Abstract
Genotype-environment interaction consists of the different response of individual genotypes resulting from changing environmental conditions. Its significance is a phenomenon that makes the breeding process very difficult. On the one hand, the breeder expects stable genotypes, i.e., yielding similarly regardless of environmental conditions. On the other hand, selecting the best genotypes for each region is one of the key challenges for breeders and farmers. The aim of this study was to evaluate genotype-by-environment interaction for grain yield in new maize hybrids developed by Plant Breeding Smolice Co. Ltd., utilizing the additive main effects and multiplicative interaction (AMMI) model. The investigation involved 69 maize (Zea mays L.) hybrids, tested across five locations in a randomized complete block design with three replications. Grain yield varied from 8.76 t ha–1 (SMH_16417 in Smolice) to 16.89 t ha–1 (SMH_16043 in Płaczkowo), with a mean yield of 13.16 t ha–1. AMMI analysis identified significant effects of genotype, environment, and their interaction on grain yield. Analysis of variance indicated that 25.12% of the total variation in grain yield was due to environment factor, 35.20% to genotypic differences, and 21.18% to genotype by environmental interactions. Hybrids SMH_1706 and SMH_1707 are recommended for further breeding programs due to their high stability and superior average grain yield.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Environmental conditions can influence genotypes in various ways, leading to genotype by environment interactions (GEI) (Grishkevich and Yanai 2013; de Leon et al. 2016; Baldassarre et al. 2023; Gan et al. 2023; Oroian et al. 2023). While some genotypes exhibit consistent phenotypic traits across different environments, others demonstrate significant variability (Williams et al. 2008; Khare et al. 2024; Kumar et al. 2024). These interactions can cause substantial variation in yield performance among different genotypes under varying environmental conditions (Shrestha et al. 2012; Oladosu et al. 2017; Chauhan et al. 2023; Pour-Aboughadareh et al. 2023). GEI can be statistically defined as the discrepancy between the observed phenotypic performance and the predicted value from a model that considers the overall mean and main effects of genotype and environment (Bocianowski and Prażak 2022; Nowosad et al. 2023). Multi-environmental studies often reveal that traits such as yield and its components vary under different environmental conditions, classifying somo varieties as unstable (Vaezi et al. 2019; Ruswandi et al. 2023; Saeidnia et al. 2023). While major environmental effects can explain of this variation, GEI also contribute significantly to the differing performances of genotypes across environments. Breeders and farmers prefer varieties that exhibit stability or minimal environmental modification (Ceccarelli 1994; Atlin et al. 2017; von Gehren et al. 2023; Abdala et al. 2024). Assessing the stability of a genotype involves testing its general and specific adaptation through a series of trails (Ahakpaz Karkaji et al. 2023; Amelework et al. 2023; do Couto et al. 2023; Fekadu et al. 2023; Taleghani et al. 2023).
One primary goal in maize breeding is to consistently increase grain yield to enhance productivity (Duvick 2005). Understanding the genetic determinants of grain yield aids breeders in managing genetic improvements (Boote et al. 2001; Swarup et al. 2021). The most desirable varieties for agriculture are those with stable, high average yields or other essential traits, adapted widely to the target region (Ceccarelli 1994; Saeidnia et al. 2023; Kwambai et al. 2024). Varieties with narrow adaptation to specific habitat conditions are less frequently preferred.
Grain yield is a complex quantitative trait influenced by genotype, environmental factors, and GEIs. The differential response of genotypes to varying environmental conditions during plant growth adds to this complexity. GEI is often analyzed using the additive main effects and multiplicative interaction (AMMI) model (Mandel 1961, 1971; Gauch 1988; Zobel et al. 1988). The AMMI model, as proposed by Crossa (1990), combines the strengths of analysis of variance (ANOVA) and principal component analysis (PCA) through singular value decomposition (SVD). Initially, AMMI analysis estimates the main effects of genotypes and environments using ANOVA (Ferraudo and Perecin 2014; Hongyu et al. 2014; Demelash 2024; Patel et al. 2024).
The objective of this study was to evaluate genotype by environment interaction for grain yield in maize (Zea mays L.) grown in South Poland using the AMMI model.
Material and methods
Plant material
The study material comprised 69 maize (Zea mays L.) genotypes, including 66 experimental hybrids bred by Plant Breeding Smolice Ltd. IHAR Group and three cultivars (NK Ravello, Ricardinio, and ES Gallery). The 66 experimental hybrids are genotypes intended for use for grain, resulting from the crossbreeding of two opposite heterotic groups, which are forms with grain type dent and forms with grain type flint. These objects are characterized by intermediate flint/dent grain type. Thirty-one of them are single-cross varieties, and 35 objects are three-way cross varieties. The earliness of the hybrids tested ranged from about FAO 210 to about FAO 280. The final FAO value is indicated by Research Center for Cultivar Testing after 2-year registration experiments based on comparison with other reference varieties. The reference varieties are registered hybrids representing different earliness groups: NK Ravello, single-cross cultivar, grain type: flint, FAO number 210, intended for grain cultivation; Ricardinio, single-cross cultivar, grain type: flint/dent, FAO number 230/240, intended for grain and whole-crop silage cultivation; and ES Gallery, single-cross cultivar, grain type: dent, FAO number 270, intended for grain cultivation.
