Abstract
A Bayesian approach to analyze complex traits is presented that can help plant eneticists and breeders in exploiting the marker and phenotypic data on pedigreed populations as available from ongoing breeding programs. The statistical model for the quantitative trait may include non-genetic and genetic components. The latter component can be divided into QTL on known marker linkage groups, major genes and a polygenic component. The full probability model, prior assumptions on model variables are presented and criterion for model selection and posterior inferences are given. Simulated data on a known pedigreed population structure of the EU project HiDRAS was used to illustrate the use of the Bayesian approach to analyze complex traits. It was shown that estimates for QTL parameters were more accurate when non-genetic factors were included in the model and when a polygenic component was included when not all linkage groups were analyzed simultaneously. The Bayesian approach has been implemented into the software package FlexQTL and allows plant breeders explore their pedigreed populations for segregating QTL alleles that are relevant in their breeding program.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breeders and geneticists have developed and applied statistical methods to identify quantitative trait loci (QTL) based on genetic marker and quantitative trait data. These methods were designed to answer basic questions concerning QTL (e.g. number, mode, and magnitude) and to map QTL on the genome to facilitate their application in breeding programs. In plant QTL mapping experiments, populations derived from specific crosses of inbred lines have predominantly been used, see e.g., Jansen in Balding et al. (2003). However, major reasons exist to study complex populations derived from multiple founders or taken from ongoing breeding programs. Here we provide three of them. Firstly, improved exploration of QTL variation: It is very likely that if a population arises from many founders multiple alleles are present, thereby increasing the probability to detect the most valuable QTL allele. Secondly, practical relevance of identified QTL alleles: Experimental line crosses often do not represent the (commercial) breeding populations. And thirdly, improved cost effectiveness of QTL mapping: Costs for marker genotyping decline rapidly; hence trait phenotyping becomes relatively more expensive. Breeding programs routinely evaluate the trait phenotypes of many progeny with replication at several locations.
The above incentives should convince plant geneticists and breeders to exploit the data on pedigreed populations as available from ongoing breeding programs. However, the analysis of this type of data has been hampered by the absence of flexible and robust statistical methods and software tools. Important criteria for QTL mapping in complex data may be summarized as:
-
1.
Robustness and flexibility with regard to tackling structures in the data, especially pedigree structures, i.e. individuals may cover several generations, the population may consist of several families, who may differ in size but may also be related.
-
2.
Incompleteness of marker information; this holds on multiple levels, i.e. an individual’s marker data may be partially or fully, dominant marker scoring, or non-informative meioses.
-
3.
Non-genetic factors may contribute to the phenotypic trait variation which, if ignored, will reduce power and accuracy of the estimates of genetic parameters. However, pre-correction for these factors may lead to biased estimates.
-
4.
The number of segregating QTL is unknown. Also, the mode of action of QTL is unknown and may interact with the genetic or environmental background in which it is expressed.
In this paper we accommodate these criteria by applying a Bayesian approach, see e.g., Gelman et al. (2004). The Bayesian approach provides practical methods for making inferences from data using probability models for quantities we observe (e.g., traits and markers) and for quantities we wish to learn about (e.g., genes). An essential characteristic is the explicit use of probability distributions to quantify uncertainty. In a Bayesian analysis the prior knowledge is integrated with the likelihood of the data and the resulting posterior distribution represents the accumulated knowledge on the parameters of interest. A Bayesian framework with Markov chain Monte Carlo (MCMC) see e.g., Gilks et al. (1996) algorithms provides a powerful tool for estimating the chromosomal location, the contribution of genes affecting complex traits and, potentially, gene-by-gene and gene-by-environment interactions. Note that we will not consider interactions here as they are beyond the scope of this paper.
In this study we assume that the quantitative trait may also be affected by non-genetic factors, e.g., year and location in which phenotypes were scored, as well as a polygenic component representing many small genes, undetectable via linked markers. After describing the probability model and its variables with their prior distributions, we will present results from the analyses of simulated data to dissect complex traits into their underlying genetic components. For the simulated data set we use the known pedigree structure of a dataset on 13 related full-sib populations that is currently produced within the EU project HiDRAS (http://www.hidras.unimi.it/).
