jannink JL, Edwards JW, Kaeppler SM (2016) Introduction to a special issue on genotype by environment interaction. Crop Sci 56:2081–2089. https://doi.org/10.2135/cropsci2016.07.0002in
Ekpa O, Palacios-Rojas N, Kruseman G, Fogliano V, Linnemann AR (2018) Sub-Saharan African maize-based foods: Technological perspectives to increase the food and nutrition security impacts of maize breeding programmes. Global Food Secur 17:48–56. https://doi.org/10.1016/j.gfs.2018.03.007
Esan VI, Oke GO, Ogunbode TO, Obisesan IA (2023) AMMI and GGE biplot analyses of Bambara groundnut [Vigna subterranea (L) Verdc] for agronomic performances under three environmental conditions. Front Plant Sci 13:997429
Farshadfar E, Sutka J (2003) Locating QTLs controlling adaptation in wheat using AMMI model. Cereal Res Comm 31(3):249–256. https://doi.org/10.1007/BF03543351
Fekonja M, Bavec F, Grobelnik-Mlakar S, Turinek S, Jakop M, Bavec M (2011) Growth performance of sweet maize under non-typical maize growing conditions. Biol Agric Hortic 27(2):147–164. https://doi.org/10.1080/01448765.2011.9756644
Galani YJH, Orfila C, Gong YY (2022) A review of micronutrient deficiencies and analysis of maize contribution to nutrient requirements of women and children in Eastern and Southern Africa. Crit Rev Food Sci Nutr 62(6):1568–1591. https://doi.org/10.1080/10408398.2020.1844636
Garg M, Sharma A, Vats S, Tiwari V, Kumari A, Mishra V, Krishania M (2021) Vitamins in cereals: a critical review of content, health effects, processing losses, bioaccessibility, fortification, and biofortification strategies for their improvement. Front Nutr 8:586815. https://doi.org/10.3389/fnut.2021.586815
Gauch HG (1992) Statistical analysis of regional trials. AMMI Analysis of factorial design, 1st edn. Elsevier, New York, p 278
Gauch HG, Zobel RW (1990) Imputing missing yield trial data. Theor Appl Genet 79:753–761
Jiménez-Viveros Y, Valiente-Banuet JI (2023) Colored shading nets differentially affect the phytochemical profile, antioxidant capacity, and fruit quality of piquin peppers (Capsicum annuum L. var. glabriusculum). Horticulturae 9(11):1240. https://doi.org/10.3390/horticulturae9111240
Katsenios N, Sparangis P, Leonidakis D, Katsaros G, Kakabouki I, Vlachakis D, Efthimiadou A (2021) Effect of genotype × environment interaction on yield of maize hybrids in Greece using AMMI analysis. Agronomy 11(3):479. https://doi.org/10.3390/agronomy11030479
Leksrisompong PP, Whitson ME, Truong VD, Drake MA (2012) Sensory attributes and consumer acceptance of sweet potato cultivars with varying flesh colors. J Sens Stud 27:59–69. https://doi.org/10.1111/J.1745-459x.2011.00367.X
Lertrat K, Pulam T (2007) Breeding for increased Sweetness in sweet corn. Int J Plant Breed 1(1):27–30
Linus RA, Olanrewaju OS, Oyatomi O, Idehen EO, Abberton M (2023) Assessment of yield stability of bambara groundnut (Vigna subterranea (L.) Verdc.) using genotype and genotype-environment interaction biplot analysis. Agronomy 13:2558
Luo J, Pan YB, Que Y, Zhang H, Grisham MP, Xu L (2015) Biplote valuation of test environments and identification of mega-environment for sugarcane cultivars in China. Sci Rep 5:1–11
Maqbool MA, Beshir Issa A, Khokhar ES (2021) Quality protein maize (QPM): importance, genetics, timeline of different events, breeding strategies and varietal adoption. Plant Breed 140:375–399. https://doi.org/10.1111/pbr.12923
Marenya P, Wanyama R, Alemu S, Westengen O, Jaleta M (2022) Maize variety preferences among smallholder farmers in Ethiopia: Implications for demand-led breeding and seed sector development. PLoS ONE 17(9):e0274262. https://doi.org/10.1371/journal.pone.0274262
Mousavi SMN, Illés A, Szabó A, Shojaei SH, Demeter C, Bakos Z, Vad A, Széles A, Nagy J, Bojtor C (2023) Stability yield indices on different sweet corn hybrids based on AMMI analysis. Braz J Biol 84:270680. https://doi.org/10.1590/1519-6984.270680
Nowosad K, Liersch A, Popławska W, Bocianowski J (2016) Genotype by environment interaction for seed yield in rapeseed (Brassica napus L.) using additive main effects and multiplicative interaction model. Euphytica 208:187–194
Nowosad K, Liersch A, Poplawska W, Bocianowski J (2017) Genotype by environment interaction for oil content in winter oilseed rape (Brassica napus L.) using additive main effects and multiplicative interaction model. Indian J Genet Plant Breed 77(2):293–297