Field experiment
Field experiments were conducted across five locations with varying weather and soil conditions: Kobierzyce, Mikulice, Płaczkowo, Radzików and Smolice (Table 1, Fig. 1) in 2017. These experiments were arranged in a randomized complete block design with three replicates, on plots of 5.01 m2 (1.5 m × 3.34 m). Sowing was conducted using a precision drill in the second (E3, E5) or third decade of April (E1, E2, E4) and harvesting took place in the first (E2, E3) or second decade of October (E1, E4, E5). Agronomic practices were in accordance with accepted agricultural practice appropriate for each location. In accordance with this agricultural practice and knowledge of the field, winter plowing and spring tillage treatments appropriate to field conditions were applied. The following treatments were carried out: spear plowing, harrowing, and pre-sowing cultivation with tillage units. The following fertilization was applied in each locality: Kobierzyce, 125 kg ha–1 N, 50 kg ha–1 P, 120 kg ha–1 K; Mikulice, 139 kg ha–1 N, 80 kg ha–1 P, 120 kg ha–1 K; Płaszkowo, 160 kg ha–1 N, 94 kg ha–1 P, 170 kg ha–1 K, Radzików, 170 kg ha–1 N, 80 kg ha–1 P, 120 kg ha–1 K; Smolice, 145 kg ha–1 N, 110 kg ha–1 P, 160 kg ha–1 K. The preceding crops were maize (E1, E4, E5), winter rapeseed (E2), and winter wheat (E3). The experiments were conducted on class II (E3, E4), III a (E1, E2), and III b (E5) soils. Grain yield was standardized to 14% moisture content, and dry matter content was measured using the dryer method. In addition to the basic agronomic traits of yield and dry matter content at harvest, the following were also observed: number of plants infested with head blight (pcs), pre-harvest lodging (pcs), and cob flowering—the number of days from sowing to the appearance of nevi in 50% of the plants in the plot, plant height (cm) and height of setting of the first cob (cm).
Statistical analysis
A two-way fixed effect model was applied to quantify the main effects of variability and their interaction on grain yield. Least-squares means were calculated using the AMMI model. Initially, the model fits additive effects for the main effects of genotypes (G) and environments (E), followed by multiplicative effects for GEI using principal component analysis (PCA). The AMMI model (Gauch and Zobel 1990) is formulated as
where yg,e,k is the grain yield mean of genotype g in environment e for replicate k, μ is the grand mean, αg is the genotypic mean deviation, βe is the environmental mean deviation, N is the number of PCA axes retained in the adjusted model, λn is the eigenvalue of the PCA axis n, γg,n is the genotype score for PCA axis n, δe,n is the score eigenvector for PCA axis n, Qg,e is the residual containing all multiplicative terms not included in the model, and ϵg,e,k is the experimental error. The expected distribution of Qg,e is normal. The interaction matrix, post-residualization captures the GEI effects, assumed to be multiplicative and estimated via PCA. The approximate F-test by Gollob (1968) was used to determine the optimal number of multiplicative terms required within the AMMI model.
The AMMI stability value (ASV) was used to assess genotype stability (Purchase et al. 2000):
where SS is the sum of squares and IPCA1 and IPCA2 are the first and the second principal component axes of the interaction, respectively. The IPCA1 and IPCA2 scores are the genotypic scores in the AMMI model. A lower ASV indicates greater genotype stability across environments.
A genotype selection index (GSI) was calculated for each genotype, combining average grain yield and ASV into a single criterion (GSIi) as (Farshadfar and Sutka 2003):
where GSIi is the genotype selection index for i-th genotype, RYi is the rank of the average grain yield for i-th genotype, and RASVi is the rank of the AMMI stability value for the i-th genotype. All analyses were performed using GenStat v. 23.1 statistical software (VSN International 2023).
Results
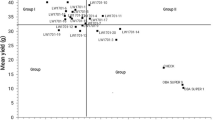
The analysis of variance revealed that the sum of squares for the main effect of genotypes constituted 35.20% of the total variation, having the most significant impact on grain yield. Environmental differences accounted for 25.12% of the total grain yield variation, while GEI effects explained 21.18% (Table 2). These components were all highly significant. The first three principal components of GEI collectively explained 84.30% of the total variation in grain yield, with the first principal component (IPCA 1) accounting for 33.65%, IPCA 2 for 29.54%, and IPCA 3 for 21.11% (Fig. 2, Table 2).
Field test results highlighted the influence of weather conditions, environment, and genotypes on grain maize yield (Table 1). The grain yield of the tested genotypes varied from 8.76 t ha–1 (SMH_16417 in Smolice) to 16.89 t ha–1 (SMH_16043 in Płaczkowo) across five locations, averaging 13.16 t ha–1 (Table 3). Line SMH_1706 had the highest average yield (14.92 t ha–1), while line SMH_16417 had the lowest (10.87 t ha–1). Average grain yield per location ranged from 11.81 t ha–1 in Smolice to 14.01 t ha–1 in Płaczkowo.