Methods
Linear regression model
Let θ denote the vector with model parameters affecting the trait of interest. Using standard regression notation, we can express the relationship between observed phenotypes to the unknown parameters in the following linear model
where y is the vector with observed phenotypes for the quantitative trait; H is the design matrix linking model parameters to the phenotypes, and ε is the environmental error. The environmental errors are assumed to be independent and identically and normally distributed, i.e., \( \varepsilon \sim N(0,\sigma ^{2}_{e} ) \).
Principle of Bayesian analysis
Gelman et al. (2004) divide the process of a Bayesian data analysis into the following steps: 1. Setting up a full probability model; 2. Calculating and interpreting the appropriate posterior distribution; and 3. Evaluating the fit of the model and the implications of the resulting posterior distribution.
Let p(θ,y) represent the joint probability of the model parameters (θ) and the data (y). The terms p(y) and p(θ) represent the marginal distributions of the data and the set of parameters, respectively. Also, let \( p(\theta \left| {\mathbf{y}} \right.) \) and \( p({\mathbf{y}}\left| \theta \right.) \) represent the conditional distributions of the parameters given the data and the reverse, respectively. Then, the joint probability distribution of θ and y is
The combination of (2) and (3) leads to \( p{\left( {\theta \left| {\mathbf{y}} \right.} \right)} = {p{\left( \theta \right)}p{\left( {{\mathbf{y}}\left| \theta \right.} \right)}} \mathord{\left/ {\vphantom {{p{\left( \theta \right)}p{\left( {{\mathbf{y}}\left| \theta \right.} \right)}} {p{\left( {\mathbf{y}} \right)}}}} \right. \kern-\nulldelimiterspace} {p{\left( {\mathbf{y}} \right)}}.\) The marginal distribution of the data, p(y) after having observed the data is a fixed constant and the conditional distribution becomes proportional to,
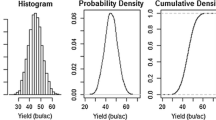
This Eq. 4 points to the well-known Bayes’ rule (Bayes 1763) that the posterior probability is proportional to the product of the prior probability and the likelihood of the data. The concept of a Bayesian analysis is shown in Fig. 1. The prior distribution is relatively flat representing vague knowledge on our parameter of interest. The posterior distribution is relatively peaked, indicating an increase in knowledge (certainty) on the parameter, and its position is intermediate between the prior distribution and the likelihood of the data being a weighted average of the two information sources.
Probability distributions representing the principle of Bayesian analysis: The posterior probability distribution (black, solid) is the product of the prior probability distribution (brown, dotted) and the likelihood distribution of the data (blue, dashed). The area of each probability distribution sums to 1.0. The 0.90 Highest Density Region (HDR) of each distribution are indicated by horizontal lines
Population characteristics
We consider diploid populations with known pedigree structure among all its individuals. These populations may either originate from fully inbred, homozygous parents or from outbred, heterozygous parents. The pedigree information specifies the two parents of every individual. The two parents of founder individuals are unknown. Next to the information on the pedigree relationships, the data consist of phenotypic trait values, y, and marker genotypes, m, for individuals in a mapping population. We assume that marker loci are organized into a linkage map with known distances and recombination rates, applying the Haldane mapping function, (Haldane 1919). The genotypes for markers are assumed to be co-dominantly scored, i.e., both alleles known.
Within the linkage map putative QTL may occur at any position, i.e., the unknown position has a continuous distribution. In general the genotypes, g, for these putative QTL are unobservable, except at completely informative markers, but their probability distribution can be inferred from the observed marker data by using a multipoint method, e.g., Jiang and Zeng (1997). This probability distribution is used as the prior distribution of QTL genotypes in the proposed Bayesian framework. The primary interest in QTL mapping is inferring the number, locations and effects of QTL on one or multiple chromosomes.
In absence of genetic marker data, the genetic component of quantitative traits may be modeled via polygenic variance component or via individual major genes (Kennedy et al., 1992; Janss et al. 1995; Pong-Wong et al. 1999; Bink 2002). We will refer to the first as the infinitesimal model (TIM) and to the latter as finite polygenic model (FPM). The assumptions for the FPM model and the QTL model are identical, except that the genes in the FPM model are anonymous, i.e., they are not linked to markers and the individuals’ genotypes are inferred from pedigree and phenotypic data. So, the transmission of alleles at a QTL can be inferred from the known transmission patterns of linked markers whereas the alleles of a FPM gene obey only the Mendelian transmission rules.