Nowosad K, Bocianowski J, Kianersi F, Pour-Aboughadareh A (2023) Analysis of linkage on interaction of main aspects (genotype by environment interaction, stability and genetic parameters) of 1000 kernels in maize (Zea mays L). Agriculture 13(10):2005. https://doi.org/10.3390/agriculture13102005
Nuss ET, Tanumihardjo SA (2010) Maize: a paramount staple crop in the context of global nutrition. Comp Rev Food Sci Food Saf 9:417–436. https://doi.org/10.1111/j.1541-4337.2010.00117.x
Olivoto T, Licio ADC, da Silva JAG, Marchioro VS, de Souza VQ, Jost E (2019) Mean performance and stability in multienvironment trials I: combining features of AMMI and BLUP techniques. Agron J 111:2949–2960
Palacios-Rojas N, McCulley L, Kaeppler M, Titcomb TJ, Gunaratna NS, Lopez-Ridaura S, Tanumihardjo SA (2020) Mining maize diversity and improving its nutritional aspects within agro-food systems. Compr Rev Food Sci Food Saf 19:1809–1834. https://doi.org/10.1111/1541-4337.12552
Patel PM (2023) Agricultural transformations in the arid, drought-prone region of Kachchh: people-led, market-oriented growth under adverse climatic conditions. Front Sustain Food Syst 7:1159011. https://doi.org/10.3389/fsufs.2023.1159011
Patel R, Parmar DJ, Kumar S, Patel DA, Memon J, Patel MB, Patel JK (2023) Dissection of genotype × environment interaction for green cob yield using AMMI and GGE biplot with MTSI for selection of elite genotype of sweet corn (Zea mays conva. Saccharata var. rugosa). Indian J Genet Plant Breed 83(1):59–68
Paterniani E (1990) Maize breeding in the tropics. Crit Rev Plant Sci 9(2):125–154. https://doi.org/10.1080/07352689009382285
Pour-Aboughadareh A, Ghazvini H, Jasemi SS, Mohammadi S, Razavi SA, Chaichi M, Ghasemi Kalkhoran M, Monirifar H, Tajali H, Fathihafshjani A, Bocianowski J (2023a) Selection of high-yielding and stable genotypes of barley for the cold climate in Iran. Plants 12:2410
Pour-Aboughadareh A, Koohkan SA, Zali H, Marzooghian A, Gholipour A, Kheirgo M, Barati A, Bocianowski J, Askari-Kelestani A (2023b) Identification of high-yielding genotypes of barley in the warm regions of Iran. Plants 12:3837
Purchase JL, Hatting H, van Deventer CS (2000) Genotype × environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. South African J Plant Soil 17:101–107
Ranum P, Peña-Rosas JP, Garcia-Casal MN (2014) Global maize production, utilization, and consumption. Ann NY Acad Sci 1312:105–112. https://doi.org/10.1111/nyas.12396
Revilla P, Anibas CM, Tracy WF (2021) Sweet Corn Research around the World 2015–2020. Agronomy 11(3):534. https://doi.org/10.3390/agronomy11030534
Ruswandi D, Yuwariah Y, Ariyanti M, Syafii M, Nuraini A (2020) Stability and adaptability of yield among earliness sweet corn hybrids in West Java, Indonesia. Int J Agron 2020:4341906. https://doi.org/10.1155/2020/4341906
Schulz VS, Schumann C, Weisenburger S, Müller-Lindenlauf M, Stolzenburg K, Möller K (2020) Row-intercropping maize (Zea mays L.) with biodiversity-enhancing flowering-partners—effect on plant growth, silage yield, and composition of harvest material. Agriculture 10(11):524. https://doi.org/10.3390/agriculture10110524
Shelton AC, Tracy WF (2015) Recurrent selection and participatory plant breeding for improvement of two organic open-pollinated sweet corn (Zea mays L) populations. Sustainability 7(5):5139–5152. https://doi.org/10.3390/su7055139
Shirke GD, Pinjarkar MS (2023) Post-harvest technology of tree spices. J Pharmacognosy Phytochemistry 12(2):88–102
Smith AB, Cullis BR, Thompson R (2005) The analysis of crop cultivar breeding and evaluation trials: an overview of current mixed model approaches. J Agric Sci 143:449–462
Soria-Lopez A, Garcia-Perez P, Carpena M, Garcia-Oliveira P, Otero P, Fraga-Corral M, Cao H, Prieto MA, Simal-Gandara J (2023) Challenges for future food systems: From the Green Revolution to food supply chains with a special focus on sustainability. Food Front 4:9–20. https://doi.org/10.1002/fft2.173
Stansluos AAL, Öztürk A, Niedbała G, Türkoğlu A, Haliloğlu K, Szulc P, Omrani A, Wojciechowski T, Piekutowska M (2023) Genotype-Trait (GT) biplot analysis for yield and quality stability in some sweet corn (Zea mays L. saccharata sturt) genotypes. Agronomy 13(6):1538. https://doi.org/10.3390/agronomy13061538