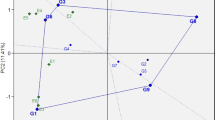
The AMMI1 biplot (Fig. 2) illustrates genotype and environment stability, along with specific GEIs. Among the genotypes, line SMH_16129 had the highest IPCA1 value of 0.630, whereas line SMH_16099 had the lowest IPCA1 value was –0.858 (Fig. 2, Table 2). Among the environments, Kobierzyce had the lowest IPCA1 value (–1.197), while Smolice had the highest (1.957) (Table 2, Fig. 2). Genotype stability, which indicates consistent performance across varying environmental conditions, was mainly influenced by climatic factors in this study. Figure 2 categorizes genotypes with specific adaptations into four groups. The stability of genotypes is further assessed by the biplot for grain yield (Fig. 3). Genotypes G59 (SMH_1723) and G69 (ES Gallery) interacted positively with Kobierzyce and Mikulice; line G14 (SMH_16145) with Radzików; G43 (SMH_1708) and G44 (SMH_1709) with Płaczkowo; and lines G01 (SMH_1682), G10 (SMH_16129), G17 (SMH_16201), and G33 (SMH_16461) with Smolice (Figs. 2 and 3). The analysis indicated that while some genotypes exhibit broad adaptation, most show specific adaptation. AMMI stability values (ASV) demonstrated variability in grain yield stability among the 69 genotypes (Table 3). A stable variety is characterized by an ASV value close to zero. Consequently, lines SMH_1706, SMH_1710, SMH_1729, SMH_1711, and SMH_1707, with ASV values of 0.124, 0.131, 0.132, 0.133, and 0.136 respectively, were the most stable. Conversely, lines such as SMH_16099 (0.982), SMH_1723 (0.919), SMH_1724 (0.919), and SMH_1720 (0.833) were the least stable (Table 3). Genotypes located at the highest point in specific sections of the graph performed best in environments within the same section (Fig. 3). On the biplot, lines SMH_16309 and SMH_1729 stood out with average grain yields of 13.14 and 13.22 t ha–1, respectively, close to the overall average of 13.18 t ha–1. Line SMH_1729 was particularly notable for its high stability. Lines SMH_1706, SMH_16043, and the cultivar ES Gallery exhibited the highest average grain yields but with different adaptations (Figs. 2 and 3): SMH_16043 showed specific adaptation to Płaczkowo, ES Gallery cultivar to Kobierzyce and Mikulice, and SMH_1706 demonstrated the greatest stability and the highest average grain yield (14.92 t ha–1). SMH_1706 also achieved the best genotype selection index of 2 (Table 3).
Discussion
Effect of environmental conditions on biological and commercial traits of maize is very significant. The results of field trials demonstrated the impact of weather conditions (especially influence of monthly rainfall), environment, and genotypes on the grain yield of maize hybrids. In 2017, the spring season was notably cool, with substantial precipitation occurring during the latter half. Grain yield in maize (Zea mays L.) is influenced by multiple genes, resulting in varying genotype performance across different environments (Singamsetti et al. 2021; Alam et al. 2022; Kimutai et al. 2023; Ljubičić et al. 2023; Sabitha et al. 2024). Alam et al. (2022) showed that the phenotypic coefficient of variance had a higher value than the genotypic coefficient of variance, indicating the influence of environment on yield expression. In addition, they found that high heritability, combined with high genetic advance, indicated additive effects of genes. Kimutai et al. (2023) identified some QTLs for maize yield under nitrogen regimes. These QTLs have significant value for further validation and possible rapid introgression into maize populations using marker-assisted selection. In this study, three sources of variation were highly significant, consistent with findings from other researchers. For instance, Yue et al. (2022) analyzed 18 advanced maize genotypes and one check hybrid over 2 years at 37 locations across seven provinces. Nzuve et al. (2013) examined the interaction of 49 genotypes in three environments, while Signor et al. (2001) reviewed data from the French Association Générale des Producteurs de Maïs trial network, including 132 hybrids and 229 environments over 12 years. They conducted GEI analysis for datasets of 1 year, for 2 consecutive years, and for a dataset of 12 years. The results indicate that the magnitude of the GEI variance was equal to or greater than the genotypic variance (Signor et al. 2001). Ljubičić et al. (2023) studied four genotypes of different maturity classes and vegetation periods at one location over two growing seasons. Matongera et al. (2023) evaluated 24 inbred lines from various nutritional categories for GEI across stress and non-stress environments over 2 years at 11 sites in Zimbabwe. do Couto et al. (2023) analyzed 13 varieties in nine locations based on their widespread cultivation. Bocianowski et al. (2019c) investigated 32 maize genotypes (19 inbred lines and their 13 F1 hybrids) in four environments (2 years in two locations) for grain yield. Dehghani et al. (2009) and Adu et al. (2013) found non-significant GEI for grain yield, with Dehghani et al. (2009) studying 12 late maize hybrids in 11 sites over two growing seasons in Iran and Adu et al. (2013) analyzing 100 genotypes (98 single-cross maize hybrids, one local three-way hybrid, Akposoe, and an open-pollinated variety, Dodzi) in three environments. Adu et al. (2013) observed a lack of significant GEI for grain yield, which they interpret as consistent expression of this trait across all environments tested. Multi-location experiments often use the AMMI biplot to visualize the main effect of genotypes in different environments. The AMMI model is frequently employed in multi-species studies (Brancourt-Hulmel and Lecomte 2003; Pacheco et al. 2005; Balestre et al. 2009; Abakemal et al. 2016; Edwards 2016; Pires et al. 2018; Bocianowski and Liersch 2022; Mousavi et al. 2024).
It is a valuable tool for identifying GEI patterns and improving response estimate accuracy. It allows for grouping genotypes based on response characteristics and detecting trends across environments (Bocianowski et al. 2019a, 2019b, 2021; Patel et al. 2023; Yang et al. 2023; Pramanik et al. 2024; Singh et al. 2024).
Researchers can identify robust varieties with competitive yields in various environments using this approach, extracting more information from GEIs (Nowosad et al. 2018). In Southern Poland, grain yield expression is significantly influenced by genotype and environment main effects, as well as GEI. The main effect of the environment was primarily due to precipitation differences between June and July in the study year, while temperature had a smaller impact. These findings align with physiological processes in grain development, where water deficit can disrupt grain formation and reduce yield, as noted by Warzecha et al. (2010).