Fixed number of QTL
The QTL is assumed to be bi-allelic, allowing three genotypes to be distinguished, i.e., AA, Aa, and aa, having genotypic values equal to +α, δ and −α, respectively. The variables α and δ represent the additive and dominance effects of a single gene. For convenience, we will assume absence of dominance, i.e., δ = 0, and omit dominance in the remainder of this section. The allele frequency of the positive allele A is denoted by f α , and may take any value between 0 and 1 with equal prior probability. To define the full probability model we expand the concise linear model (1) into several factors that may affect our trait of interest,
where β is a vector containing an overall mean (μ) and all non-genetic variables affecting the trait of interest, which may include year and location effects. The vectors α qtl and α fpm represent the additive genetic contributions of a gene in the QTL or FPM models, respectively. The vector u contains the polygenic effect of individuals, accounting for joint contribution of small genes not captured by the QTL or FPM models. The incidence matrices X, W, V, and Z connect the phenotypes to non-genetic variables, QTL, FPM, and TIM, respectively. The elements of matrices W and V depend on the genotype assigned to each individual. For an additive model, the elements in W and V are equal to +1, 0, or −1, when an individual has the positive homozygous genotype AA (increasing the phenotypic value), the heterozygous genotype Aa (aA), or the negative homozygous genotype aa (decreasing the phenotypic value), respectively. The dimensions of matrices W and V depend on the number of QTL and FPM genes, respectively, in the model.
Random number of QTL
The number of QTL and FPM genes are treated as random variables in our Bayesian analysis, similar to previous studies by e.g., (Fisch et al. 1996; Heath 1997; Sillanpaa and Arjas 1998; Uimari and Sillanpaa 2001; Bink et al. 2002). Treating the number of genes as a random variable in a Bayesian framework can be facilitated by the use of the Reversible Jump sampler (Green 1995; Waagepetersen and Sorensen 2001).
Prior assumptions
The non-genetic variables are assumed to follow a Normal distribution a priori. The variance of this Normal distribution is unknown and this random variable is assumed to follow a scaled inverse chi-square distribution, e.g., Sorensen and Gianola (2002, p. 85). In case of the overall mean, always present in the model, the mean of the Normal prior was data-dependent, \( \hat{\mu } = {\mathbf{\ifmmode\expandafter\bar\else\expandafter\=\fi{y}}} = \tfrac{1} {n}{\sum\nolimits_i^n {y_{i} } } \). The prior distribution for the residual variance \( {\left( {\sigma ^{2}_{e} } \right)} \) is taken to be a scaled inverse chi-square distribution. Let matrix A denote the matrix of additive genetic relationships, e.g., Lynch and Walsh (1998) given the known pedigree of all individuals. Then, the prior for the polygenic effects can be taken as
where \( \sigma ^{2}_{u} \) is the additive genetic variance, which is assumed to follow a scaled inverse chi-square distribution as well. The additive effects of the QTL (and FPM genes) are assumed to follow a univariate Normal distribution where the variance assumed to be both data-dependent and dependent on the number of QTL (or FPM genes) in the model, as previously proposed by (Yi 2004; Yi et al. 2005). Let \( \hat{\sigma }^{2}_{y} = \tfrac{1} {n}{\sum\nolimits_i^n {{\left( {y_{i} - \ifmmode\expandafter\bar\else\expandafter\=\fi{y}} \right)}} }^{2} \) denote the estimate of the phenotypic variance of the trait, then,
where \( \sigma ^{2}_{{\alpha _{{(N_{{QTL}} )}} }} = {\sigma ^{2}_{\alpha } } \mathord{\left/ {\vphantom {{\sigma ^{2}_{\alpha } } {N_{{QTL}} }}} \right. \kern-\nulldelimiterspace} {N_{{QTL}} } \) and \( {\sigma ^{2}_{\alpha } } \mathord{\left/ {\vphantom {{\sigma ^{2}_{\alpha } } {{\left( {2.0 \times \hat{\sigma }^{2}_{y} } \right)}}}} \right. \kern-\nulldelimiterspace} {{\left( {2.0 \times \hat{\sigma }^{2}_{y} } \right)}}\sim Beta{\left( {2,10} \right)} \). This implies that the variance of the normal distribution shrinks when the number of genes in the model increases and vice versa. The number of QTL is here assumed to have a Poisson distribution with mean κ, e.g., Heath (1997), Sillanpaa and Arjas (1998). The prior mean for the number of QTL affecting the trait was equal to 1.0 in this study. The influence of the value for κ on the posterior for the number of QTL can be examined by applying different values. The position for the j th QTL is specified in centiMorgan (Haldane 1919), and denoted by λ, and we assume that the position of a QTL takes a uniform prior distribution along the entire genome. The variance explained by all QTL jointly is calculated as
where Hardy Weinberg equilibrium is assumed in the initial founder population (Falconer 1989) and linkage equilibrium among QTL. The variance explained by all FPM genes jointly can be calculated similarly.