Swapna G, Jadesha G, Mahadevu P (2020) Sweet corn: a future healthy human nutrition food. Int J Curr Microbiol App Sci 9(07):3859–3865. https://doi.org/10.20546/ijcmas.2020.907.452
Szempliński W, Dubis G (2011) Wstępne badania nad plonowaniem i wydajnością energetyczną wybranych roślin uprawianych na cele biogazowe. Fragm Agron 28(1):77–86
Tayyab M, Yang Z, Zhang C, Islam W, Lin W, Zhang H (2021) Sugarcane monoculture drives microbial community composition, activity and abundance of agricultural-related microorganisms. Environ Sci Pollut Res 28:48080–48096. https://doi.org/10.1007/s11356-021-14033-y
Vaezi B, Pour-Aboughadareh A, Mohammadi R, Mehraban A, Hossein-Pour T, Koohkan E, Ghasemi S, Moradkhani H, Siddique KHM (2019) Integrating different stability models to investigate genotype × environment interactions and identify stable and highyielding barley genotypes. Euphytica 215:63
VSN International (2023) VSN international genstat for windows, 23rd edn. VSN International, Hemel Hempstead
Vukadinović J, Srdić J, Tosti T, Dragicević V, Kravić N, Mladenović Drinić S, Milojković-Opsenica D (2022) Alteration in phytochemicals from sweet maize in response to domestic cooking and frozen storage. J Food Compos Anal 114:104637. https://doi.org/10.1016/j.jfca.2022.104637
Waligóra H, Skrzypczak W, Weber A, Szulc P (2010) Plonowanie i długość okresu wegetacji kilku odmian kukurydzy cukrowej w zależności od warunków pogodowych. Nauka Przyr Technol 4(1):5
Wang R, Wang H, Huang S, Zhao Y, Chen E, Li F, Qin L, Yang Y, Guan Y, Liu B, Zhang H (2023) Assessment of yield performances for grain sorghum varieties by AMMI and GGE biplot analyses. Front Plant Sci 14:1261323
Yactayo-Chang JP, Boehlein S, Beiriger RL, Resende MFR, Bruton RG, Alborn HT, Romero M, Tracy WF, Block AK (2022) The impact of post-harvest storage on sweet corn aroma. Phytochem Lett 52:33–39. https://doi.org/10.1016/j.phytol.2022.09.001
Yadav OP, Hossain F, Karjagi CG, Kumar B, Zaidi PH, Jat SL, Chawla JS, Kaul J, Hooda KS, Kumar P, Yadava P, Dhillon BS (2015) Genetic improvement of maize in India: retrospect and prospects. Agric Res 4:325–338. https://doi.org/10.1007/s40003-015-0180-8
Yadav D, Singh DV, Singh RJ, Babu S, Sharma NK, Singh D, Kumawat A, Kumar P, Mishra M (2022) Evaluation & development of cultivation techniques of rainfed maize + sweet potato inter-cropping under Indian North-Western Himalaya. Food Sci Eng 3(1):43–55
Yan W, Kang MS, Ma B, Woods S, Cornelius PL (2007) GGE biplot vs. AMMI analysis of genotype-by-environment data. Crop Sci 47:643–653
Ye D, Chen J, Yu Z, Sun Y, Gao W, Wang X, Zhang R, Zaib-Un-Nisa SuD, Atif Muneer M (2023) Optimal plant density improves sweet maize fresh ear yield without compromising grain carbohydrate concentration. Agronomy 13(11):2830. https://doi.org/10.3390/agronomy13112830
Yu X, Xiao S, Yan T, Chen Z, Zhou Q, Pan Y, Yang W, Lu M (2023) Interspecific competition as affected by nitrogen application in sweet corn-soybean intercropping system. Agronomy 13(9):2268. https://doi.org/10.3390/agronomy13092268
Zhang B, Li K, Cheng H, Hu J, Qi X, Guo X (2023) Effect of thermal treatments on volatile profiles and fatty acid composition in sweet corn (Zea mays L). Food Chem X 18:100743. https://doi.org/10.1016/j.fochx.2023.100743
Zystro J, Peters TE, Miller KM, Tracy WF (2021) Inbred and hybrid sweet corn genotype performance in diverse organic environments. Crop Sci 61:2280–2293. https://doi.org/10.1002/csc2.20457
Acknowledgements
The authors are grateful for support provided by the Poznan University of Life Sciences, Poland, including technical support and materials used for field experiments.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization, J.B., H.W. and L.M.; methodology, J.B. and H.W.; software, J.B.; validation, J.B., L.M. and H.W.; formal analysis, J.B.; investigation, L.M.; resources, H.W.; data curation, L.M.; writing—original draft preparation, J.B.; writing—review and editing, L.M.; visualization, J.B.; supervision, H.W.; project administration, J.B.; funding acquisition, J.B. and L.M. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bocianowski, J., Waligóra, H. & Majchrzak, L. Genotype by year interaction for selected traits in sweet maize (Zea maize L.) hybrids using AMMI model. Euphytica 220, 89 (2024). https://doi.org/10.1007/s10681-024-03352-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-024-03352-z