AMMI analyses can detect genotypes best suited to specific environmental conditions by estimating genotype interaction effects in each environment (Alizadeh et al. 2020; Zulfqar et al. 2021; Qasemi et al. 2022; Arinaitwe et al. 2023; Shojaei et al. 2023; Okla et al. 2023; Oghan et al. 2024). For grain yield, a significant GEI was demonstrated using AMMI analysis. Ranking of first four AMMI selections per environment selections per environment for grain yield is as follows: G69, G41, G49, G11 (in Kobierzyce); G69, G41, G04, G06 (in Mikulice); G04, G41, G06, G51 (in Płaczkowo); G08, G06, G32, G14 (in Radzików); and G36, G41, G44, G25 (in Smolice).
High genotype stability is related to the AMMI stability value, which evaluates the main effects of genotype, environment, and significant GEIs displayed on a GE biplot. The importance of these facts was pointed out in their studies, among others: Khan et al. (2021), Hailemariam Habtegebriel (2022), and Greveniotis et al. (2023). AMMI models effectively measure the importance of environments, genotypes, and their interactions, providing a value that indicates genotype stability across all environments, given grain yield (Sabaghnia et al. 2008; Mohammadi et al. 2018, 2023; Pour-Aboughadareh et al. 2022).
Conclusions
This research demonstrates the use of the AMMI model to analyze the impact of genotype by environment interactions on maize grain yield. Sixty-nine maize hybrids were evaluated across five different environments. The analysis of variance revealed a highly significant influence of genotypes, environments, and their interactions on grain yield. The stability analysis identified line SMH_1729 as having high stability. Additionally, lines SMH_1706 and SMH_16043, along with the variety ES Gallery, exhibited the highest average grain yields but with varying environmental adaptations. Specifically, line SMH_16043 was best adapted to the conditions in Płaczkowo, while the ES Gallery variety was best suited to Kobierzyce and Mikulice. In contrast, line SMH_1706 demonstrated the greatest stability and highest average grain yield across environments. This line had the optimal genotype selection index and is recommended for further inclusion in breeding programs and production. AMMI analysis most often describes the response of genotypes to varying environmental conditions using a limited number of multiplicative components, usually based on the first two or three principal components for GEI effects. In the present study, the percentage of interaction variation explained by the first two and three principal components was 63.19% and 84.30%, respectively. This meant a loss of information of 36.81% and 15.70%, respectively.
Data availability
The datasets generated during and analyzed during the current study are available from the corresponding author on reasonable request.
References
Abakemal D, Shimelis H, Derera J (2016) Genotype-by-environment interaction and yield stability of quality protein maize hybrids developed from tropical-highland adapted inbred lines. Euphytica 209:757–769
Abdala LJ, Otegui ME, Mauro GD (2024) On-farm soybean genetic progress and yield stability during the early 21st century: a case study of a commercial breeding program in Argentina and Brazil. Field Crops Res 308:109277. https://doi.org/10.1016/j.fcr.2024.109277
Adu GB, Akromah R, Abdulai MS, Obeng-Antwi K, Kena AW, Tengan KML, Alidu H (2013) Assessment of genotype by environment interactions and grain yield performance of extra-early maize (Zea mays L.) Hybrids. J Biol Agricul Healthcare 3(12):7–15
Ahakpaz Karkaji F, Majidi Hervan E, Roustaii M, Bihamta M, Mohammadi S (2023) Comprehensive stability analysis of wheat genotypes through multi-environmental trials. J Agric Sci 29(1):317–334. https://doi.org/10.15832/ankutbd.999060
Alam MA, Rahman M, Ahmed S, Jahan N, Khan MA-A, Islam MR, Alsuhaibani AM, Gaber A, Hossain A (2022) Genetic variation and genotype by environment interaction for agronomic traits in maize (Zea mays L.) hybrids. Plants 11(11):1522. https://doi.org/10.3390/plants11111522
Alizadeh B, Pasban Eslam B, Rezaizad A, Yazdandoost Hamadani M, Mostafavirad M (2020) Yield Stability assessment of winter oilseed rape lines in cold regions of Iran using AMMI model. J Plant Prod Res 27(3):85–96. https://doi.org/10.22069/jopp.2020.16557.2509
Amelework AB, Bairu MW, Marx R, Laing M, Venter SL (2023) Genotype × environment interaction and stability analysis of selected cassava Cultivars in South Africa. Plants 12(13):2490. https://doi.org/10.3390/plants12132490
Arinaitwe U, Clay SA, Nleya T (2023) Growth, yield, and yield stability of canola in the Northern Great Plains of the United States. Agron J 115:744–758. https://doi.org/10.1002/agj2.21269
Atlin GN, Cairns JE, Das B (2017) Rapid breeding and varietal replacement are critical to adaptation of cropping systems in the developing world to climate change. Global Food Secur 12:31–37. https://doi.org/10.1016/j.gfs.2017.01.008