Joint posterior distribution
Let P and M denote the pedigree and marker data, respectively, and let \( \theta = {\left( {\beta ,{\mathbf{u}},\alpha _{{QTL}} ,\alpha _{{FPM}} ,\sigma ^{2}_{e} } \right)} \), then the joint posterior distribution of all unknowns can be written as (omitting matrices X and Z),
where the first term is the conditional distribution of the phenotypic data given all unknowns from Eq. 5. The second term is the probability distribution of QTL genotypic states (genotypes) conditional on the number and locations of QTL, the QTL allele frequencies, and the pedigree and marker data. The third term is the probability distribution of FPM genotypic states conditional on the number of FPM genes, the allele frequencies and the pedigree data. The final term in Eq. 9 is the joint prior distribution of the model variables.
Posterior computations
The calculation of the above joint posterior distribution is analytically intractable, and we apply a Markov chain Monte Carlo (MCMC) approach (Gilks et al. 1996) to obtain draws from the joint posterior distribution. Different MCMC sampling algorithms are used , i.e., the Gibbs sampler (Gelman et al. 1995; Sorensen and Gianola 2002) when the full conditional sampling distributions have a recognizable kernel, and the Metropolis Hastings algorithm (Gelman et al. 1995) when the sampling distribution has an unknown kernel. To allow changes in model dimension, i.e., increase or decrease the number of QTL or FPM genes in the model, we use the reversible jump MCMC method (Green 1995). The probabilities of proposals for an increase or a decrease were both equal to 0.40 at a given cycle of the Markov chain, if neither an increase nor a decrease was proposed all variables of the current model were updated.
Posterior inference and model selection
The draws obtained from the joint posterior distribution are used to calculate the marginal posterior distributions for the variables of interest. These draws are used to calculate point estimates such as the mean, mode, and standard deviation, but also to summarize the distribution by a region of the sample space covering a specific probability. The highest density regions (HDR) are those regions that occupy the smallest possible volume in the sample space or in other words, every point in the region has probability density at least as large as every point outside that region (Hyndman 1996). In case of a unimodal symmetric distribution, e.g., a Normal, an HDR coincides with the usual probability region symmetric around the mean. However, in case of a multimodal distribution, an HDR often consists of several disjoint subregions. In Bayesian analysis, HDR’s are applied to the posterior distributions and are called ‘credible sets’, ‘plausible sets’, ‘Bayesian confidence sets’ or ‘highest posterior density regions’. We will use the Highest Posterior Density regions covering 0.90 probability (HPD90) in the posterior inferences. In Fig. 1 it can be seen that the HPD90 (of the posterior) is relatively small compared to the 90% credible regions of the prior distribution, indicating that the knowledge increased substantially after including the information from the data. The number of QTL and their positions in a certain chromosome is of main interest. The chromosome is divided into small intervals (bins) and the number of QTL per bin per cycle is used to calculate the posterior QTL intensity (Sillanpaa and Arjas 1998). For the posterior inference on the chromosomal positions of the QTL we use 0.90 Highest Posterior Intensity (HPI90).