Baldassarre L, Reitzel AM, Fraune S (2023) Genotype–environment interactions determine microbiota plasticity in the sea anemone Nematostella vectensis. PLoS Biol 21(1):e3001726. https://doi.org/10.1371/journal.pbio.3001726
Balestre M, Von Pinho RG, Souza JC, Oliveira RL (2009) Genotypic stability and adaptability in tropical maize based on AMMI and GGE biplot analysis. Genet Mol Res 8:1311–1322
Bocianowski J, Liersch A (2022) Multidimensional analysis of diversity in genotypes of winter oilseed rape (Brassica napus L.). Agronomy 12(3):633. https://doi.org/10.3390/agronomy12030633
Bocianowski J, Prażak R (2022) Genotype by year interaction for selected quantitative traits in hybrid lines of Triticum aestivum L. with Aegilops kotschyi Boiss. and Ae. variabilis Eig. using the additive main effects and multiplicative interaction model. Euphytica 218(2):11. https://doi.org/10.1007/s10681-022-02967-4
Bocianowski J, Księżak J, Nowosad K (2019a) Genotype by environment interaction for seeds yield in pea (Pisum sativum L.) using additive main effects and multiplicative interaction model. Euphytica 215:191. https://doi.org/10.1007/s10681-019-2515-1
Bocianowski J, Niemann J, Nowosad K (2019b) Genotype-by-environment interaction for seed quality traits in interspecific cross-derived Brassica lines using additive main effects and multiplicative interaction model. Euphytica 215:7. https://doi.org/10.1007/s10681-018-2328-7
Bocianowski J, Nowosad K, Tomkowiak A (2019c) Genotype – environment interaction for seed yield of maize hybrids and lines using the AMMI model. Maydica 64:M13
Bocianowski J, Radkowski A, Nowosad K, Radkowska I, Zieliński A (2021) The impact of genotype-by-environment interaction on the dry matter yield and chemical composition in timothy (Phleum pratense L.) examined by using the additive main effects and multiplicative interaction model. Grass Forage Sci 76(4):463–484. https://doi.org/10.1111/gfs.12551
Boote KJ, Kropff MJ, Bindraban PS (2001) Physiology and modelling of traits in crop plants: implications for genetic improvement. Agric Syst 70(2–3):395–420. https://doi.org/10.1016/S0308-521X(01)00053-1
Brancourt-Hulmel M, Lecomte C (2003) Effect of environmental varieties on genotype x environment interaction of winter wheat. A comparison of biadditive factorial regression to AMMI. Crop Sci 43:608–617. https://doi.org/10.2135/cropsci2003.6080
Ceccarelli S (1994) Specific adaptation and breeding for marginal conditions. Euphytica 77:205–219. https://doi.org/10.1007/BF02262633
Chauhan P, Shrivastava MK, Kumar V, Patel N, Biswal M (2023) Stability analysis in wheat (Triticum aestivum L.) genotypes under different environmental conditions. Int J Plant Soil Sci 35(21):1218–1223. https://doi.org/10.9734/ijpss/2023/v35i214099
Crossa J (1990) Statistical analyses of multilocation trials. Adv Agron 44:55–85. https://doi.org/10.1016/S0065-2113(08)60818-4
de Leon N, Jannink J-L, Edwards JW, Kaeppler SM (2016) Introduction to a special issue on genotype by environment interaction. Crop Sci 56:2081–2089. https://doi.org/10.2135/cropsci2016.07.0002in
Dehghani H, Sabaghnia N, Moghaddam M (2009) Interpretation of genotype-by-environment interaction for late maize hybrids’ grain yield using a biplot method. Turk J Agric for 33(2):5. https://doi.org/10.3906/tar-0712-25
Demelash H (2024) Genotype by environment interaction, AMMI, GGE biplot, and mega environment analysis of elite Sorghum bicolor (L.) Moench genotypes in humid lowland areas of Ethiopia. Heliyon 10(5):E26528. https://doi.org/10.1016/j.heliyon.2024.e26528
do Couto DP, Oliveira WBDS, de Oliveira JS, Guilhen JHS, Bernardes CdO, Posse SCP, Ferreira MFDS, Ferreira A (2023) Analysis of the effect of the interaction of genotype and environment on the yield stability of maize varieties; genetic resources for breeding. Agronomy 13(8):1970. https://doi.org/10.3390/agronomy13081970
Duvick DN (2005) The contribution of breeding to yield advances in maize (Zea mays L.). Adv Agron 86:83–145. https://doi.org/10.1016/S0065-2113(05)86002-X
Edwards JW (2016) Genotype×environment interaction for plant density response in maize (Zea mays L.). Crop Sci 56:1493–1505
Farshadfar E, Sutka J (2003) Locating QTLs controlling adaptation in wheat using AMMI model. Cereal Res Commun 31:249–256
Fekadu W, Mekbib F, Lakew B, Haussmann BIG (2023) Genotype × environment interaction and yield stability in barley (Hordeum vulgare L.) genotypes in the central highland of Ethiopia. J Crop Sci Biotechnol 26:119–133. https://doi.org/10.1007/s12892-022-00166-0
Ferraudo GM, Perecin D (2014) Mixed Model, AMMI and Eberhart-Russel comparison via simulation on genotype × environment interaction study in sugarcane. Appl Math 5(14):48155. https://doi.org/10.4236/am.2014.514205
Gan Y, Wang Y, Yu F, Xiao Q, Luo X, Han Z, Ke J, You W, Ke C (2023) Genotype by environment interactions for productive traits of purebred and crossbred abalone strains under different rearing modes. Aquaculture 563(1):738966. https://doi.org/10.1016/j.aquaculture.2022.738966
Gauch HG (1988) Model selection and validation for yield trials with interaction. Biometrics 44:705–715. https://doi.org/10.2307/2531585
Gauch HG, Zobel RW (1990) Imputing missing yield trial data. Theor Appl Genet 79:753–761
Gollob HF (1968) A statistical model which combines features of factor analytic and analysis of variance techniques. Psychometrika 33:73–115. https://doi.org/10.1007/BF02289676
Greveniotis V, Bouloumpasi E, Zotis S, Korkovelos A, Kantas D, Ipsilandis CG (2023) Genotype-by-environment interaction analysis for quantity and quality traits in faba beans using AMMI, GGE models, and stability indices. Plants 12(21):3769. https://doi.org/10.3390/plants12213769
Grishkevich V, Yanai I (2013) The genomic determinants of genotype × environment interactions in gene expression. Trends Genet 29(8):479–487. https://doi.org/10.1016/j.tig.2013.05.006
Hailemariam Habtegebriel M (2022) Adaptability and stability for soybean yield by AMMI and GGE models in Ethiopia. Front Plant Sci 13:950992. https://doi.org/10.3389/fpls.2022.950992
Hongyu K, García-Peña M, Borges de Araújo L, dos Santos DT (2014) Statistical analysis of yield trials by AMMI analysis of genotype × environment interaction. Biom Letters 51(2):89–102. https://doi.org/10.2478/bile-2014-0007
Khan MMH, Rafii MY, Ramlee SI, Jusoh M, Al Mamun M (2021) AMMI and GGE biplot analysis for yield performance and stability assessment of selected Bambara groundnut (Vigna subterranea L. Verdc.) genotypes under the multi-environmental trials (METs). Sci Rep 11:22791. https://doi.org/10.1038/s41598-021-01411-2
Khare V, Shukla RS, Pandey S, Singh SK, Singh C (2024) Exploring the genotype-environment interaction of bread wheat in ambient and high-temperature planting conditions: a rigorous investigation. Sci Rep 14:2402. https://doi.org/10.1038/s41598-024-53052-w
Kimutai C, Ndlovu N, Chaikam V, Ertiro BT, Das B, Beyene Y, Kiplagat O, Spillane C, Prasanna BM, Gowda M (2023) Discovery of genomic regions associated with grain yield and agronomic traits in Bi-parental populations of maize (Zea mays L.) Under optimum and low nitrogen conditions. Front Genet 14:1266402. https://doi.org/10.3389/fgene.2023.1266402
Kumar A, Sharma N, Kumar A, Kumar D, Jnanesha AC, Gupta AK, Chanotiya CS, Bisht D, Lal RK (2024) Differential responses of genotype × environment interaction on agronomic interventions affect the yield and quality of essential oil of interspecific basil hybrid of Ocimum basilicum L. × Ocimum kilimandscharicum. Ecol Genet Genomics 30:100217. https://doi.org/10.1016/j.egg.2024.100217
Kwambai TK, Griffin D, Struik PC, Stack L, Rono S, Brophy C, Nyongesa M, Gorman M (2024) Seed quality and variety preferences amongst potato farmers in North-Western Kenya: lessons for the adoption of new varieties. Potato Res 67:185–208. https://doi.org/10.1007/s11540-023-09626-8
Ljubičić N, Popović V, Kostić M, Pajić M, Buđen M, Gligorević K, Dražić M, Bižić M, Crnojević V (2023) Multivariate interaction analysis of Zea mays L. genotypes growth productivity in different environmental conditions. Plants 12(11):2165. https://doi.org/10.3390/plants12112165
Mandel J (1961) Non-additivity in two-way analysis of variance. J Am Stat Assoc 56:878–888. https://doi.org/10.1080/01621459.1961.10482132
Mandel J (1971) A new analysis of variance model for non-additive data. Technometrics 13(1):1–8. https://doi.org/10.1080/00401706.1971.10488751
Matongera N, Ndhlela T, van Biljon A, Labuschagne M (2023) Genotype x environment interaction and yield stability of normal and biofortified maize inbred lines in stress and non-stress environments. Cogent Food Agric 9(1):2163868. https://doi.org/10.1080/23311932.2022.2163868
Mohammadi R, Armion M, Zadhasan E, Ahmadi M, Amri A (2018) The use of AMMI model for interpreting genotype × environment interaction in durum wheat. Exp Agric 54(5):670–683. https://doi.org/10.1017/S0014479717000308
Mohammadi R, Jafarzadeh J, Poursiahbidi MM, Hatamzadeh H, Amri A (2023) Genotype-by-environment interaction and stability analysis for grain yield in durum wheat using GGE biplot and genotypic and environmental covariates. Agric Res 12:364–374. https://doi.org/10.1007/s40003-023-00661-y
Mousavi SMN, Illés A, Szabó A, Shojaei SH, Demeter C, Bakos Z, Vad A, Széles A, Nagy J, Bojtor C (2024) Stability yield indices on different sweet corn hybrids based on AMMI analysis. Braz J Biol 84:e270680. https://doi.org/10.1590/1519-6984.270680
Nowosad K, Tratwal A, Bocianowski J (2018) Genotype by environment interaction for grain yield in spring barley using additive main effects and multiplicative interaction model. Cereal Res Commun 46(4):729–738