So, an essential characteristic of a Bayesian analysis is the explicit use of probability to quantify uncertainties in posterior inferences based on statistical data analysis. The model described above includes different genetic components, i.e., QTL and major gene bi-allelic effects and polygenic effects. Also, one or more non-genetic effects can be included when these explain substantial parts of the phenotypic variation of the quantitative trait of interest. The first model selection criterion will be proportion of phenotypic variance explained. When the HPD90 of the marginal posterior distribution includes the value zero for a particular variable, one may exclude this variable from the model.
The Bayesian analysis offers the ability to utilize data containing unbalanced structures and to study and select among complex models. Whether model selection will be successful depends on the quality and quantity of the data, in absence of these, the posterior inference will reflect the prior knowledge. We use Bayes factors (Kass 1993; Kass and Raftery 1995) as a measure of evidence coming from the data for different QTL models (Table 1).
Data
Simulated data is used to demonstrate the applicability of our Bayesian approach to analyze genetic components underlying quantitative traits. The simulation is based on a subset of the pedigreed apple population of the EU-project HiDRAS (http://www.hidras.unimi.it/). The core of the population is an incomplete di-allel design (Fig. 2), where the 604 individuals of the 13 full sib mapping populations are connected to each other through their 15 parents. However, the pedigrees of the parents are known from breeding records (see Fig. 3) and trace up to four generations back in time. Note that in the EU-project DNA was still available for most of the ancestors (here we assume all individuals will have marker data available).
Pedigree structure of the simulated dataset. The 13 Full Sib populations (orange boxes) are at the lower part of pedigree; their 15 parents (red boxes) are related through their intermediate (light blue) and founder (dark blue) ancestors. The Pedimap software (R. Voorrips, pers. comm.) was used to produce this figure
We simulated 2 chromosomes of 100 cM with each 11 markers equi-distantly spaced. We allowed 2 alleles per locus, alleles were equally probable to be assigned to the 26 founders assuming linkage equilibrium among loci. Note that only 2 chromosomes were simulated for reasons of conciseness and clearness.
The quantitative trait was affected by 4 QTL all similar in size, positioned at 25 and 75 cM on chromosomes 1 and 2. The heritability of the 4 QTL jointly was approximately 0.32. A small polygenic variance was simulated, i.e., heritability equal to 0.03. No FPM gene effects were simulated. Furthermore, we simulated a substantial contribution of a non-genetic factor (NGF), i.e., explaining 0.32 of the total phenotypic variance. This non-genetic factor was simulated by randomly assigning 21 classes (e.g., 7 locations × 3 years) to all individuals. The effects pertaining to these 21 classes were randomly drawn from a Normal distribution. Note that this non-genetic factor does not involve any interaction structure with the genetic factors and only the variance due to the (single) non-genetic factor is of importance.
Before analyzing the data using different models, the phenotypic trait records were scaled such that the phenotypic variance was equal to 1.0.
Results
Models without marker data
The analysis of pedigree and phenotypic data on the quantitative trait, ignoring marker data, revealed clearly a genetic component (Fig. 4). Fitting only an overall mean, resulted in an posterior mode (and mean) estimate for the error variance of 1.00 with a 0.90 probability interval equal to 0.90 to 1.10. Fitting the non-genetic factor with a Normal prior resulted in a clear decrease of the estimated of the error variance while the posterior credible region for the NGF variance was considerably large with a long right tail. The posterior distribution for the NGF variance was very stable across all different models considered in this study (see also Fig. 5). When allowing both components TIM and FPM, the TIM was favored to explain the variance of the phenotypic trait, while the FPM variance had most probability mass at or close to zero (Fig. 4). This seems counter-intuitive as the QTL and the FPM models have similar assumptions. The TIM model was able to fit all genetic variance, including the 4 QTL, as previously shown by (Bink 2002) for simulation scenarios without selection. These results may change dramatically when the simulated dataset included selection, in that case the trait variance may be better explained by the FPM (Bink 2002).
Marginal posterior densities of variance components of non-QTL models. Components are error variance (ERR), non-genetic factor variance (NGF), the infinitesimal model polygenic variance (TIM), and finite polygenic model variance (FPM). The models were (a) overall mean; (b) overall mean + NGF; (c) overall + NGF + TIM; (d) overall mean + NGF + TIM + FPM. The estimates for the posterior mean and 0.90 highest posterior density region (between brackets) are given for the variance components
Marginal posterior densities of variance components for QTL models. Components are error variance (ERR), non-genetic factor variance (NGF), the infinitesimal model polygenic variance (TIM), and Quantitative Trait Loci variance (QTL). The models were (a) overall mean + QTL; (b) overall mean + NGF + QTL; (c) overall + NGF + QTL (chromosome 2 excluded); (d) overall mean + NGF + TIM + QTL (chromosome 2 excluded)
Models with marker data, all linkage groups
Fitting the model with a QTL for the analysis of pedigree, marker and phenotypic data resulted in clear evidence for one or more QTL explaining the phenotypic variation of the quantitative trait (Fig. 5a). Accounting for the non-genetic factor clearly improved the estimated posterior density for the QTL and error variance components. The HPD90 regions were much smaller, and posterior mode estimates were close to the values used in the simulation of the data set (Fig. 5a, b). The posterior mode of the error variance shifted from 0.72 to 0.34 and HPD90 region decreased in length from 0.16 down to 0.12. The inclusion of a polygenic component (TIM) did not further improve the estimates for the QTL parameters, which may be due to the small value for this component in the simulated data set (results not shown). The estimates for the Bayes factors provided strong evidence for the 2 QTL model for chromosome 1 (Table 2), which was the model used in the simulation. Including the non-genetic variable into the model significantly improves the model selection, i.e., the estimates of the Bayes factor become much more decisive (Table 1).
Models with marker data, excluding linkage group 2
When analyzing only 1 chromosome, the variance explained by QTL clearly decreased and the error variance increased (Fig. 5b, c). The estimation of the posterior distribution of the non-genetic variable was very similar in all analyses. For the analysis of the first chromosome solely, the inclusion of the TIM component resulted in a sharper posterior density for the QTL variance, i.e., the length of the HPD regions decreased from 0.20 down to 0.16. Also the estimated posterior mode decreased from 0.18 down to 0.14, which may indicate that there was some overestimation of the QTL variance for a single linkage group when not accounting for QTL on other linkage groups. The posterior density for the error variance shifted to smaller values but its credible region became larger, indicating that there seemed to be some difficulty to dissect the polygenic and error variance components (Fig. 6).
Marginal posterior intensity, prior intensity and highest posterior intensity regions (HPI90) for the position of the QTL on chromosome 1. The models were (a) overall mean + QTL; (b) overall mean + NGF + QTL; (c) overall + NGF + QTL (chromosome 2 excluded); (d) overall mean + NGF + TIM + QTL (chromosome 2 excluded), (cf. Fig. 5)
The prior mean for the number of QTL at this single linkage group was equal to 1.0. The estimated posterior mean for the number of QTL were 3.5 and 2.4 for the models excluding and including the TIM component into the model. This suggests that there was an over-fitting of QTL on chromosome 1 when the model cannot account for the QTL on chromosome 2. These latter QTL may be accommodated via the inclusion of a TIM as the number of QTL fitted onto chromosome 1 become more consistent with the simulated number (2) of QTL. This was also reflected in the Bayes factor estimates, models with more QTL on chromosome 1 were favored when ignoring the QTL on chromosome 2 (Table 2).
Discussion
In this study we presented a Bayesian approach for dissecting the phenotypic variation of a quantitative trait into genetic and non-genetic components. Given the pedigree and phenotypic data the evidence for a genetic component of the quantitative trait can be asserted. The next step would then be to include genetic marker data into the analysis to map QTL to known marker linkage groups. The approach taken here (only 2 chromosomes in the simulation) is also applicable to more realistic plant breeding situations in which the number of chromosomes and genome size is much larger.
Accurate knowledge on the positions of QTL and their contributions to the important traits opens the possibilities for marker assisted breeding and selection. Including pedigree structures also means that it becomes feasible to utilize data from breeding programs and more importantly to accumulate data over time and to improve the accuracy of parameter estimates for QTL and polygenic background which likely improves the efficiency of the breeding program (Podlich et al. 2004).