Nowosad K, Bocianowski J, Kianersi F, Pour-Aboughadareh A (2023) Analysis of linkage on interaction of main aspects (genotype by environment interaction, stability and genetic parameters) of 1000 kernels in maize (Zea mays L.). Agriculture 13(10):2005. https://doi.org/10.3390/agriculture13102005
Nzuve F, Githiri S, Mukunya DM, Gethi J (2013) Analysis of genotype x environment interaction for grain yield in maize hybrids. J Agric Sci 5(11):75–85. https://doi.org/10.5539/jas.v5n11p75
Oghan HA, Bakhshi B, Rameeh V, Tabrizi HZ, Faraji A, Ghodrati G, Fanaei HR, Askari A, Kiani D, Payghamzadeh K, Sadeghi H, Danaei AK, Kazerani NK, Afrouzi MAAN, Dalili A (2024) Comparative study of univariate and multivariate selection strategies based on an integrated approach applied to oilseed rape breeding. Crop Sci 64:55–73. https://doi.org/10.1002/csc2.21104
Okla MK, Saleem MH, Saleh IA, Zomot N, Perveen S, Parveen A, Abasi F, Ali H, Ali B, Alwasel YA, Abdel-Maksoud MA, Oral MA, Javed S, Ercisli S, Sarfraz MH, Hamed MH (2023) Foliar application of iron-lysine to boost growth attributes, photosynthetic pigments and biochemical defense system in canola (Brassica napus L.) under cadmium stress. BMC Plant Biol 23:648. https://doi.org/10.1186/s12870-023-04672-3
Oladosu Y, Rafii MY, Abdullah N, Magaji U, Miah G, Hussin G, Ramli A (2017) Genotype × Environment interaction and stability analyses of yield and yield components of established and mutant rice genotypes tested in multiple locations in Malaysia. Acta Agric Scand Sect B 67(7):590–606. https://doi.org/10.1080/09064710.2017.1321138
Oroian C, Ugruțan F, Mureșan IC, Oroian I, Odagiu A, Petrescu-Mag IV, Burduhos P (2023) AMMI analysis of genotype × environment interaction on sugar beet (Beta vulgaris L.) Yield, sugar content and production in Romania. Agronomy 13(10):2549. https://doi.org/10.3390/agronomy13102549
Pacheco RM, Duarte JB, Vencovsky R, Pinheiro JB, Oliveira AB (2005) Use of supplementary genotypes in AMMI analysis. Theor Appl Genet 110:812–818. https://doi.org/10.1007/s00122-004-1822-6
Patel R, Parmar DJ, Kumar S, Patel DA, Memon J, Patel MB, Patel JK (2023) Dissection of genotype × environment interaction for green cob yield using AMMI and GGE biplot with MTSI for selection of elite genotype of sweet corn (Zea mays conva. Saccharata var. rugosa). Indian J Genet Plant Breed 83(1):59–68
Patel MK, Tiwari D, Sharma V, Singh D (2024) Assessment of genotype x environment interaction and seed yield stability in sesame (Sesamum indicum L.) using AMMI analysis. J Sci Res Rep 30(1):63–70. https://doi.org/10.9734/jsrr/2024/v30i11825
Pires LP, Garcia Von Pinho R, Balestre M (2018) AMMI Bayesian models to study stability and adaptability in maize. Agron J 110:1765–1776. https://doi.org/10.2134/agronj2017.11.0668
Pour-Aboughadareh A, Khalili M, Poczai P, Olivoto T (2022) Stability indices to deciphering the genotype-by-environment interaction (GEI) effect: an applicable review for use in plant breeding programs. Plants 11(3):414. https://doi.org/10.3390/plants11030414
Pour-Aboughadareh A, Barati A, Gholipoor A, Zali H, Marzooghian A, Koohkan SA, Shahbazi-Homonloo K, Houseinpour A (2023) Deciphering genotype-by-environment interaction in barley genotypes using different adaptability and stability methods. J Crop Sci Biotechnol 26:547–562. https://doi.org/10.1007/s12892-023-00199-z
Pramanik K, Sahu GS, Acharya GC, Tripathy P, Dash M, Koundinya AVV, Jena C, Kumar DS, Mohapatra PP, Pradhan J, Karubakee S, Moharana DP (2024) Estimating phenotypic stability for relevant yield and quality traits in French bean (Phaseolus vulgaris L.) using AMMI analysis. Heliyon 10(5):e26918. https://doi.org/10.1016/j.heliyon.2024.e26918
Purchase JL, Hatting H, van Deventer CS (2000) Genotype × environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. S Afr J Plant Soil 17:101–107
Qasemi SH, Mostafavi K, Khosroshahli M, Bihamta MR, Ramshini H (2022) Genotype and environment interaction and stability of grain yield and oil content of rapeseed cultivars. Food Sci Nutr 10:4308–4318. https://doi.org/10.1002/fsn3.3023
Ruswandi D, Maulana H, Karuniawan A, Mansyur IA, Maxiselly Y, Fauzan MR, Abdullah MA, Yuwariah Y (2023) Multi-traits selection of maize hybrids under sole-crop and multiple-crops with soybean. Agronomy 13(10):2448. https://doi.org/10.3390/agronomy13102448
Sabaghnia N, Sabaghpour SH, Dehghani H (2008) The use of an AMMI model and its parameters to analyse yield stability in multi-environment trials. J Agric Sci 146(5):571–581. https://doi.org/10.1017/S0021859608007831
Sabitha N, Reddy DM, Reddy DL, Sudhakar P, Reddy BR (2024) Association analysis over seasons among morphological, physiological and yield components with kernel yield in maize (Zea mays L.). J Adv Biol Biotechnol 27(5):151–156. https://doi.org/10.9734/jabb/2024/v27i5774
Saeidnia F, Taherian M, Nazeri SM (2023) Graphical analysis of multi-environmental trials for wheat grain yield based on GGE-biplot analysis under diverse sowing dates. BMC Plant Biol 23:198. https://doi.org/10.1186/s12870-023-04197-9