An ongoing discussion in modeling QTL is the number of alleles that may be expected in plant populations. Here, we used the bi-allelic model, which allows relatively simple extensions to dominance and epistatic actions of the QTL (Yi 2004; Yi et al. 2005). A single bi-allelic QTL model was shown to be less robust to situations with a simulated single multi-allelic QTL than the reverse scenario (Hoeschele et al. 1997). However, the latter case has been successfully accommodated by allowing multiple closely-linked bi-allelic QTL where the number of QTL is a random variable (results not shown). Alternatively to the bi-allelic QTL is the model assuming two unique allelic effects for every founder of the mapping pedigree, as may be done in a regression context (Jansen et al. pers. comm.). Another approach might be to include the number of alleles for a single QTL as a random variable in the model as was recently proposed (Jannink and Wu 2003). The estimation of the number of alleles may be hampered by the fact that only a limited number of founder alleles will be transmitted to the mapping populations and the differences in size of the allelic effects may not be large enough to distinguish all of them (Jannink and Wu 2003).
In our study we applied model selection for the number of QTL by allowing jumps between models differing in the number of QTL, i.e., a reversible jump sampler (Green 1995) was implemented to add or delete a QTL in the Markov chain simulation. The estimates for the Bayes factors (Kass 1993; Kass and Raftery 1995) were used to examine the evidence from data on the number of QTL affecting the quantitative trait. The use of Bayes factors seems more appropriate than the use of posterior probabilities for inference on the number of QTL as the latter are more sensitive to the prior specification (results not shown). Another Bayesian approach for QTL model selection may be to include all markers along the genome into the linear model and applying shrinkage to the individual marker contributions (Meuwissen et al. 2001; Xu 2003; ter Braak et al. 2005). That approach may be appealing as a fixed model space is assumed and the use of a reversible jump sampling algorithm can be avoided. However, the interpretation of the estimates for the QTL effects may be complicated due to shrinkage. A somewhat intermediate method is the use of a composite model space (Yi et al. 2005; Yi 2004) where the model parameter space is fixed but latent variables are introduced to indicate whether a putative QTL contributes to the quantitative trait.
In conclusion, we have presented a Bayesian approach that can be very flexible in modeling complex traits, allowing both genetic and environmental factors. The Bayesian approach automatically provides useful information on the remaining uncertainty on the estimates of the genetic variables, accounting for the uncertainty in other variables. The approach is very well suited for utilizing data from ongoing breeding programs as it automatically accounts for relationships among all individuals by including the known pedigree information. The Bayesian approach has been implemented into the software package FlexQTLTM (www.flexqtl.nl).
Abbreviations
- QTL:
-
Quantitative trait loci
- FPM:
-
Finite polygenic model
- TIM:
-
The infinitesimal model
- MCMC:
-
Markov chain Monte Carlo
- HPD:
-
Highest posterior density
- HPI:
-
Highest posterior intensity
References
Balding DJ, Bishop M, Cannings C (eds) (2003) Handbook of statistical genetics, 2nd edn. John Wiley & Sons
Bayes T (1763) An essay towards solving a problem in the doctrine of chances. Philoph Trans Roy Soc 330–418
Bink MCAM (2002) On flexible finite polygenic models for multiple-trait evaluation. Genet Res 80:245–256
Bink MCAM, Uimari P, Sillanpaa MJ, Janss LLG, Jansen RC (2002) Multiple QTL mapping in related plant populations via a pedigree-analysis approach. Theor Appl Genet 104:751–762
Falconer DS (1989) Introduction to quantitative genetics. Longman [etc.], Harlow [etc.]