Shojaei SH, Mostafavi K, Ghasemi SH, Bihamta MR, Illés Á, Bojtor C, Nagy J, Harsányi E, Vad A, Széles A, Mousavi SMN (2023) Sustainability on different canola (Brassica napus L.) cultivars by GGE biplot graphical technique in multi-environment. Sustainability 15(11):8945. https://doi.org/10.3390/su15118945
Shrestha S, Asch F, Dusserre J, Ramanantsoanirina A, Brueck H (2012) Climate effects on yield components as affected by genotypic responses to variable environmental conditions in upland rice systems at different altitudes. Field Crops Res 134:216–228. https://doi.org/10.1016/j.fcr.2012.06.011
Signor CE-L, Dousse S, Lorgeou J, Denis J-B, Bonhomme R, Carolo P, Charcosset A (2001) Interpretation of genotype × environment interactions for early maize hybrids over 12 years. Crop Sci 41:663–669. https://doi.org/10.2135/cropsci2001.413663x
Singamsetti A, Shahi JP, Zaidi PH, Seetharam K, Vinayan MT, Kumar M, Singla S, Shikha K, Madankar K (2021) Genotype × environment interaction and selection of maize (Zea mays L.) hybrids across moisture regimes. Field Crops Res 270:108224. https://doi.org/10.1016/j.fcr.2021.108224
Singh SB, Kumar S, Kumar R, Kumar P, Yathish KR, Jat BS, Chikkappa GK, Kumar B, Jat SL, Dagla MC, Kumar B, Kumar A, Kasana RK, Kumar S (2024) Stability analysis of promising winter maize (Zea mays L.) hybrids tested across Bihar using GGE biplot and AMMI model approach. Indian J Genet Plant Breed 84(1):73–80
Swarup S, Cargill EJ, Crosby K, Flagel L, Kniskern J, Glenn KC (2021) Genetic diversity is indispensable for plant breeding to improve crops. Crop Sci 61:839–852. https://doi.org/10.1002/csc2.20377
Taleghani D, Rajabi A, Saremirad A, Fasahat P (2023) Stability analysis and selection of sugar beet (Beta vulgaris L.) genotypes using AMMI, BLUP. GGE biplot and MTSI. Sci Rep 13:10019. https://doi.org/10.1038/s41598-023-37217-7
Vaezi B, Pour-Aboughadareh A, Mohammadi R, Mehraban A, Hossein-Pour T, Koohkan E, Ghasemi S, Moradkhani H, Siddique KHM (2019) Integrating different stability models to investigate genotype × environment interactions and identify stable and high-yielding barley genotypes. Euphytica 215:63. https://doi.org/10.1007/s10681-019-2386-5
von Gehren P, Bomers S, Tripolt T, Söllinger J, Prat N, Redondo B, Vorss R, Teige M, Kamptner A, Ribarits A (2023) Farmers feel the climate change: variety choice as an adaptation strategy of european potato farmers. Climate 11(9):189. https://doi.org/10.3390/cli11090189
VSN International (2023) Genstat for Windows, 23rd edn. VSN International, Hemel Hempstead
Warzecha T, Adamski T, Kaczmarek Z, Surma M, Goliński P, Perkowski JM, Chełkowski J, Wiśniewska H, Krystkowiak K, Kuczyńska A (2010) Susceptibility of hulled and hulless barley doubled haploids to Fusarium culmorum head blight. Cereal Res Commun 38:220–232
Williams RM, O’Brien L, Eagles HA, Solah VA, Jayasena V (2008) The influences of genotype, environment, and genotype × environment interaction on wheat quality. Aust J Agric Res 59(2):95–111. https://doi.org/10.1071/AR07185
Yang X, Soliman AA, Hu C, Yang F, Lv M, Yu H, Wang Y, Zheng A, Dai Z, Li Q, Tang Y, Yang K, Zhang Y, Niu W, Wang L, He Y (2023) Yield adaptability and stability in field pea genotypes using AMMI, GGE, and GYT biplot analyses. Agriculture 13(10):1962. https://doi.org/10.3390/agriculture13101962
Yue H, Gauch HG, Wei J, Xie J, Chen S, Peng H, Bu J, Jiang X (2022) Genotype by environment interaction analysis for grain yield and yield components of summer maize hybrids across the Huanghuaihai Region in China. Agriculture 12(5):602. https://doi.org/10.3390/agriculture12050602
Zobel RW, Wright MJ, Gauch HG (1988) Statistical analysis of yield trial. Agron J 80:388–393
Zulfqar M, Mustafa HSB, Ejaz-Ul-Hasan SS, Qamar R, Gill AN, Mahmood T, Ud D, Ahsan M, Kalyar MTA, Ali S, Hameed A, Salim J, Wakeel A (2021) Quantitative evaluation of commercial canola cultivars through G × E analysis under different agro-climatic conditions. Plant Cell Biotechnol Mol Biol 22(71–72):469–480
Author information
Authors and Affiliations
Contributions
Conceptualization: J.B. and K.N. Methodology: J.B., K.N., and D.R. Software: J.B. Validation: J.B., K.N., and D.R. Formal analysis: J.B. Investigation: J.B., K.N., and D.R. Resources: J.B., K.N., and D.R. Data curation: D.R. Writing—original draft preparation: J.B. and K.N. Writing—review and editing: J.B., K.N., and D.R. Visualization: J.B. Supervision: K.N. Project administration: J.B. Funding acquisition: K.N. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Communicated by: Izabela Pawłowicz
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bocianowski, J., Nowosad, K. & Rejek, D. Genotype-environment interaction for grain yield in maize (Zea mays L.) using the additive main effects and multiplicative interaction (AMMI) model. J Appl Genetics (2024). https://doi.org/10.1007/s13353-024-00899-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13353-024-00899-4