Fisch RD, Ragot M, Gay G (1996) A generalization of the mixture model in the mapping of quantitative trait loci for progeny from a biparental cross of inbred lines. Genetics 143:571–577
Gelman A, Carlin JB, Stern HS, Rubin DB (1995) Bayesian data analysis. Chapman & Hall, London
Gelman A, Carlin JB, Stern HS, Rubin DB (2004) Bayesian data analysis, 2nd edn. Chapman & Hall, London
Gilks WR, Richardson S, Spiegelhalter DJ (1996) Markov chain monte carlo in practice. Chapman & Hall, London
Green PJ (1995) Reversible jump Markov chain Monte Carlo computation and Bayesian model determination. Biometrika 82:711–732
Haldane JBS (1919) The combination of linkage values and the calculation of distances between the loci of linked factors. J Genet 8:299–309
Heath SC (1997) Markov chain Monte Carlo segregation and linkage analysis for oligogenic models. Am J Hum Genet 61:748–760
Hoeschele I, Uimari P, Grignola FE, Zhang Q, Gage KM (1997) Advances in statistical methods to map quantitative trait loci in outbred populations. Genetics 147:1445–1457
Hyndman RJ (1996) Computing and graphing highest density regions. Am Statist 50:120–126
Jannink JL, Wu XL (2003) Estimating allelic number and identity in state of QTLs in interconnected families. Genet Res 81:133–144
Janss LLG, Thompson R, Vanarendonk JAM (1995) Application of Gibbs sampling for inference in a mixed major gene-polygenic inheritance model in animal populations. Theor Appl Genet 91:1137–1147
Jiang C, Zeng ZB (1997) Mapping quantitative trait loci with dominant and missing markers in various crosses from two inbred lines. Genetica 101:47–58
Kass RE (1993) Bayes factors in practice. Statistician 42:551–560
Kass RE, Raftery AE (1995) Bayes factors. J Amer Statist Assoc 90:773–795
Kennedy BW, Quinton M, Vanarendonk JAM (1992) Estimation of effects of single genes on quantitative traits. J Anim Sci 70:2000–2012
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits, 1 edn. Sinauer Associates, Sunderland, MA
Meuwissen TH, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829
Podlich DW, Winkler CR, Cooper M (2004) Mapping as you go: an effective approach for marker-assisted selection of complex traits. Crop Sci 44:1560–1571
Pong-Wong R, Haley CS, Woolliams JA (1999) Behaviour of the additive finite locus model. Genet Select Evol 31:193–211
Sillanpaa MJ, Arjas E (1998) Bayesian mapping of multiple quantitative trait loci from incomplete inbred line cross data. Genetics 148:1373–1388
Sorensen D, Gianola D (2002) Likelihood, Bayesian, and MCMC methods in quantitative genetics. Springer-Verlag, New York
ter Braak CJF, Boer MP, Bink MCAM (2005) Extending Xu’s Bayesian model for estimating polygenic effects using markers of the entire genome. Genetics 170:1435–1438
Uimari P, Sillanpaa MJ (2001) Bayesian oligogenic analysis of quantitative and qualitative traits in general pedigrees. Genet Epidemiol 21:224–242
Waagepetersen R, Sorensen D (2001) A tutorial on reversible jump MCMC with a view toward applications in QTL-mapping. Int Stat Rev 69:49–61
Xu S (2003) Estimating polygenic effects using markers of the entire genome. Genetics 163:789–801
Yi NJ (2004) A unified Markov chain Monte Carlo framework for mapping multiple quantitative trait loci. Genetics 167:967–975
Yi NJ, Yandell BS, Churchill GA, Allison DB, Eisen EJ, Pomp D (2005) Bayesian model selection for genome-wide epistatic quantitative trait loci analysis. Genetics 170:1333–1344
Acknowledgments
This paper has been carried out with financial support from the Commission of the European Communities, specific research program “Quality of Life and Management of Living Resource”, QLK5-2002-01492 “High quality Disease Resistant Apple for a Suistanable Agriculture”, coordinated by L. Gianfranceschi from the University of Milan (It.). This manuscript does not necessarily reflect the Commission’s views and in no way anticipates its future policy in this area. Its content is the sole responsibility of the publishers. We especially acknowledge the contributions of F. Laurens, C.-E. Durel, and A. Kouassi from INRA (Fr) and the helpful comments from two anonymous reviewers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License ( https://creativecommons.org/licenses/by-nc/2.0 ), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Bink, M.C.A.M., Boer, M.P., ter Braak, C.J.F. et al. Bayesian analysis of complex traits in pedigreed plant populations. Euphytica 161, 85–96 (2008). https://doi.org/10.1007/s10681-007-9516-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-007-9516-1